Abstract
Although the treatment of liver cancer has made great progress, the mechanism of its occurrence is not completely clear. miR-155 plays an important regulatory role in tumorigenesis and development, including survival, proliferation, migration and invasion. However, the role and regulatory mechanism of miR-155 in liver cancer has rarely been reported. We analyzed miR-155 expression in liver cancer tissue samples and cell lines by qRT-PCR. The expression of miR-155 was measured by qRT-PCR before and after miR-155-mimic and sh-miR-155 transfection. CCK-8 and clonogenic assays were used to detect the proliferation of liver cancer cells. Cell scratch and invasion assays were used to detect migration and invasion. RNA-seq was used to detect the difference in RNA expression in liver cancer cells. SRPK1 expression was detected in liver cancer cells before and after transfection by qRT-PCR and western blotting. We observed that miR-155 was downregulated in liver cancer tissues compared with normal tissues. Furthermore, we demonstrated that liver cancer cell proliferation, migration and invasion are markedly suppressed by miR-155. Importantly, we also demonstrated that SRPK1 is directly regulated by miR-155 during the process of liver cancer cell proliferation and metastasis. Finally, the overexpression of miR-155 inhibits malignant biological behavior of human liver cancer cells. We report the abnormal expression of the miR-155 cluster in liver cancer cells, which inhibits cancer cell proliferation and metastasis. In addition, we identified SRPK1 as a target gene of miR-155 during the process of liver cancer cell proliferation and metastasis.
Introduction
The incidence of liver cancer worldwide is increasing annually and is a great hazard to the health of people. 1,2 As a malignant tumor disease, studies have shown that a variety of factors are involved in the complex development of liver cancer, 3 and its highly metastatic and invasive nature is a multistep process. 4 However, the cause of the disease is still unclear. Studies have shown that miRNAs have a variety of biological functions related to cancer progression, including cell survival, proliferation, and migration, and are closely related to the development of various tumors. 5 It has been reported that miR-494 can aggravate oral squamous cell carcinoma by targeting the type I collagen alpha 1 gene (collagen type I alpha 1, COL1A1). 6 MiR-155 inhibits the viability and invasion of colon cancer cells by targeting the nuclear transcription factor GATA6. 7,8 At the same time, it is reported in the literature that miR-155 can act as a tumor suppressor in a variety of metastatic cancers and that it can directly target the forkhead box protein C1 (FOXC1) in pituitary adenomas. 9 -11 However, there is no clear evidence of the relationship between miR-155 and liver cancer. Studies have shown that SRPK1 can act as a tumor promoter to regulate tumorigenesis and development and is closely related to tumors and other diseases.12,13 The expression level of SRPK1 in lung cancer cells increased significantly, and the high expression of SRPK1 resulted in worse overall survival after surgery. 14
Therefore, this study focused on whether miR-155 can inhibit the malignant biological behaviors of liver cancer cells, further studied its relationship with SRPK1 and revealed the influence of the interaction between the 2 on the underlying mechanism of liver cancer progression.
Methods
Human Samples
From February 2017 to December 2018, hepatocellular carcinoma tissues and adjacent liver cancer tissues were obtained from 48 patients with liver cancer who underwent laparoscopic precision hepatectomy at the Affiliated Hospital of Shaanxi University of Traditional Chinese Medicine. The patient information, including age, sex, tumor size, pTNM category and differentiation, is summarized in Table 1. The study was approved by the Ethics Committee of Shaanxi University of Traditional Chinese Medicine(No:2019ST341). Inform all patients of the experimental method and purpose, and sign the patient’s informed consent.
Clinicopathological features of the patients with liver cancer.

