Abstract
Differentiation between small-cell lung cancer (SCLC) from non-small-cell lung cancer (NSCLC) brain metastases is crucial due to the different clinical behaviors of the two tumor types. We propose the use of a deep learning and transfer learning approach based on conventional magnetic resonance imaging (MRI) for non-invasive classification of SCLC vs. NSCLC brain metastases. Sixty-nine patients with brain metastasis of lung cancer origin were included. Of them, 44 patients had NSCLC and 25 patients had SCLC. Classification was performed with EfficientNet architecture on crop images of lesion areas and based on post-contrast T1-weighted, T2-weighted and FLAIR imaging input data. Evaluation of the model was carried out in a 5-fold cross-validation manner, and based on accuracy, precision, recall, F1 score and area under the receiver operating characteristic curve. The best classification results were obtained with multiparametric MRI input data (T1WI+c+FLAIR+T2WI), with a mean overall accuracy of 0.90 ± 0.04, and F1 score of 0.92 ± 0.05 for NSCLC and 0.87 ± 0.08 for SCLC for the validation data and an accuracy of 0.87 ± 0.05, with an F1 score of 0.88 ± 0.05 for NSCLC and 0.85 ± 0.05 for SCLC for the test dataset. The proposed method provides an automatic noninvasive method for the classification of brain metastasis with high sensitivity and specificity for differentiation between NSCLC vs. SCLC brain metastases. It may be used as a diagnostic tool for improving decision-making in the treatment of patients with these metastases. Further studies on larger patient samples are required to validate the current results.
Introduction
Primary tumors of the lung are the most common cause of brain metastases, with as many as up to 65% of patients with lung cancer ultimately developing brain metastases. 1 The main subtypes of lung cancer are small-cell lung carcinoma (SCLC) and non-small-cell lung carcinoma (NSCLC), accounting for ∼20% and 80% respectively, of all lung cancers. 2,3 The differentiation between SCLC and NSCLC brain metastases is critical since SCLC tends to disseminate earlier than NSCLC in the course of the disease, is clinically more aggressive and is usually treated non-surgically. NSCLC, however, is managed by a combination of surgery and adjuvant therapy 2,4,5 with newer treatment opportunities and much better prognosis. Currently, the two tumor types can be differentiated solely based on histologic tissue characterization following tissue biopsy. A way for noninvasively arriving at accurate preoperative diagnosis may allow improvement of the decision-making process toward more personalized treatment strategies.
Convolutional neural networks (CNN), comprising a class of deep learning architectures, has become the state-of-the-art approach in various visual computer tasks, with extensive applications in medical image analysis tasks as image classification, object detection, segmentation, registration etc. 6 -9
Several studies have shown the feasibility of deep learning for classification of NSCLC and SCLC cancer based on histopathology images. 10,11 Only one study used deep learning based imaging data of computerized tomographic (CT), achieving an 85.71% accuracy in identifying tumor types 12 and demonstrating a promising potential analytic approach for differentiation between SCLC and NSCLC based on radiology data.
In this work, we propose the use of an EfficientNet deep learning model and a transfer learning approach for the differentiation between SCLC and NSCLC brain metastases based on preoperative magnetic resonance imaging (MRI) data.
Materials and Methods
Dataset
This retrospective study analyzed 102 tumors obtained from 69 patients with brain metastasis of lung origin. The MRI data were collected retrospectively from patients’ routine clinical assessment performed at different sites, with different MRI vendors and systems and various acquisition parameters. 44 scans were performed on GE systems, 55 on Siemens systems and 2 scans on Philips MRI systems. Detailed description of MRI acquisition parameters are given in Supplementary Materials. The mean tumor volume, as measured by commercial software (AnalyzeDirect 11.0) was 18.3 ± 22.9cc3 (SCC = 19.4 ± 30.9 cc3, NSCC = 16.8 ± 15.3 cc3), with no significant difference between groups (
All scans included post-contrast T1-weighted image (T1WI+c), FLAIR images and T2-weighted image (T2WI). The study was approved by the local institutional review board which waived informed consent.
Data Preprocessing and Annotation
Preprocessing was performed in a MATLAB R2019b environment and included coregistration (realigned and resized) of the FLAIR and T2WI to the T1WI+c images by applying the SPM registration tool, brain extraction, bias field correction using intensity inhomogeneity correction algorithm implemented in SPM 13 and intensity normalized by the equation
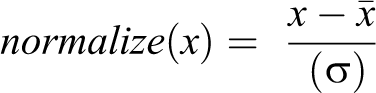
where

Input data. Cropped T1WI+c (A), FLAIR (B), T2WI (C) and concatenates (D) images. SCC, small-cell lung cancer; NSCC, non-small-cell lung cancer.
