Abstract
Drugs have the potential of causing adverse reactions or side effects and prior knowledge of these reactions can help prevent hospitalizations and premature deaths. Public databases of common adverse drug reactions (ADRs) depend on individual reports from drug manufacturers and health professionals. However, this passive approach to ADR surveillance has been shown to suffer from severe under-reporting. Social media, such as online health forums where patients across the globe willingly share their drug intake experience, is a viable and rich source for detecting unreported ADRs. In this paper, we design an ADR Detection Framework (ADF) using Natural Language Processing techniques to identify ADRs in drug reviews mined from social media. We demonstrate the applicability of ADF in the domain of Diabetes by identifying ADRs associated with diabetes drugs using data extracted from three online patient-based health forums: askapatient.com, webmd.com, and iodine.com. Next, we analyze and visualize the ADRs identified and present valuable insights including prevalent and less prevalent ADRs, age and gender differences in ADRs detected, as well as the previously unknown ADRs detected by our framework. Our work could promote active (real-time) ADR surveillance and also advance pharmacovigilance research.
Keywords
Introduction
Adverse Drug Reactions (ADRs) are harmful events, side effects, or allergic reactions caused by medication. 1 Adverse drug reactions contribute to rising cases of morbidity and mortality globally. 2 For example, research has shown that ADRs cause approximately 197,000 deaths in Europe yearly 3 and responsible for 13% of hospital readmissions. 4 Through pharmacovigilance, ADRs can be detected, assessed, understood, and prevented. Over the years, public databases of common ADRs (e.g., FDA Adverse Event Reporting System, 5 SIDER–Side Effect Resource, 6 etc.) have been created to inform people about reported side-effects of drugs. However, these databases are usually based on individual reports of ADRs from sources such as drug manufacturers and health professionals, hence suffer from severe under-reporting.7,8
Social media data, such as online drug reviews from patients, have been considered as a viable source of ADRs to compensate for the shortcomings of passive surveillance which relies on reports from health professionals and manufacturers.9,10 Such patient reviews are available on online health forums (e.g., askapatient.com) where patients can interact and also share their experiences with respect to the side effects of their medications so that others can learn from them. 11 These online communities not only empower patients to actively participate in the pharmacovigilance process and become co-creators of the values embedded in it 12 but also serve as complementary sources of ADRs. 13 A recent study shows that ADR reports on social media share similarities with the traditional public databases of adverse events. 14 Also, clinicians’ failure to track the side effects self-reported by patients (on social media) has been linked to drug non-compliance and preventable adverse events. 15 Consequently, researchers are increasingly mining social media data to collect evidence for adverse drug events, drug compliance, and drug effectiveness to promote drug safety and treatment management in recent years.16,17 Hence, extracting and analyzing patients’ drug reviews posted on social media platforms will promote active ADR surveillance and uncover previously unknown side effects that should be considered by health professionals when prescribing medications to patients. In addition, drug regulators will be able to engage pharmaceutical industries on ways to address newly discovered ADRs in a timely and effective manner.
Therefore, in this paper, we present the ADR Detection Framework (ADF) for extracting ADRs from online patient-authored drug reviews on social media. The framework applies natural language processing (NLP) techniques 18 including named entity recognition (NER) to identify ADRs in the reviews. NLP is a well-established computational method for parsing, segmenting, extracting, or analyzing text data, 19 and has been widely used to process health-related texts in various domains including disease management,20,21 mental health,22,23 and pharmacovigilance. 24 In contrast to supervised machine learning-based NLP approaches10,25, ADF does not require annotated data to acquire knowledge and could identify ADRs in drug reviews across multiple health domains (including Diabetes which is our case study). In other words, our framework addresses two main shortcomings evident in supervised machine learning (ML) approaches: (i) dependency on annotated data which could be very expensive and hard to obtain, and (ii) ADRs identified are limited to those learned from the data on which the ML models have been trained, hence cannot extend to diverse health domains.
The ADF is implemented as a Java application programming interface (API) that receives online drug reviews obtained from one or more sources as input and automatically extracts ADRs mentioned in the data. Behind the scene, the ADF’s interpreter component utilizes the SPECIALIST lexicon 26 which consists of over 500,000 Biomedical and English terms, 27 as well as the NLP toolkit – MetaMap. 28 To categorize biomedical terms or concepts detected in patient reviews, ADF uses the Unified Medical Language System (UMLS) Semantic Network 26 which consists of 15 semantic groups and 127 semantic types. Based on existing research which states that signs/symptoms and disorders/diseases are potential side effects, 29 the framework considers concepts from semantic types under the “Disorders” group as ADRs. Next, to investigate the framework’s applicability, we mined diabetes-related drug reviews from three large and popular online patient-based health forums: AskaPatient (askapatient.com), WebMD (webmd.com), and Iodine (iodine.com), and then used the framework to extract ADRs from the reviews. We focus on Diabetes as our domain of interest for demonstrating ADF due to its increased global prevalence: 387 million diabetic patients, rapid increase among children, adolescents, and young adults, as well as five-fold increase in mortality rate. 30 In addition, a recent study found that 46% of diabetic patients reported adverse drug events, 31 hence detecting previously unknown or unreported ADRs of diabetes drugs will be an important contribution to knowledge. Next, we performed comparative analysis of the ADRs identified from the three data sources. We also compared the ADRs across gender and age groups to uncover interesting insights. Finally, we benchmarked our framework (ADF) against the SIDER database 6 to uncover previously unreported ADRs detected by ADF.
Our work offers 3 main contributions. First, we designed and implemented ADF which is capable of detecting ADRs in online drug reviews from patients across multiple health domains (e.g., Diabetes) using NLP techniques. Second, we demonstrated ADF using an example in which ADRs within diabetes-related drug reviews were automatically identified, and then analyzed and visualized to uncover valuable insights such as prevalent and less prevalent ADRs, age and gender differences in ADRs detected, as well as previously unknown ADRs identified by our framework after benchmarking against SIDER. Our findings could guide healthcare professionals, drug regulators, and manufacturers in making data-driven decisions, and also foster further research in pharmacovigilance. Finally, besides detecting ADRs, the framework can be extended to identify other interesting concepts in health-related conversations or documents from any source by modifying the semantic types. Hence, the framework could be useful for building interactive and intelligent health systems including early warning systems, as well as medication management and medical diagnosis systems.
Related work
Previous research in ADR surveillance using social media adopted various approaches (e.g., lexicon-based entity recognition) in detecting ADRs mentioned in user posts or reviews. Leaman et al., 32 for instance, built a custom lexicon using four resources: COSTART (one of the UMLS biomedical vocabularies), SIDER 6 (a database containing side effects), MedEffect (the Canada Drug Adverse Reaction database), and a set of colloquial phrases collated manually. Having applied the NLP technique to preprocess the social media comments, extracted tokens that match some words in the lexicon were selected as ADRs. The volume of ADRs that can be discovered is limited by the number of lexical terms available in the lexicon. Thus, instead of selecting only one of the UMLS Metathesaurus vocabularies, our work leveraged the nearly 200 available vocabularies 33 to ensure wider coverage of ADRs. In contrast with Leaman et al.’s approach, Liu et al. 34 built their lexicons using ADRs extracted from user reviews for statin drugs through heuristics and NLP. Their objective is to detect side effects that are strongly associated with statin drugs. They used the log-likelihood ratio test to determine the likelihood that a side effect appears more frequently in statin reviews than non-statin reviews. A side effect that appears more frequently in statin reviews will have a p-value < .05, thereby making it strongly associated with statin drugs. This approach is rigid since lexicons can only be generated when reviews are available. Similarly, Li et al.’s work constructed a lexicon that contains ADRs detected from drug reviews 35 . They selected tokens that meet a certain threshold based on weights assigned through their information gain model, and then add them to the lexicon as potential ADRs. This lexicon is then utilized in identifying ADRs present in user reviews for Glucophage (i.e., Metformin). This approach is also not flexible since lexicons can only be generated when user reviews are provided.
Other approaches applied the lexicon-based approach in conjunction with statistical learning, 36 machine learning, 37 rule-based 29 or association rule mining 38 techniques to determine drug-ADR relationships (or cause-effect pairs) within health-related texts. These approaches are applicable to cases where the user is not reviewing a preselected drug, but rather expressing his/her experience with any medication(s) on social media.
Methods
In this section, we describe the ADF architecture (see Figure 1) which is made up of the data ingestion, data preprocessing, semantic types definition, and the ADR detection components. We also demonstrate applicability of the framework in the area of diabetes pharmacovigilance. ADF utilizes natural language processing (NLP) techniques, as described in subsequent subsections. The ADR detection framework (ADF).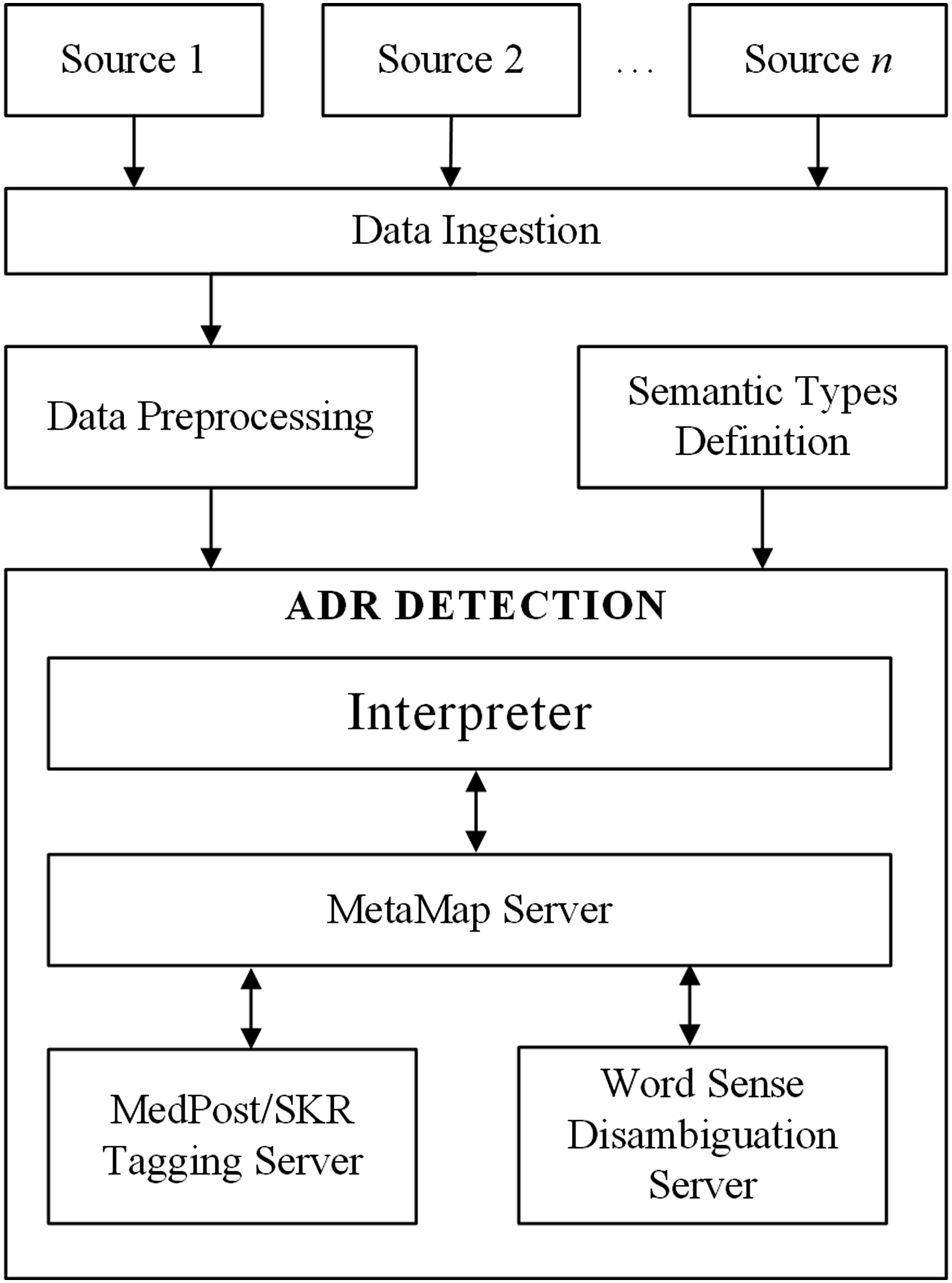
Data ingestion
Total drug reviews from three social networks.

