Abstract
Keywords
Introduction
Lung cancer is one of the most common malignant tumors in the world, and its morbidity and mortality rank first among all malignant tumors. 1 Approximately 80% of lung cancer types are nonsmall-cell lung cancer (NSCLC), and more than 50% of NSCLC patients have adenocarcinomas. 2 Although chemotherapy still plays a dominant role in NSCLC, the OS of most patients is less than a year after chemotherapy, and chemotherapy has a high grade 3-5 toxicity. Most patients stop treatment because they cannot tolerate the toxicity of chemotherapy. 3
However, in the past decade, molecular targeted therapy based on patient genomes has made a major breakthrough and completely changed the prospects of treatment for advanced NSCLC. 4 Epidermal growth factor receptor (EGFR) tyrosine kinase inhibitors (TKIs) are the first-line treatment choice for advanced NSCLC, and advanced NSCLC has the most common form of EGFR mutation: deletion of exon 19 or exon 21 L858R. 5 Gefitinib, erlotinib, icotinib, osimertinib, and other targeted drugs have become first-line treatment standards for EGFR mutations in advanced NSCLC. 6 However, drug resistance is inevitable in EGFR-TKIs, and a number of clinical trials have shown that7–9 approximately 5% to 10% of NSCLC patients with EGFR-sensitive mutations are mainly resistant to EGFR-TKI therapy. Meanwhile, some mechanisms of anti-EGFR-TKIs have been investigated, such as EGFR T790M mutation, MET gene amplification, PIK3CA mutation, ERBB2 amplification, and so on.
MYC is an out-of-control oncogene in human cancer and was first discovered in 1983. 10 MYC is one of the most highly amplified oncogenes in many human cancers, 11 and it supports tumorigenesis and progression and is often found in cervical, colon, breast, gastric, and lung cancer. 12 MYC is one of the most commonly amplified genes in lung cancer, 13 and some studies have shown that amplification of the MYC gene is related to the loss of cell differentiation. 12 Other studies have shown that MYC gene amplification is associated with poor prognosis in NSCLC and small cell lung cancer (SCLC).14–16 Some studies have shown that MYC family genes, including MYC, MYCL, and MYCN, as carcinogenic drivers may constitute new therapeutic targets in small cell lung cancer. 17 However, there is no approved treatment for this kind of mutation. Similarly, TP53 is the most frequently altered gene in human cancer. 18 Many studies have shown that TP53 mutation is a negative prognostic factor for the prognosis of patients with NSCLC and may lead to drug resistance to EGFR-TKIs.19,20 However, there are many different kinds of TP53 mutations, and the effect of each mutation on prognosis is heterogeneous. 3
The relationship between MYC and TP53 genes was first proposed Gazzeri et al. 21 used Southern and Northern blotting techniques and found that there may be a correlation between TP53 mutation and MYC activation in NSCLC, but the correlation was not significant. At the same time, TP53 and MYC gene changes are very important, representing independent factors in the development of lung cancer. In recent reports, researchers have observed a highly significant co-occurrence of MYC gene amplification and TP53 gene mutations in breast cancer and later in NSCLC. 11 However, the prognosis and predictive value of MYC/TP53 comutation in patients with EGFR mutation-positive advanced NSCLC treated with oral EGFR-TKIs are still unclear, although some studies have suggested that MYC gene amplification is associated with possible EGFR-TKI resistance.14,16 However, there is not a large amount of clinical data to confirm this hypothesis. For this reason, we studied the correlation between MYC/TP53 comutation and the therapeutic effect of EGFR-TKIs in Chinese patients with late EGFR mutation-positive NSCLC to determine whether MYC/TP53 comutation indicates a poor prognosis and whether it is related to primary drug resistance to EGFR-TKIs.
Materials and Methods
Patients Recruitment and Sample Collection
The study is a single center retrospective study, a total of 65 patients with stage IIIA-IV NSCLC diagnosed in the Department of Oncology, Northern Jiangsu People's Hospital from 2016-08 to 2020-10 were retrospectively collected, all the patients enrolled signed a written informed consent form that their tissue samples were analyzed by next-generation sequencing (NGS) and the study have obtained approval from the Ethics Committee of Northern Jiangsu People's Hospital (Approval: ID2017005). The reporting of this study conforms to STROBE guidelines. 22 The inclusion criteria were as follows: (1) EGFR positive patients were analyzed by NGS. (2) Eastern Cooperative Oncology Group (ECOG) score 23 0-2 points, and there were no serious heart, lung, and other basic diseases. (3) NSCLC staged according to CSCO guidelines as stage IIIA-IV. (4) Oral EGFR-TKI therapy was used at all times throughout the course of the disease, including three kinds of EGFR-TKI drugs gefitinib, alphatinib, or osimertinib. Exclusion criteria: (1) Patients with wild-type EGFR; (2) The general situation was poor, ECOG > 2; (3) Age > 75 years; (4) Patients who were not treated with TKI were selected.
