Abstract
Keywords
Introduction
Lung cancer is the most commonly diagnosed cancer, with almost 2.2 million (11.4%) new cases and 1.8 million (18.0%) deaths reported in 2020.1,2 Nonsmall-cell lung cancer (NSCLC), which accounts for 85% of patients with lung cancer, remains challenging for treatment due to the complexity of the variation.3,4 There are several approaches to nonsmall-cell lung carcinoma therapy: surgery, radiotherapy, chemotherapy, immunotherapy, and targeted therapy. Although surgery is the most effective first-line treatment for NSCLC, 5-year survival rates are still poor.5,6 In this case, the importance of immunotherapy is beyond question.
In the targeted therapy of NSCLC, genetic alterations of epidermal growth factor receptor (EGFR), anaplastic lymphoma kinase (ALK), ROS proto-oncogene 1 (ROS1), human epidermal growth factor receptor 2 (HER2), KIT proto-oncogene (KIT), and B-Raf proto-oncogene (BRAF) have been proved to be effective prognostic biomarkers. 7 However, acquiring enough tissue for a biopsy may be problematic in clinical practice as most advanced NSCLC patients are diagnosed pathologically using microscopic biopsy specimens obtained via bronchoscopy or percutaneous needle biopsy. Insufficient samples are a central stumbling point and about 20% to 30% of patients only rely on cytology specimens for diagnosis.8,9 Besides, cell morphological uncertainty caused by quick sample processing, staining, and freezer burns at the sample's periphery could also impact the tissue's molecular profiles.
Bronchoalveolar lavage (BAL) may be an alternate source of more sensitive lung cancer indicators as a minimally invasive, easy, and cost-effective molecular diagnostics tool.10,11 Many scholars have paid attention to the importance of BAL in lung diseases. Mills NE et al found excellent sensitivity and concordance in detecting Kirsten rat sarcoma viral oncogene homolog (KRAS) gene mutations between BAL and tissue. 12 Another study demonstrated that extracellular vehicles (EVs) in BAL could be used for EGFR genotyping in NSCLC patients. 13 EV DNAs obtained from BAL even had a higher sensitivity than those from blood 14 and could be used to detect cancer-related mutations.10,15 However, there is limited knowledge on the yield of performing capture-based targeted ultradeep sequencing on BAL from patients with advanced NSCLC.
Gene mutation landscapes and TMB values have not been investigated simultaneously in paired tissue and BAL samples from patients with advanced NSCLC. In this study, we retrospectively collected the tissue and BAL samples and evaluated the utility of BAL for reflecting the genetic profiles of patients with advanced lung cancer.
Methods
Patients and Sample Collection
All the BAL, tissue biopsies, and plasma samples were retrospectively collected from the lung cancer patients. The study period was from June 2021 to September 2022 at Peking University Shenzhen Hospital. We confirm that all patient details have been de-identified in this cohort study. The Lavage acquisition and processing process has been described in previous publications. 16 In summary, 100 mL of sterile saline was instilled into either the middle lobe or the lingula and then extracted. The BAL was subsequently filtered through sterile gauze and centrifuged at 462g at 4 °C to yield a supernatant and a cell pellet. The cell pellets were collected for nucleic acid extraction and sequencing. 16 All the BAL, tissue biopsies, and plasma samples were collected before treatment. Of the 20 patients, 18 patients provided paired BAL and tissue samples, as shown in Supplemental Table S1. The other 2 patients provided paired BAL, tissue, and plasma samples. Meanwhile, we also gathered all patients’ epidemiological and clinicopathologic information (age, sex, smoking status, TNM stage, etc). The reporting of this study conforms to STROBE guidelines.17,18 All the patients have signed the written informed consent.
