Abstract
Objectives
We retrospectively analyzed the next-generation sequencing (NGS) results from diagnosed NSCLC patients to identify and compare genomic alterations of NSCLC between Moroccan patients and the Cancer Genome Atlas (TCGA). We also aimed to investigate the distribution and frequency of concurrent genomic alterations.
Methods
From December 2022 to December 2023, a retrospective study of 76 formalin-fixed paraffin-embedded (FFPE) samples have been profiled using the Oncomine™ Precision Assay on the Ion Torrent™ Genexus™ Integrated Sequencer across the panel of 50 key genes that are applicable for the selection of targeted therapy.
Results
Seventy of the 76 FFPE sequenced samples carried at least one genetic alteration in the tested genes. The study identified 234 genetic alterations in 18 genes. Targetable genetic alterations in EGFR, KRAS, MET, BRAF, ALK, RET and ROS1 were identified in 84.3% of tumors. EGFR and KRAS mutations were frequently reported, occurring in 24.3% and 22.9% of cases, respectively. The untargetable genetic alterations were found in 74.3% of the specimens in FGFR3, TP53, ERBB2, PIK3CA, CDKN2A, PDL1, FGFR1, PTEN, CHEK2 and ERBB3. There were additional uncommon/rare mutations in EGFR, BRAF, RET and ROS1. Comparing the prevalence of selected mutated genes in the NSCLC patients from the TCGA database identified substantial differences in EGFR (24.3%, vs14.97%), KRAS (22.9%, vs 25.99%), and TP53 (34.3%, vs 50.94%). ALK, ROS1, and RET gene rearrangements were detected in 4.3% of the 70 tumors tested. The ALK/RET/MET/ROS1/EML4 fusions were detected in 11.4% of samples. Co-alterations occurred in 67.1% of specimens. Co-occurring driver gene mutations were observed in 44.3%. TP53 mutations co-occurred driver gene mutations in 30% of tumors. Three cases (4.3%) harbored concurrent FGFR3, TP53, and PIK3CA alterations.
Conclusion
Our results regarding the proportion of samples with actionable mutations demonstrate the value of NGS testing for NSCLC patients in a real-world clinical diagnostic setting.
Keywords
Introduction
Lung cancer remains one of the most lethal cancers, contributing significantly to global mortality rates and being a leading cause of cancer-related deaths. 1 It was expected that over 1.8 million cancer-related deaths and over 2.2 million new cancer cases would have occurred globally in 2020, accounting for 18.0% of all cancer-related deaths and 11.4% of all cancer diagnoses, respectively. It is the foremost cause of cancer-related death in men and the second in women, following breast cancer. 1
Lung cancer has a significantly lower 5-year survival rate (7%–25%) compared to other major malignancies. 2 By 2035, it is projected to cause up to 3 million deaths globally. It is expected that the number of deaths from lung cancer will increase twofold for both males (from 1.1 million in 2012 to 2.1 million in 2035) and women (from 0.5 million in 2012 to 0.9 million in 2035), with the gender gap continuing to exist. Especially for the East Mediterranean and African regions, which are predicted to see the fastest rate of growth. 3 In North Africa, lung cancer incidence is rising rapidly, becoming the leading cancer among men in Morocco (20.8%), Algeria (13.6%), Tunisia (20.2%), and Libya (18.9%). The mortality and age-standardized incidence rates are 9.9 and 11.1 per 100,000, respectively. 4 In 2020, Morocco reported the highest incidence and mortality rates in Northern Africa, with 7353 new cases and 6551 deaths.5,6
Moroccan's lung cancer clinical and histopathological pattern is characterized by a decrease in the age at diagnosis, an increase in female cases, and an advanced tumor stages. The most significant histological subtype is adenocarcinoma. 7 Histologically, there are two primary forms of lung cancer: non-small cell lung cancer (NSCLC), which constitutes 85% of all lung cancers, and small cell lung cancer (SCLC), accounting for the remaining 15% of all lung cancers. 8 NSCLC is further subdivided into three histologic subtypes: adenocarcinoma (50-60%), squamous cell carcinoma (20-30%), and large-cell carcinoma (15%) of lung cancer diagnoses. 9 Adenocarcinoma, a peripheral type of cancer, is more common among women and non-smokers. It often tests positive for targetable driver mutations, including those in EGFR, ALK, ROS1, BRAF, NTRK, MET and RET. Actionable mutations, gene amplifications, or oncogenic fusion transcripts are found in more than 65% of patients with NSCLC. Over 65% of NSCLC patients exhibit actionable mutations, gene amplifications, or oncogenic fusion transcripts. 10
The emergence of immunotherapies and molecularly targeted treatments has revolutionized the management of NSCLC with oncogenic drivers, significantly enhancing patient overall survival (OS) compared to traditional chemotherapy.11,12 Tyrosine kinase inhibitors (TKI) against the driver mutations, as well as immune checkpoint blocking therapy, represented by inhibitors of PD-1/PD-L1 and CTLA-4, may replace or supplemented chemotherapy in eligible patients. 13 Recently, the US Food and Drug Administration (FDA) approved a targeted therapy for KRAS G12C-mutant NSCLC, a driver mutation present in approximately 13% of NSCLC patients. Further new targeted therapies are being developed to address gene alterations associated with ERBB2 (HER2), MET, NRG1, and FGFR. 14
