Abstract
Purpose:
Esophageal cancer is one of the most common malignancies worldwide. Ubiquitin-dependent degradation of regulatory proteins reportedly plays a central role in diverse cellular processes. This study investigated the expression levels of ubiquitin in esophageal squamous cell carcinoma tissues and the functions of ubiquitin in the context of esophageal squamous cell carcinoma progression.
Methods:
The expression of ubiquitin in esophageal squamous cell carcinoma and normal esophageal samples was determined via immunohistochemistry. Serum ubiquitin levels were determined by enzyme-linked immunosorbent assay. The association between serum ubiquitin level and clinicopathological factors was analyzed. Real-time PCR analysis was employed to measure the mRNA levels of the ubiquitin coding genes
Results:
Ubiquitin expression in esophageal squamous cell carcinoma tissues was markedly higher than that in normal and tumor adjacent tissues. The levels of ubiquitin in esophageal squamous cell carcinoma serum samples were significantly higher than those in healthy controls. Serum ubiquitin levels were correlated with tumor stage and lymph node metastasis. To silence the expression of ubiquitin, we knocked down the ubiquitin coding genes
Conclusion:
Ubiquitin expression is upregulated in esophageal squamous cell carcinoma tissues and serum samples. Serum ubiquitin levels were correlated with tumor stage and lymph node metastasis. Downregulation of ubiquitin suppresses the aggressive phenotypes of esophageal squamous cell carcinoma cells by complex mechanisms; ubiquitin may represent a novel target for the treatment of esophageal squamous cell carcinoma.
Keywords
Introduction
Esophageal cancer is the seventh most common cancer and the sixth most common cause of cancer-related mortality. In 2018, 572,034 new cases of esophageal cancer and 508,585 esophageal cancer-related deaths were reported; more than half of these deaths occurred in China. 1 There are two main histological types of esophageal cancer: esophageal squamous cell carcinoma (ESCC) and esophageal adenocarcinoma. In China, approximately 95% of esophageal cancer cases are of the ESCC type. Despite continuous advances in the management of cancer, esophageal cancer still has an unfavorable prognosis because of its late diagnosis and rapid metastasis. During the past few years, the incidence of esophageal cancer in China has declined slightly. 1 -3 However, with the large population base of China, the national cancer burden continues to rise, and the underlying molecular mechanisms of ESCC remain unclear.
The stability and correct degradation of intracellular proteins is essential for maintaining normal cell functions. It is well accepted that cellular proteins are degraded by three main proteolytic pathways: the ubiquitin-proteasome proteolytic pathway, lysosome-mediated proteolysis, and the caspase hydrolysis pathway.
4
Ubiquitin/proteasome system (UPS)-mediated degradation of intracellular proteins influences numerous cellular processes, including signal transduction, cell cycle progression, development, apoptosis, and metabolic regulation.
5
-7
Lack of proper control over the UPS may contribute to pathological conditions, including cancer.
8
Proteasome inhibitors, such as bortezomib and carfilzomib, have emerged as novel and promising anticancer drugs in clinical applications.
9,10
Ubiquitin is a small 76-amino acid protein (8.6 kDa) that is highly conserved among eukaryotes.
11
Ubiquitin is encoded by four different genes:
By governing proteolysis, ubiquitin indirectly plays crucial roles in a variety of cellular processes and housekeeping functions. Altered ubiquitin levels may have a positive relationship with several cancer types. Choongseob Oh et al. found that the downregulation of ubiquitin inhibited the proliferation of breast cancer and prostate cancer cells and that ubiquitin may serve as an anticancer target. 13 Studies have shown that the expression level of ubiquitin is significantly higher in hepatocellular carcinoma (HCC) than in other chronic liver diseases, which may be due to the lack of poorly differentiated HCC. 14 We have previously reported that ubiquitin expression was significantly upregulated in human lung cancer cells and that silencing ubiquitin genes inhibited the proliferation and radioresistance of H1299 xenografts by promoting apoptosis. 15 These findings suggest that ubiquitin may act as an inhibitor of cell growth in certain types of cancer cells.