Some of the reviews posted by drug users (or patients).

aOnline reviews are included verbatim throughout the paper, including spelling and grammatical mistakes. Reviews are available for public use but are anonymized to protect user privacy.
Data preprocessing
Prior to ADR detection, we cleaned the extracted data using Python by following the step-by-step process outlined below:
(1) We removed special characters, such as newline, tabs, carriage returns, double quotes, underscore, and extra spaces.
(2) We expanded contractions (for instance, don’t becomes do not, could’ve becomes could have, etc.).
(3) We removed duplicate entries.
In addition to the above, we retained some punctuation marks, such as comma (,) and period (.) since they are important for sentence segmentation, tokenization, and semantic analysis. Stop words (such as and, have, for, is, that, what, with, etc.) were also retained to preserve the semantics of phrases and sentences.
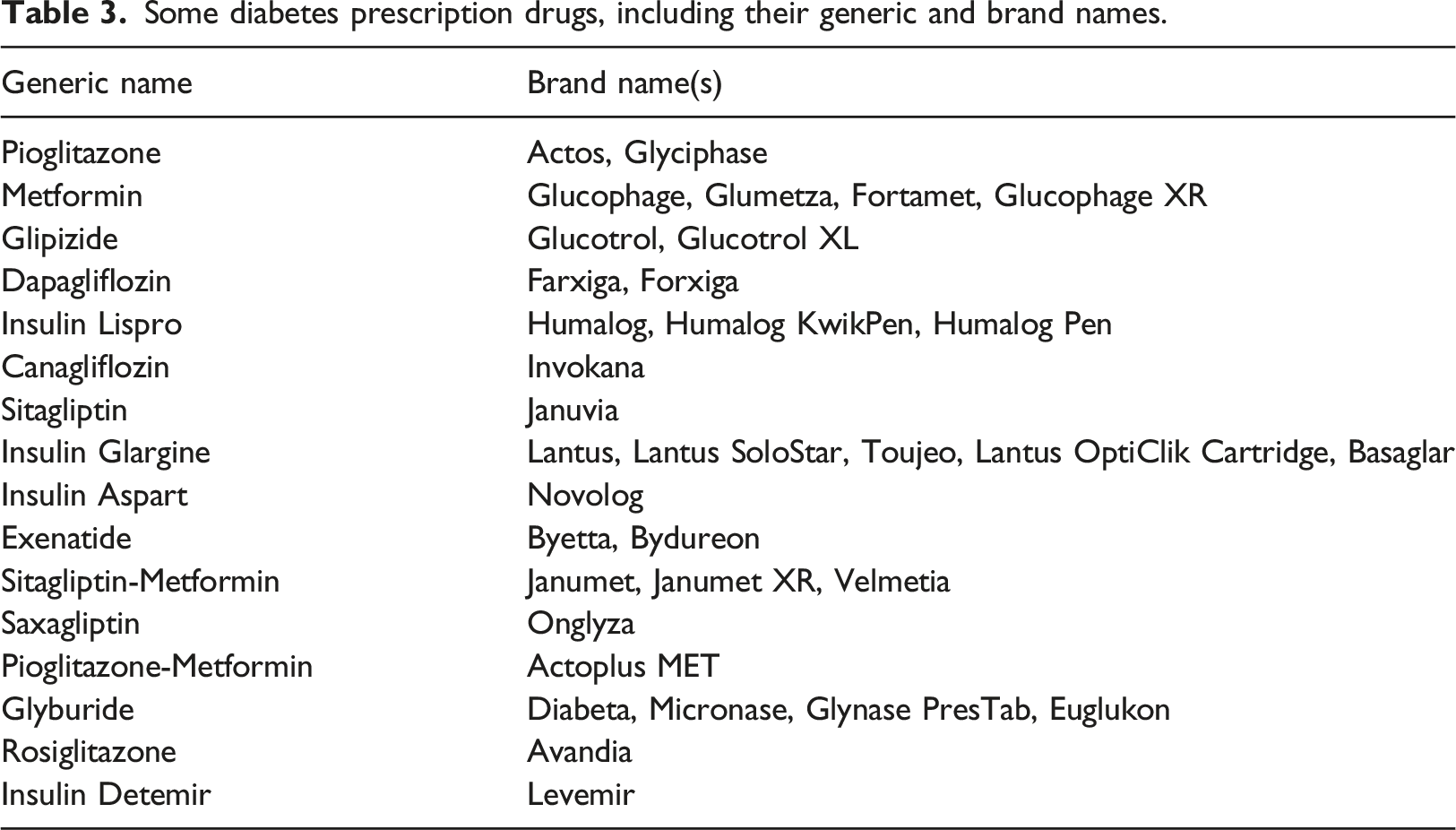
Some diabetes prescription drugs, including their generic and brand names.
Semantic types definition
The Semantic Network, accessible through MetaMap, provides a consistent categorization of all biomedical and health-related concepts in the UMLS Metathesaurus.26,39
There are 15 semantic groups and 127 semantic types in the UMLS Semantic Network. The semantic groups include Activities and Behaviors, Anatomy, Chemicals and Drugs, Concepts and Ideas, Devices, Disorders, Genes and Molecular Sequences, Geographic areas, Living beings, Objects, Occupations, Organizations, Phenomena, Physiology, and Procedures, with each group linked to different semantic types. We focused on biomedical concepts from semantic types under the
Detecting adverse drug reactions
To detect ADRs in each drug review, we developed an Interpreter using Java programming language to communicate with MetaMap by providing drug reviews and semantic types as parameters in the required format and then interpreting (or parsing) the output from MetaMap. The parsing activity involves retrieving the ADRs (and other metadata) and associating them with the drugs, as well as storing, harmonizing, and filtering the ADRs.
Figure 2, which is a granular version of the ADR detection component depicted in Figure 1, shows the various steps involved in extracting the ADRs. The Part-of-Speech Tagging and Word Sense Disambiguation (WSD) tasks are handled by MedPost/SKR Tagging server
40
and WSD server
41
respectively, while other tasks are handled by the MetaMap server. All three servers were set up locally to listen for incoming requests on different ports. Adverse drug reactions (ADR) detection stages.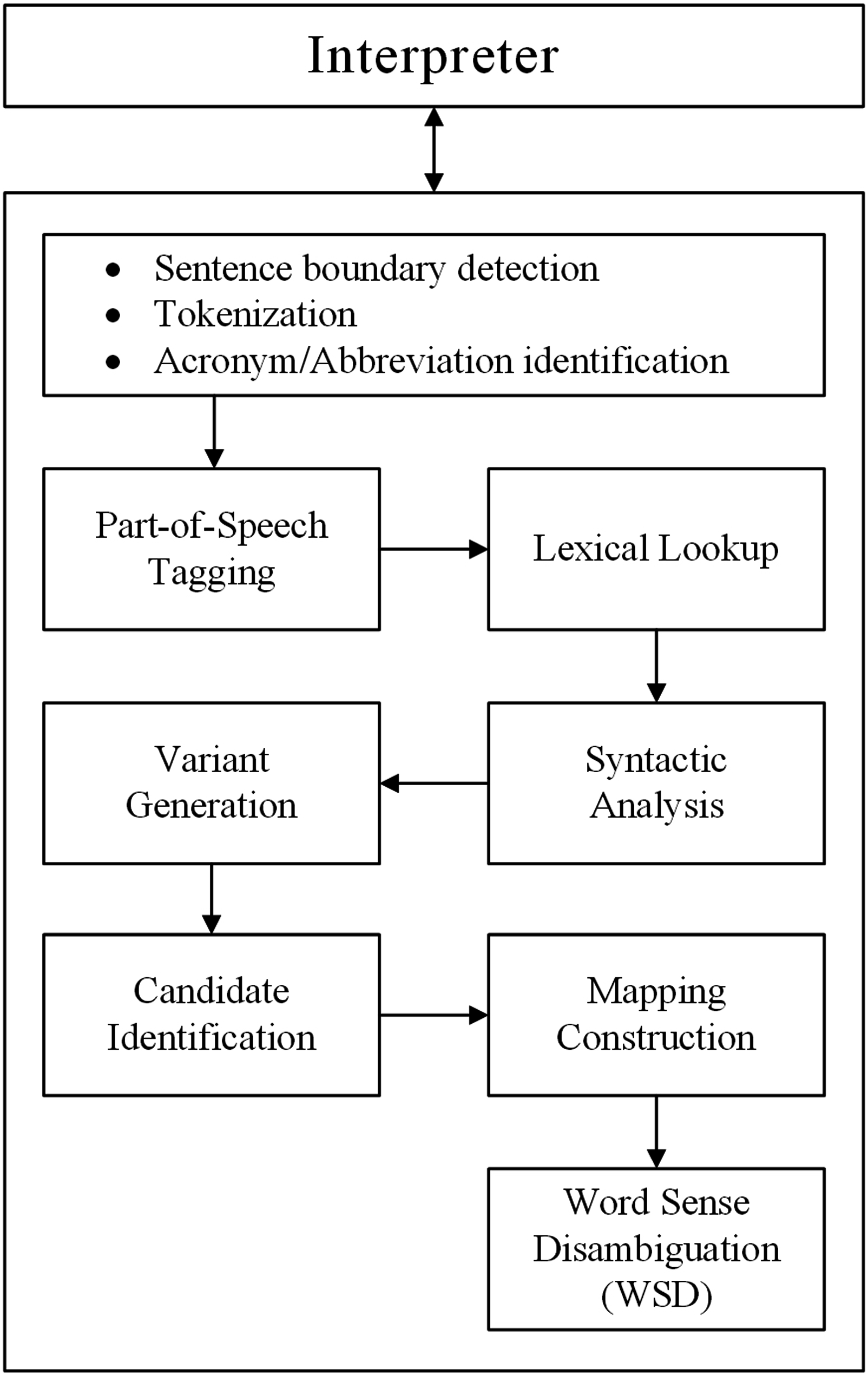
Sentence boundary detection, Tokenization, and Acronym/Abbreviation (AA) identification
Upon receiving an input text, MetaMap breaks the text into sentences using end-of-sentence marker (e.g., period) and it is robust enough to handle ill-formed texts where the sentence boundary is difficult to detect. Afterwards, each sentence is tokenized into meaningful phrases. For instance, the following review is from a patient who took Exenatide: “My first dose and I feel bad. It was almost instant tiredness and sick feeling. I wish I had seen this site before I took the dose. I may have just changed my mind” – [R19] (Drug brand: Byetta)
Sentences and corresponding tokens for the sample drug review.