The practice of single-gene testing for NSCLC despite guidelines recommending comprehensive genomic profiling (CGP) remains common in various countries and regions. For instance, in the United States, up to 40% of patients with advanced NSCLC still receive single-gene testing rather than CGP. 15 Single-gene tests are frequently used to test for specific mutations such as EGFR and ALK. However, these tests are less likely to identify fusion abnormalities such as RET, MET, and NTRK. Additionally, single-gene testing is prevalent in many community practices where CGP is less accessible. This practice variation can lead to incomplete molecular profiling and delayed treatment decisions. 16 Single-gene tests are selectively used by oncologists because they are individually less expensive, simpler to interpret, and have a faster turnaround time (TAT) compared to CGP. Nevertheless, these tests lack the sensitivity of CGP due to inherent methodological limitations, which can cause them to miss clinically relevant variations. 17 This practice variation can lead to incomplete molecular profiling and delayed treatment decisions, ultimately impacting patient outcomes.18,19 Timely identification of actionable driver oncogenes is crucial for implementing biomarker-driven therapies and enhancing OS in patients with NSCLC. 20
Multigene Next Generation Sequencing (NGS) testing is pivotal in precision medicine, offering personalized therapy and a comprehensive molecular profile for NSCLC. Compared to single-gene sequencing, multigene NGS testing identifies genomic alterations in a higher number of NSCLC patients. This advanced technology allows for the simultaneous analysis of multiple genes across numerous samples, significantly reducing both costs and TAT. 21
Here, we applied the Oncomine™ Precision Assay (OPA) on the Ion Torrent™ Genexus™ Sequencer in NSCLC patients in order to investigate the prevalence of genomic alterations which indicate sensitivity or resistance to targeted therapies. In addition, we compared the results with The Cancer Genome Atlas (TCGA) NSCLC dataset, which involves mainly Western populations. We also evaluated the incidence and patterns of concurrent genomic aberrations which contribute to the heterogeneity of oncogenic driver-defined subgroups of NSCLC.
Materials and Methods
Patients
We retrospectively collected a cohort of 76 formalin-fixed paraffin-embedded (FFPE) samples of Moroccan patients with NSCLC. The patients were selected consecutively from the Department of Pathology, the Nations-Unites Pathology Center of Rabat between December 2022 and December 2023. The study size was determined based on the availability of the FFPE blocks during these specified time periods. All tissue sections were evaluated after staining using hematoxylin/eosin (H&E) by a histopathologist.
To minimize potential sources of bias, eligible patients were newly histologically diagnosed with lung cancer according to the World Health Organization (WHO) Classification of Tumors, sixth edition, published in 2021. 22 The sixth edition of the WHO criteria provides updated and comprehensive guidelines for the histological classification of lung cancer, ensuring consistency and accuracy in diagnosis. This classification system categorizes lung cancer into two broad histological subtypes: small-cell lung cancer (SCLC) and non-small-cell lung cancer (NSCLC). Participants were selected to represent a diverse demographic, covering various ages, genders, tumor types, tumor locations, and tumor grades. Patients who received neoadjuvant chemotherapy, immunotherapy, targeted therapy, or radiotherapy or were diagnosed with other lung cancers were excluded.
We have followed relevant Equator guidelines, and the reporting of this study conforms to the Strengthening the Reporting of Observational Studies in Epidemiology (STROBE) checklist. 23 Written informed consent was signed and obtained from all participants before their inclusion in the study. And we have de-identified all patient details. This study was approved by the Ethics Committee for Biomedical Research (CERB) of the Faculty of Medicine and Pharmacy in Rabat, with approval number; CERB 36–23 and approval date; January 22, 2024.
Sequencing Protocol
The targeted sequencing of 76 FFPE samples was performed on the Ion Torrent™ Genexus™ Integrated Sequencer (Thermo Fisher Scientific, Waltham, MA, USA) using the (OPA, Thermo Fisher Scientific).
A completely automated NGS system that combines library preparation, template preparation, sequencing, and data analysis is the Ion Torrent™ Genexus™ Integrated Sequencer. Genomic alterations were detected on the same platform via Ion semiconductor sequencing (Ion Torrent™ Technology, Thermo Fisher Scientific).
We utilized the Oncomine™ Precision Assay GX (OPA, catalog number A46291), which is designed to detect actionable alterations. This panel covers 78 variants, including 45 mutations, 14 copy number variations (CNVs), 19 fusion variants, and various hotspot mutations (substitutions, insertions, and deletions) across 50 key genes relevant for targeted therapies. The panel also includes a 5’/3’ expression imbalance caller for identifying novel fusions. The specific genes analyzed are listed in Table 1.
Oncomine Precision Gene List and the Associated Biological Processes and Molecular Functions.

*: DNA Hotspot, !: CNV, #: Inter genic fusions, +: intra genic fusions, NA: not available.
Somatic variants and fusion transcripts were called using Ion Reporter with specific Browser Extensible Data (BED) files and appropriate filters. We verified the identified variants using Integrative Genomic Viewer version 2.3 (or higher) and identified short reads with potential mispriming events.