However, the expression of ubiquitin and its function in the progression of ESCC have not been fully elucidated. In this study, we investigated the expression of ubiquitin in ESCC and normal tissue and serum samples. We further demonstrated the role of ubiquitin in driving cell proliferation and in shaping the proteomic landscape in ESCC cells.
Materials and Methods
Tissue Samples
The ESCC samples were collected from patients (n = 120) who had not received either chemotherapy or radiotherapy prior to surgery. For these patients, if preoperative ultrasonography or computed tomography (CT) did not show any enlarged cervical lymph nodes (minor axis < 0.5 cm), patients underwent a subtotal esophagectomy with two-field lymphadenectomy through a right thoracotomy, followed by a laparotomy. If enlarged cervical lymph nodes were found by preoperative ultrasonography or CT, patients underwent radical esophagectomy with cervico-thoraco-abdominal three-field lymphadenectomy through a right thoracotomy, followed by a laparotomy and cervical incision. The resection included nodes along the cervical part of the esophagus and deep cervix. Tumor adjacent tissues were collected 3 cm away from the tumor. Fifteen normal esophageal tissue samples were obtained from upper endoscopies of trauma patients, which were performed by the same operator. These tissues were obtained postoperatively between 2010 and 2012 from the Gastrointestinal Center, Jiangyin People’s Hospital, the Jiangyin Clinical College of Xuzhou Medical University, (Jiangyin, China) as reported previously. 16 All patients provided signed and informed consent for their tissue samples to be used for scientific research. Ethical approval for the study was obtained from the Jiangyin People’s Hospital, the Jiangyin Clinical College of Xuzhou Medical University. All diagnoses were based on pathological and/or cytological evidence. The histological features of the specimens were evaluated by a senior pathologist according to the classification criteria from the World Health Organization. 17 Tissue samples were obtained prior to chemotherapy and radiation therapy and were immediately fixed in 10% neutral buffered formalin prior to immunohistochemistry analysis.
Immunohistochemistry Staining (IHC)
The tissue samples were fixed in 10% neutral buffered formalin and embedded in paraffin. Three-micrometer thick paraffin sections were deparaffinized and heat-treated with citrate buffer (pH 6.0) for 7 min as an epitope retrieval protocol. Endogenous peroxidase was blocked with 3% hydrogen peroxide for 15 min at room temperature and tissue non-specific-binding sites were blocked with 4% skim milk for 30 min. Sections were then incubated with the ubiquitin antibody (Santa Cruz Biotechnology, Santa Cruz, CA) for 1 h and 2% skim milk to reduce the non-specific staining before being incubated with the biotinylated secondary antibody for 30 min. IHC staining was visualized with a substrate solution containing diaminobenzidine (DAB) and hydrogen peroxide. Hematoxylin was used for counter-staining. All steps were performed at room temperature. The negative controls consisted of tissue sections treated with similar staining procedures in the absence of the primary antibody. The criteria for scoring the stained sections were as follows: negative, 0 < 10% of the whole tissue section stained positive; weakly positive (+ 1), 10-25% of the whole tissue section stained positive; moderately positive (+ 2), 25-75% of the whole tissue section stained positive; and strongly positive (+ 3), 75% of the tissue section stained positive. Tissues with score 0 or 1 were considered ubiquitin negative, while those with score 2 or 3 were considered as ubiquitin positive.