Part-of-Speech (POS) tagging, lexical lookup, and syntactic analysis
For each word, the part of speech is determined by the MedPost/SKR Tagger and the SPECIALIST parser (accessible through the MetaMap server) searches the SPECIALIST lexicon 42 for a match. The SPECIALIST lexicon keeps the syntactic, morphological, and graphemic information for biomedical terms and English words. The SPECIALIST parser performs further syntactic processing that determines the syntactic structure supporting the semantic interpretation of each phrase (such as finding the head noun and post-modifying prepositional phrase). 39 For instance, in the phrase “almost instant tiredness” (see Table 4), the head noun is tiredness.
Variant generation, candidate identification, mapping construction, and Word Sense Disambiguation
MetaMap looks up the variants of each phrase word through the Lexical Variant Generator (lvg) program, based on relationships already defined in the UMLS Metathesaurus. 26
The intermediate results consisting of Metathesaurus strings or candidates are combined (or mapped) and evaluated to produce a result that best matches the input phrase. MetaMap, through the WSD server, favours mappings involving concepts that are semantically consistent with the surrounding text. 28 In case there are no candidates, the concept that best matches the phrase words is returned. For instance, no candidate was generated for the phrase “sick feeling” (see Table 4) but sick and feeling both matched the Metathesaurus concept “Feeling Sick”. Similarly, the phrase “almost instant tiredness” produces no candidates but tiredness matched the Metathesaurus concept “Fatigue”. Hence, the ADRs detected from the input text (see [R19]) are Feeling Sick and Fatigue.
Furthermore, MetaMap can identify negations (or negated concepts) in phrases using its custom NegEx algorithm. 28 As a result, a negated ADR is considered invalid since the reviewer did not experience it.
Finally, since the Metathesaurus also tracks similar concepts, MetaMap exposes two names as part of its output: concept name and preferred concept name. The preferred concept name is the harmonized or generic name shared by similar concepts. For instance, “fatigue” is the preferred concept name while “tiredness” is the concept name. Another benefit of the preferred concept name is that it shows the expanded name in case the concept name is an abbreviation/acronym (AA). For instance, if the concept name is “UTI”, the preferred concept name shows “Urinary Tract Infection”.
Harmonizing ADRs
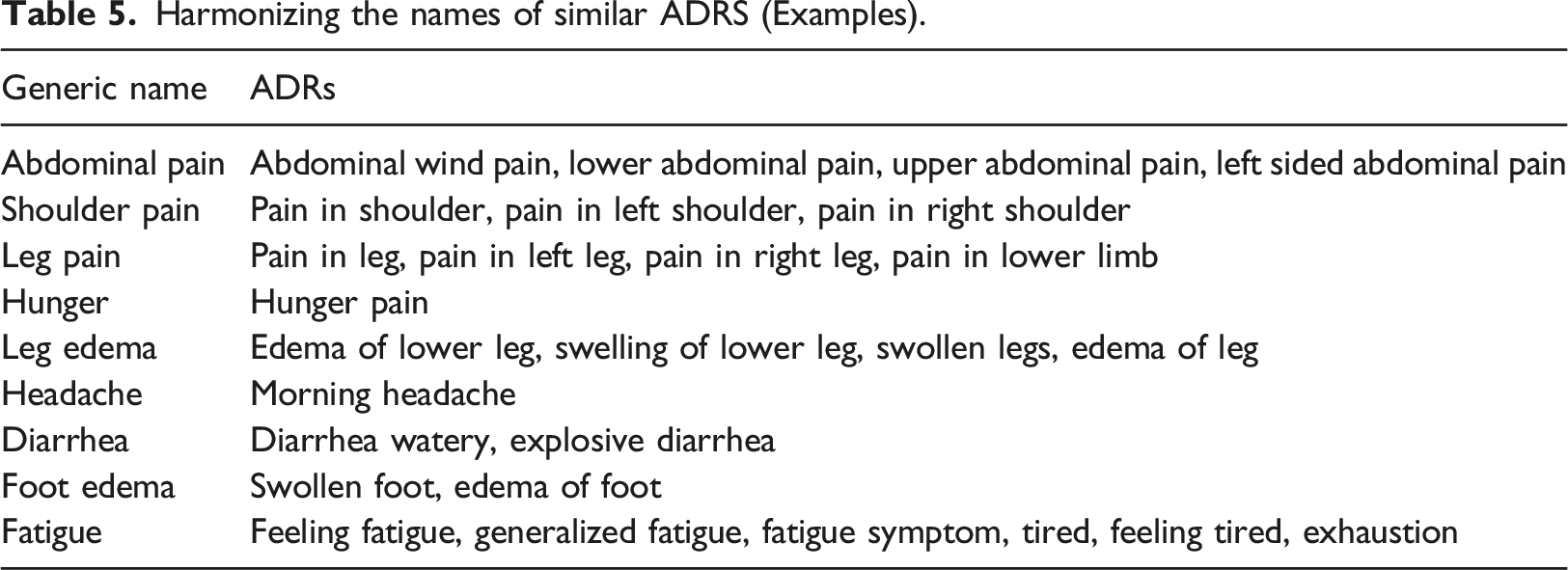
Harmonizing the names of similar ADRS (Examples).
Filtering ADRs
To exclude terms wrongly selected as ADRs, certain filters were applied on the ADR data. For instance, the word “bed” in the input text was wrongly matched to the ADR concept “BED” which is the acronym for “Binge eating disorder”, hence we added “Binge eating disorder” to the list of ADRs to be excluded from the ADR data. The filtering task is also handled automatically by the Interpreter.
Results
From the 6797 reviews extracted, a total of 2572 ADRs associated with 48 drugs were detected. The “2572” total is inclusive of ADRs that are associated with more than one drug. There are 684 unique ADRs in the data. We discuss further details in subsequent subsections.
High-level comparison of drugs and ADRs
The ScatterText
43
plot in Figure 3 shows the comparison of diabetes drugs across the three sources based on prevalence or frequency (i.e., number of reviews associated with each drug). We compared the corpus of WebMD (Y-axis), which is the largest, with the corpus of both AskAPatient and Iodine (X-axis). Drugs closer to the top-right corner (and closer to the Frequent label on both X and Y axes) got more reviews than those closer to the bottom-left. Drugs comparison based on frequency (i.e., number of reviews per drug).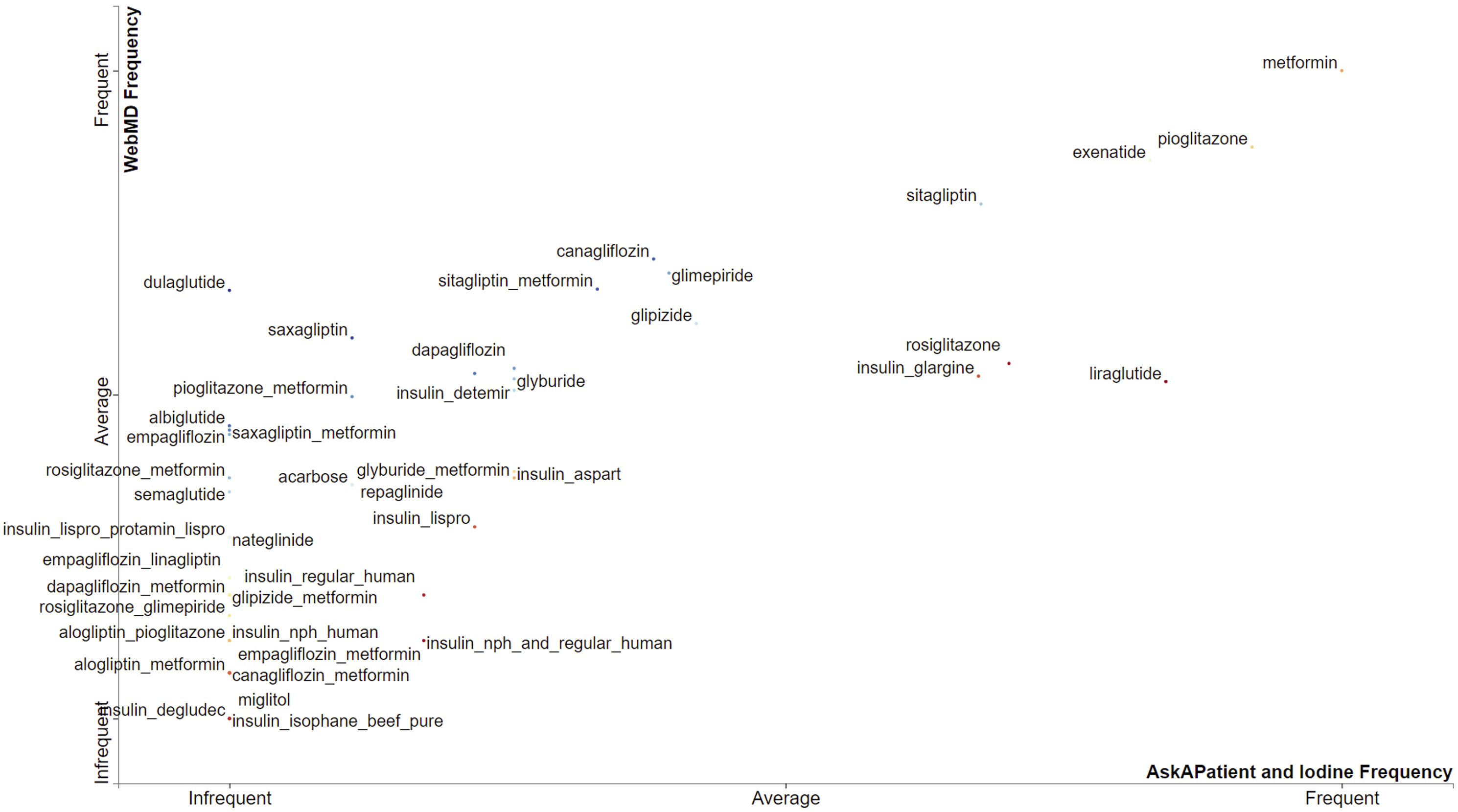
As shown in Figure 3, Metformin is “frequent” on both axes which means it got the highest number of reviews across all sources. Overall, Metformin is the most prevalent drug, followed by Pioglitazone, Exenatide, and Sitagliptin. Some drugs (such as Canagliflozin, Glimepiride, Dulaglutide, Glipizide, etc.) are prevalent on WebMD but below average on both AskAPatient and Iodine. Majority of the drugs got a small number of reviews across the three sources, hence are below the average mark on both X and Y axes (see Figure 3).