DNA Extraction, Library Preparation, and Target Capture Sequencing
Genomic DNA (50-200 ng) was extracted from formalin-fixed paraffin-embedded (FFPE) using the QIAamp DNA FFPE tissue kit (Qiagen, Hilden, Germany). Cell-free DNA (10-50 ng) was isolated from BAL and plasma using the QIAamp Circulating Nucleic Acid kit (Qiagen, Hilden, Germany). According to the manufacturer's instructions, qualified DNA samples were prepared to construct cDNA libraries (Integrated DNA Technologies, Coralville, IA, USA). Four IDT custom-designed probe panels, that is, 688-, 13-, 20-, or 50-gene panels, were used to capture their respective gene target (Integrated DNA Technologies, Coralville, IA, USA). The specific target gene list is in Supplemental Table S2. All genes are associated with solid tumors including lung cancer. To filter germline mutations, DNA from white blood cells was utilized to generate a control library. All cDNA libraries were sequenced on an MGISEQ-2000 platform (MGI, Shenzhen, China).
Sequencing Data Analysis
Oseq, a genome analysis toolkit, was used to analyze sequencing data to detect single nucleotide variants (SNVs), small insertions, deletions, copy number variations (CNVs), and gene fusions. 19 The genomic reference sequence used was genome GRCh37. Variants were annotated based on NCBI (www.ncbi.nlm.nih.gov/) annotation release 104, 1000 Genomes Project (release number 20130502), dbSNP137, Clinvar (www.ncbi.nlm.nih.gov/clinvar/), and Catalog of Somatic Mutations in Cancer v91 (cancer.sanger.ac.uk/cosmic). Interpretation of the variants in genes relevant to clinically available targeted treatment choices was performed according to the standards and guidelines for the interpretation and reporting of sequence variants in cancer, 20 in which somatic variations were categorized into a three-tiered system: tier I, variants with strong clinical significance; tier II, variants with potential clinical significance; and tier III, variants of unknown clinical significance. The TMB was obtained by calculating the number of mutations (allele frequency > 1.5%) in nondriver genes per Mb in each sample. MSIsensor and MANTIS were used to detect the status of microsatellite instability (MSI).21,22
Survival Analysis
The overall survival (OS) months were counted starting from the diagnosis. The follow-up survival status of the patients was obtained from their medical records and the consultation conducted by the hospital. The Kaplan-Meier (KM) curve for OS was present by treatment strategy. The Cox proportional hazards method was performed for multiple variant analyses with patient clinical data as additional input. The selected clinical variants for the Cox analysis were patient gender, age in years, and smoker age.
Statistical Analysis