Sample Collection and the Evaluation of Clinical Efficacy
The patient underwent imaging examination and at least one repeated radiological examination. Baseline characteristics of patients (age, sex, smoking history, histology, Eastern Cancer Cooperation Group (ECOG) performance status, current survival status, etc.) and results after continuous and regular TKI treatment were obtained using medical and radiological records and patient follow-up information. The significance of regular review during the administration of EGFR-TKIs is that patients receive chest and abdominal CT, brain MRI, and other imaging examinations every 2-3 months to evaluate the potential therapeutic effect of EGFR-TKIs, according to the RECIST (solid tumor efficacy evaluation criteria) 24 standard to determine the clinical effect. To ensure the privacy of patients, all patient details have been deidentified. The main endpoints of this study were to determine the difference in PFS or OS among the three groups of patients after EGFR-TKI treatment according to the mutation status of MYC and TP53: double wild-type tumors (WT/WT), MYC wild-type and TP53 mutant tumors (TP53/WT) and double mutant tumors (MYC/TP53). The PFS of EGFR-positive NSCLC patients after EGFR-TKI treatment was the time from the beginning of oral EGFR-TKI treatment to disease progression, and OS was the time from oral EGFR-TKI treatment to death or the last follow-up time in September 2021. Secondary endpoints included ORR and DCR. ORR is the proportion of patients whose tumor size reduced to a predetermined value and can maintain the minimum time limit, which is the sum of complete response (CR) and partial response (PR), ORR = CR + PR. DCR is the percentage of evaluable cases with response (PR + CR) and stable disease (SD) after treatment, DCR = CR + PR + SD.
Targeted Panel NGS
A total of 168 genes closely related to the pathogenesis and targeted therapy of lung cancer were extracted from tumor tissue and plasma. The important exon regions of 168 genes and the hot intron regions of eight genes were detected by probe hybridization and high-throughput sequencing. Comprehensive and accurate detection of lung cancer-related gene mutation, copy number variation and rearrangement (fusion) and other mutations was performed. For detection, target area probe capture technology and second-generation high-throughput sequencing technology (NGS) based on the Illumina sequencing platform were used to analyze the samples. This technology was independently developed, analyzed and verified by Burning Stone Medicine. The Institute of Burning Stone Medicine has completed the verification of the technical platform according to CLIA'88 and relevant technical guidelines at home and abroad and has passed the interroom quality assessment of high-throughput sequencing for tumor diagnosis and treatment in the Clinical Test Center of the Health Commission. This test can cover the single nucleotide variation (SNV), short fragment insertion or deletion variation (INDEL), gene copy number variation (CNV) and gene rearrangement (rearrangement/fusion) in the capture exons and + /−20 bp flanking region of each exon. The evidence used to distinguish between benign and malignant variants comes from OncoDB, 25 the internal database of burning stone, and refers to public databases such as NCCN guidelines, 26 ACMG guidelines, 27 and OncoKB. 28
DNA Extraction Information
Using the DNeasy Blood and Tissue kit (Qiagen) and QIAamp DNA FFPE Tissue Kit (Qiagen) in accordance with the manufacturer's instructions, genomic DNA was extracted from fresh tissue and FFPE samples. 29 Tissue DNA was sheared using Covaris M220 (Covaris, MA, USA), followed by end repair, phosphorylation and adaptor ligation. Fragments between 200 and 400 bp from the sheared tissue DNA were purified (Agencourt AMPure XP Kit, Beckman Coulter, CA, USA), followed by hybridization with capture probes baits, hybrid selection with magnetic beads, and PCR amplification. The quality and the size of the fragments were assessed by high sensitivity DNA kit using Bioanalyzer 2100 (Agilent Technologies, CA, USA). Indexed samples were sequenced on Nextseq500 (Illumina, Inc., USA) with paired-end reads and target sequencing depth of 1,000× for tissue samples.
Bioinformatics Pipeline Information
Sequence data were mapped to the reference human genome (hg19) using Burrows–Wheeler Aligner v.0.7.10. 30 Local alignment optimization, duplication marking and variant calling were performed using Genome Analysis Tool Kit v.3.2, 31 and VarScan v.2.4.3. 32 Variants were filtered using the VarScan fpfilter pipeline, loci with depth less than 100 were filtered out. Base-calling in tissue samples required at least 8 supporting reads for single nucleotide variations (SNV) and five supporting reads for insertion-deletion variations (INDEL). Variants with population frequency over 0.1% in the ExAC, 1000 Genomes, dbSNP or ESP6500SI-V2 databases were grouped as single nucleotide polymorphisms (SNP) and excluded from further analysis. Remaining variants were annotated with ANNOVAR (February 01, 2016 release) 33 and SnpEff v.3.6. 34 Analysis of DNA translocation was performed using Factera v.1.4.3. 35 Copy number variations (CNVs) were analyzed based on the depth of coverage data of capture intervals. Coverage data were corrected against sequencing bias resulting from GC content and probe design. The average coverage of all captured regions was used to normalize the coverage of different samples to comparable scales. Copy number was calculated based on the ratio between the depth of coverage in tumor samples and average coverage of an adequate number (n > 50) of samples without CNV as references per capture interval. CNV is called if the coverage data of the gene region was quantitatively and statistically significant from its reference control. The limit of detection for CNVs is 1.5 and 2.64 for deletions and amplifications, respectively.