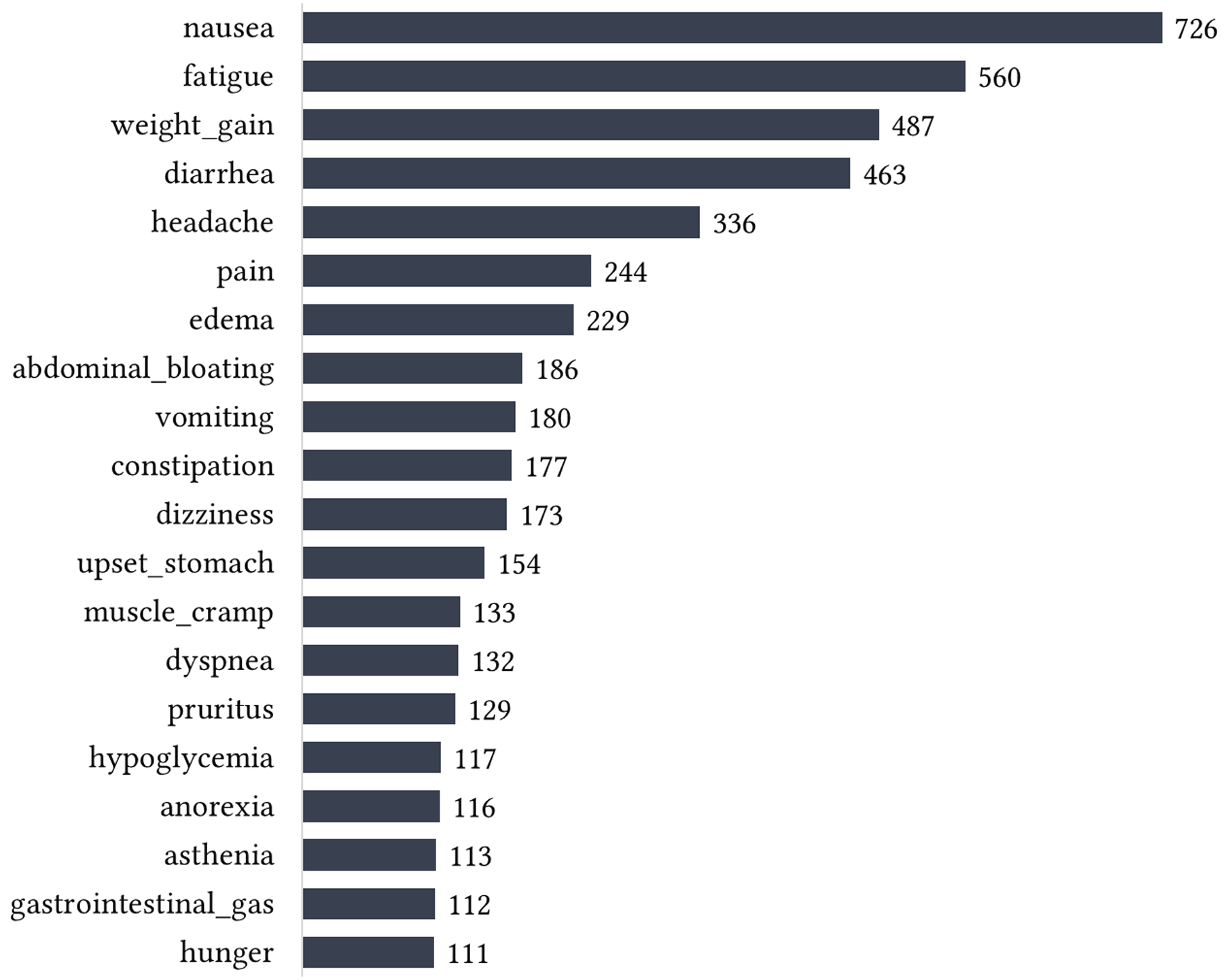
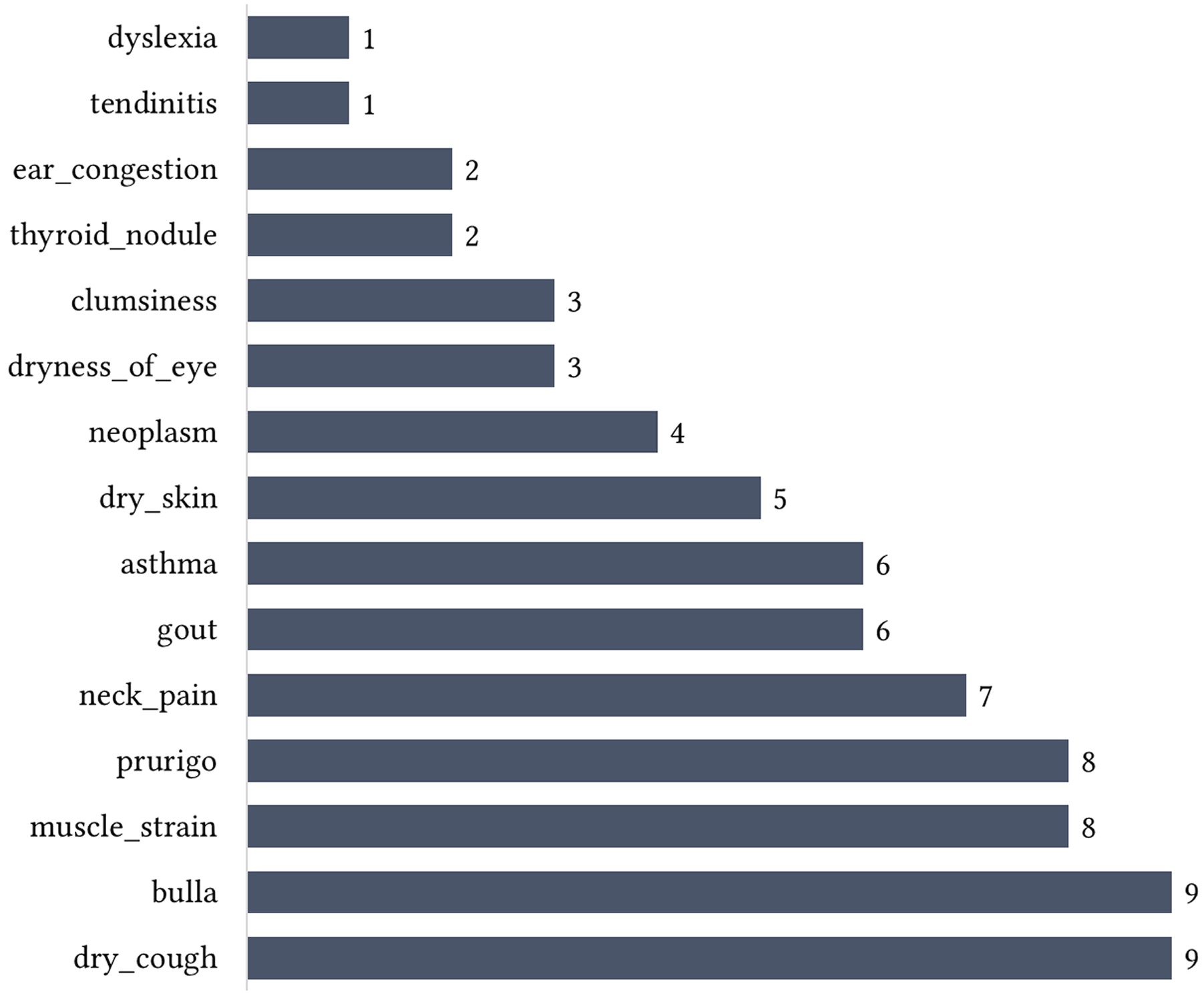
Similarly, Figure 4 shows the distribution of ADRs based on their frequency (i.e., number of reviews mentioning each ADR) across the sources, while Figure 5 shows top 20 ADRs in terms of frequency. Overall, nausea has the highest frequency (n = 726), hence the most prevalent ADR. Other ADRs with high frequency include fatigue (n = 560), weight gain (n = 487), diarrhea (n = 463), and headache (n = 336). Others are pain (n = 244), edema (n = 229), abdominal bloating (n = 186), vomiting (n = 180), constipation (n = 177), dizziness (n = 173), upset stomach (n = 154), muscle cramp (n = 133), dyspnea (n = 132), pruritus (n = 129), and so on. Majority of the ADRs (80.3%) have frequency of less than 10. Some of these less frequently highlighted ADRs are shown in Figure 6. Adverse drug reactions comparison based on frequency (i.e., number of reviews mentioning each ADR). Top 20 ADRs by frequency. Examples of less frequent ADRs.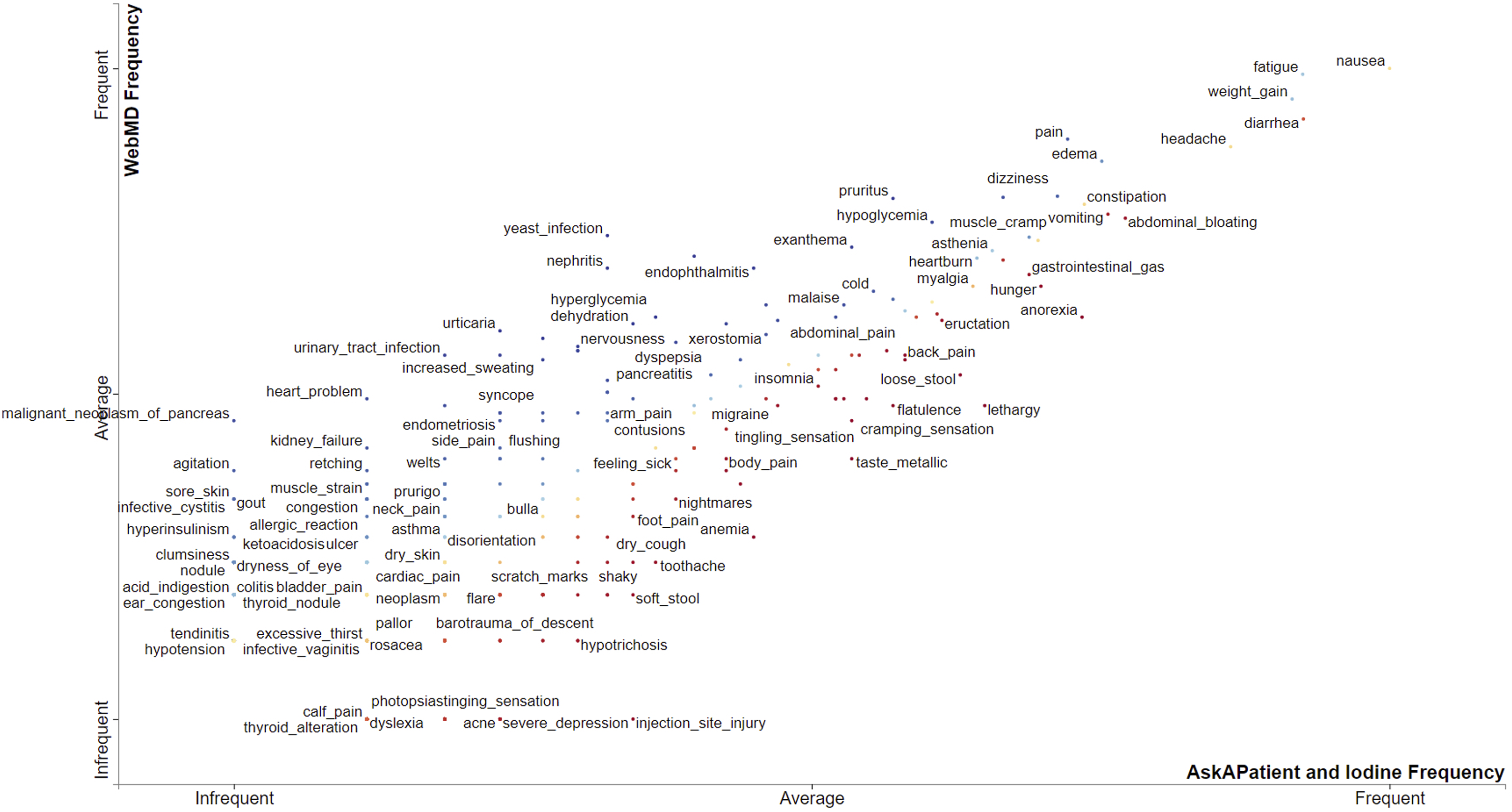