Visualization and statistical analyses were performed using R (v4.1.0) and Oviz 23 (https://bio.oviz.org/). Shapiro-Wilk tests were employed to examine for normality for a small sample size (< 50 samples). Then, a t-test (normal data) or Wilcoxon signed-rank test (nonnormal data) were performed to analyze the differences between BAL and tissue samples. Pearson's chi-squared test and Fisher's exact test were performed to analyze the difference in variants between paired BAL and tissue samples. Power analysis was performed using the pwr package (https://cran.r-project.org/web/packages/pwr). P < .05 is regarded as statistically significant.
Results
Clinical Characteristics and Mutation Spectrum of Patients
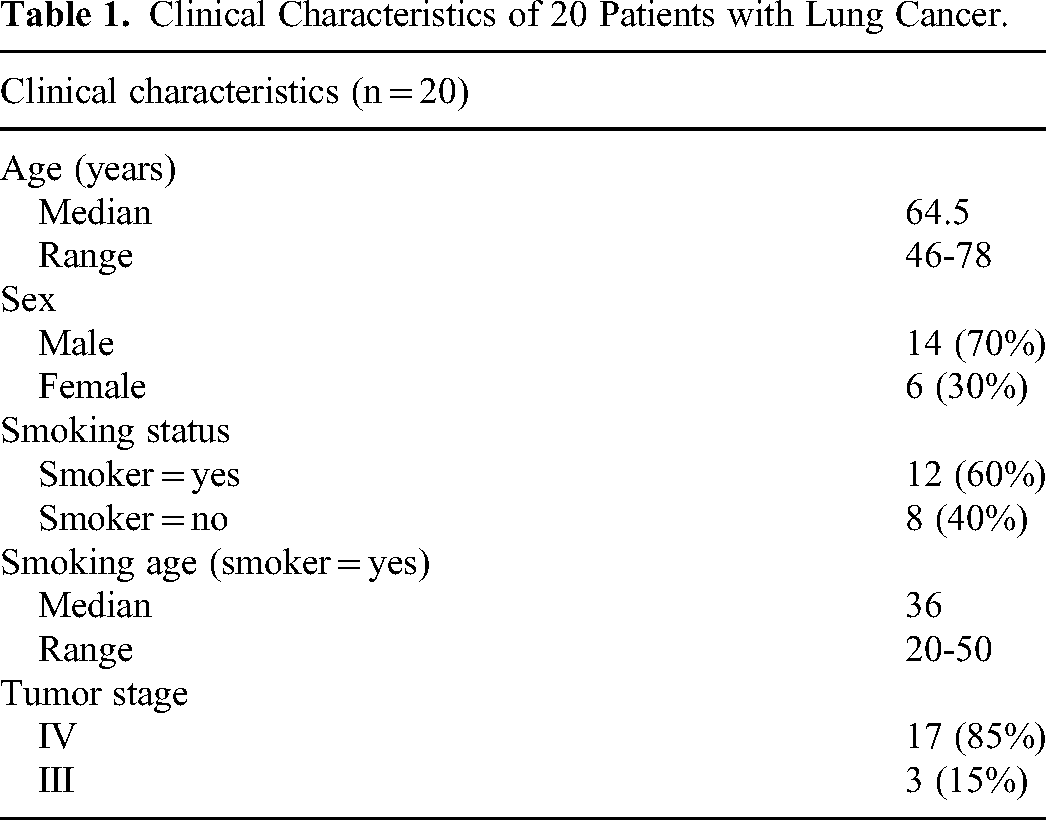
This study included 20 advanced NSCLC patients (6 female and 14 male) with a total of 42 samples (20 BAL, 20 tissue, and 2 plasma samples) collected for targeted sequencing. The median age of the patients was 64.5 years old (range: 46-78). 85% (17/20) of patients were in stage IV and 15% (3/20) were in stage III. 60% (12/20) of patients had a history of smoking and 40% (8/20) were never-smokers. More details on clinical characteristics can be found in Table 1.
Clinical Characteristics of 20 Patients with Lung Cancer.
Among the 42 sequenced samples, 15 BAL, 15 tissues, and 1 plasma sample were screened using a 688-gene panel, and the remaining were screened using 13-, 20-, or 50-gene panels. There was a total of 12 genes shared in these panels, that is, EGFR, ALK, BRAF, Erb-B2 receptor tyrosine kinase 2 (ERRB2), MET proto-oncogene (MET), ret proto-oncogene (RET), ROS1, NRAS proto-oncogene (NRAS), mitogen-activated protein kinase 1 (MAPK1), neurotrophic receptor tyrosine kinase 1 (NRTK1), fibroblast growth factor receptor 3 (FGFR3), KRAS. The mean depth of targeted gene coverage of BAL, tissue, and plasma was 2255.93 ×, 2203.48 ×, and 4570.52 ×, respectively. In total, 99.57% (range: 98.21%-100%) of the targeted gene were covered with more than one sequencing read and the average base sequencing Q30 ratio is 87.87% (range: 82.60%-91.73%) in Supplemental Table S3. Additionally, the average sequencing depth (t-test, P = .45), coverage (Wilcoxon signed-rank test, P = .73), and base sequencing Q30 ratio (t-test, P = .76) did not differ between BAL and tissue, which could exclude the interference of sequencing.
As a result, 795 high-confidence somatic mutations were identified in the 42 samples, including 669 SNVs, 97 InDel (deletion, insertion, duplication), 23 copy number variations (CNVs), and 6 gene fusions. In total, 90% (18/20) of the patients had at least one potentially actionable variant. We showed the top 45 gene mutations with the highest frequency in Figure 1A. The most common mutations in 42 samples were tumor protein 53 (TP53) followed by zinc finger homeobox 4 (ZFHX4), LDL receptor-related protein 1B (LRP1B), CUB and sushi multiple domains 3 (CSMD3), EGFR, mucin 16 (MUC16), ryanodine receptor 2 (RYR2), lysine methyltransferase 2C (KM2TC), FAT atypical cadherin 3 (FAT3), and ALK. Full details of variants for all patients are available in Supplemental Table S4.