For the final report, sequences were aligned with the reference genome hg19 using Oncomine™ Knowledgebase Reporter Software (Thermo Fisher Scientific). We set the mutation frequency threshold at 2%, with a median coverage depth exceeding 1000X.
The TAT covered the entire process, starting from the acquisition of FFPE samples and culminating in the electronic validation of the NGS report using hospital software. This comprehensive workflow included steps such as deparaffinization, nucleic acid extraction, sequencing, genomic alteration analysis, and post-analytical procedures.
Statistical Analysis
The statistical analysis was performed using the χ2 test or Fisher's exact tests, with a significance threshold set at p ≤ 0.05. In our analyses, sex was treated as a categorical variable, with patients grouped into females and males. This categorization was based on the established understanding that the risk of NSCLC is more frequent in males. This approach allowed us to investigate potential sex-based differences in NSCLC characteristics. All analyses were conducted using the SPSS (Statistical Package for the Social Sciences) version 23 (IBM Corp.)
Results
Patient Characteristics
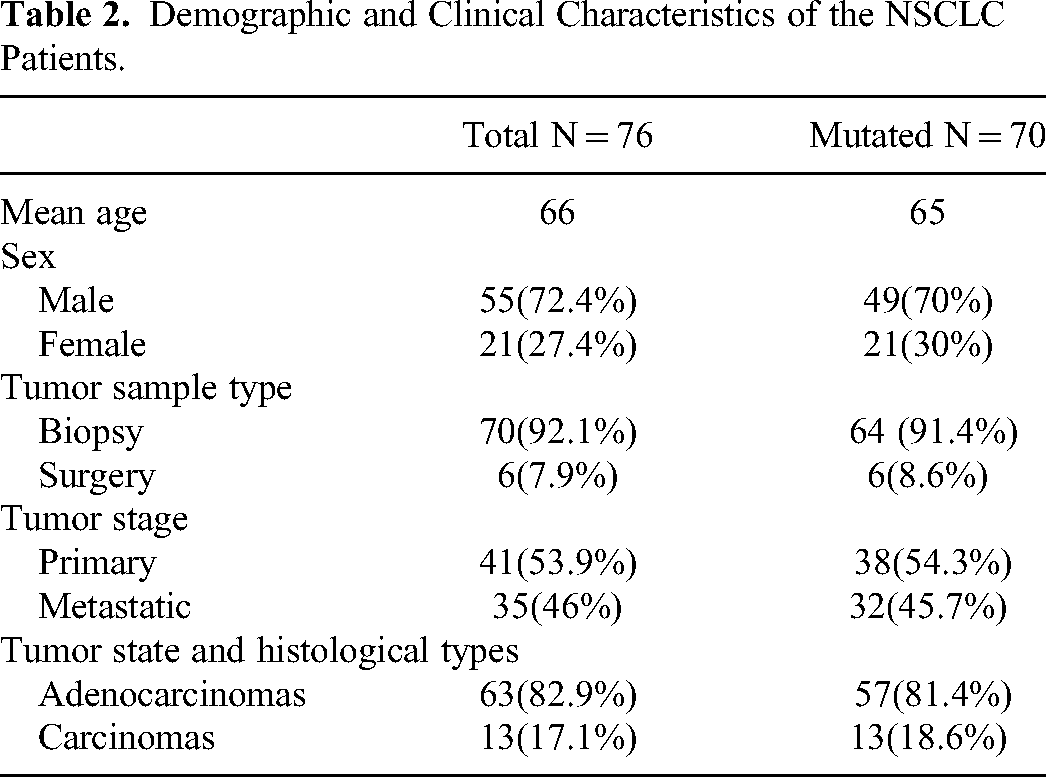
A total of 76 FFPE samples from NSCLC patients were successfully sequenced in this study. The mean age of diagnosis was 65 years with a range of 40–90 years. There were 18.4% (n = 14) patients aged 65 years or younger with a median age of 45.5 (range 40-51 years) and 81.6% (n = 62) patients were 52 years or older with a median age of 74.5 (range 49-90 years) at diagnosis. The overall male-to-female ratio was 2.6:1 (72.4%, n = 55 vs 27.4%, n = 21, p = 0.72) with a p-value of 0.72, determined using the χ² test (or Fisher's exact test). The majority of tumors with known histology were classified as adenocarcinomas (82.9%, n = 63). Baseline characteristics are displayed in Table 2.
Demographic and Clinical Characteristics of the NSCLC Patients.
Overall Genetic Alterations status Detected
Mutation analysis of the 50 gene NGS panel revealed the presence of at least one genetic alteration in 92.1% (n = 70) of the tumors tested. The 70 NSCLC patients showed a median number of three genetic alterations (range 1 to 12). A total of 234 genetic alterations were found in 18 genes. Among the 18 genes, 18.6% (13/70) of NSCLC patients were found to have a single alteration and 81.4% (57/70) harbored multiple alterations: 27.1% (19/70) had double alterations, 17.1% (12/70) had triple alterations, and 37.1% (26/70) had more than three alterations. Distributions of altered genes in 70 patients with NSCLC are shown in Figure 1.

Genomic alterations detected in 70 NSCLC patients.