Data Splitting
The entire dataset was split at the in stratified manner—at the subject level into 90% training and validation and 10% testing datasets, and proportional to group’s size, and ensuring that all images belonging to a given patient would be allocated to the same group. The training and validation dataset were split into 80% training and 20% validation in a five-fold cross-validation manner.
Network Training
Deep learning model training and evaluation were performed using the Fast.ai framework built on top of the PyTorch environmen. 14
Input Data
The input data were the cropped MRI images, resized to 64 × 64 image size. The networked performance were tested based on four input configurations: T1WI+c images (Figure 1a), FLAIR images (Figure 1b), T2WI (Figure 1c) and the concatenated images (T1WI+c+FLAIR+T2WI) (Figure 1d).
Data Augmentation
In order to increase the dataset size and variance, data augmentation was performed and included random rotations, zooming and lightning. In addition, mixup augmentation was applied (11) which combining training samples using their linear combinations. 15
Networks Architecture
EfficientNet 16 convolutional neural network was used (see Supplementary Materials for detailed description of the used network architecture). Network training was performed using cross-entropy loss function and a batch size of 16. In order to cope with the imbalanced datasets, data oversampling was employed, enable sampling of the two classes in roughly equal amounts. Optimization of the network hyper parameters (EfficientNet architecture [bo-b7], learning rate, number of epochs, the metric for evaluation of the model during training and input imaging data), was performed on the training and validation data, in five-fold cross-validation manner.
Transfer Learning
Due to the relatively small data size that was available for this study, the network was trained by a pre-trained model based on EfficientNet-b0 pre-trained weights that had been trained on ImageNet data set as previously described. 17,18
Evaluation of the Results
Evaluation of the classification results was performed on the validation and test datasets, for each one of the input datasets, and for each one of the 5-folds, using accuracy, precision, recall, F1 score and area under the receiver operating characteristic curve (ROC AUC).
Results
Patients
A total of 296 axial slices of 102 tumors from 69 patients with brain metastasis from lung cancer origin were included for analysis. Forty-five of those patients had NSCLC (24 males, age ± standard deviation 63.7 ± 10.5 years) and 25 had SCLC (17 males, age 64.4 ± 11.6 years). No significant age or sex differences were detected between the two groups. All patients had pathological diagnosis of the primary lung tumor or pathology diagnosis from the brain metastases. Patient characteristics are given in Table 1.
Patient Characteristics.

Abbreviations: NSCLC, non-small-cell lung carcinoma; SCLC, small-cell lung carcinoma.
a Mean ± standard deviation.
Classification Results
Following optimization, network training was carried out with an EfficientNet-b0 model, with learning rate of 0.001, with accuracy as the metric for model evaluation during training, and with total of 100 epochs, while preserving the model which achieved the best level of accuracy during training.
Classification results for differentiation between NSCLC and SCLC for the different input datasets and across the five validation datasets are given in Tables 2 and 3.
Classification Results for the Validations Datasets.
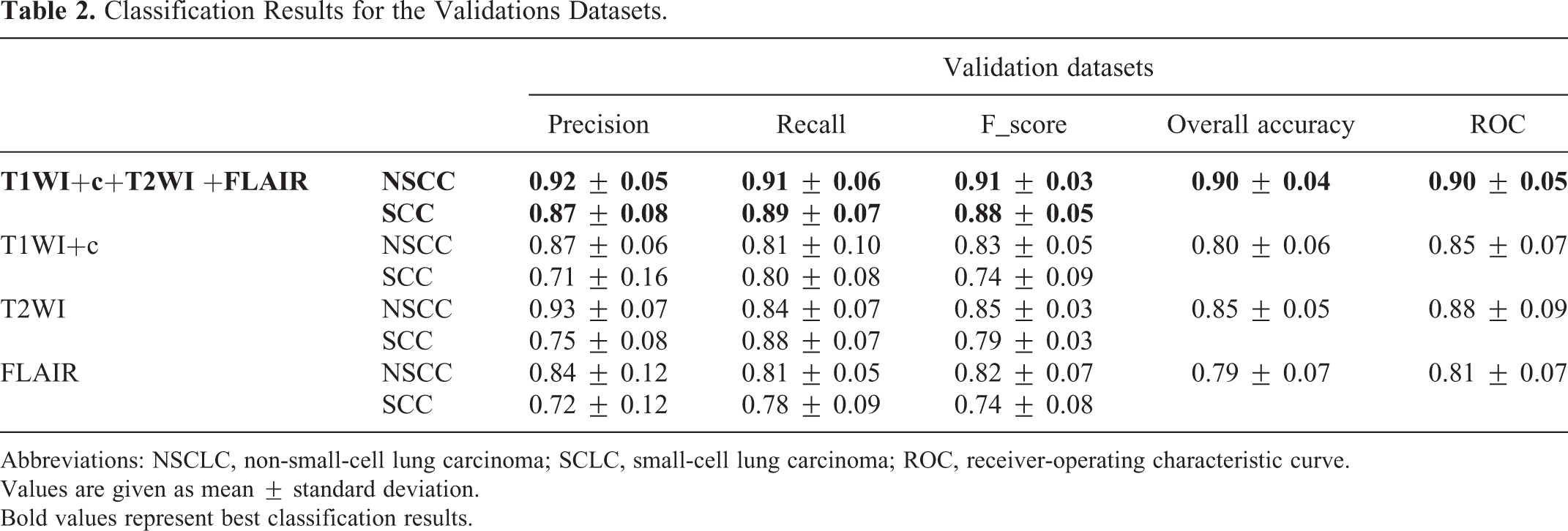
Abbreviations: NSCLC, non-small-cell lung carcinoma; SCLC, small-cell lung carcinoma; ROC, receiver-operating characteristic curve.