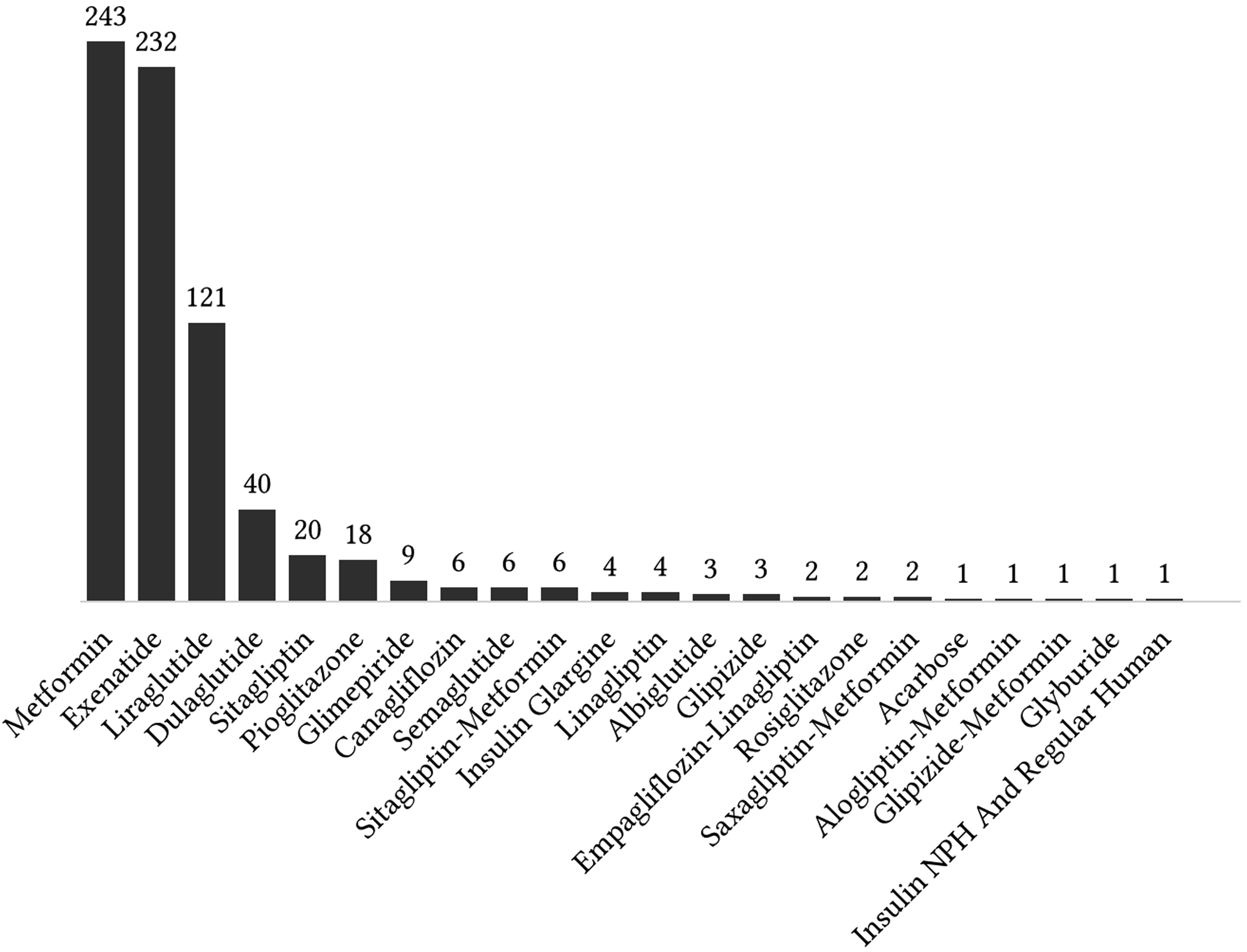
Drugs and the associated ADRs
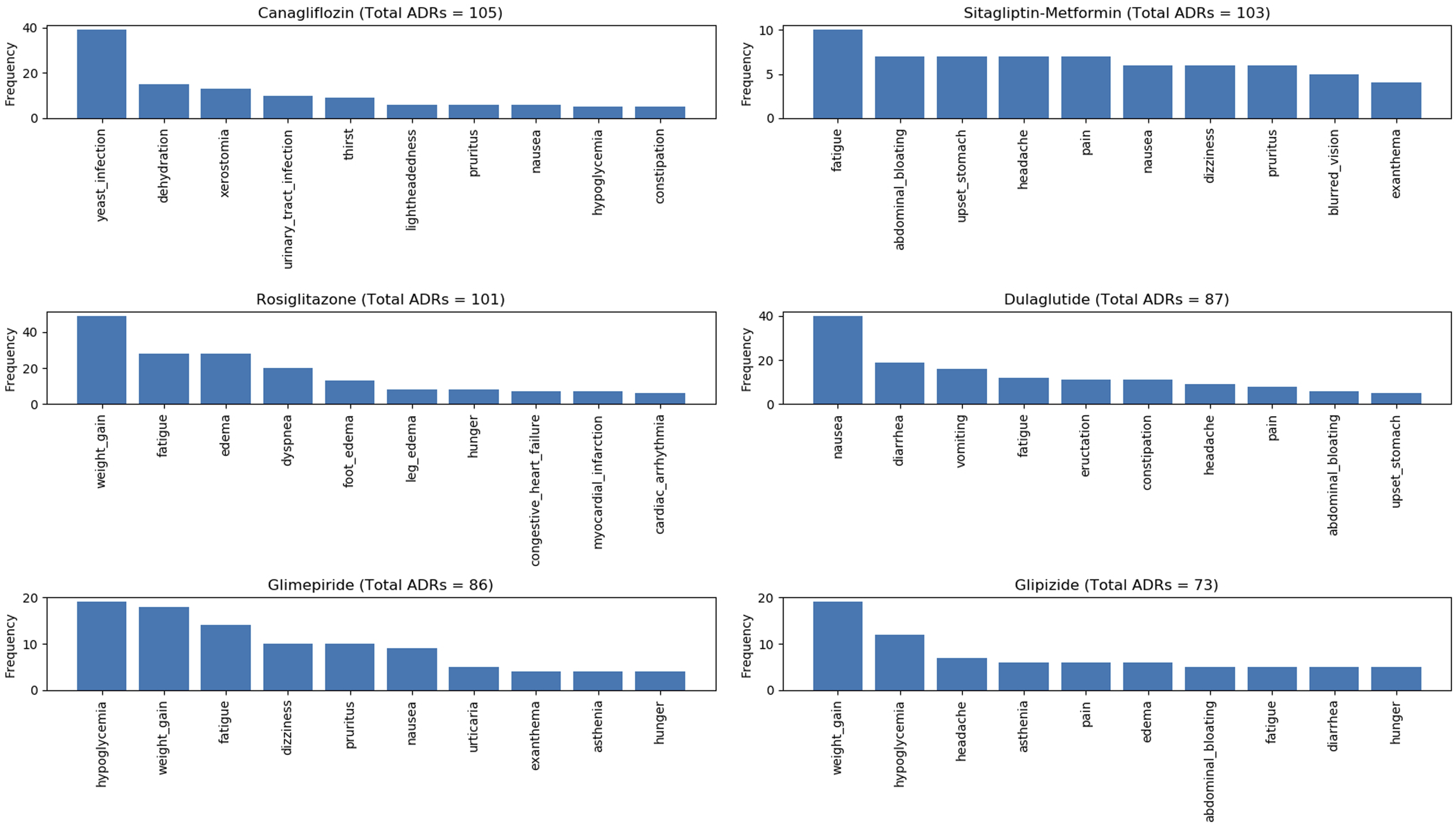
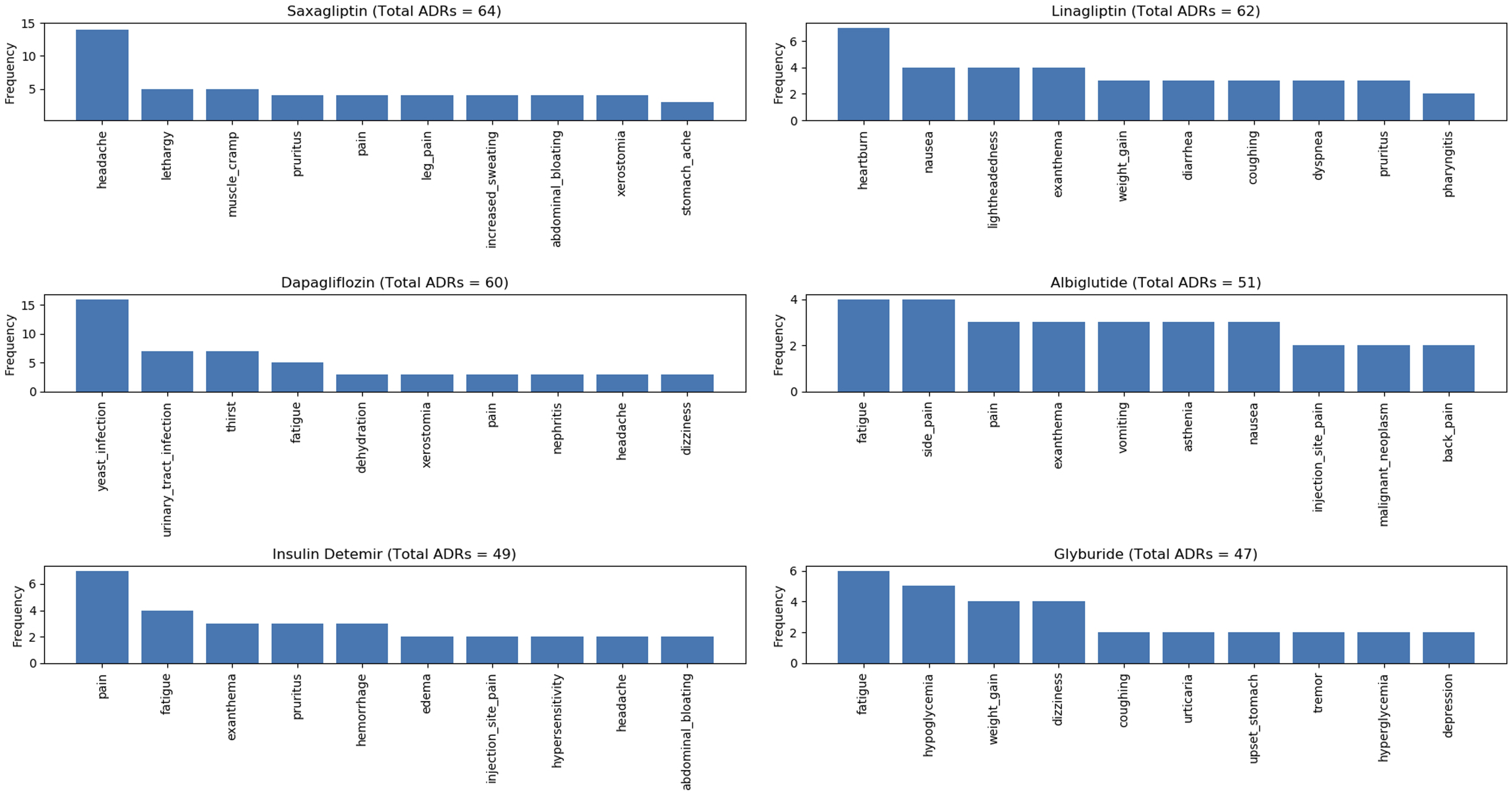
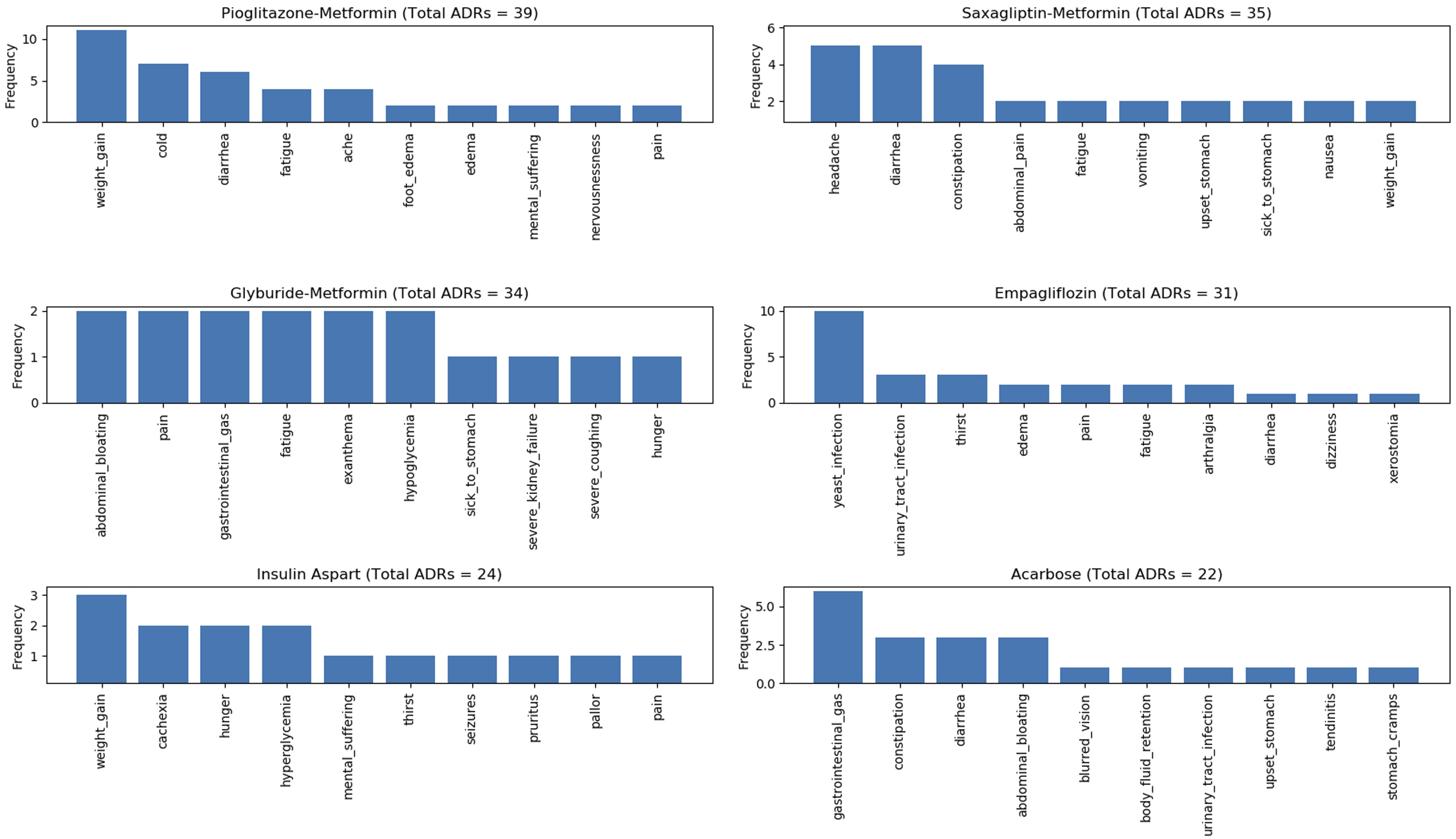
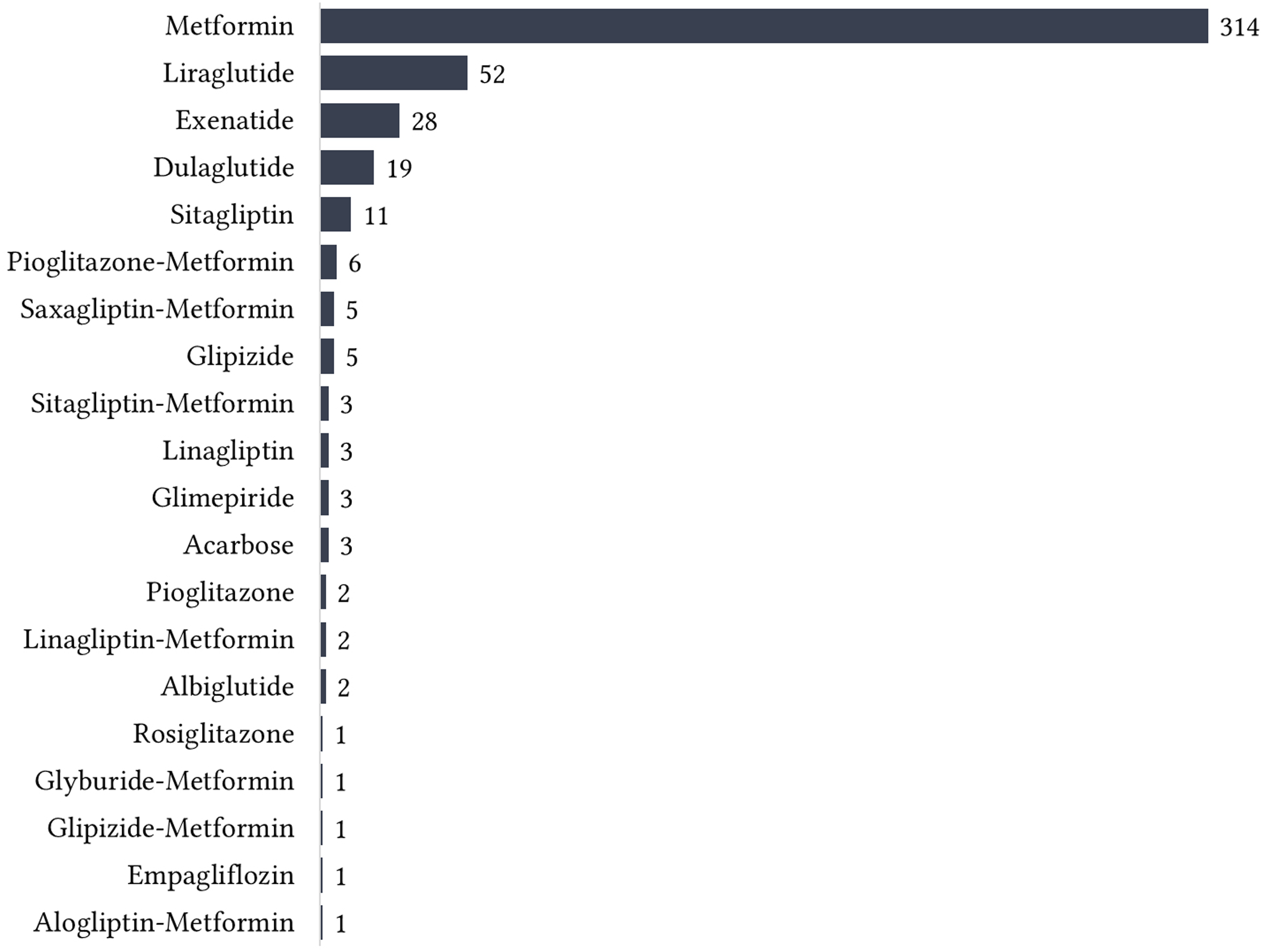
Figure 7, Figure 8, Figure 9, and Figure 10 show the top 10 ADRs associated with some of the diabetes drugs. The total number of ADRs associated with each drug is indicated in the chart title. In Figure 11, the diabetes drugs associated with nausea are shown and ranked by frequency. Metformin (n = 243) is a major cause of nausea, closely followed by Exenatide (n = 232). Moreover, Metformin (n = 314) is a major cause of diarrhea, followed by Liraglutide (n = 52), Exenatide (n = 28), etc. as shown in Figure 12. Top 10 ADRs associated with Metformin, Sitagliptin, Liraglutide, Pioglitazone, Exenatide, and Insulin Glargine. Top 10 ADRs associated with Canagliflozin, Rosiglitazone, Glimepiride, Sitagliptin-Metformin, Dulaglutide, and Glipizide. Top 10 ADRs