Targetable genetic alterations in EGFR, KRAS, MET, BRAF, RET, ALK, and ROS1 were identified in 84.3% (n = 59/70) of NSCLC patients. There were no alterations detected in the NTRK gene. EGFR and KRAS mutations were reported most frequently (24.3%, n = 17/70 and 22.9%, n = 16/70, respectively), followed by MET (12.8%, n = 9/70), BRAF (7.1%, n = 5/70), RET (7.1%, n = 5/70), ALK (5.7%, n = 4/70) and ROS1 (4.3%, n = 3/70).
Regarding common EGFR driver mutations, 14.3% (n = 10/70) were deletions in exon 19 or point mutations in exon 21 [10% (n = 7/70) and 4.3% (n = 3/70), respectively]. Mutations in exons 18 and 20 were also detected [1.4% (n = 1/70) and 8.6% (n = 6/70), respectively]. Amplification of EGFR was identified in 7.1% (n = 5/70). The uncommon/rare EGFR mutations were A289 T, E709_T710 delinsD, R836C, and which are found in exons 7, 18 and 21, respectively. These three mutations account for 17.6% of EGFR mutation-positive NSCLC and 4.3% (n = 3/70) of all NSCLC patients. The T790 M resistance mutation was found in 8.6% (n = 6/70) of all NSCLC mutant samples. The majority of KRAS genomic aberrations were single nucleotide variations (SNVs), accounting for 28.6% (n = 20/70) of all KRAS alterations. KRAS amplifications were the second most common type of alteration, accounting for a less proportion of only 5.7% (n = 4/70). Among all patients included, 7.1% (n = 5/70) carried G12C mutation whereas 14.3% (n = 10/70) harbored non-G12C mutations. Ratio of G12C versus non-G12C mutations was 1:2. MET amplification and exon 14 skipping mutations were comprised 11.4% (n = 8/70) and 1.4% (n = 1/70), respectively. BRAF changes included 2.8% (n = 2/70) V600 and 2.8% (n = 2/70) non-V600 mutations. Uncommon/rare BRAF exon 15 mutation V600_K601delinsE was observed in 1.4% (n = 1/70) of NSCLC tumors. RET R912Q, RET R912 W and ROS1 D2033N mutations, relatively uncommon/rare, were identified in 4.2% (n = 3/70) of patient with NSCLC. ALK/ROS1/RET gene rearrangements appeared three times out of 70 mutated tumors. The ALK/RET/MET/ROS1/EML4 fusions were detected in 11.4% (n = 8/70) of NSCLC patients including 2.8% (n = 2/70) EML4-ALK, 2.8% (n = 2/70) ALK, 1.4% (n = 1/70) CD74-ROS1, 1.4% (n = 1/70) MET-MET, 1.4% (n = 1/70) RET-KIF13A and 1.4% (n = 1/70) RET. There were two mutations hotspots, c.527G > T (p.C176F) observed in TP53 and c.35G > A (p.G12D), observed in KRAS. Table 3 displayed the prevalence of genetic alterations that driver NSCLC.
Frequencies and Mutation Characteristics of Detected Driver Oncogenes in the 70 NSCLC-Mutated Samples.

NA: not available.
Several other genetic alterations with no available targeted drugs were found in 74.3% (52/70) NSCLC patients: FGFR3 (40%, n = 28/70), TP53 (34.3%, n = 24/70), ERBB2 (12.9%, n = 9/70), PIK3CA (8.6%, n = 6/70), CDKN2A (5.7%, n = 4/70), PDL1 (5.7%, n = 4/70), FGFR1 (4.3%, n = 3/70), PTEN (2.8%, n = 2), CHEK2 (1.4%, n = 1/70) and ERBB3 (1.4%, n = 1/70).
Comparing the prevalence of 18 mutated genes in the NSCLC patients from the TCGA database identified substantial differences were found in the following genes; EGFR (24.3%, vs14.97%), KRAS (22.9%, vs 25.99%), and TP53 (34.3%, vs 50.94%). The Comparison of the frequencies of selected mutated genes in NSCLC patients between the two cohorts is represented in the Figure 2.