Values are given as mean ± standard deviation.
Bold values represent best classification results.
Classification Results for the Test Datasets.

Abbreviations: NSCLC, non-small-cell lung carcinoma; SCLC, small-cell lung carcinoma; ROC, receiver-operating characteristic curve.
Values are given as mean ± standard deviation.
Bold values represent best classification results.
The best classification results were obtained for the multiparametric MRI input data (T1WI+c+FLAIR+T2WI), results with mean overall accuracy of 0.90 ± 0.04, with an F1 score for the validation data of 0.92 ± 0.05 for NSCLC and 0.87 ± 0.08 for SCLC and an accuracy of 0.87 ± 0.05, with an F1 score of 0.88 ± 0.05 for NSCLC and 0.85 ± 0.05 for SCLC for the test dataset. Representative classification results are given in Figure 2. The high performance obtained across the five datasets for both the validation and test data demonstrated the robustness of the obtained model.

Representative classification results. SCC, small-cell lung cancer; NSCC, non-small-cell lung cancer.
The mean ROC areas were 0.90 ± 0.05, 0.85 ± 0.07, 0.88 ± 0.09 and 0.81 ± 0.07 for the T1WI+c+T2WI+FLAIR, T1WI+c, T2WI and FLAIR input data, respectively. The ROC curve that was derived from differentiating between NSCLC and SCLC given T1WI+c + FLAIR + T2WI is given in Figure 3.
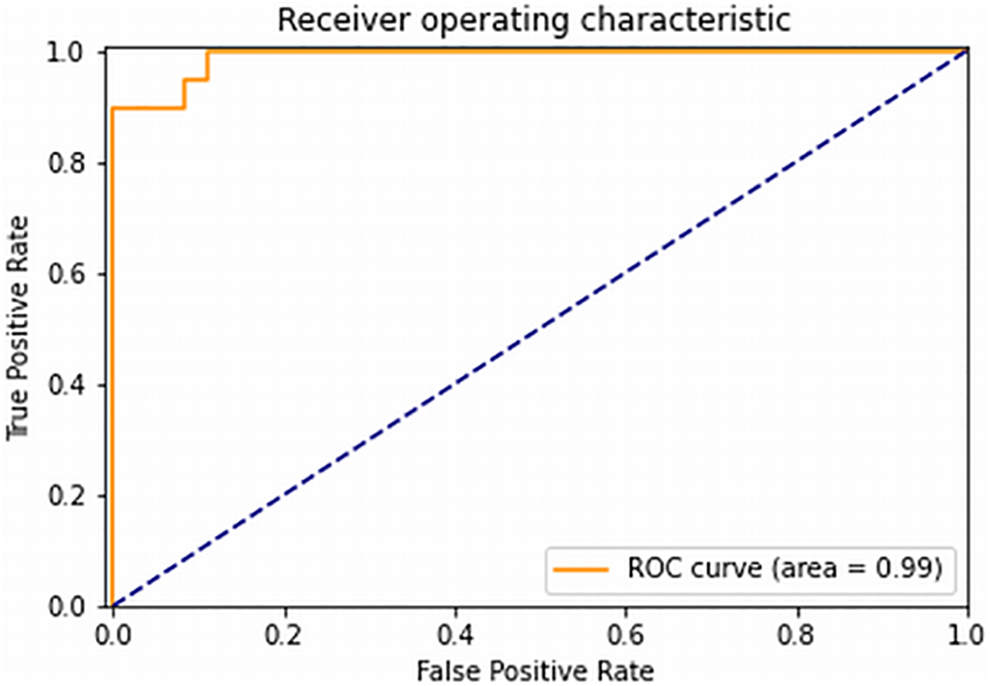
ROC curve from differentiating between NSCLC and SCLC given T1WI+c + FLAIR + T2WI input data.