Ubiquitin Detection by Enzyme-Linked Immunosorbent Assay (ELISA)
The ESCC serum samples (n = 99) were collected from the Gastrointestinal Center, Jiangyin People’s Hospital, the Jiangyin Clinical College of Xuzhou Medical University, (Jiangyin, China) prior to any treatment. The control group (n = 49) consisted of age- and sex-matched normal healthy donors with no history of major disease, and their specimens were collected from the Health Examination Center, Jiangyin People’s Hospital, Jiangyin Clinical College of Xuzhou Medical University (Jiangyin, China). All patients provided signed and informed consent for their serum samples to be used for scientific research. Ethical approval for the study was obtained from the Jiangyin People’s Hospital, the Jiangyin Clinical College of Xuzhou Medical University. Serum samples were collected between 2018 and 2019. These samples were collected and stored in aliquots at -80°C until use. Ubiquitin in the serum of ESCC patients and healthy controls was measured using a commercial human Ubiquitin ELISA kit (Elabscience, Wuhan, China). The ubiquitin concentration in the serum was calculated using a standard curve obtained from measuring the absorbance of known concentrations of highly purified recombinant human ubiquitin and expressed in ng/mL. Briefly, 0.5 mL of serum was centrifuged at 12,000 g for 10 min at 4°C, and the supernatant was stored at -70°C for 72 h. Thawed supernatant (100 μL) was used in the ELISA according to the manufacturer’s instructions. The optical absorbance of the samples was measured at 450 nm. A serial dilution of human recombinant ubiquitin was included in each assay to obtain a standard curve from which the sample concentration of ubiquitin could be calculated in ng/mL using the measured absorbance values.
Cell Culture and Transfection
The human esophageal cancer cell lines TE-1 and Eca-109 were maintained in DMEM supplemented with 10% FBS and antibiotics (Gibco, Grand Island, NY). Cells were grown at 37°C with 5% CO2.
The shRNAs targeting
Tissue Collection and RNA Extraction
TRIzol reagent was used to extract the total RNA from the ESCC cells according to the manufacturer’s protocols. Total RNA was then eluted using 100 µL of nuclease-free water. Total RNA was quantified using a NanoDrop ND-2000 spectrophotometer (Thermo Scientific, Wilmington, MA), and RNA integrity was assessed using an Agilent Bioanalyzer 2100 (Agilent Technologies, Santa Clara, CA).
Real-Time PCR Analysis
Total RNA from ESCC cells was reverse transcribed to cDNA using an oligo(dT)12 primer and Superscript II reverse transcriptase (Invitrogen). SYBR green dye (Takara, Japan) was used for amplification of cDNA. mRNA levels of the targeted genes as well as those of the internal standard, glyceraldehyde 3-phosphate dehydrogenase (
Cell Viability Assay
Cell viability was evaluated by using the 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyl-2H-tetrazolium bromide (MTT) assay. Cells were plated in 96-well plates. The next day, cells were treated with or without 100 nM cisplatin, according to the experimental design. The cells were then incubated with 20 µL MTT (5 mg/mL) for 4 h. The medium was removed, 100 µL of DMSO was added, and the optical density (OD) at 490 nm was measured using a Microplate Reader (Bio-Rad, Hercules, CA). The viability index was calculated as the experimental OD/control OD. Three independent experiments were performed in quadruplicate.
Focus Formation
ESCC cells transfected with the indicated vectors were plated at low density (1,000 cells per 6-cm plate), incubated for 10 days, fixed, and stained with crystal violet. Foci and colonies containing more than 50 cells were counted using a microscope.
In Vitro Migration Assay
Migration was assayed using a modified Boyden chamber (Corning-Costar, Cambridge, MA) that contained a polycarbonate Transwell membrane filter (6.5 mm diameter, 8 µm pore size). The upper chamber was plated with 1 × 105 TE-1 or Eca-109 cells in serum-free DMEM. The lower chamber contained 600 µL medium from vector- or shRNA-transfected cells. After 5 h incubation at 37°C in 5% CO2, the filter was removed. Cells on the upper surface of the filter were wiped off, while the migrated cells on the bottom surface were fixed with methanol, stained with Giemsa, and counted under a microscope.
Protein Extraction for Proteomic Analysis
shRNA-NC and shRNA-
Tryptic Digest
The protein solution was reduced with 10 mM DTT for 1 h at 37°C and alkylated with 20 mM IAA for 45 min at room temperature in the dark. The protein sample was diluted with 100 mM TEAB to a final urea concentration of less than 2 M. Finally, trypsin was added at a 1:50 trypsin-to-protein ratio for the first overnight digestion and a 1:100 trypsin-to-protein ratio for the second 4 hdigestion. Approximately 100 µg of protein from each sample was digested with trypsin for subsequent analysis.