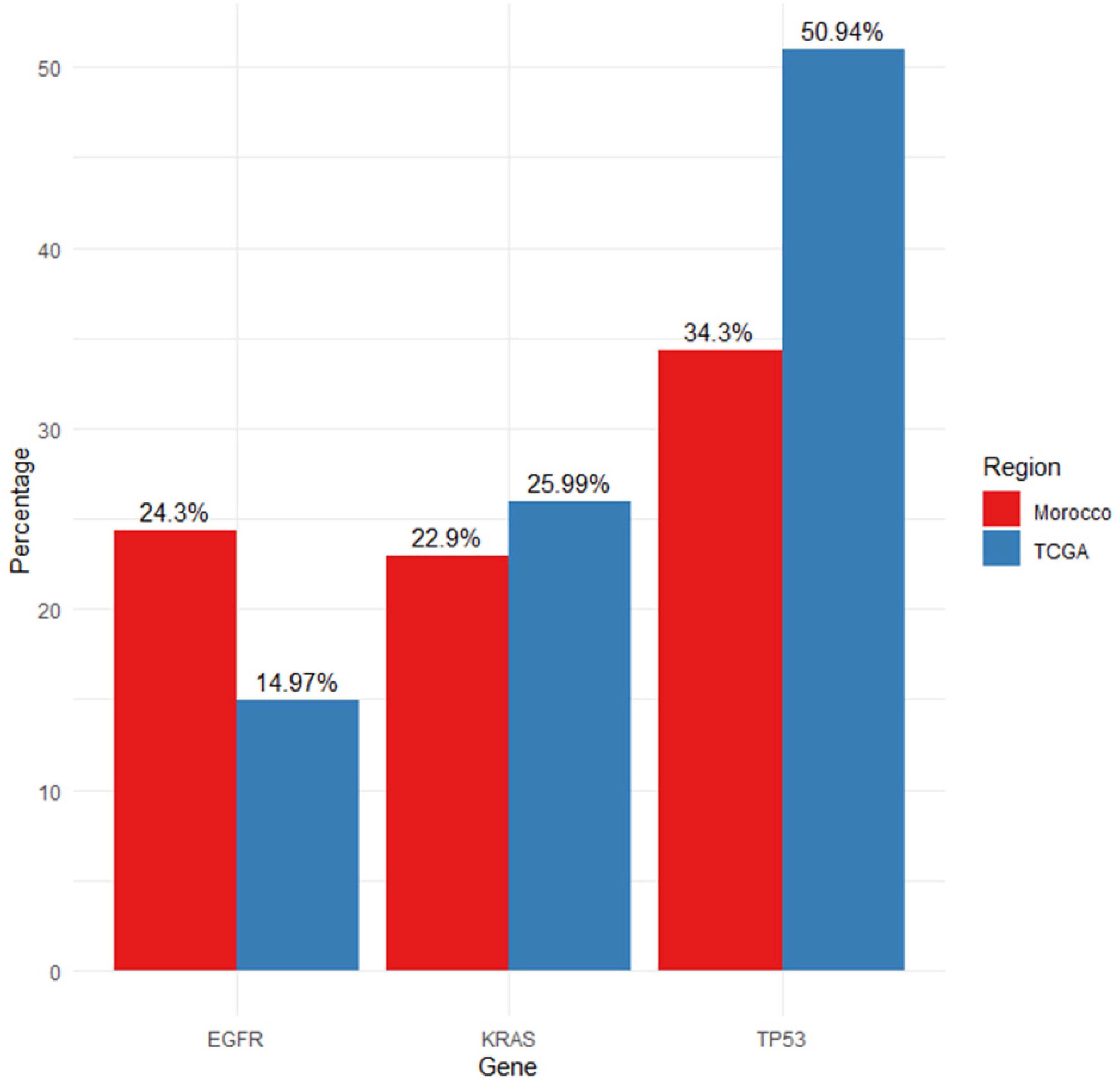
Comparison of the frequencies of selected mutated genes in NSCLC patients between the Moroccan and TCGA cohorts.
Co-Occurring Genetic Alterations
Co-occurring genetic alterations were present in 67.1% (n = 47/70), including 27.1% (n = 28/70) and 54.3% (n = 19/70) of tumors samples had 1–2 and ≥ 3 co-occurring genomic alterations, respectively. Genes with the highest co-occurrence genetic aberrations were FGFR3 (55.7%, n = 39/70), followed by TP53 (51.4%, n = 36/70). Co-occurring driver gene mutations were observed in 44.3% (n = 31/70) of patients. TP53 mutations co-occurred driver gene mutations in 30% (n = 21/70). EGFR and KRAS mutant genes were mostly co-occurring with other genes in 40% (n = 28/70) and 32.9% (n = 23/70) respectively. T790 M in EGFR and G12C in KRAS mutations co-occurring in (11.4%, n = 8/70) and (10%, n = 7/70), respectively. Three cases (4.3%, n = 3/70) harbored concurrent, TP53, FGFR3 and PIK3CA alterations; FGFR3 and PIK3CA amplifications co-ocurred with TP53 (V157F), TP53 (C176 W, C176G, P177_C182del) and TP53 (V216 M). The details of the co-occurrence of altered genes identified in 70 mutated NSCLCs are displayed in Table 4.