Discussion
The differences in clinical behavior and treatment approaches to SCLC and NSCLC metastases to the brain bestow critical importance to the differentiation between them. In this study, we assessed the feasibility to differentiate between NSCLC and SCLC brain metastases by means of a deep learning approach. Deep learning had been shown to outperform other machine learning methods in multiple various visual computer tasks, including those involving medical image analysis. 6 However, reaching an optimal level of performance while avoiding model overfitting requires a large training dataset. This often proves to be problematic in some fields of medical imaging in which the training datasets are often small. In order to cope with the small size of the data in our own study, we trained our deep learning model by applying transfer learning, using EfficientNet-b0 pre-trained weights that had been trained on an ImageNet data set. Transfer learning had been shown to be the preferable method of choice for training a deep learning model in the setting of a small database, providing more robust and higher levels of performance compared to training models starting from scratch (i.e., randomly initiated the convolutional neural network filters). 9,19 -21
In addition, classification was perform on 2D space, and using up to three slices per patient, enabling to increase the data set size while maintaining the database splitting at the subject level. We further increased the data size and various by using conventional data augmentation, which includes random brightness, contrast flipping, rotation and scaling transformations, as well as mixup augmentation, which trains a neural network in combinations of pairs of examples and their labels. These augmentations have shown to substantially improve deep learning model performance on several visual computer tasks. 15,22
We classified imaging findings into either NSCLC or SCLC by using EfficientNet-b0 architecture. Tan and Le. 16 demonstrated that balancing network depth, width and resolution can lead to better performance, and those authors proposed that the EfficientNets family of models uniformly scales all model dimensions (depth, width and resolution) by using a simple yet highly effective compound coefficient. EfficientNets family contains eight models that are structurally similar and follow certain scaling rules for adjustment to larger image sizes. EfficientNet model achieved state-of-the-art performance on ImageNet challenge and also transferred well to several datasets, including those involved in medical tasks, such as for COVID-19 diagnosis based on X-ray images 23 and CT scans. 24
We tested the network performances on different configurations of input data. As expected, the best classification results were obtained by using multiparametric MRI data. Combining multiple imaging contrasts which reflect different aspects of pathophysiological processes, such as tissue permeability, T1 and T2 relaxometry and water content, may provide more insight than what can be obtained using a single imaging approach. 25
Due to its outstanding performance, deep learning is expected to improve the current standard diagnostic imaging techniques by providing a non-invasive tool for tumor classification. Our results demonstrate clinical feasibility of using deep learning to distinguish between NSCLC and SCLC brain metastases based on MRI, and may indicate the promising potential of this technology for also differentiating SCLC from NSCLC primary site, and for differentiating brain metastases from lung cancer and from other tumor types. However further research is needed to substantiate this hypothesis.
The major limitation on this study is the relatively small sample size available to us for testing the model’s robustness and generalization ability using deep learning approach. At the same time, it is important to demonstrate the clinical feasibility of using deep learning to distinguish between NSCLC and SCLC brain metastases based on MRI. Future studies should include a larger sample size, preferably from multiple centers.
Conclusion
An EfficientNet deep learning model is proposed for noninvasive classification of NSCLC vs. SCLC brain metastases. Our results demonstrate that it has high sensitivity and specificity for the differentiation between the two tumor types. The proposed method may be used as a diagnostic aid to improve decision-making in the treatment of patients with NSCLC and SCLC brain metastases, although further studies on larger sample size are required to validate the results of our work.
Supplemental Material
Supplemental Material, sj-pdf-1-tct-10.1177_15330338211004919 - Differentiating Small-Cell Lung Cancer From Non-Small-Cell Lung Cancer Brain Metastases Based on MRI Using Efficientnet and Transfer Learning Approach
Supplemental Material, sj-pdf-1-tct-10.1177_15330338211004919 for Differentiating Small-Cell Lung Cancer From Non-Small-Cell Lung Cancer Brain Metastases Based on MRI Using Efficientnet and Transfer Learning Approach by Rachel Grossman, Oz Haim, Shani Abramov, Ben Shofty and Moran Artzi in Technology in Cancer Research & Treatment
Footnotes
Authors’ Note
Our study was approved by the Tel Aviv Sourasky Medical Center review board (IRB approval number 0200-10), with waiver of informed consent. No informed consent was required by the IRB for this retrospective study on anonymous data. All procedures were carried out in accordance with relevant guidelines and regulations.
Abbreviations
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