Statistical Analysis
The Kruskal–Wallis test or Fisher's exact test was used to evaluate the differences in clinical covariables among the three groups (WT/WT, TP53/WT, and MYC/TP53). The PFS and OS of the patients were analyzed by the Kaplan–Meier method. The Cox proportional hazard regression model was used to evaluate the correlation between clinical covariables and PFS/OS, and the risk ratio and 95% confidence interval (CI) were obtained. All the reported P values were double-tailed, and P values less than 0.05 were considered to be statistically significant. All statistical analyses were performed using SPSS25.0 for statistical analysis and GraphPadPrism version 7 to generate drawings.
Results
Baseline Characteristics
We identified 65 patients with EGFR-positive NSCLC (Table 1). Among 65 patients, 17 had TP53 and MYC wild-type mutations (WT/WT), 36 had TP53 mutant and MYC wild-type mutations (TP53/WT), and 12 had coexisting MYC/TP53 mutations (MYC/TP53) (Table 2). There was no significant difference in median age (P = .194), sex (P = .783), clinical stage (P = .597), smoking history (P = .242) or EGFR mutation type (P = .358) among the three groups. Compared with other groups, there were more patients with MYC/TP53 wild-type tumors (WT/WT) with an ECOG score of 0-1 (82% WT/WT vs 56% TP53/WT vs 50% MYC/TP53; P = .044), with significant histological differences between groups (P = .029).
Baseline Characteristics.

Baseline Patient and Tumor Characteristics by Mutation Subgroup.

Note: *P < .05.
Abbreviations: TP53/WT, group of patients with NSCLC harboring TP53 mutation but no MYC mutation; MYC/TP53, group of patients with NSCLC harboring MYC and TP53 mutations; WT/WT, group of patients with NSCLC harboring no TP53 or MYC mutation.
Univariate and Multivariate Analysis of the Relationship between Clinical Characteristics and Prognosis in Patients with EGFR Mutation NSCLC
We summarized the results of survival and response analysis of 65 patients with NSCLC who received EGFR-TKIs at any stage by Cox regression analysis. In univariate analysis (Table 3), Gene mutation status (WT/WT, TP53/WT, and MYC/TP53) was a significant predictor of PFS (HR: 0.596, 95% CI: 0.393-0.903, P = .015) and OS (HR: 0.427, 95% CI: 0.274-0.665, P < .001). In addition, clinical stage (IIIA, IIIB, and IV) and ECOG score (0-1, 2) were also predictors of PFS (HR: 0.131, 95% CI: 0.040-0.430, P = .004, HR 3.013, 95% CI: 1.876-4.839, P < .001) and OS (HR: 0.216, 95% CI: 0.067-0.703, P = .010, HR: 2.493, 95% CI: 1.638-3.794, P < .001). In the multivariate analysis adjusted for other variables (age, sex, smoking, and histology) (Table 4), we observed a significant correlation between PFS and clinical stage (HR: 0.126, 95% CI: 0.034-0.461, P = .002), ECOG score (HR: 2.556, 95% CI: 1.530-4.270, P < .001), and Gene mutation status (HR: 0.622, 95% CI: 0.389-0.996, P = .048). Similarly, OS was significantly correlated with ECOG score (HR:1.899, 95% CI: 1.221-2.954, P = .004) and Gene mutation status (HR: 0.427, 95% CI: 0.338-0.947, P = .030).
Univariate Regression Analysis of PFS and OS in EGFR-Positive NSCLC.