Cell Culture and Transfection
Human lung cancer cell lines, including HepG2, Huh-7, SK-Hep1, SMMC-7721 and Hep-3B (ATCC, MA), were incubated in DMEM with 10% fetal bovine serum. MiR-155-mimics, miR-155-NC, shmiR-155 and shControl were designed and synthesized in JIkai (Shanghai, China). The cells were inoculated at a density of 1 x 105 cells/well into 24-well plates at a cell confluence of approximately 50% to prepare for transfection. For transfection, 1 µl of Lipofectamine 2000 (Invitrogen, USA) and 50 µl of Opti-MEMI were mixed and added to serum-free medium (Invitrogen, USA) and allowed to stand at room temperature for 10 minutes. Two microliters of FAM-siRNA and 50 µl of Opti-MEMI were mixed and added to the serum-free medium. After incubation for 5 minutes at room temperature, the diluted FAM-siRNA and Lipofectamine 2000 were mixed for 20 minutes. The cells were then incubated for 6 hours at 37°C and 5% CO2 and then the medium was replaced with conventional medium.
Quantitative Real-Time PCR (RT-qPCR)
TRIzol reagent is used to extract total RNA. The RNA was reverse transcribed into cDNA using a reverse transcription kit. The reaction conditions were set as follows: predenaturation at 95° C for 10 minutes; degradation at 95° C for 10 seconds; annealing at 60° C for 20 seconds and extension at 72° C for 35 seconds. The 2-ΔΔt method was used to calculate the relative mRNA expression of the gene of interest. The primer sequences are shown in Table 2.
Primer sequences used in qRT-PCR.

Immunohistochemistry Analysis
The tissue samples harvested from clinical patients after operation were fixed with formalin, embedded in paraffin, and finally analyzed by immunohistochemistry. The 4-μm-thick slides were dewaxed and hydrated; the antigen was recovered, and then the slides were washed with phosphate buffered saline (PBS) and incubated with primary antibodies overnight at 4°C. The slides were then washed in PBS and co-incubated with secondary antibodies containing horseradish peroxidase (Santa, USA) for 1 hour at room temperature.
Western Blotting
The cells were lysed on ice for 30 minutes and then centrifuged at 3500 rpm for 3 hours at 4°C, and the supernatant was extracted as total protein. The sample was then mixed with loading buffer and boiled at 100°C for 5 minutes, and the prepared SDS gel was used for separation. After separation, the proteins on the gel were transferred to a PVDF membrane and incubated with the primary antibody at 4°C overnight. The membrane was incubated with a skimmed milk solution for 1 hour the next day, and then the labeled secondary antibody was blocked for 1 hour.
Cell Proliferation Assay (CCK-8)
The number of cells was adjusted to 4 × 105 cells/well and plated at 37°C in a 5% CO2 incubator for 1-4 days. Afterward, 20 µl of CCK-8 reagent was added to each well, protected from light for 3 hours, and then placed on a shaker for 15 minutes. The OD value of the cell CCK-8 mixture was measured by a microplate reader at 450 nm for viability analysis.
Colony Formation Assay
Logarithmic growth phase cells (300 cells/well on a 6-well plate) were used for the colony formation assay with trypsin digestion. A total of 2 mL of culture solution was used. The cells were cultured for 15 days, washed with PBS, air-dried at room temperature, stained with crystal violet, and counted.
Scratch Assay
The cells in the logarithmic growth phase were trypsinized, inoculated into a 6-well plate, cultured at 37°C and 5% CO2 with 2 mL of DMEM per well. When the cell density reached 80%, the cells were scratched. The serum-free medium was replaced and photos were taken at 0 h and 24 h.
Invasion Assay
The cell invasion ability was determined using a transwell chamber coated with Matrigel. After being suspended in serum-free medium, cells (5 × 104) were seeded into the upper chamber. The lower chamber was filled with complete medium containing 10% FBS. Following incubation at 37°C for 24 h, the migrated/invaded cells on the lower side of the membrane were fixed with 100% ethanol and stained with 0.5% crystal violet (Beyotime Biotechnology, Shanghai, China) at room temperature for 30 min.
RNA Transcriptome Sequencing
First, RNA samples were treated with DNase I to degrade any possible DNA contamination. Then, oligomeric (dT) beads were used to enrich the mRNA. After mixing with fragment buffer, the RNA fragments were transformed into short fragments. The first strand of the DNA was then synthesized using random hexamer primers. The second chain was synthesized by adding buffer, dNTP, RNase H and DNA polymerase I. Double-stranded DNA was purified by magnetic beads. End repair and the addition of 30-terminal mononucleotide alpha (adenine) were performed.