Tandem Mass Tag (TMT) Labeling
After digestion, the tryptic peptides were desalted using a Strata X C18 SPE column (Phenomenex, Torrance, CA, USA) and vacuum-dried. Peptides were reconstituted in 0.5 M TEAB and processed with a 6-plex TMT kit according to the manufacturer’s protocol. One unit of each TMT reagent (defined as the amount of reagent required to label 100 µg of protein) was thawed and reconstituted in 24 µL acetonitrile (ACN) and added to each sample. Next, the peptide mixtures were incubated for 2 h at room temperature. The mixtures were pooled, desalted, and dried by vacuum centrifugation.
HPLC Fractionation
The labeled peptide sample was then fractionated by high pH reverse-phase HPLC using an Agilent 300 Extend C18 column (5 µm particles, 4.6 mm ID, and 250 mm length). Peptides were separated with a gradient of 2% to 60% acetonitrile in 10 mM ammonium bicarbonate (pH 10) for over 80 min, into 80 fractions. These were then combined into 18 fractions and dried by vacuum centrifugation.
Quantitative Proteomic Analysis by LC-MS/MS
Peptides were dissolved in 0.1% FA and directly loaded onto a reverse-phase pre-column (Acclaim PepMap 100, Thermo Scientific, Rockford, IL, USA). Next, these peptides were separated using a reverse-phase analytical column (Thermo Scientific, Rockford, IL, USA). The gradient began with an increase from 8% to 22% in solvent B (0.1% FA in 98% ACN) over 26 min, followed by an increase from 22% to 35% over 6 min, finally rising to 80% over 4 min, and holding at 80% for the last 4 min, all at a constant flow rate of 300 nl/min on an EASY-nLC 1000 UPLC system. Spectra were obtained using a Q ExactiveTM plus hybrid quadrupole-Orbitrap mass spectrometer (Thermo, Woburn, MA, USA).
The peptides were subjected to NSI (Nano Spray Ionization) source ionization followed by tandem mass spectrometry (MS/MS) in a Q ExactiveTM plus that was coupled online to the UPLC. Intact peptides were detected at a resolution of 70,000. Peptides were fragmented for MS/MS using an NCE (Nano Capillary Electrospray) of 33, and the resulting ion fragments were detected at a resolution of 17,500. A top-20 data-dependent method was applied to the top 20 precursor ions above a threshold ion count of 2E4 in the MS survey scan, with a 30.0 s. dynamic exclusion. The electrospray voltage applied was 2.0 kV. The automatic gain control (AGC) setting of 5E4 ions was used to prevent overfilling of the ion trap. For MS scans, the m/z scan range was 350 to 1800. A fixed first mass was set at 100 m/z.
Database Search
The resulting spectra were processed using the Mascot search engine (v.2.3.0). Tandem mass spectra were searched against the Swiss Prot
We first checked the mass error of all the identified peptides. The distribution of mass error was near zero, and usually less than 0.02 Da. Most of the identified peptides were between 8 and 16 amino acids in length, which agrees with the expected length of tryptic peptides and indicates that tryptic digestion was efficient.
KEGG Annotation
KEGG analysis was performed to determine the biological roles of the differentially expressed mRNAs based on the latest KEGG (Kyoto Encyclopedia of Genes and Genomes) database (http://www.genome.jp/kegg/). The
Statistical Data Analysis
Data are expressed as the mean ± standard error of the mean (SEM) of at least three independent experiments. Standard error bars are included for all data points. The data were first analyzed with the Kolmogorov-Smirnov test for data distribution normality. The data were then analyzed using Student’s
Results
Ubiquitin Expression Is Upregulated in ESCC Tissues
To investigate the role of ubiquitin in ESCC, we first evaluated the expression of ubiquitin in ESCC tissue samples, tumor adjacent tissues and normal esophageal tissues by IHC analysis. Fifteen paraffin-embedded normal esophageal tissue samplesand 120 ESCC tissues, 52 of which had corresponding adjacent tissues, were collected. Ubiquitin staining was significantly stronger in ESCC samples than in tumor adjacent or normal tissue samples. As shown in Figure 1A and B, ubiquitin expression was noticeably more pronounced in the nucleus and cytoplasm of ESCC cells. These results indicated that ubiquitin was overexpressed in ESCC and might play important roles in the progression of ESCC.