Co-Occurring Gene Alterations Identify in 70 Mutated NSCLC

Discussion
The most prevalent kind of lung cancer, NSCLC, has a high incidence of acquired somatic abnormalities. Progression-free survival (PFS) and OS rates in NSCLC patients have increased as a consequence of the use of targeted treatments, which have significantly improved patient outcomes. 24 The new guidelines from the National Comprehensive Cancer Network (NCCN), the European Society of Medical Oncology (ESMO), the American Society of Clinical Oncology-Ontario Health (ASCO-OH), and the International Association for the Study of Lung Cancer/College of American Pathology (IASLC/CAP) all suggest extensive testing using NGS technical platforms to identify immunochemistry for PD-L1 and clinically relevant genomic alterations in EGFR, ALK, ROS1, BRAF, HER2, KRAS, MET, NTRK, and RET.25–28
In general, when diagnosing NSCLC within different populations, it is critical to compare the diagnostic yields of various methodologies to underscore the effectiveness of the approach used. Single-gene tests are frequently used for specific mutations in NSCLC, such as EGFR and ALK. However, these tests are less likely to identify fusion abnormalities such as RET, MET, and NTRK, leading to a lower diagnostic yield. For example, yields from single-gene testing in NSCLC are typically around 10%–20%, depending on the prevalence of the mutation and the specific group being tested. 29 While small target gene panels improve yields by testing for multiple actionable mutations simultaneously in NSCLC (eg, EGFR, ALK, KRAS, and BRAF). Diagnostic yields with small panels vary between 30% and 50%, depending on the specific genes included and the population being examined. 30 Large target gene panels encompassing hundreds of genes greatly enhance the successful diagnosis rate, frequently reaching 50%–75%. This method enables a more comprehensive detection of genetic changes in NSCLC, particularly those that are infrequent or uncommon. 30 Whole Exome/Genome Sequencing (WES/WGS) is increasingly used as an important technique for NSCLC because it has a thorough method of identifying genetic modifications with diagnostic yields reaching 70–90%. WES and WGS can find SNVs, CNVs, and structural rearrangements like gene fusions. These genetic alterations are crucial in driving targeted therapy decisions for NSCLC. 29 For Moroccan population, molecular testing for alterations in driver mutation genes and targeted therapies has become a standard procedure. Advanced techniques identify rare mutations, comprehensively understanding the NSCLC genetic landscape. 31 In advanced Moroccan NSCLC patients, multigene NGS testing has become increasingly important for detecting driver mutation genes. This study demonstrates the efficacy of the current approach in providing a robust and comprehensive diagnostic tool for NSCLC in the Moroccan population.
In our study, Targetable genetic alterations in EGFR, KRAS, MET, BRAF, RET, ALK, and ROS1 were identified in 84.3% (n = 59/70) of NSCLC patients. There were no alterations detected in the NTRK gene. EGFR and KRAS genetic alterations were the most frequent (24.3% and 22.9%, respectively), followed by MET (12.8%), BRAF (7.1%), RET (7.1%), ALK (5.7%), and ROS1 (4.3%). Our data corroborate the genetic heterogeneity of NSCLC worldwide, showing that EGFR mutation prevalence in the Moroccan population is slightly higher than that seen in NSCLC patients of Caucasian ethnicity (≈15%) but is lower than that identified in Asian NSCLC patients (≈50%).32,33 Additionally, a comparison of driver gene mutations in our NSCLC patients with the TCGA dataset showed that EGFR was mutated at a much higher frequency in our Moroccan cohort compared with Caucasians.
In contrast, KRAS and TP53, the most common mutated genes in Caucasians, were found in only 22.9%, and 34.3%, respectively, of our Moroccan patients with NSCLC. Concerning the mutational frequencies of other druggable mutations also showed notable differences compared to the TCGA cohort. Specifically, MET, BRAF, RET, ALK and ROS1 mutations were found in 12.8%, 7.1%, 7.1%,5.7% and 4.3%, respectively of our Moroccan NSCLC patients, which is not reported in the TCGA dataset for Caucasian patients. These discrepancies can be attributed to racial/ethnic backgrounds and environmental factors such as smoking and exposure to pollutants that have been linked to variations in mutational profiles of NSCLC across different populations, as supported by several studies.31,33–37
The geographical features of Moroccan NSCLC patients, including genetic diversity and environmental exposures, play a crucial role in the observed mutational profiles. Studies have shown that the prevalence of certain mutations, such as EGFR, differs from region to region, ranging from 21.7% to 23.5%. Otherwise, there is a significant lack of data from other regions of our country, indicating a possible absence of regular mutation testing and subsequent availability of EGFR-targeted drugs. This indicates the necessity for additional research efforts. 38 Morocco lacks comprehensive regional cancer records, particularly in the southern regions. According to Harkati et al, there are notable differences in the prevalence of lung cancer and its relationship with lifestyle factors between the populations of the Southern and Northern regions. These differences can be attributed to variations in customs, rituals, and behaviors. Harkati et al, also discovered differences in exposure to risk factors for lung cancer across different regions, which help to explain the varying incidence rates in Southern Morocco particularly those related to lifestyle, food consumption, and the professional environment. 39 Another local study has shown that these differences across regions are caused by many factors such as delayed diagnosis, ongoing reliance on conventional treatments, and limited access to advanced therapies like tyrosine kinase inhibitors, targeted therapy, and immunotherapy. As a result, traditional chemotherapy continues to be the primary option. 40 These differences highlight the importance of considering regional genetic variations when developing targeted therapies. Understanding these geographical features is essential for optimizing treatment strategies and improving patient outcomes in the Moroccan population.
To fill our research gap, our study focuses on analyzing the molecular profile of NSCLC patients from Morocco to identify the possible genetic modifications associated with the disease. Although the sample size is limited and there is a lack of clinical data, the findings could enhance a comprehensive genetic database for tailored treatment strategies.