Statistical Analysis
The data were analyzed using GraphPad Prism 8 software. The data from 2 groups were statistically compared using unpaired Student’s t-test. One-way ANOVA with Bonferroni’s correction was used for multiple comparisons. The in vitro experiments and the data are presented as the means ± standard deviations (SDs) from at least 3 independent experiments. P values ≤0.05 were considered significant for all analyses.
Results
Expression of miR-155 in Cancer and Normal Tissues and Different Liver Cancer Cell Lines
The 48 pairs of cancer tissues and normal tissues obtained from different liver cancer patients were analyzed by qRT-PCR. The results showed that the expression of miR-155 in cancer tissues was significantly lower than that in normal tissues (Figure 1A). This finding suggests that the low expression of miR-155 in liver cancer tissues may be related to the occurrence of liver cancer. Subsequently, we examined the expression of miR-155 in 5 liver cancer cell lines (HepG2, Huh-7, SK-Hep1, SMMC-7721, and Hep-3B) and in a normal human liver cell line (HL-7702) and found that the expression of miR-155 in liver cancer cells was lower than that in Huh7 cells, and the expression in HepG2 cells was the lowest (Figure 1B). Therefore, it can be concluded that miR-155 is closely related to the occurrence of liver cancer.

MiR-155 is downregulated in clinical liver cancer tissues and differentially expressed in liver cancer cell lines. (A) The relative mRNA level of miR-155 in clinical liver cancer tissues and in normal tissues; n = 48. (B) The mRNA expression of miR-155 in a series of liver cancer cell lines was detected by qRT-PCR.
MiR-155 Inhibits Liver Cancer Cell Viability In Vitro
First, miR-155-mimics, miR-155-NC, sh-miR-155 and sh-NC were separately transfected into HepG2 and Huh-7 cells for 48 h. RNA was extracted, and qRT-PCR was used to detect the expression of miR-155 in the cells before and after transfection. The results showed that after miR-155-mimic transfection, the expression of miR-155 was significantly increased in the HepG2 cells, while the expression of miR-155 was