Abstract
Background:
Approximately one third of type 2 diabetes mellitus (T2DM) cases present with diabetic nephropathy (DN), the leading cause of end-stage renal disease. Inflammation plays an important role in T2DM disease and DN pathogenesis. NLRP3 inflammasomes are complexes that regulate interleukin-1B (IL-1B) and IL-18 secretion, both involved in inflammatory responses. Activation of NLRP3 is associated with DN onset and progression. Here, we explore whether DN is associated with variants in genes encoding key members of the NLRP3 inflammasome pathway.
Methods:
Using genome-wide association data, we performed a pilot case-control association study, between 101 DN-T2DM and 185 non-DN-T2DM cases from the Hellenic population across six NLRP3 inflammasome pathway genes.
Results:
Three common CARD8 variants confer decreased risk for DN, namely rs11665831 (OR = 0.62, p = 0.016), rs11083925 (OR = 0.65, p = 0.021), and rs2043211 (OR = 0.66, p = 0.026), independent of sex or co-inheritance with an IL-1B variant.
Conclusion:
CARD8 acts as an NLRP3, NF-κB and caspase-1 inhibitor; perhaps, alterations in the cross-talk between CARD8, NF-κB, and NLRP3, which could affect the pro-inflammatory environment in T2DM, render diabetic carriers of certain common CARD8 variants potentially less likely to develop T2DM-related pro-inflammatory responses followed by DN. These preliminary, yet novel, observations will require validation in larger cohorts from several ethnic groups.
Introduction
Diabetic Nephropathy (DN), a serious microvascular complication of type 2 diabetes mellitus (T2DM), affects one third of T2DM patients and is the leading cause of end-stage renal disease (ESRD) worldwide. The incidence rate of the disease is still rising, 1 highlighting the fact that the pathogenic mechanisms implicated in DN are complex and not fully elucidated.
Inflammasomes are cytoplasmic protein complexes that detect different stress signals; they control the secretion of pro-inflammatory cytokines, and are, thus, important members of the immune inflammatory reactions. The NLRP3 inflammasome, specifically, consists of the protein products of NLRP3, PYCARD, and CASP1 genes. In its resting state, NLRP3 activation by oligomerization is attenuated by the adaptor protein caspase recruitment domain family member-8, encoded by CARD8.2,3 NLRP3 is involved in caspase-1 activation, which controls the cleavage of precursor IL-1B and IL-18 into bioactive cytokines which trigger downstream inflammatory responses. 4 Pro-inflammatory cytokines, such as IL-1B, are significantly increased in T2DM and inflammation seems to play an important role in T2DM pathogenesis. 5
NLRP3 inflammasome has been implicated in the pathogenesis of several renal conditions including DN. 4 NLRP3 inflammasome is expressed in human renal cells and its activation could be implicated in DN onset and progression. 6 Animal studies have also shown that renal inflammation promotes DN. 7 The NLRP3 inflammasome is activated by several signaling pathways, including NF-κB. 4 The negative regulator of NLRP3, namely CARD8, was found to indirectly inhibit NF-κB, presumably via the direct inhibition of IKK kinase subunit IKKγ/NEMO, 8 and to directly inhibit caspase-1, 9 thus controlling a pro-inflammatory cytokine response. Perturbations in these pathways could also promote structural changes causing renal cell injury, and compromise cell survival, leading to suboptimal kidney function. However, the exact underlying mechanisms of inflammasome activation in DN remain poorly understood.
Genetic variations in NLRP3 inflammasome have been previously associated with obesity, insulin resistance and diabetes. 10 Their association with microvascular diabetic complications remain largely unknown, despite the documented role of inflammation in DN. 7 In the present study, we explore whether variations in NLRP3, PYCARD, CASP1, and CARD8, as well as IL-1B and IL-18, associate with the development of inflammatory responses in T2DM and specifically with DN.
Methods
Study subjects
A group of 286 unrelated subjects, originally recruited through the GR-DIAGENES Initiative for the Hellenic population, 11 were included in the study. Inclusion criteria for T2DM were according to the American Diabetes Association (2010). The sample consisted of two subgroups: (a) 101 T2DM patients with DN in all five stages of chronic kidney disease, persistent proteinuria (⩾500 mg/24h), albuminuria (⩾300 mg/24h), random protein/creatinine ratio ⩾0.5 g/day (or ⩾500 mg/g or ⩾50 mg/mmol) or random urine albumin/creatinine ratio ⩾0.3 g/day (or ⩾300 mg/g or ⩾30 mg/mmol), and diabetic retinopathy, and (b) 185 T2DM patients as non-DN controls with normoalbuminuria and an estimated GFR >60 mL/min/1.73m2. Patients with (clinical or laboratory) evidence of non-diabetic nephropathy, hematuria, urinary tract disease, acute illness or diabetes caused by liver dysfunction were excluded. Laboratory measurements were performed as previously described. 12 All participating patients gave written informed consent. The study was approved by the Ethics Committee of the Alexandroupoli University General Hospital and was in accordance with the Helsinki Declaration of Human Rights.
