Abstract
Background
Radiology requests and reports contain valuable information about diagnostic findings and indications, and transformer-based language models are promising for more accurate text classification.
Methods
In a retrospective study, 2256 radiologist-annotated radiology requests (8 classes) and reports (10 classes) were divided into training and testing datasets (90% and 10%, respectively) and used to train 32 models. Performance metrics were compared by model type (LSTM, Bertje, RobBERT, BERT-clinical, BERT-multilingual, BERT-base), text length, data prevalence, and training strategy. The best models were used to predict the remaining 40,873 cases’ categories of the datasets of requests and reports.
Results
The RobBERT model performed the best after 4000 training iterations, resulting in AUC values ranging from 0.808 [95% CI (0.757–0.859)] to 0.976 [95% CI (0.956–0.996)] for the requests and 0.746 [95% CI (0.689–0.802)] to 1.0 [95% CI (1.0–1.0)] for the reports. The AUC for the classification of normal reports was 0.95 [95% CI (0.922–0.979)]. The predicted data demonstrated variability of both diagnostic yield for various request classes and request patterns related to COVID-19 hospital admission data.
Conclusion
Transformer-based natural language processing is feasible for the multilabel classification of chest imaging request and report items. Diagnostic yield varies with the information in the requests.
Introduction
Radiology reports are the primary communication method between radiologists and referring physicians and contain valuable information that impacts patient care.1,2 Most radiology reports are for single use, and physicians use the information from them to treat their patients. Data science can prove valuable by providing information from aggregated data; this is certainly true for radiology reports. 3
Radiology dashboards are suitable for monitoring and predicting radiology volumes and resource utilization;4,5 however, these systems do not provide in-depth information about referrals or diagnostic findings. Therefore, these systems are less suitable for providing insight into aggregate data about, for example, imaging appropriateness. 6 Reasons for referrals (or the input for the radiology process) can be retrieved from the content of the requests. 7 The percentage of positive findings, the diagnostic yield, can be calculated from the information in the radiology report. The diagnostic yield provides insight into disease prevalence, which not only informs referring clinicians but also can impact patient management. 8 Insight into appropriateness (input) and diagnostic yield (output) is not routinely assessed, even though this type of information is valuable—it contributes to effective resource utilization9–12 and allows for the identification of factors related to overtesting. 13
To leverage the great potential of aggregated data, the retrieval of information from radiology requests and reports must be automated. This is particularly the case when documents need to be classified for multiple co-occurring items. Instead of obtaining this information by manually categorizing radiology requests and reports, data mining using natural language processing (NLP) offers a promising alternative.14,15 In the field of chest imaging, traditional NLP has been successfully applied and can identify pulmonary nodules, 16 pneumonia, 17 and pulmonary embolisms 18 from radiology reports. Deep learning-based NLP methods perform equally well or better than traditional NLP models and can also be used to classify chest radiology reports. 19 High-performance transformer-based NLP algorithms, such as Bidirectional Encoding Representations for Transformers (BERT), have been applied to medical texts and are available open-source in several languages.20–22 The strengths of transformer-based NLP models are that they are pretrained on large datasets and account for a word’s context.20,23 These pretrained models can be applied to various NLP tasks after fine-tuning; that is, additional training with data from a specific task. Transformer-based models have been used for different NLP-tasks in radiology such as report section segmentation, 24 assessing spational information in radiology reports, 25 dectection of actionable findings, 26 image annotation, 27 and radiology report generation. 28
In our previous work, we proved BERT’s superior performance during a single-label classification task compared to other deep learning NLP methods, such as the convolutional neural network (CNN) and the long short-term memory (LSTM) neural network. 29 To our knowledge, the application of transformer-based models for the multilabel classification of a combination of chest imaging requests and reports (which can contribute to increased insights into the role and performance of chest imaging by assessing the diagnostic yield of different categories) has not yet been reported by other authors. Both the development and evaluation of a multilabel classification method are prerequisites for further research about the application of NLP to the evaluation of clinical care and the leveraging of data for predictive purposes.
In the current study, we hypothesise that (1) transformer-based NLP models that have been pretrained with Dutch language data will perform better compared to multilingual or English models when fine-tuned for radiology requests and reports written in Dutch; (2) multilabel classification with transformer-based NLP will perform equally well among different request and report categories and will be comparable to single-label classification; and (3) diagnostic yield will vary depending on the information in the request. Our research objectives are as follows: 1. Develop a deep learning-based NLP pipeline for the multilabel classification of radiology requests and reports. 2. Apply the pipeline to train and test five transformer-based models from a chest imaging dataset sample, and then compare performance metrics pertaining to model type, training method, and text characteristics. 3. Use the best-trained model to predict the classifications of requests and reports to demonstrate the feasibility of applying NLP to an extensive dataset in a proof-of-concept study of chest imaging, and analyse the relationships among the referral reason, diagnostic yield, and variability of requests over time.