significantly decreased in the sh-miR-155–transfected Huh-7 cells (Figure 2A and D). Subsequently, CCK-8 detected the proliferation of HepG2 and Huh-7 cells before and after transfection. The results showed that miR-155 overexpression significantly inhibited the proliferation of HepG2 cells, while miR-155 knockdown significantly promoted the proliferation ability of Huh-7 cells (Figure 2B and E). The colony formation experiment showed the same result: the number of clones of HepG2 cells after miR-155 overexpression was significantly lower than that of the control group, and the number of Huh-7 cell clones was significantly higher than that of the control group (Figure 2D and F). This finding indicates that miR-155 can effectively inhibit liver cancer cell proliferation and clonality.
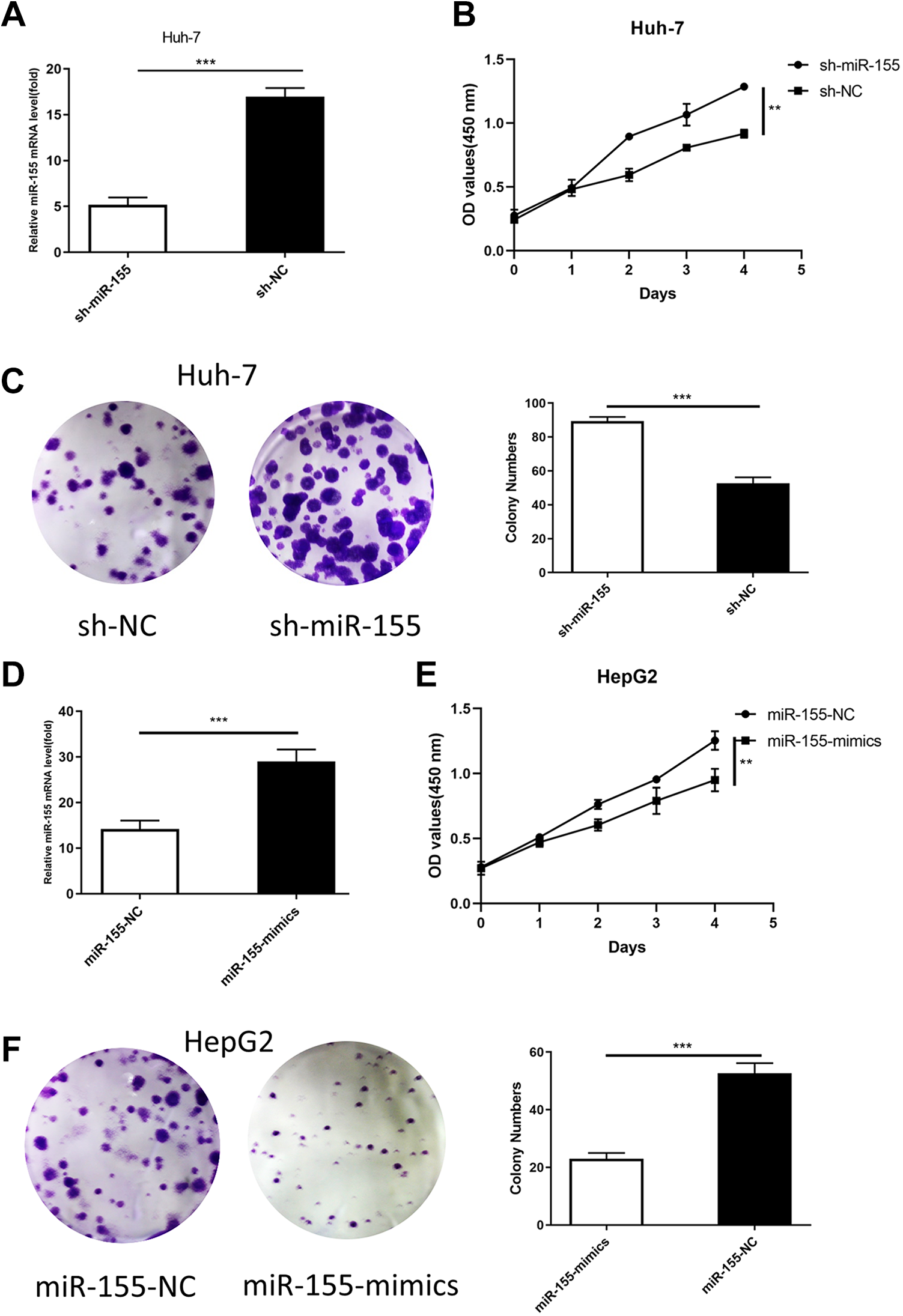
MiR-155 inhibited liver cancer cell viability in vitro. (A, D) A specific shRNA against miR-155 (sh-miR-155) and miR-155-mimics were used. (B, E) A CCK-8 assay was performed to assess the effects of miR-155 knockdown in Huh-7 cells and overexpression in HepG2 cells on cell viability. (C, F) During the 15-day colony formation assay, colony staining was used to visualize and quantify colony formation and is displayed on the column.
MiR-155 Inhibits Cell Metastasis in Liver Cancer Cells
The cell migration and invasion ability of liver cancer cells were detected before and after transfection using a cell scratch assay and invasion assay. miR-155 also promoted cell migration and invasion after knockdown of Huh-7 cells (Figure 3A and C) but inhibited cell migration and invasion after overexpression of HepG2 cells (Figure 3B and D). This finding indicates that miR-155 has an inhibitory effect on the migration and invasion of liver cancer cells.

MiR-155 inhibited cell metastasis in liver cancer cell lines. (A-B) Wound healing was measured with Huh-7 and HepG2 cells. Sh-miR-155 was transfected into Huh-7 cells, and miR-155-mimics was transfected into HepG2 cells before testing. Representative images were taken 24 hours after serum-free migration. (C-D) Transwell invasion assays were performed.