Genotyping and bioinformatics analysis
Genomic DNA was extracted from peripheral whole blood. Whole-genome genotyping was performed on the Illumina Infinium PsychArray-24 v.1.1, which targets 603k SNPs, of which 560k are described in dbSNP. We performed standard genome-wide quality control steps to filter for sub-standard markers and samples that could introduce biases. Population stratification was evaluated by Principal Component Analysis (PCA) using EIGENSOFT. We extracted the subset of markers that reside within the genes of interest, using a flanking window of 20 kb per gene. Genomic coordinates were according to GRCh37/hg19 assembly. Quality control steps and data analysis were conducted using PLINK.
Statistical analysis
Association testing was performed using PLINK. Standard chi square test was used for single-marker association analysis in order to compare allele and genotype frequencies between patient groups, followed by permutation testing. The level of statistical significance was set to 0.05. For inter-group comparisons of categorical variables we used chi square test, whereas for continuous variables we used either Student’s t test or Mann-Whitney U test for normally or not normally distributed data, respectively. Data were tested for normality by Kolmogorov–Smirnov test.
Results
In the present study we performed a pilot case-control association study of six genes encoding key protein members of the NLRP3 inflammasome pathway, in order to explore potential genetic associations with DN across a group of 101 T2DM patients with DN and 185 non-DN T2DM patients. DN subjects were followed for over a period of at least 7 years 12 ; their average duration of T2DM was 19.7 ± 8.5 years. The non-DN patients had a documented history of T2DM for an average of 16.7 ± 11 years, and at least 5 years of disease diagnosis. The proportion of males exceeded that of females among the DN cases; sex distribution was almost equal for the non-DN group (p = 0.01). There were no statistically significant differences between DN versus non-DN T2DM cases regarding age at diagnosis (50.6 ± 10.9 years vs 51.6 ± 10.3 years, p = 0.44), BMI (31.7 ± 5.5 kg/m2 vs 31.2 ± 5.4 kg/m2, p = 0.39), waist circumference (106.8 ± 11.0 cm vs 103.7 ± 14.7 cm, p = 0.083), HbA1c [7.53 ± 1.1% vs 7.37 ± 1.2% (or 58.8 ± 12.4 mmol/mol vs 57.1 ± 13.2 mmol/mol), p = 0.92] and FPG [162.7 ± 73.2 mg/dL vs 150.8 ± 48.2 mg/dL (or 9.0 ± 4.1 mmol/L vs 8.4 ± 2.7 mmol/L), p = 0.46].
PCA showed no discernible stratification issues for separation into cases and controls. Association tests were performed for a total of 170 biallelic SNPs; of these, 73 were polymorphic and 97 were monomorphic for the Hellenic population and, thus, excluded from further analysis. No variants in NLRP3, PYCARD, CASP1, and IL-18, reached statistical significance. Three variants in CARD8 and one in IL-1B were statistically significantly associated with DN, but after permutation the association remained significant only for CARD8 (Table 1). Specifically, rs11665831 [OR = 0.62 (95% CI, 0.43–0.9), pperm = 0.016], rs11083925 [OR = 0.65 (95% CI, 0.44–0.95), pperm = 0.021], and rs2043211 [OR = 0.66 (95% CI, 0.45–0.96), pperm = 0.026] seem to confer protection against DN (Table 1). The IL-1B rs4447608 was marginally associated with an increased risk for DN [OR = 1.436 (95% CI, 1.02–2), pperm = 0.057] (Table 1). When considering the genotype status for the aforementioned variants, DN was significantly associated under a dominant model of inheritance of the minor allele only for rs11665831 (TC+CC, pperm = 0.023) (Table 1). Associations of CARD8 variants with disease status were independent of sex or co-inheritance with the rs4447608 IL-1B variant.
Association of CARD8 and IL-1B variants with DN in T2DM cases of the Hellenic population on allelic and genotypic level, corrected for multiple testing.