Overexpression of ubiquitin in ESCC tissues and serum samples. (A) Representative images of immunohistochemical (IHC) staining for ubiquitin in normal esophageal tissues, tissues adjacent to esophageal squamous-cell carcinoma (ESCC), and ESCC tissues (magnification, 200×). (B) IHC staining score for ubiquitin in normal esophageal tissues (n = 15), ESCC tissues (n = 120), and tissues adjacent to the ESCC (n = 52), *
Serum Ubiquitin Levels Are Elevated in ESCC Patients
We next examined the serum levels of ubiquitin in 99 ESCC patients and 49 healthy controls by ELISA, and the demographic features of the patients are summarized in Table 1. The results revealed that the serum levels of ubiquitin in the ESCC group were notably higher than those in the control group. The mean serum levels of ubiquitin were 78.869 ± 7.418 ng/ml in the ESCC group, and 42.946 ± 4.098 ng/ml in the control group (
Patient Demographic Features for the Serum Ubiquitin Analysis.

Association of Serum Ubiquitin Level With Clinicopathological Factors
The relationship between serum ubiquitin level and clinicopathological factors is shown in Table 2. The serum ubiquitin level was correlated with an advanced tumor stage (P = 0.026). The serum ubiquitin level was found to be significantly higher in stage III-IV than in stage I-II tumors (94.219 ± 11.888 vs. 61.18 ± 27.536,
Relationship Between Clinicopathological Features and Serum Ubiquitin Levels.
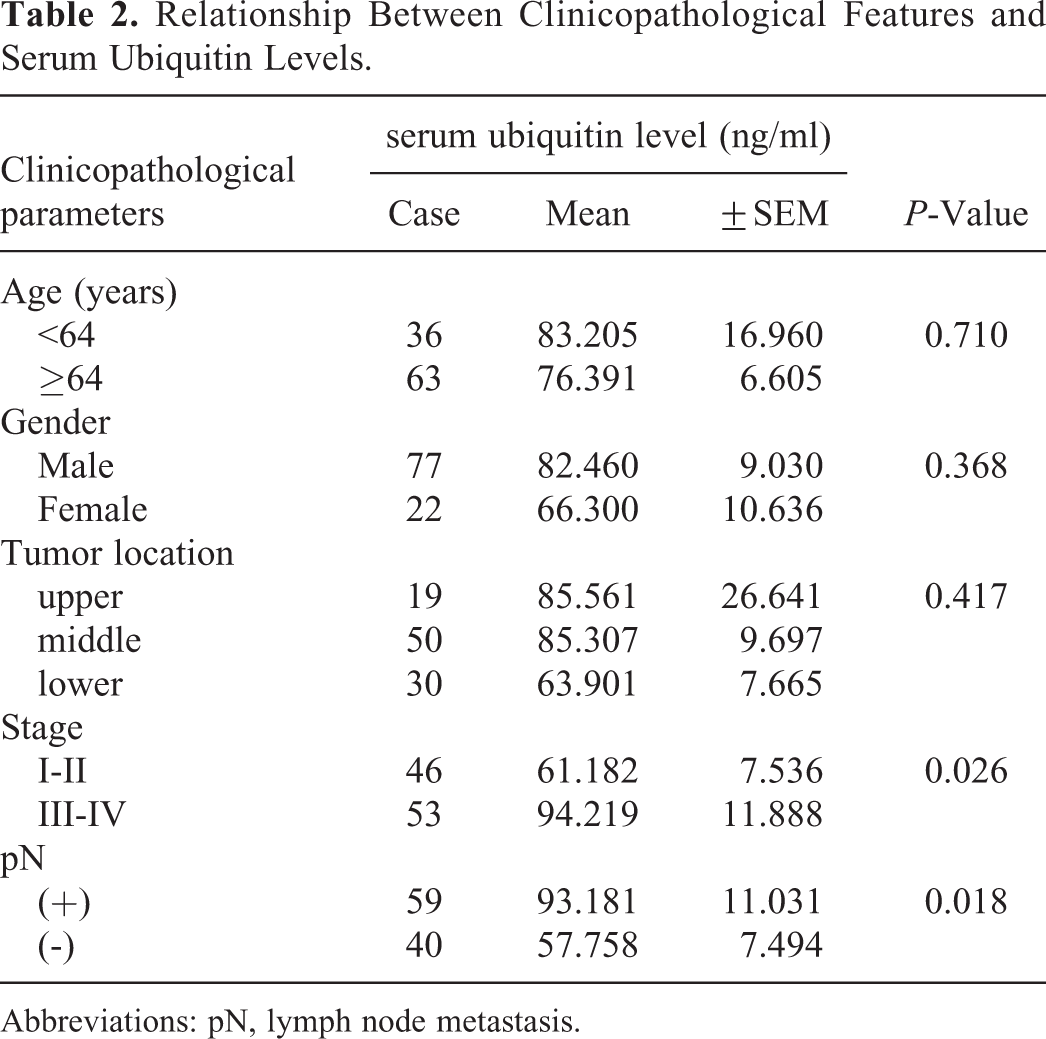
Abbreviations: pN, lymph node metastasis.
Knockdown of Ubiquitin Attenuated the Aggressive Phenotypes in ESCC Cells
To determine whether ubiquitin is associated with aggressive phenotypes in ESCC cells, we knocked-down ubiquitin in Eca-109 and TE-1 cell lines. The cells were transfected either with the shRNA control (shNC) or with 4 shRNAs targeting

Knockdown of ubiquitin coding genes (
We first evaluated the effect of the knockdown of ubiquitin coding genes on the viability of ESCC cells. As shown in the MTT assay (Figure 3A and B), knockdown of

Downregulation of
Proteomic Analysis of Dysregulated Proteins in Ubiquitin Silenced ESCC Cells