(A) Mutations profile and clinical characteristics of 20 patients. The mutation landscape shows the variation of the top 45 genes with the highest mutation frequency in 20 patients. Each square shows the mutation of each sample provided by the patient in one gene. Different colors denote 5 types of mutations. The sidebars on the left represent the percentage of patients with a specific mutation. The bar chart on the top demonstrates the distribution of TMB. (B) Mutation spectrum in tissue and BAL samples. Each column denotes one patient and each row indicates a gene. Different colors mean 3 tiers of mutations. Darker colors represent the same mutations, while lighter colors represent the different mutations. (C) Lollipop plots of all identified mutations in EGFR, comparing BAL samples with tissue samples.
Mutation Spectrum in Paired Tissue and BAL Samples
To evaluate the detection efficacy for somatic mutations in BAL samples by targeted sequencing, we chose 17 paired BAL and tissue samples (patients 3, 5, and 13 with no variants detected were excluded). In total, 426 and 363 genetic aberrations were identified in tissue and BAL samples, respectively. Most mutations (68.56%, 302/487) were found in both BAL and tissue samples. Besides, the genetic aberrations spanned 233 genes for tissue samples (362 SNVs, 7 insertions, 35 deletions, 5 duplications, 15 CNVs, and 2 gene fusions) and 209 genes for BAL samples (309 SNVs, 6 insertions, 32 deletions, 7 CNVs, 6 duplications, and 3 gene fusions). A few discrepancies existed and are marked with light color in Figure 1B. Collectively, 185 discordant mutations (ie, 5 tier I, 33 tier II, and 147 tier III) were found between BAL and their matched tissue sample. In terms of mutations in specific genes, we illustrated 2 genes for example. EGFR showed only one different mutation (Figure 1C), while TP53 has the most discordant mutations (n = 6), indicating heterogeneity between BAL and tissue samples.
For variants detection rate, there's no statistically significant difference between paired BAL and tissue samples (Pearson's chi-squared test, P = .591 for variant tier; Fisher's exact test, P = .409 for variant type) in Table 2, based on the 17 paired BAL and tissue samples. Tier I variants (96.2% vs 84.6%) and gene fusions (75% vs 50%) had higher detection rates in BAL compared with tissue samples. However, for other variants, such as tier II (89.6% vs 76.0%), tier III (87.1% vs 72.6%), SNV (89.6% vs 76.5%), InDel (74.6% vs 69.8%), and CNV (93.8% vs 43.8%), tissue samples had better variants detection rates than BAL (Table 2).
The Difference in Variants Between 17 Paired BAL and Tissue Samples.

Abbreviations: BAL, bronchoalveolar lavage; SNV, single nucleotide variant; CNV, copy number variation.
Pearson's chi-squared test.
Fisher's exact test.
Clinical Implication in Tissue and BAL Samples
To further compare clinical significance in multiple dimensions, we analyzed key indicators in cancer, including tumor mutational burden (TMB) and allele frequency heterogeneity (AFH). TMB of BAL (aTMB) and tissue (tTMB) from 14 patients (the TMB index of patients 1, 4, and 6 was missing) showed a strong correlation (R2 = 0.96, P < .001; Figure 2A). However, aTMB values were lower than tTMB values (t-test, P = .004; Figure 2A). The AFH of BAL and tissue also showed good correlations (R2 = 0.87, P < .001; Figure 2B). In advanced NSCLC patients, a poor prognosis may be indicated when AFH < 10%. 24 The results of BAL and tissue samples were consistent in predicting whether the patient's condition would deteriorate, with a consistency rate of 71.43% (10/14). There were contrasts in TMB among males, smokers, and age ≥ 70 (P < .05), but not in females, nonsmokers, and age < 70 (Figure 2C). To test the validity of this conclusion, we performed a power calculation. For TMB, under the prior significance level of 0.05, the power was 69.91%. Therefore, for 14 paired sample sizes (n = 28), the accuracy of measuring the difference between 2 samples is ∼ 70%. The TMB difference between the 2 samples can be accurately inferred with 90% certainty when the number reached 25 for patients with paired samples.