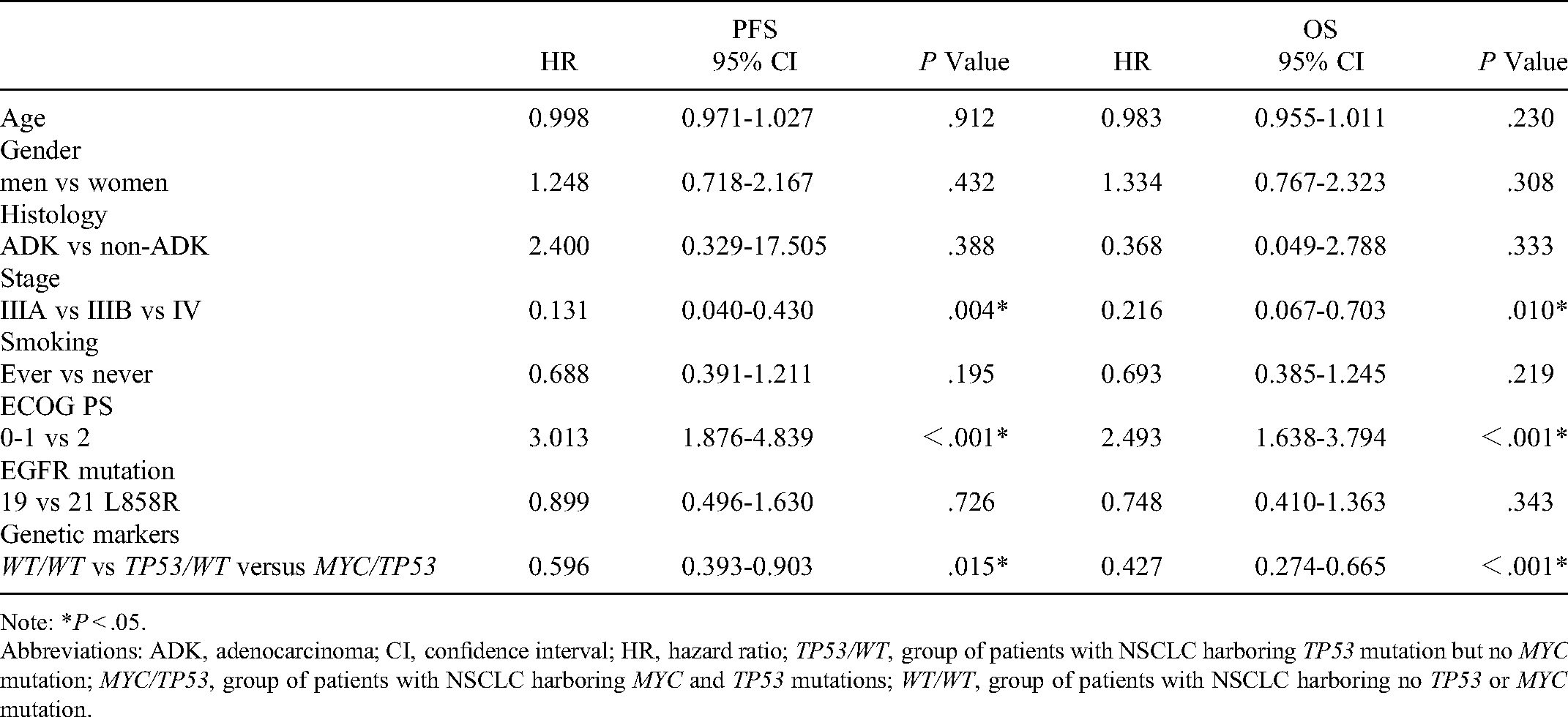
Note: *P < .05.
Abbreviations: ADK, adenocarcinoma; CI, confidence interval; HR, hazard ratio; TP53/WT, group of patients with NSCLC harboring TP53 mutation but no MYC mutation; MYC/TP53, group of patients with NSCLC harboring MYC and TP53 mutations; WT/WT, group of patients with NSCLC harboring no TP53 or MYC mutation.
Multivariate Regression Analysis of PFS and OS in EGFR-Positive NSCLC.

Note: *P < .05.
Abbreviations: ADK, adenocarcinoma; CI, confidence interval; HR, hazard ratio; TP53/WT, group of patients with NSCLC harboring TP53 mutation but no MYC mutation; MYC/TP53, group of patients with NSCLC harboring MYC and TP53 mutations; WT/WT, group of patients with NSCLC harboring no TP53 or MYC mutation.
Analysis of the Survival status of the Three Groups of Patients
Because both univariate and multivariate analyses showed that the state of gene mutation (WT/WT, TP53/WT, and MYC/TP53) is a predictor of PFS and OS, we stratified the patients with EGFR-positive WT/WT (MYC, TP53 wild-type mutation), TP53/WT (MYC wild-type, TP53 mutation positive), and MYC/TP53 comutations and compared the survival time of the three groups by Kaplan–Meier analysis (Figure 1). When 12 patients with MYC/TP53 comutation were compared with the other two groups (TP53/WT and WT/WT), mPFS and mOS were significantly lower than those in the other two groups (mPFS: 4.1 months vs 6.0 months, 12.3 months, HR: 0.769, 95% CI: 4.592–7.608, P = .047. mOS: 14.6 months vs 24.1 months, 31.5 months, HR: 3.170, 95% CI: 18.786–31.214, P < .001).