associated with Saxagliptin, Dapagliflozin, Insulin Detemir, Linagliptin, Albiglutide, and Glyburide. Top 10 ADRs associated with Pioglitazone-Metformin, Glyburide-Metformin, Insulin Aspart, Saxagliptin-Metformin, Empagliflozin, and Acarbose. Drugs associated with Nausea. Drugs associated with Diarrhea.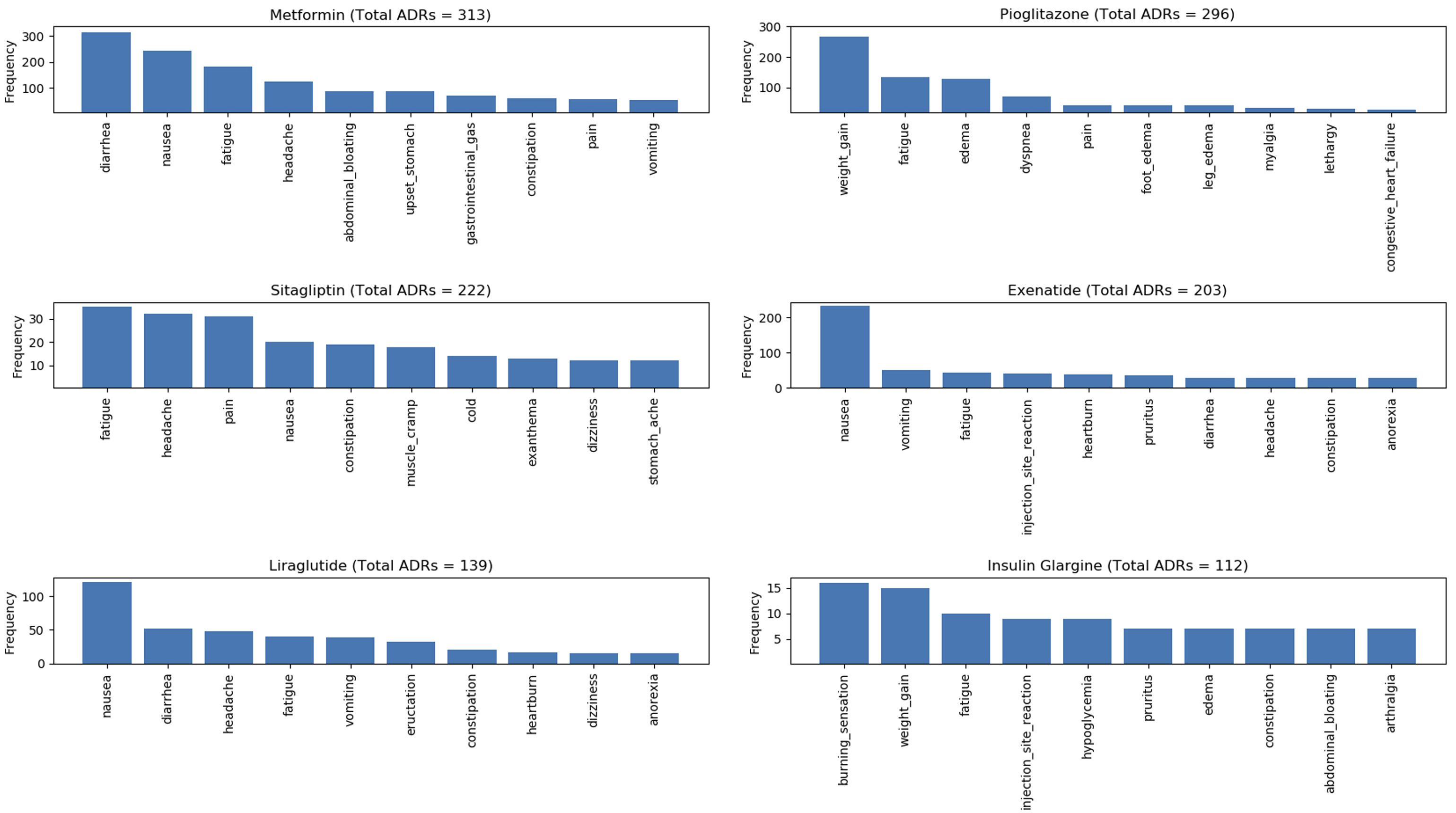
Comparing ADRs by gender
From the top-right corner of the plot in Figure 13, it is obvious that nausea, fatigue, weight gain, diarrhea, headache, pain, edema, dizziness, constipation, upset stomach, abdominal bloating, anorexia, vomiting, muscle cramp, hypoglycemia and other ADRs that surpass the average mark on both axes occur in both male and female more frequently (above average). Comparing ADRs by gender.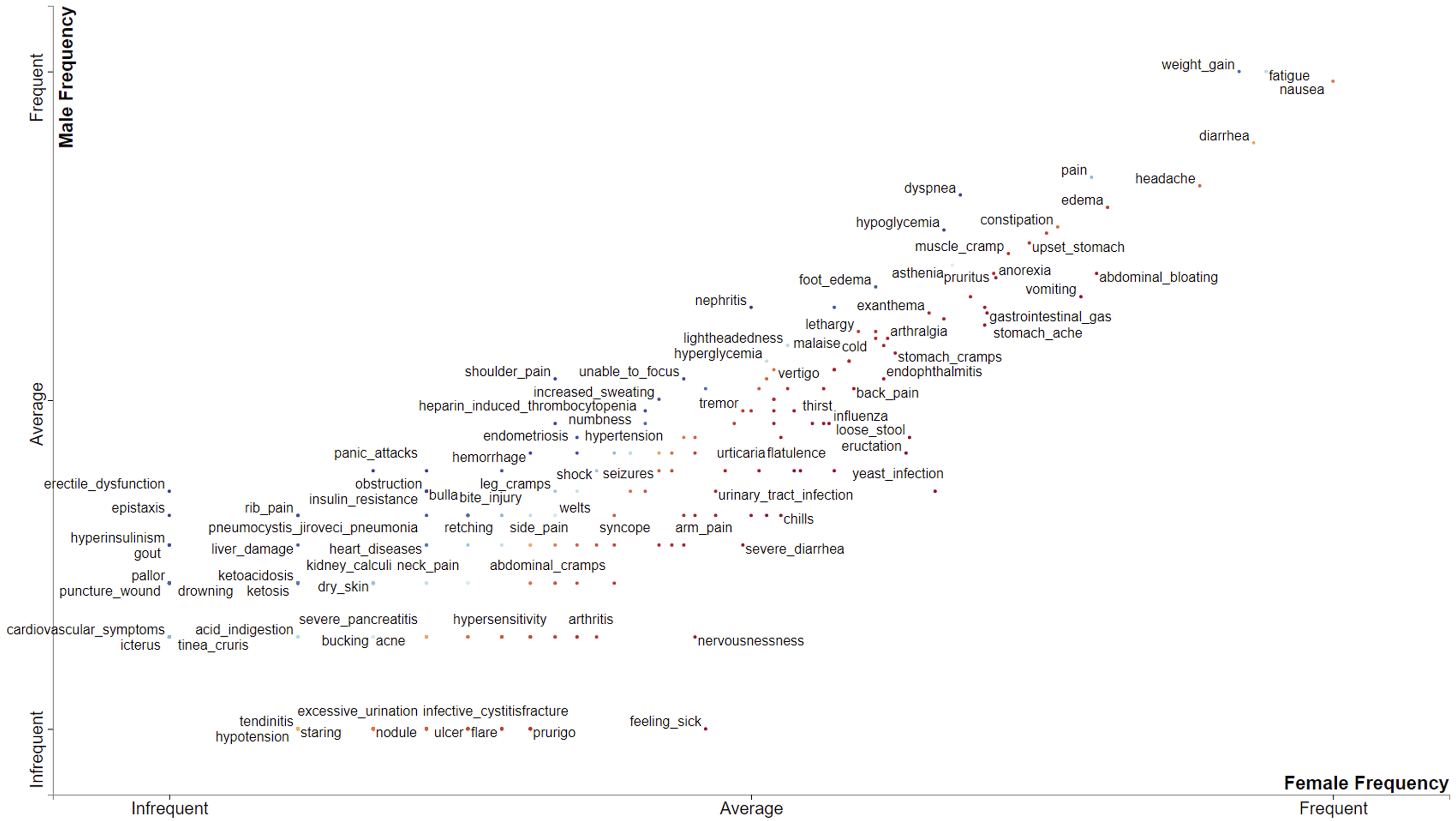
ADRs in females
Adverse drug reactions that exceed the average mark on the Female Frequency axis, but not surpassing that of the Male Frequency axis occur more frequently in female patients than male patients. Such ADRs include yeast infection, eructation, loose stool, chills, etc.
ADRs in males
On the other hand, ADRs that exceed the average mark on the Male Frequency axis, but not surpassing that of the Female Frequency axis occur more frequently in Male patients than Female patients. They include shoulder pain, unable to focus, increased sweating, etc.
Comparing ADRs by age
Figure 14 shows various word clouds (generated using Python) illustrating the prevalence of ADRs by age group. Nausea is prevalent among 19–64 years age group, though more prevalent among adults between 25 and 44 years old. On the other hand, diarrhea is more prevalent among 13–34 years age group than 35–44 years age group. Fatigue, however, is prevalent among patients above 74 years old and those between 35 and 44 years old. Furthermore, weight gain is found to be prevalent among adults above 44 years old. Finally, headache is prevalent among adolescents between 13 and 18 years old, followed by young adults between 19 and 24 years old. Comparing ADRs by Age group.