Since ubiquitin is involved in protein ubiquitination and subsequent protein degradation,
5
-7
we therefore explored the differences in protein dysregulation between parental and ubiquitin-silenced ESCC cells by proteomic analysis. In total, 4,774 proteins were quantified from the two groups of esophageal cancer cells. When the

Silencing of ubiquitin coding genes affected the protein profiles. (A) Volcano diagram and bar diagram showing significant differentially expressed proteins in the control and
Differentially Expressed Proteins in Ubiquitin Knockdown ESCC Cells (Top 20).

Discussion
Ubiquitin-dependent degradation of regulatory proteins is involved in key cellular processes, including cell cycle progression, cell proliferation, apoptosis, and DNA damage response. 5,6,19 Dysfunction of ubiquitin has been implicated in a vast array of human diseases, including cancer. Ishibashi Y et al reported that the levels of free ubiquitin (FUb) and multi-ubiquitin chains (MUC) were significantly higher in colorectal cancer tissues than in normal tissues. 19 The level of FUb in cancerous tissues was also correlated with the depth of invasion, while the level of MUC demonstrated a positive relationship with blood or lymphatic vessel invasion. Studies have shown that the labeling index of ubiquitination is significantly higher in human hepatocellular carcinoma than in other chronic liver diseases and tend to increase in proportion with the extent of undifferentiation. 20 This study, for the first time to our knowledge, illustrates the upregulation of ubiquitin in ESCC tissue and serum samples. In this study, using IHC, we detected ubiquitin in both the nucleus and cytoplasm of ESCC cells. The protein level of ubiquitin in ESCC tissues was significantly higher than that in normal and tumor adjacent tissues. Furthermore, the serum levels of ubiquitin were markedly elevated in ESCC samples compared with healthy controls. We found that serum ubiquitin levels were correlated with tumor stage and lymph node metastasis. We therefore suggest that ubiquitin may be involved in esophageal oncogenesis and tumor development.