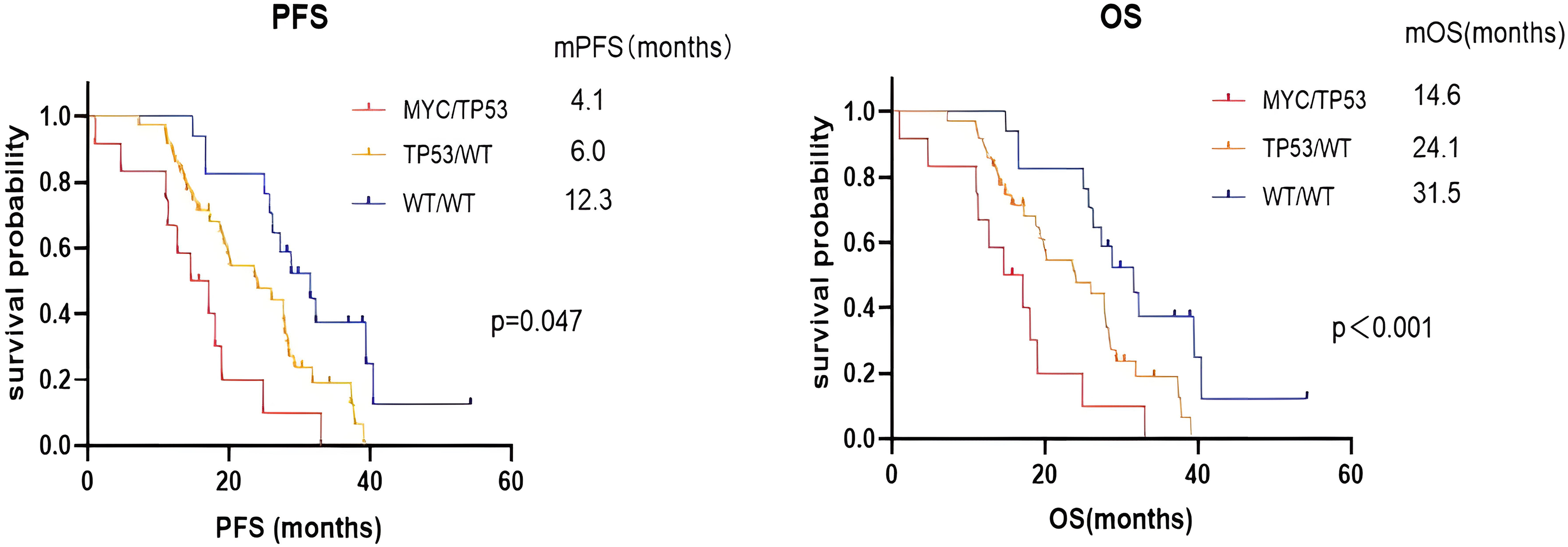
Analysis of the survival status of the three groups of patients. Abbreviations: PFS, progression-free survival; OS, overall survival; TP53/WT, group of patients with NSCLC harboring TP53 mutation but no MYC mutation; MYC/TP503, group of patients with NSCLC harboring MYC and TP53 mutations; WT/WT, group of patients with NSCLC harboring no TP53 or MYC mutation.
In addition, in patients whose PFS was less than 3 months, 33.3% (4/12) had MYC/TP53 comutations, 22.2% (8/36) had TP53/WT mutations, and only 5.9% (1/17) had WT/WT mutations. Similarly, 50% of patients with OS less than 15 months were MYC/TP53 comutants, 33.3% (12/36) had TP53/WT mutations, and 5.9% (1/17) had WT/WT mutations, which suggests that individuals with MYC/TP53 comutations have poorer clinical outcomes and are more prevalent in patients with early resistance to EGFR-TKI treatment.
We also analyzed the secondary outcome ORR, DCR of 65 patients in these three groups (WT/WT, TP53/WT, and MYC/TP53) (Figure 2). The ORR, DCR of patients with MYC/TP53 comutation was lower than that of the other two groups (ORR, 25% vs 44.4%, 70.6%, P = .045. DCR, 58.3% vs 72.2%, 82.4%, P = .365) (Table 5). There was no statistical significance in DCR among the three groups, but ORR showed statistical significance. Thus, patients with MYC/TP53 comutations are more likely to develop drug resistance after early treatment with EGFR-TKIs and have a worse clinical outcome.