(A) The correlations of TMB between BAL and the tissue samples. (B) Correlations between BAL and tissue samples of AFH. (C) The distribution of TMB in BAL and tissue samples under different clinical features (ie, age, sex, and smoking status). (D) Survival fraction in patients with targeted therapy (n = 8) or standard of care (n = 9). n.s.: nonsignificant, * P < .05, ** P < .001.
Survival and Prognostic Factors
Among the 20 patients, 5 (25%) died. The median survival was 15.5 months (standard deviation [SD]: 5.6 months). Figure 2D shows the KM curve for overall survival for 17 patients (patients 12, 17, and 19 on immunotherapy were excluded). Although the 8 patients on targeted therapy appeared to have a better survival curve than the standard of care (SOC, n = 9) group, there was no statistical significance (P = .19). The mean survival for patients under treatment targeted on BAL mutations was 14.8 months (SD: 3.5 months) when compared with SOC group (mean ± SD, 12.6 ± 7.1 months). In the multivariant analysis using the Cox regression, none of the variants remained significant, as shown in Table 3. The Hazard Ratios for gender, age in years, and smoking time were 2.12, 1.02, and 1.01, respectively.
Multiple Variant Analysis of Prognostic Factors.

Discussion
Although tissue samples remain the preferred starting material for NGS analysis, obtaining them can be difficult in patients with advanced-stage NSCLC. 25 BAL has received a lot of attention in academic research for its easy availability11,26 and its potential to detect mutated cancer genes.13‐15 However, limited data have been published on the use of BAL for capture-based targeted ultradeep sequencing in patients with advanced lung cancer. It has been reported that for advanced NSCLC, cell-free DNA (cfDNA)-NGS derived from blood can detect additional tier-1 variants compared to tissue alone, increasing the tier-1 detection rate by 46%. 27 In this study, we found that BAL could also detect additional tier I variants with a slightly higher overall detection rate compared to tissue samples. This suggests that BAL may be used as a supplement for advanced NSCLC patients. However, for tier II and tier III variants, tissue samples appeared to have a higher detection rate, which is consistent with studies that suggest tissue is the gold standard for variant detection. 25 In our study, the genetic aberrations were also much higher (426 vs 363) identified in tissue than in BAL samples.
We demonstrated no discernible difference in the performance of capture-targeted sequencing between paired BAL and tissue samples. Since intra-tumor heterogeneity exists in lung cancer, 13 in our study, for TP53, there are 6 differential mutations between BAL and tissue samples, revealing that using paired BAL and tissue samples for clinical practice could do more favor for lung cancer patients. Another result also showed the high sensitivity and specificity of BAL EGFR genotyping. 13 In our study, the detection rate for BAL CNV was quite lower than tissue, which was consistent with the sensitivity of the plasma cfDNA CNV detection. 24 Although DNAs in BAL consist of large-sized genomic DNAs (10 kb or longer) and tumor-specific oncogenic mutant DNAs, unlike the fragmented cfDNAs (180 bp),14,28 the detection rate of CNV in the BAL was still not high. The heterogeneity in lung cancer may be responsible for mutations that are exclusively detected in blood samples but not in tumor biopsy samples, 24 biopsy samples may also exhibit a variety of mutations that originate from distinct regions within a tumor. 29 In contrast, the ctDNA allele fraction in plasma is lower than in tumor tissues due to dilution by cfDNA from normal cells. 30 In this study, we used targeted sequencing for mutation detection, only part of the information would be captured from the whole genome. The low detection rate of CNV in the BAL might also be due to the limited and diluted DNA information captured from BAL by targeted sequencing. Actually, the genetic aberrations were also low in BAL samples in our study.
In lung adenocarcinoma, EV DNAs obtained from BAL have been shown to have higher sensitivity than blood-derived cfDNAs. 14 It has also been demonstrated that EV DNAs are longer in size and have higher stability than cfDNA, although their isolation is technically more complex. 14 Additionally, BAL-derived EV DNAs have been shown to be a reliable DNA source compared to tissue samples.31,32 In our study, we collected the cell pellet derived from BAL for targeted sequencing and found no significant difference in the number of detected variants between BAL and tissue samples. To date, there's no comparison of the mutation detection sensitivity of EV DNAs and cell pellet-derived DNA from BAL in NSCLC. As the isolation of EV DNAs is technically sophisticated, 14 more work should be done in the future to compare the detection samples for clinical use.