Cancer cells acquire their malignant capabilities through uncontrolled cell proliferation and genomic instability. The downregulation of ubiquitin inhibits numerous ubiquitination-mediated pathways and is an effective way to inhibit cell proliferation in multiple cancers, including neuroblastoma, gynecological cancer, liver hepatocellular carcinoma and lung cancer.
15,20
-22
Silencing ubiquitin induces apoptotic cell death, indicating that ubiquitin might be a potential target for cancer treatment.
21,22
Consistent with previous studies, we found that knock-down of
To detect novel biomarkers and molecular targets in ESCC tissues, we identified 159 differentially expressed proteins (92 upregulated and 67 downregulated) between ESCC and normal esophageal tissues with an iTRAQ-based proteomic approach. The results revealed that multiple proteins were significantly upregulated, including ferritin light chain, ferritin heavy chain, CRABP2 and CDT1. Ferritin is a major driving force in iron delivery, cell proliferation and angiogenesis. 23 It is overexpressed in a variety of cancer cells, including breast cancer, glioblastoma, liver cancer, lung cancer, osteosarcoma, and head and neck squamous cell carcinoma and is correlated with poor prognosis. 24 -30 Wu T et al reported that expression of ferritin light chain (FLT) was elevated in glioblastoma, and downregulation of FLT inhibited the proliferation of glioblastoma cells via the GADD45/JNK pathway. 28 DNA replication factor 1 (CDT1) is overexpressed in hepatocellular carcinoma and breast cancer and contributes to oncogenesis. 31,32 Among the upregulated proteins, cellular retinoic acid-binding protein 2 (CRABP2) is reported to be associated with tissue differentiation and proliferation 33 and modulate both oncogenic and anti-cancer functions. Yang Q et al. demonstrated that the protein and mRNA levels of CRABP2 were markedly lower in ESCC specimens and correlated with TNM stage, tumor size, pathology, cell differentiation, infiltration depth and occurrence position. 34 Upregulation of CRABP2 strikingly suppressed cell proliferation, induced cell apoptosis and blocked cell metastasis; thus, it may serve as a tumor suppressor. We hypothesized that ubiquitin may mediate the biological behavior of esophageal cancer through CRABP2. Choongseob Oh et al reported that the downregulation of ubiquitin inhibited the cell proliferation and induced apoptotic cell death in cancer cells. 35 In previous studies, we found that silencing ubiquitin genes inhibited the expression of anti-apoptotic protein Bcl-2 in lung cancer H1299 cells, and silencing ubiquitin genes suppressed the proliferation and radioresistance of H1299 transplanted xenografts by promoting apoptosis. 15 Further studies are needed to explore the role of these proteins in ESCC tissues. Tumor growth and invasion involve numerous genes and pathways. Our study mainly focuses on the biological function of ubiquitin. And we’ll further explore the pathways affected by ubiquitin in subsequent studies.
In summary, our study demonstrates that ubiquitin expression is upregulated in ESCC tissues and serum samples. Serum ubiquitin levels were correlated with tumor stage and lymph node metastasis. The downregulation of
Footnotes
Abbreviations
Authors’ Note
These authors contributed equally to this work.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Statement
This study was approved by the Ethics Committee of the Jiangyin People’s Hospital, the Jiangyin Clinical College of Xuzhou Medical University, China (approval number: 2018002). All patients provided written informed consent prior to enrollment in the study.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by National Natural Science Foundation of China (31670857, 81703022 and 31700737), the Natural Science Foundation of Jiangsu Province (BK20161152 and BK20171157), Jiangsu Province Key Youth Talents Project (QNRC2016262, QNRC2016136 and QNRC2016137), and the Scientific Research Program of Wuxi Municipal Health Commission (Z201808, CXTD004, QNRC092 and QNRC082).