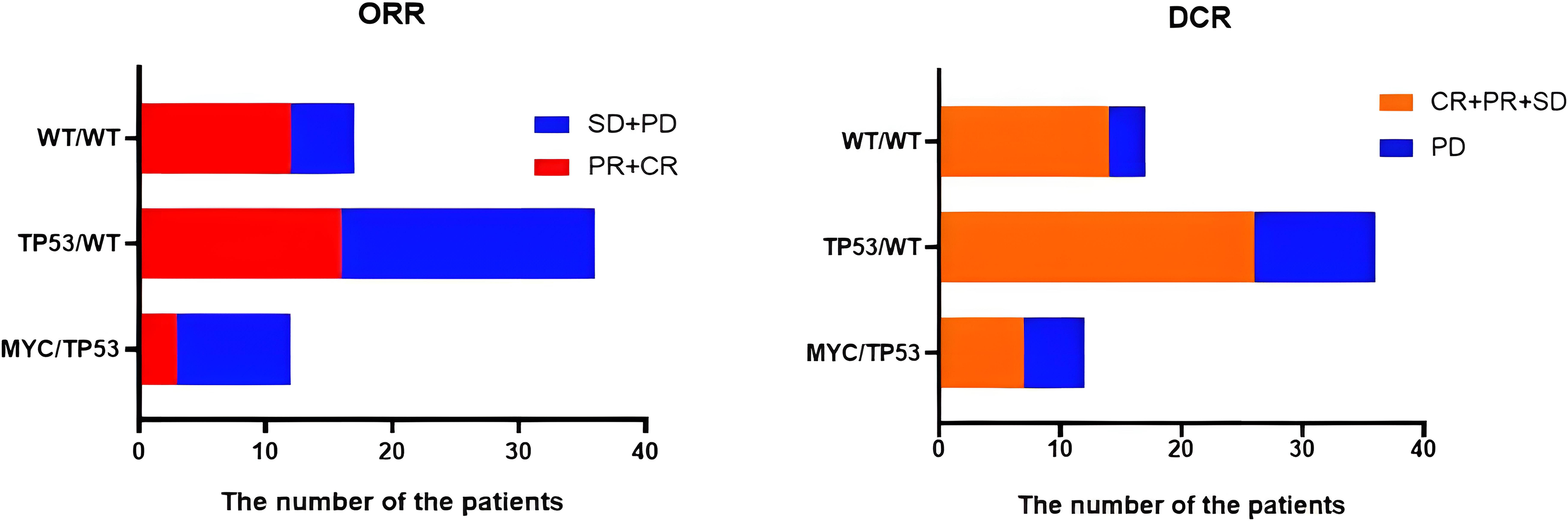
ORR and DCR of the three groups of patients. Abbreviations: ORR, objective response rate; DCR, disease control rate; CR, complete response; PR, partial response; SD, stable disease; PD, progressive disease; TP53/WT, group of patients with NSCLC harboring TP53 mutation but no MYC mutation; MYC/TP53, group of patients with NSCLC harboring MYC and TP53 mutations; WT/WT, group of patients with NSCLC harboring no TP53 or MYC mutation.
Chart of Response After EGFR TKI Treatment in Three Subgroups of Patients(WT/WT, TP53/WT, MYC/TP53).

Note: *P < .05.
Abbreviations: ORR, objective response rate; DCR, disease control rate; PFS, progression-free survival; OS, overall survival; TP53/WT, group of patients with NSCLC harboring TP53 mutation but no MYC mutation; MYC/TP53, group of patients with NSCLC harboring MYC and TP53 mutations; WT/WT, group of patients with NSCLC harboring no TP53 or MYC mutation.
Evaluation of the Difference in PFS and OS between MYC/TP53 and TP53/WT
We found that there were differences in PFS and OS among the three groups of patients (WT/WT, TP53/WT, and MYC/TP53). To further analyze the nature of the differences, we made a pairwise comparison among the three groups of patients. (Figure 3A, B). There was no statistical significance in PFS between the patients with MYC/TP53 and TP53/WT (4.1 months vs 6.0 months, HR: 0.775, 95% CI: 0.3858-1.557, P = .25), but there was statistical significance in OS (14.6 months vs 24.1 months, HR: 0.658, 95% CI: 0.3274-1.321, P = .015). OS was an independent predictor of prognosis in MYC/TP53 and TP53/WT.

The survival curve of pairwise comparison of three subgroups (WT/WT, TP53/WT, and MYC/TP53). Abbreviations: PFS, progression-free survival; OS, overall survival; TP53/WT, group of patients with NSCLC harboring TP53 mutation but no MYC mutation; MYC/TP53, group of patients with NSCLC harboring MYC and TP53 mutations; WT/WT, group of patients with NSCLC harboring no TP53 or MYC mutation.
Evaluation of the Difference in PFS and OS Between MYC/TP53 and WT/WT
We then performed Kaplan–Meier analysis of PFS and OS in the MYC/TP53 and WT/WT groups (Figure 3C, D). There were significant differences in PFS and OS between the patients with MYC/TP53 and WT/WT (PFS: 4.1 months vs 12.3 months, HR: 0.378, 95% CI: 0.1668-0.8568, P = .012. OS: 14.6 months vs 31.5 months, HR: 0.5032, 95% CI: 0.222-1.14, P < 0.001). The results showed that both PFS and OS are independent predictors for the prognosis of MYC/TP53 and WT/WT.
Effect of MYC Mutation Site on Prognosis
We then studied the effects of different MYC mutation sites on EGFR positive Chinese patients with advanced NSCLC, and Table 6 listed the MYC mutation sites as well as PFS and OS in 12 patients with MYC/TP53 comutation. It can be seen that the PFS and OS of MYC short variant are longer than MYC amplification (mPFS: 7.0 months vs 3.4 months, HR: 2.059, 95% CI: 0.6027 to 7.033, P = .21. mOS: 24.8 months vs 11.3 months, HR: 2.195, 95% CI: 0.6425 to 7.497, P = .031) (Figure 4). The results showed that the patients with MYC amplification had a worse prognosis. Due to the small total sample size, a larger sample size needs to be collected in the future to confirm this conclusion.