The results demonstrated significant correlations between TMB and AFH in paired BAL and tissue samples. AFH has been identified as an independent predictor for poor prognosis. 33 There is no significant difference between BAL and tissue samples, which means AFH of BAL < 10% may also have the strongest prognostic value for advanced NSCLC patients. Otherwise, tissue TMB has been added as a biological marker for Nivolumab ± Ipilimumab immunotherapy in NCCN Guidelines for NSCLC. 34 Our study revealed significant differences between aTMB and tTMB, as has been observed in blood TMB (bTMB). Although tTMB and bTMB are positively correlated, there can be variations between different tests.28,35‐38 At present, there is no consensus on the application of aTMB in clinical tumor treatment, and further research is necessary to establish a consensus on the optimal use of TMB in clinical tumor treatment.
Additionally, our prognostic and overall survival analysis suggested that patients with therapy targeted at BAL mutations appeared to have a better survival curve, although statistical significance was not observed. This might be due to the small sample size and short follow-up period. In this observational study, patients were consecutively collected only if they had sequencing results from both BAL and tissue. Due to the strict inclusion criteria, only 20 patients were included in the study, which limited the generalizability of the conclusions. However, we observed significant correlations of TMB between BALs and paired tissues. Additionally, the results showed no statistical difference in variant detection between BAL and tissue samples, indicating the potential usability of BAL in advance NSCLC. For future studies, more patients could be enrolled and longer follow-up time could be applied for further investigation.
Conclusion
In conclusion, we demonstrated the usability of BAL for detecting mutations with strong clinical significance. Furthermore, BAL was found to be highly related to tissue samples in mutation detection, TMB, and AFH imputation. However, due to the lower aTMB values, aTMB cannot be substituted for tTMB directly in clinical practice. Overall, BAL may be used as a supplement for the diagnosis, treatment, and prognosis evaluation of NSCLC, particularly for patients with insufficient tissue biopsy.
Supplemental Material
sj-xlsx-1-tct-10.1177_15330338231202881 - Supplemental material for Bronchoalveolar Lavage as Potential Diagnostic Specimens to Genetic Testing in Advanced Nonsmall Cell Lung Cancer
Supplemental material, sj-xlsx-1-tct-10.1177_15330338231202881 for Bronchoalveolar Lavage as Potential Diagnostic Specimens to Genetic Testing in Advanced Nonsmall Cell Lung Cancer by Xuwen Lin, Yazhou Cai, Chenyu Zong, Binbin Chen, Di Shao, Hao Cui, Zheng Li and Ping Xu in Technology in Cancer Research & Treatment
Footnotes
Abbreviations
Acknowledgements
This work was supported by the National Natural Science Foundation of China (grant no 81972773), Natural Science Foundation of Guangdong Province (2023A1515012460), Shenzhen Science and Technology Innovation Commission Foundation (grant no. JCYJ20190809103203711, grant no. JCYJ20210324105411031), The Program for Clinical Research at Peking University Shenzhen Hospital (grant nos. LCYJ2021022, LCYJ2021008). We thank all patients who participated in this study and their families.
Author Contributions
Conception and design: PX and ZL. Administrative support: PX and ZL. Provision of study materials or patients: PX, XL, and YC. Collection and assembly of data: PX, XL, and YC. Data analysis and interpretation: BC, DS, and HC. Manuscript writing: All authors. Final approval of manuscript: All authors.
Declaration of Conflicting Interests
All author(s) declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Program for Clinical Research at Peking University Shenzhen Hospital, Natural Science Foundation of Guangdong Province, Shenzhen Science and Technology Innovation Commission Foundation, National Natural Science Foundation of China, (grant numbers LCYJ2021008, LCYJ2021022, 2023A1515012460, JCYJ20190809103203711, JCYJ20210324105411031, and 81972773).
Ethical Statement
The studies involving human participants were reviewed and approved by the Ethics Committee of the Peking University Shenzhen Hospital (grant no. 2021-316). The patients provided their written informed consent to participate in this study. The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Data Availability Statement
The data that supports the findings of this study are available in the Supplemental material of this article.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