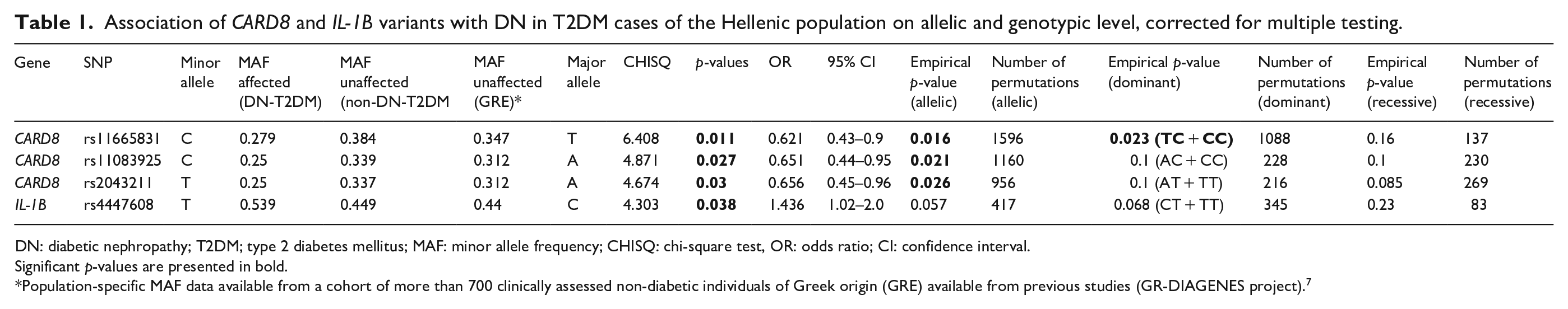
DN: diabetic nephropathy; T2DM; type 2 diabetes mellitus; MAF: minor allele frequency; CHISQ: chi-square test, OR: odds ratio; CI: confidence interval.
Significant p-values are presented in bold.
Population-specific MAF data available from a cohort of more than 700 clinically assessed non-diabetic individuals of Greek origin (GRE) available from previous studies (GR-DIAGENES project). 7
Discussion
Apart from lifestyle factors, poor glycemic control, arterial hypertension and hyperlipidemia, the manifestation of T2DM complications, such as DN, in some patients, might be partly attributed to the genetic background. A great number of genome-wide association (GWAS) and exome sequencing studies have tried to unravel the genetic components of DN predisposition, mostly in the context of T1DM; a number of those that did report on T2DM did not include Caucasian populations, presumably also because ESRD incidence and prevalence is higher in African-Americans and other ethnic groups compared to Caucasians.
In the present study we explore the association of common variants of the NLRP3 inflammasome-related genes with predisposition for the development of DN in the context of T2DM. Specifically, we highlight a likely protective role of three CARD8 variants against DN (Table 1) in T2DM carriers. Apart from being a direct NLRP3 inhibitor,2,3 CARD8 was previously shown to indirectly inhibit NF-κB activation 8 and to directly inhibit caspase-1 9 and, thus, impede pro-IL-1B expression and mature IL-1B production, respectively. Yet the mechanism of CARD8-mediated caspase-1 inhibition may not be active in non-inflammatory cells secreting IL-1B, which seems to be the case in smooth muscle cells, 13 highlighting the still controversial role of CARD8 across different cell types.
Even though CARD8 variant rs2043211 is exonic, its biological consequence remains dependent on the alternative transcript expressed in each tissue (https://gnomad.broadinstitute.org/variant/19-48737706-A-T?dataset=gnomad_r2_1). The minor allele T, which is actually the common allele for East Asians, was recently associated with increased CARD8 plasma levels and atherosclerosis in Chinese, but showed a protective role against diabetes. 14 Due to its coding position, rs2043211 has been extensively studied in association with predisposition to inflammatory conditions, triggered by increased caspase-1-mediated IL-1B secretion. rs2043211 was not associated with macro- or microvascular complications in Caucasian T2DM patients; however, the sample was small, consisting of only 34 cases with microvascular complications. 15 All other associated variants in our study are intronic, thus a direct biological role for these cannot be deduced. These common SNPs could be, however, in linkage with rare, potentially gain-of-function, CARD8 variants in non-DN cases.
NLRP3 inflammasome is activated in kidney cells, thus making it an attractive candidate for human kidney disease.4,6 Inflammasome activation was associated with the onset of DN in humans and diabetic mice, whereas inhibition of Nlrp3 ameliorated DN in rodents. 7 Overall, it has become clear that exciting data on NLRP3 inflammasome regulation and signalling point for more intensive research toward its targeting, which may provide novel therapeutic potential for DN treatment. 4
In summary, we showed that common variants in CARD8, coding for a negative regulator of NLRP3,2,3 but simultaneously acting as an NF-κB and caspase-1 inhibitor,8,9 show a protective role against the development of DN, independent of sex. CARD8 might be an important negative regulator of the pro-inflammatory cytokine response by acting at both the IL-1β generation and NF-κB activation levels. 9 Perhaps, alterations in the cross-talk between CARD8 and the NF-κB and NLRP3 pathways affect the pro-inflammatory local environment and influence the threshold of inflammasome activation in T2DM. Our preliminary observations will require validation in larger cohorts from several ethnic groups characterized by different allelic frequencies and prevalence of T2DM-related DN.
Footnotes
Acknowledgements
Patients are warmly thanked for consenting to participate in the study. We are also thankful to Dr Iordanis Karagiannidis for his work with patient samples.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by the European Social Fund and Greek funds through the Operational Programme “Education and Lifelong Learning” of the National Strategic Reference Framework–Research Funding Programme: THALES 2012–2015 (“GR-DIAGENES: The genetic architecture of type 2 diabetes mellitus in the Greek population”) and an ΙΚΥ fellowship of excellence for post-graduate studies in Greece (Siemens program).