Survival curve of the effect of MYC mutation site on prognosis. Abbreviations: PFS, progression-free survival; OS, overall survival.
MYC Mutation Site and Corresponding Survival Time.

Abbreviations: PFS, progression-free survival; OS, overall survival; MYC/TP53, group of patients with NSCLC harboring MYC and TP53 mutations
Discussion
The MYC oncogene is a common transcription factor, and it is one of the most highly amplified oncogenes in a variety of human cancers. 12 In an earlier study, Yoo et al. 36 used IHC to report the overexpression of MYC in 147 cases of NSCLC, which showed that the overexpression of MYC accounted for 16% of NSCLC (24 of 147 cases). Other studies have shown that 14% of patients with TP53 mutations in NSCLC have MYC gene amplification. 37 The conclusions of these studies are basically consistent with ours (18.5%, 12 of 65 cases). In our study, we compared the prognostic value of three groups of EGFR-positive NSCLC patients with WT/WT (MYC and TP53 wild type mutation), TP53/WT (MYC wild type, TP53 mutation positive) and MYC/TP53 comutations and verified that the EGFR-positive MYC/TP53 comutation was related to the significant shortening of PFS and OS and the decrease of DCR and ORR after EGFR-TKI treatment. This indicates that MYC/TP53 comutation EGFR-positive NSCLC patients are more likely to develop drug resistance after early treatment with EGFR-TKI and have a poor clinical outcome. Subsequently, we further analyzed the prognosis of two subtypes of MYC mutations (MYC short variant and MYC amplification) in 12 patients with MYC/TP53 comutation. We come to conclusion that the OS and PFS of MYC short variant are longer than MYC amplification, there was no significant difference in PFS between the two groups (P = .21). However, the molecular mechanism of this phenomenon is still unclear.
The results of Hall et al 38 show that the activation of MYC leads to the accumulation of cholesterol ester (CEs) stored in lipid droplets, the subsequent inactivation of MYC leads to a further increase in CEs, and the accumulation of CEs leads to the further enhancement of the invasiveness of NSCLC. Seo et al. 16 used in situ fluorescence hybridization to determine that c-MYC gene amplification is an independent poor prognostic factor for DFS and OS in stage I lung cancer and may also be an independent poor prognostic factor for EGFR mutant adenocarcinoma. This study pointed out that the increase in the c-MYC 8 chromosome was often related to smoking history, male sex, and/or lymphatic invasion, but the significance of the correlation may be limited because the number of cases with chromosome eight increases is relatively small, with only five patients. However, this study only observed patients with IA-IIIB, and in our study, we extended these results to patients with IIIA-IV.
The MYC gene is widely studied in lymphoma, prostate cancer, colorectal cancer and small cell lung cancer,39–42 and several studies have shown that overexpression of MYC is associated with drug resistance.43,44 Brägelmann et al. 17 pointed out that as a carcinogenic driving factor, MYC may constitute a new therapeutic target in small cell lung cancer, and its core role in tumor maintenance provides a new opportunity for targeted therapy. In addition, Sabattini et al. 45 used the FISH detection method and identified MYC amplification as a diagnostic or prognostic marker for malignant lymphoma. However, there are few studies on MYC gene mutations in NSCLC, which may be related to the fact that the MYC gene promotes tumor progression and leads to the transformation of NSCLC into more malignant small cell lung cancer. 46
TP53 is a common tumor suppressor gene, but because of its inoperability, TP53 mutations are not always detected in the early NSCLC molecular spectrum. 47 Therefore, the incidence of comutations leading to TP53 and other driving gene mutations may be underestimated. Similarly, the same is true of MYC mutations, and even when detecting MYC and TP53 mutations, the frequency of comutations with MYC and TP53 is not always reported, so there are few studies on the interaction of MYC and TP53 comutations in NSCLC.