Regarding common EGFR driver mutations, deletions in exon 19 and the L858R substitution in exon 21 are the two most common mutations in EGFR, accounting for 58.8% of EGFR mutation-positive NSCLC and 14.3% of all NSCLC patients. This increase has important clinical implications because in EGFR-mutant NSCLC harboring exon 19 deletion or L858R mutations, EGFR TKIs are the current standard of care for first-line treatment. The first-generation reversible EGFR TKIs Gefitinib and Erlotinib and the irreversible second-generation multi-targeted agents Afatinib and Dacomitinib were found to be highly active compared with standard chemotherapy in NSCLCs harboring EGFR-sensitive mutations.41,42
Besides common EGFR driver mutations, uncommon/rare mutations in EGFR gene were detected in 17.6% of EGFR mutation-positive NSCLC and 4.3% (n = 3/70) of all NSCLC patients. Uncommon/rare EGFR mutations included A289 T, E709_T710 delinsD, and R836C in exons 7, 18 and 21, respectively. EGFR A289 T is present in 0.07% of American association for cancer research (AACR) GENIE cases, with conventional glioblastoma multiforme, glioblastoma, anaplastic astrocytoma, lung adenocarcinoma, and astrocytoma having the greatest prevalence. 43 The finding of uncommon/rare EGFR mutations confers the challenge of treatment decision. Studies on the delE709_T710insD mutation in exon 18 and its response to EGFR-TKIs, including Gefitinib, Erlotinib, Afatinib, and Dacomitinib have been reported in recent years.44,45 Generally, NSCLC patients with EGFR exon 18 mutations respond slightly worse to first-generation EGFR-TKI compared to patients with TKI-sensitizing EGFR mutations,46–48 previous studies suggested that R836C showed inconsistent responses to Gefitinib49,50 or modest survival. 51
The prevalence of uncommon/rare EGFR mutations varied between 1.0% and 18.2% of EGFR-mutant NSCLC. 52 The frequency of uncommon/rare mutations in EGFR gene is completely dependent on the methodology across studies including differences in mutation detection methods and classification. In our study, T790 M and exon 20 insertions were excluded as uncommon EGFR mutations because these are known to be associated with first-generation TKI resistance but third-generation TKI sensitivity.53–56 In a large cohort of Brazilian patients with NSCLC, EGFR mutations were observed in 24.2% of samples, including exon 19 (12.8%), exon 21 L858R (6.9%) and exon 20 insertion (1.6%). All others were considered uncommon/rare mutations and were observed in 18.5% of all mutated patients and 4.0% of the total sample (2.3%-18.7% depending on the sequencing method). 57 In addition, the type of method used, directed or non-directed against specific mutations, influences the analysis, particularly for uncommon mutations. 57
Therapies targeting driver mutations have provided promising outcomes in relevant populations, including those with mutations in KRAS gene. G12C mutation is the most frequent KRAS point mutation in NSCLC and has been proven to be a predictive biomarker for direct KRAS G12C inhibitors in advanced solid cancers. KRAS G12C accounts for approximately 50% of KRAS mutations in NSCLC, and approximately 14% of patients with NSCLC have a KRAS G12C mutation. 58 In our study, 7.1% of patients with NSCLC harbor the KRAS G12C mutation and are therefore potentially responsive to the new anti-KRAS agents. The KRAS G12C mutation has become a druggable target in NSCLC. Sotorasib and Adagrasib have received approval from the FDA for treating NSCLC patients with KRAS G12C mutation after progression to initial therapy.59,60 As reported in numerous publications, Sotorasib achieved an objective response rate (ORR) of 32.2% (19 of 59) a median PFS of 6.3 months in a phase 1 cohort, an ORR of 37.1% (46 of 124), and a median PFS of 6.8 months in a phase 2 cohort of previously treated patients with locally advanced or metastatic NSCLC carrying the KRAS G12C mutation.61,62 Additionally, Adagrasib demonstrated a 96% disease control rate (49 of 51) and an ORR of 45% (23 of 51) in a cohort of NSCLC patients enrolled in a phase 1/2 clinical trial for solid tumor patients with a KRAS G12C mutation. 63 These two relevant inhibitors, Sotorasib and Adagrasib, inhibited the oncogenic signals and allowed the wild-type KRAS that remained to function normally by locking the exon 2 KRAS G12C mutation in an inactive state.61,64,65
BRAF mutations have been reported in about 4% of NSCLC. 66 The most frequent BRAF mutation in NSCLC and other cancers is the V600E mutation in exon 15, which accounts for ≈ 50% of BRAF-mutated NSCLC. BRAF V600E mutations specifically occur in ≈ 1–2% of patients with NSCLC. In our study, BRAF aberrations changes included 2.8% (n = 2/70) V600 and 2.8% (n = 2/70) non-V600 mutations. BRAF V600E represents an important biomarker that guides treatment decision-making. The ESMO guidelines recommend the BRAF inhibitor-Dabrafenib-combined with the MEK inhibitor-Trametinib-for BRAF-inhibitor naive patients with BRAFV600-mutated NSCLC. The combination of Dabrafenib and Trametinib is the preferred first-line treatment option in patients with advanced BRAF V600-mutated NSCLC, with an ORR of 64%, a median PFS of 10.2 months, a median overall survival of 24.6 months, and a median duration of response of 10.4 months, according to a pivotal phase 2 study. These outcomes remain inferior to those achieved with other targeted therapies in advanced NSCLC with other driver alterations. 67 Theoretically, many non-BRAF V600 mutations are kinase-impaired and, as a result, inappropriate for treatment targeting RAF. Because BRAF non-V600E mutant NSCLC is a rare mutation and there aren't many clear guidelines about how to treat it, Abouali et al suggested that NSCLC patients with BRAF non-V600E mutations should get front-line checkpoint immunotherapy, with or without platinum-based chemotherapy in the absence of any other driver mutations. 68 Furthermore, V600_K601delinsE, a rare BRAF exon 15 mutation, was discovered in one NSCLC patient. Thyroid gland papillary carcinoma, cutaneous melanoma, lung adenocarcinoma, pancreatic adenocarcinoma, and colon adenocarcinoma have the highest frequencies of BRAF V600_K601delinsE, which is detected in 0.05% of AACR GENIE cases. 43 Two other patients with metastatic melanoma who carried rare BRAF exon 15 mutations, A598_T599insV and V600_K601delinsE, were reported by Rogiers et al to have benefited clinically and shown a radiological response to inhibitors that target the mitogen-activated protein kinase pathway. 69 This emphasizes how crucial it is to employ testing methods that identify both V600E/K and non-V600E/K BRAF mutations in patients with uncommon, but potentially actionable, BRAF exon 15 mutations to maintain the option of targeted therapy treatment.