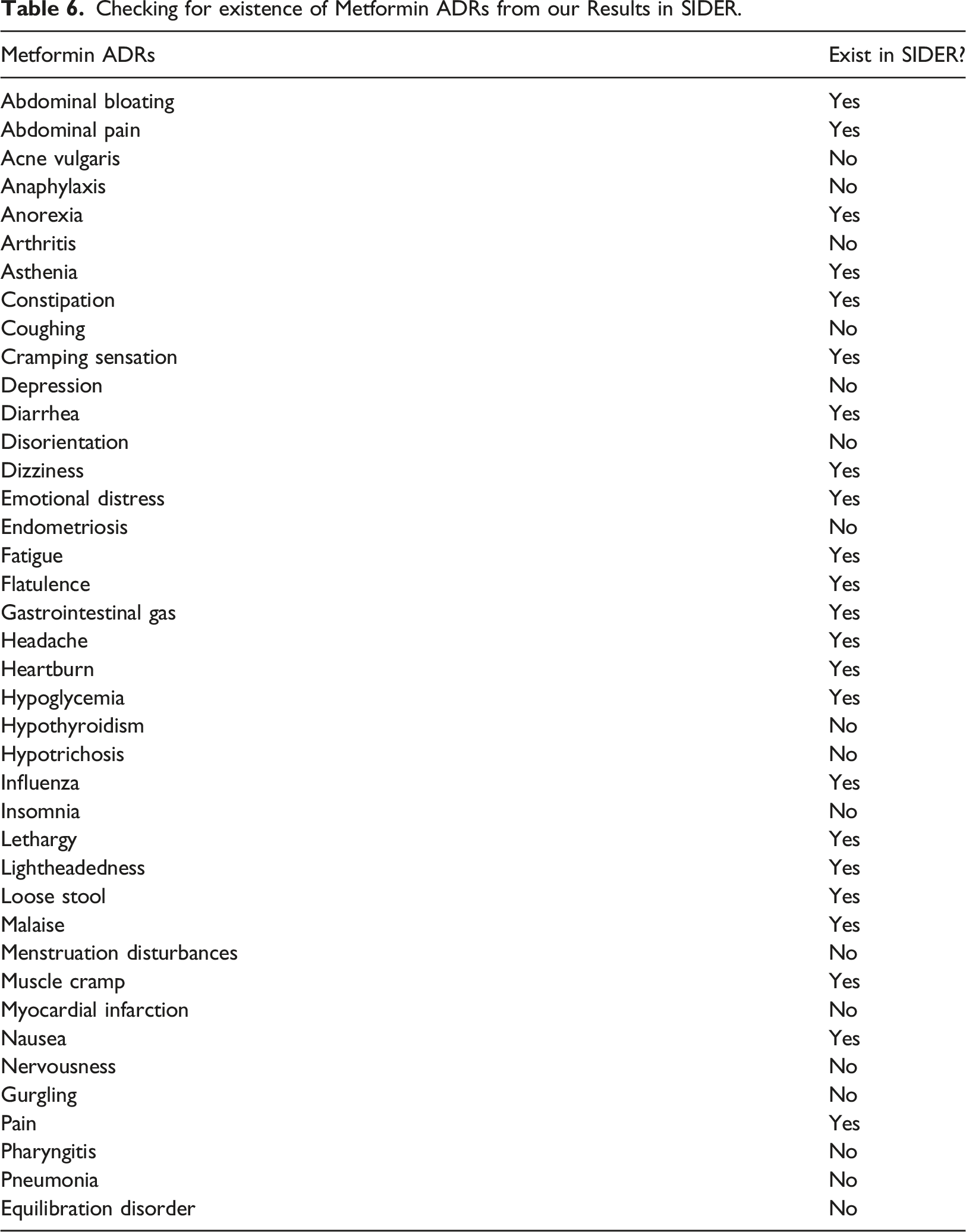
Comparing ADRs from our results with those in SIDER
Checking for existence of Metformin ADRs from our Results in SIDER.

Discussion
Social media has been shown to offer potential opportunities for real-time surveillance of ADRs, extensive ADR reporting, and expedited signal detection 44 enhanced partly by the coverage, diversity, volume, and timeliness of available data.16,45 In this work, we leveraged social media data collected from three sources to develop and evaluate our ADR Detection Framework (ADF). Despite the unstructured, non-standard, or informal reporting of adverse events on social media,16,17,44 the ADF applied NLP techniques to preprocess, harmonize, and analyze the data to uncover meaningful relations between drugs and ADRs. Furthermore, while majority of similar works limit the scope of their research to specific medications or sources,35,46 the ADF generalizes the sources and medications. As a result, the framework can identify other interesting concepts from health-related conversations or documents using relevant semantic types. Thus, the framework can be used in developing interactive and intelligent health systems that support patients by providing more information about their medications, assessing current symptoms, helping with first-level diagnosis, answering health-related questions, mediating conversations with healthcare professionals, and so on. It can also be used to develop early warning systems for pharmacovigilance targeting diverse health conditions 44 (such as Diabetes which is our use case in this work). This kind of automation is made possible by the framework’s integration with UMLS and MetaMap, thereby unlocking unrestricted access to a robust and rich backend that houses a lot of biomedical concepts along with relationships and definitions, coupled with advanced and extensible NLP capabilities.
Our case study on diabetes medications demonstrated the framework’s contribution to pharmacovigilance. The resulting data comprises both known ADRs captured in public databases and unknown ADRs that are yet to be captured. We validated our findings using the popular and publicly accessible side effect database, called SIDER, 6 using Metformin drug as basis for comparison. Metformin’s most common side effect in SIDER is diarrhea which also agrees with our result (see Figure 7). Majority of other Metformin’s ADRs such as nausea, abdominal bloating, vomiting, anorexia, headache, constipation, flatulence, asthenia, etc. are already captured in SIDER, but some ADRs were still missing in SIDER such as retrograde ejaculation, insomnia, pneumonia, gurgling, equilibration disorder, myocardial infarction, etc.
The implication of these findings is that patients and healthcare professionals relying on databases such as SIDER for knowledge of new ADRs will need additional resources in order to stay up to date. We have shown, to a reasonable degree, that publicly available data can be a complementary source of information about ADRs. The ADF can be employed to analyze and automatically share ADRs (associated with various drugs across multiple sources) with public databases (such as SIDER, FAERS, MedEffect, etc.) in a real-time manner. Through real-time access to ADRs being reported by patients globally on social media, healthcare professionals, pharmaceutical companies, and researchers will become aware of previously unknown side effects of drugs which in turn benefits patients, fosters novel drug research, and enhances ongoing research in the area of pharmacovigilance.
Furthermore, our results agree with (and extend) those reported by existing ADR research. For instance, in different studies conducted, patients placed on Metformin reported diarrhea,47–51 nausea,47–49,51 anorexia or decreased appetite,47,49–51 vomiting,49,51 headache,47,48,51 tiredness or fatigue,47,51 abdominal pain,49,51 dizziness,47,51 constipation, 51 stomach discomfort or upset stomach, 51 hyperglycemia 48 as major ADRs. In addition, a research based on social media also reported diarrhea, nausea, dizziness, and vomiting as top ADRs for Metformin. 35 These findings support our results for Metformin (see Figure 7) and further corroborates the quality of our approach for detecting ADRs on a much larger scale.
Finally, our ADR analysis and visualizations reveal interesting and valuable insights which can enable caregivers, healthcare professionals, pharmaceutical industry regulators, and drug producers in making good decisions. For instance, caregivers and healthcare professionals will be able to determine how best to administer diabetes medications to patients considering the possible ADRs that may occur due to their age and gender. Also, pharmaceutical industry regulators and drug producers can leverage the analytics in their drug-related research.
Limitations
Despite the success of the framework in detecting a lot of ADRs, both known and unknown ones, there is a need to extend its functionality to address some limitations. In its current state, the ADF cannot detect ADRs expressed in highly informal terms or slangs. For instance, comments like “generally feeling crappy” and “a lot of pooping” will not produce ADRs, even though they are supposed to produce feeling sick and diarrhea/loose stools as potential ADRs. Another limitation is the framework’s inability to detect certain abbreviations. For instance, “wt” in “20 lbs of wt gain. Hated it. Still have bad effects.” will not expanded to “weight” during parsing, hence “wt gain” is not detected as an ADR. A considerable solution is to further enrich the SPECIALIST lexicon by adding as many slang words as possible including spelling and inflectional variants. We can retrieve slang terms from the urban dictionary, for instance, which has a large collection of slang words and phrases. English abbreviations can also be retrieved from online sources and added to the SPECIALIST lexicon in a similar manner.
Since the experiment conducted in this work involves identifying ADRs within online patient-based drug reviews, we acknowledge that side effects or ADRs reported by patients may not constitute solid evidence for a potential ADR. For example, patients experiencing co-morbidities may assume that an ADR is linked to a particular medication, but this may not be the case. In addition, patients may mistakenly report side effects. Also, some causal assessment may require personal data including clinical information of patients which are usually not available on social media platforms. Therefore, while our findings and framework advance pharmacovigilance research, they should be applied with caution.
Finally, the ADF was evaluated against the SIDER database and found to be effective in identifying previously unknown ADRs in addition to known ones. We also presented valuable analytics/insights based on the framework’s output. Yet, we need to assess ADF’s reliability by comparing its output with those from similar frameworks or models in the literature using the same dataset. We plan to conduct this assessment as part of future work.
Conclusion and future work
Social media is a viable and rich source of health-related information shared directly by patients in an open and contextual fashion. To identify ADRs mentioned in drug reviews posted on social media, we designed a framework called ADF which automatically detects and extracts ADRs using Natural Language Processing (NLP) techniques. We demonstrated ADF’s applicability in real-life scenarios by mining diabetes-related drug reviews from three large and popular online patient-based health forums and feeding the data into ADF to automatically extract ADRs. We further analyzed the ADRs identified and presented our results including prevalent and less prevalent ADRs, age and gender differences in ADRs detected, as well as the previously unknown ADRs detected by our framework. Our findings showed that ADF is able to identify previously unknown ADRs after benchmarking with the SIDER database.
Our work has practical implications for health professionals, drug regulators, pharmaceutical companies, and pharmacovigilance research in the following ways: (i) The ADF generalizes its data sources and medications, hence not limited to a specific drug or health domain. This is important for active (real-time) ADR surveillance across multiple domains, (ii) Previously unknown ADRs identified through the framework would aid data-driven decisions with respect to awareness of possible side effects prior to drug prescription, and also complement the traditional adverse events reporting databases such as FDA Adverse Event Reporting System (FAERS), SIDER, and MedEffects, (iii) By changing the semantic types, the framework can be adapted to identify other interesting concepts from health-related conversations or documents; hence could be useful for developing intelligent systems that can support patients in different areas such as medication management, symptoms detection and support, real-time response to health-related questions, etc.
As part of our future work, we will enrich the SPECIALIST lexicon with slangs and many more abbreviations/acronyms to improve its coverage. Also, we will extend the ADF to create a hybrid framework that is capable of detecting cause and effect relations within patient medication reviews while employing more advanced NLP techniques including machine learning, as well as conducting more sophisticated analysis (such as network analysis) to depict the relations. We will also compare the ADF’s performance against similar frameworks/models in the literature. Finally, we will release the first version of ADF as a Representational State Transfer Application Programming Interface (REST API) that can be invoked by external applications for the purpose of identifying/extracting ADRs from patient reviews or clinical texts in real-time.
Footnotes
Acknowledgements
This research was undertaken, in part, thanks to funding from the Canada Research Chairs Program. We acknowledge the support of the Natural Sciences and Engineering Research Council of Canada (NSERC) through the Discovery Grant. We also thank the National Library of Medicine (NLM) for providing unrestricted license to access UMLS resources.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by the Canada Research Chairs Program and Natural Sciences and Engineering Research Council of Canada (NSERC) through the Discovery Grant.