At present, the clinical data on the predictive effect of EGFR-positive MYC/TP53 comutation NSCLC on the efficacy of EGFR-TKIs are still limited, and it is still controversial whether early drug resistance will be caused by EGFR-TKI treatment in EGFR-positive NSCLC patients with MYC/TP53 comutation. In a recent case report, researchers found that a patient with EGFR-positive NSCLC developed drug resistance after treatment with EGFR-TKIs. After drug resistance, new MYC gene amplification, RB1 mutation and TP53 mutation were found in the patient's tissue. 48 It is well known that the RB1 gene is considered as an important genetic marker for the transformation of NSCLC into small cell lung cancer, and researchers have shown that lung cancer with EGFR/TP53/RB1 mutations has a unique risk of histological transformation, with 25% showing new SCLC or final small cell lung cancer transformation. 49 However, for NSCLC patients with EGFR-positive MYC/TP53 comutation, the mechanism of this phenomenon is not clear. Studies have shown that proto-oncogenes in the MYC family, such as MYC, MYCN, and MYCL, are amplified and/or overexpressed in SCLC tumors, and they affect the tumor phenotype by controlling the dedifferentiation process from neuroendocrine cells to nonneuroendocrine cells 50 so that NSCLC can be transformed into small cell lung cancer. It is worth noting that in the data we studied, 3 of the 12 EGFR-positive patients with MYC/TP53 comutation experienced the transformation of small cell lung cancer during EGFR-TKI treatment, which may provide a new research direction for the drug resistance mechanism of MYC/TP53 comutation genes. Another study showed that osimertinib rapidly and sustainably reduced c-MYC levels mainly by enhancing protein degradation in EGFR-mutated NSCLC cell lines and xenotransplanted tumors, resulting in acquired drug resistance to osimertinib, 51 which is basically consistent with the conclusion of our study and can provide a scientific basis for the mechanism of drug resistance to EGFR-TKIs. Zhong et al. 52 studied the mechanism of primary drug resistance by NGS sequencing, analyzed the genetic changes of 11 patients with primary drug resistance to EGFR-TKIs and 11 patients sensitive to EGFR-TKIs after taking EGFR-TKIs, and found that MYC gene amplification was found in two sensitive patients and one patient with primary drug resistance. It was also found that the number of MYC copies in patients with primary drug resistance (13.6 times) was significantly higher than that in sensitive patients (4.8 times and 2.9 times). Thus, they concluded that the combination of MYC inhibitors and EGFR-TKIs may be a promising strategy to overcome the primary drug resistance of lung cancer to EGFR-TKIs. However, this observation needs to be confirmed because there were only 11 patients with EGFR-TKI primary drug resistance in this study, of whom only one patient had MYC gene amplification. Our study analyzed the overall prognostic levels of PFS and OS in 12 patients with MYC/TP53 comutations and not only found that these patients were prone to early drug resistance when using EGFR-TKIs but also had a shorter survival time. Another study showed that EGFR-TKIs affect the expression of PD-L1 in NSCLC through the c-MYC pathway to promote drug resistance. Eun Young Kim et al introduced siRNA targeting MYC into H60 and H2009 cells with high expression of PD-L1 to study the relationship between MYC and PD-L1. They concluded that there was a significant positive correlation between PD-L1 and MYC expression (γ = 0.210). Compared with double-positive patients, patients with double-negative tumors had better PFS (31.1 months vs 7.1 months, P = .011) and OS (56.1 months vs 14.4 months, P = .032). 12 In addition, Alidousty et al. 37 by observing ChIP-Seq data, showed that the MYC binding site was located in the EML4 promoter region, and the overexpression of MYC in TP53 mutant cells led to the upregulation of EML4-ALK, suggesting that there may be a mechanism of MYC-dependent drug resistance in patients with increased copies of yeast. These studies provide scientific ideas and methods for the mechanism of drug resistance in EGFR-positive MYC/TP53 comutation patients and have been used to guide the follow-up use of drugs.
Our research has several limitations. First, the number of EGFR-positive patients with coexisting MYC/TP53 mutations was relatively small, which limits the comparative analysis, so we have considered expanding the sample size for future research. Second, this study is a single-center retrospective study, and additional prospective studies are needed to validate the results. Finally, although this study compared the three groups (WT/WT, TP53/WT, and MYC/TP53) of patients’ PFS, OS, ORR, and DCR, molecular experiments are still needed to explore the possible mechanisms in larger-scale experimental studies. It is worth noting that we are the first to study the prognostic and predictive value of MYC/TP53 comutation in Chinese patients with advanced EGFR-positive NSCLC treated with oral EGFR-TKIs. We believe that our research contributes to the understanding of primary drug resistance and to the development of more personalized treatments. These findings must be confirmed in a larger patient population in the future.
Conclusions
Patients with MYC/TP53 comutations with EGFR-positive advanced NSCLC are more likely to develop drug resistance after early treatment with EGFR-TKIs and have a worse clinical outcome.
Supplemental Material
sj-docx-1-tct-10.1177_15330338221138213 - Supplemental material for Evaluate the Prognosis of MYC/TP53 Comutation in Chinese Patients with EGFR-Positive Advanced NSCLC Using Next-Generation Sequencing: A Retrospective Study
Supplemental material, sj-docx-1-tct-10.1177_15330338221138213 for Evaluate the Prognosis of MYC/TP53 Comutation in Chinese Patients with EGFR-Positive Advanced NSCLC Using Next-Generation Sequencing: A Retrospective Study by Jin Cao, Juan J Gu, Yichen Liang and Buhai Wang in Technology in Cancer Research & Treatment
Footnotes
Abbreviations
Acknowledgements
The authors thank the patients and family members who gave their consent on presenting the data in this study, and the investigators and research staff involved in this study.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval
The authors state that they have obtained approval from the Ethics Committee of Northern Jiangsu People's Hospital on January 4, 2017 (Approval: ID2017005). In addition, written informed consent has been obtained from the participants involved.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