RET gene is a new type of proto-oncogenic driver gene, which has been proved to be closely related to the occurrence and development of various tumors including lung cancer. Alterations involving RET are rare, occurring in less than 1% of most cancers. But they are more common in lung cancer and occur in 10% to 20% of thyroid cancers. The most common alterations in RET are RET Mutation (2.05%), RET Fusion (0.36%), RET Loss (0.33%), RET Codon 918 Missense (0.11%), and RET M918 T (0.11%). 43 Selective RET inhibitors (sRETi), namely Pralsetinib and Selpercatinib, are targeted drugs approved by the US FDA for treating RET-mutated tumors. Pralsetinib and Selpercatinib have shown the ability to overcome resistance to multikinase inhibitors. Furthermore, ongoing clinical trials are investigating several second generation sRETis that are specifically designed to target solvent front mutations, which pose a challenge for first-generation sRETis. The effective screening of patients is the first crucial step in the clinical application of RET-targeted therapy. In our study, genomic RET mutations was found in two patients with NSCLC. These mutations were: R912Q and R912 W, which are rare mutations and none of them, have been reported in TCGA before. The R912Q and R912 W mutations were found in exon 16, predominantly occurring in the cysteine-rich domain and tyrosine kinase domains.
ROS-1 gene alterations have been identified in various cancers, including NSCLC. ROS-1 gene alterations are driver mutations in 1–2% of patients with NSCLC. 70 Alterations in ROS-1 are also found in other types of cancer. ROS-1 is currently targeted by several specific TKIs, but only two of these, Crizotinib and Entrectinib, have received US FDA approval. In our study, A ROS-1 D2033N mutation was detected in one NSCLC patient. ROS-1 D2033 is the most common resistance mutation in ROS1-rearranged NSCLC. ROS-1 D2033N is present in 0.04% of AACR GENIE cases, with lung adenocarcinoma having the greatest prevalence. 43 ROS-1 D2033N lies within the protein kinase domain of the ROS-1 protein. ROS-1 D2033N has been demonstrated to confer resistance to ROS-1 TKIs and increase cell migration and invasion in the context of ROS-1 fusions in culture but has not been individually characterized and therefore, its effect on ROS-1 protein function is unknown. 71
It is important to point out that the current study has some limitations. Firstly, it is a single-center analysis of the genomic profiling of NSCLC; it doesn't represent the overall situation in Morocco, which may lead to regional bias. Secondly, this was a retrospective study with a small sample size and lacking information on clinic-pathological features. However, this study also has important strengths. Using a multigene NGS panel allows for a comprehensive analysis of multiple gene mutations simultaneously, providing detailed insights into the genetic landscape of NSCLC in Moroccan cohort. This study marks a significant advancement in understanding and treating NSCLC in Moroccan patients.
Conclusion
Our results regarding the proportion of samples with actionable mutations demonstrate the value of NGS testing for NSCLC patients in a real-world clinical diagnostic setting. The detection of multiple actionable mutations contributes significantly to determining priorities in NSCLC treatment strategies in Moroccan population. Uncommon/rare alterations may constitute new druggable targets based on the molecular background of NSCLC tumors. Our findings suggest that co-occurring alterations should be considered more in the diagnosis and treatment of NSCLC patients.
Footnotes
Abbreviations:
Acknowledgements
This study was supported by the team of the Department of Pathology, Nations-Unites Pathology Center, Rabat, Morocco, whom we thank a lot.
Authors’ Contributions
SElZ, AL, MG, designed, conceived the study, wrote the manuscript, made the analysis and the interpretation of the data. SElZ, MG, AB, FK, contributed to the collection of samples. SElZ, YB, SB, HLA, HElA, RA, MRT, SElK, BElM, ElAB, contributed to the manuscript's critical assessment. SElZ, IAL, RAElH, KE, revised the final draft of the document. All authors read and approved the final manuscript.
Availability of Data and Material
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethics Approval
This study was approved by the Ethics Committee for Biomedical Research (CERB) of the Faculty of Medicine and Pharmacy in Rabat, with approval number; CERB 36–23 and approval date; January 22, 2024.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Informed Consent
Written informed consent was signed and obtained from all participants before their inclusion in the study.