MiR-155 Inhibits the Expression of SRPK1 in Liver Cancer Cells
RNA-seq technology was used to detect the difference in expression of genes at the RNA level before and after the transfection of Huh-7 cells. The upregulation multiples of SRPK1, ING2 and DPC4 were the highest (doubled), and they were all related to the tumor proliferation pathway by KEGG pathway enrichment analysis (Figure 4A). qRT-PCR was used to verify the expression in Huh-7 cells in the sh-miR-155 group. The results were consistent with the RNA-seq results, and SRPK1 was upregulated the most. According to the results of the RNA-seq assay, we used qRT-PCR and western blotting to detect the expression of SRPK1 in Huh-7 and HepG2 cells before and after transfection. SRPK1 significantly increased with decreasing miR-155 in Huh-7 cells (Figure 4C and E) but decreased with increasing miR-155 in HepG2 cells (Figure 4D and F).
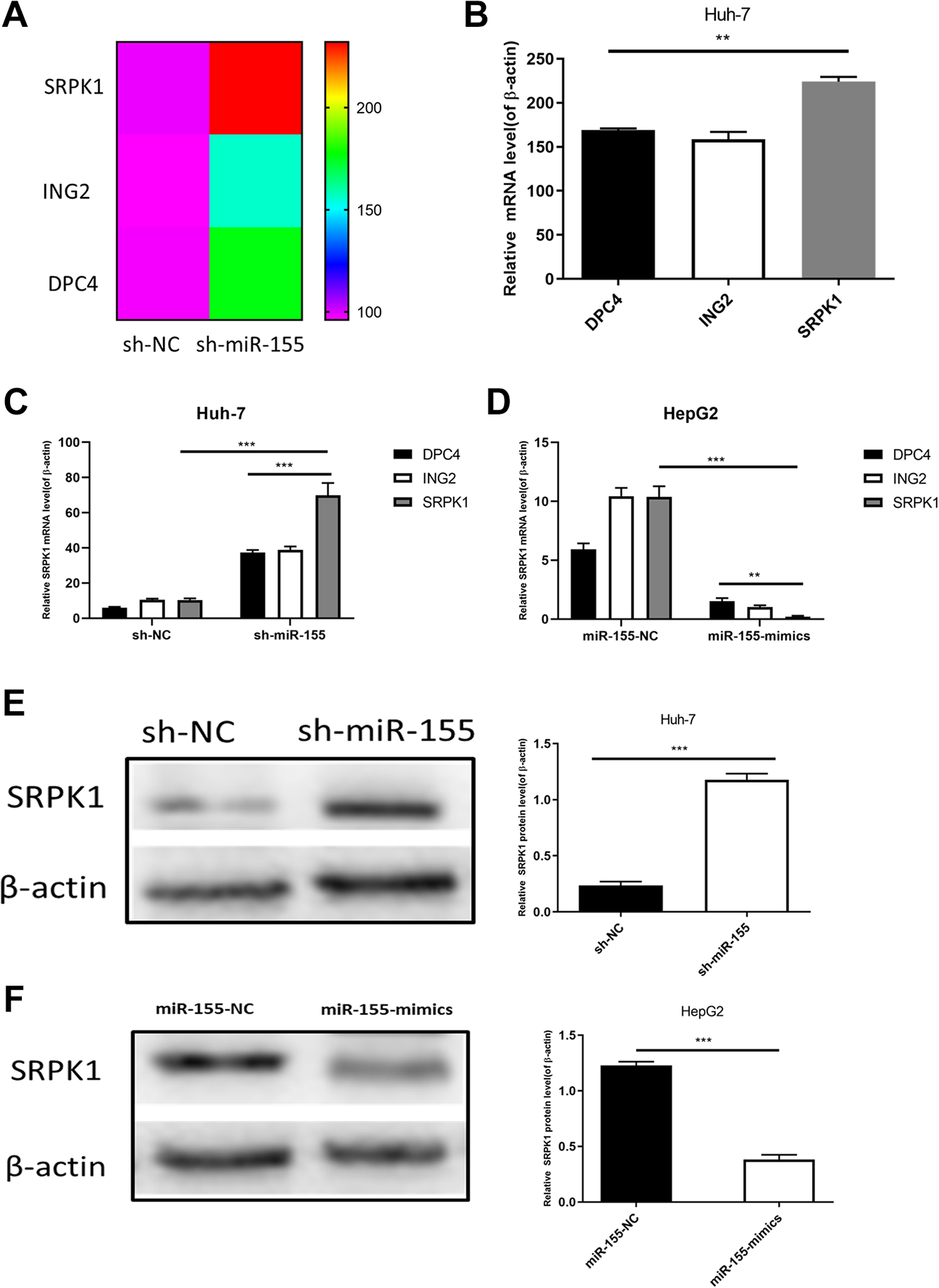
MiR-155 inhibited SRPK1 in Huh-7 and HepG2 liver cancer cells. (A) RNA-seq detected the expression of different genes in sh-miR-155–transfected Huh-7 cells. (B) qRT-PCR was used to detect the expression of different genes in Huh-7 cells. (C, D) qRT-PCR was used to detect the expression of SRPK1 in Huh-7 cells and HepG2 cells before and after transfection. (E, F) Western blotting was used to analyze SRPK1 expression in miR-155-depleted Huh-7 cells and miR-155-overexpressing HepG2 cells.
Discussion
Liver cancer is considered to be one of the most malignant diseases of the digestive diseases, and the incidence of liver cancer is increasing year by year in China. 3 Although the treatment of liver cancer has improved, due to its low survival rate, the current research hotspots are more focused on new targeted therapies for liver cancer. 15 As a new strategy, gene therapy has achieved effective results. In this study, we focused on the functional role of miR-155 and SRPK1 in liver cancer. Based on the experimental results, we concluded that miR-155 can inhibit malignant biological behaviors such as the proliferation, migration and invasion of liver cancer cells by targeting SRPK1.
Studies have reported that miR-155 acts as a tumor suppressor in various types of malignancies to inhibit tumors, especially in the gastrointestinal system. 16 Some studies demonstrated that miR-155 can effectively inhibit the cell cycle progression of gastric cancer. 17 At the same time, some results showed that miR-155 can promote tumor cell apoptosis and inhibit cell proliferation and migration in bladder cancer. 10 To further investigate the role and molecular mechanism of miR-155 in liver cancer, we first tested the expression of miR-1355 in liver cancer tissues and cell lines by qRT-PCR. The results showed that miR-155 in liver cancer tissues and cell lines was significantly downregulated, the highest fold was achieved in Huh-7 cells, and the expression in HepG2 was the lowest.
In other types of human malignancies, such as esophageal cancer, lung cancer, bladder cancer and colorectal cancer, the data and analyses indicate that miR-155 reduces the proliferation, invasion and migration of tumor cells. 18 -20 In addition, CCK-8 and colony formation assays showed that miR-155 overexpression significantly inhibited the proliferation of HepG2 cells, while miR-155 knockdown significantly promoted the proliferation of Huh-7 cells. miR-155 also promoted cell migration and invasion after the knockdown of Huh-7 cells but inhibited cell migration and invasion after the overexpression of HepG2 cells. These results indicate that miR-155 acts as a tumor suppressor during the development of liver cancer.
As for the further mechanism, we hypothesized that miR-155 downregulates the malignant biological capacity of liver cancer cells by modulating the corresponding genes. We used RNA-seq to detect the expression of genes at the RNA level in Huh-7 cells before and after miR-155 downregulation and found that SRPK1, ING2 and DPC4 were upregulated after miR-155 decreased. Then, verification by qRT-PCR was performed, and the results are consistent with the results of RNA-seq, where SRPK1 has the highest upregulation. Then, qRT-PCR and western blotting were used to detect changes in SRPK1 at the RNA and protein levels before and after miR-155 transfection. The results showed that SRPK1 decreased significantly with the overexpression of miR-155, indicating that miR-155 has an inhibitory effect on SRPK1 and has the strongest targeting effect.
Overall, this study reveals that miR-155 inhibits malignant biological behaviors such as the proliferation and migration of liver cancer cells by directly targeting SRPK1 and decreasing its expression. Through a large number of detection and bioinformatics analyses, this study provides a certain understanding of the mechanism of liver cancer at the gene level as well as a feasible targeted therapy strategy for the clinical treatment of liver cancer.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Scientific Research Fund of Shaanxi Provincial Department of Science and Technology (NO.2016SF-234, NO.2018KJXX-093).