Methods
The NLP pipeline, organised into sections of Python code.
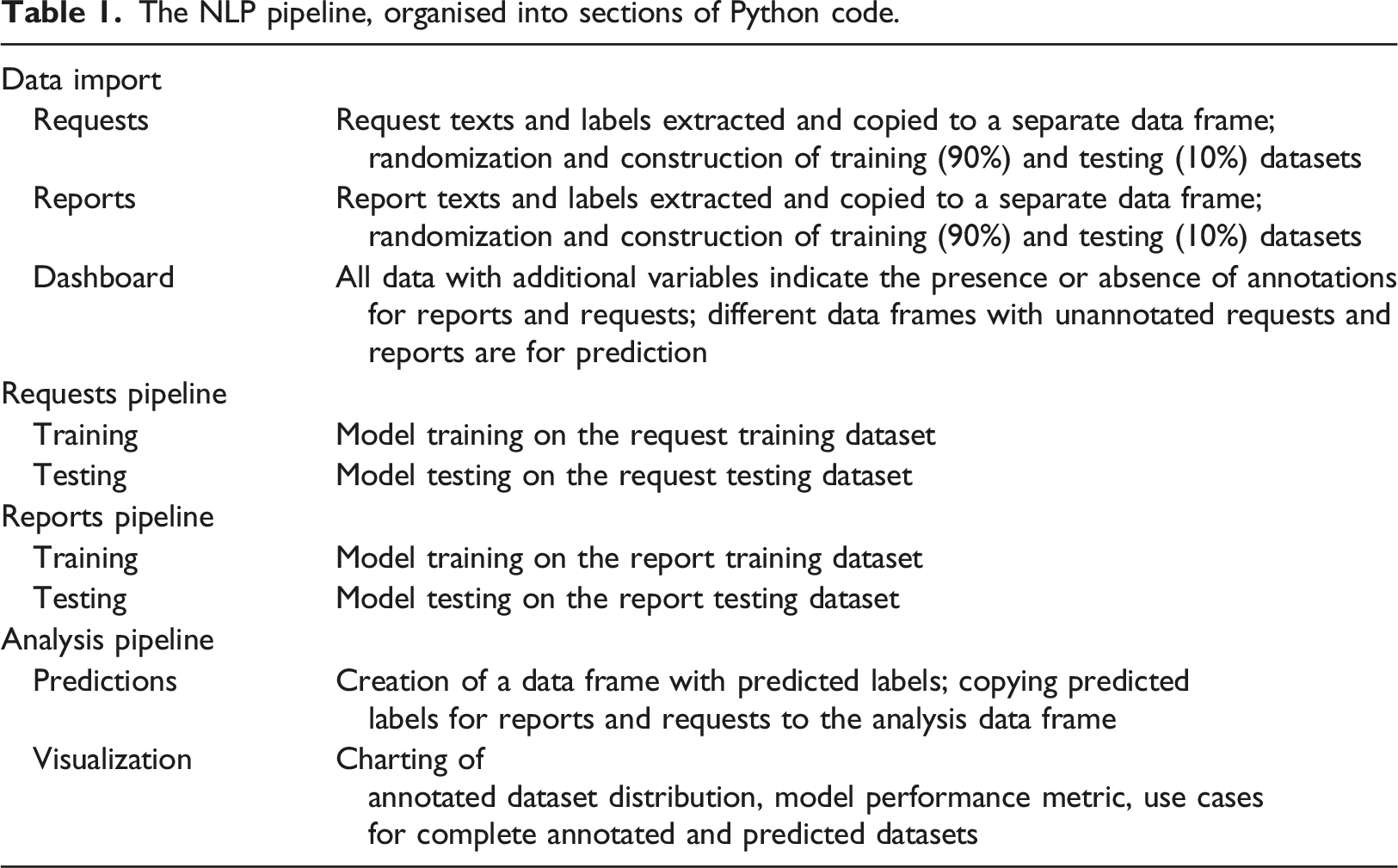
In a retrospective study, we used annotated datasets of radiology requests and reports to train and test five transformer-based NLP models for multilabel classification: BERTje, 30 RobBERT, 31 BERT-multilingual, 32 BERT-clinical, 33 and BERT-base. 23 An LSTM model was trained as a baseline for comparison.
The best-performing models were used to predict the classification of an unannotated dataset of requests and reports. According to Dutch law regarding medical research on humans, no informed consent was needed because of the nature of the retrospective chart review. The project was approved by the local research committee and the hospital’s board of directors.
Data
All requests for and reports of chest radiographs (CR) and computed tomography (CT) from January 2020 to September 2021 (n = 43,129) were retrieved from the PACS of a general hospital in the northern part of the Netherlands (Treant Healthcare Group).
The dataset was pseudonymized and stored in both the hospital IT system and secure cloud storage. Figure 1 shows the data processing flowchart. Data processing flowchart.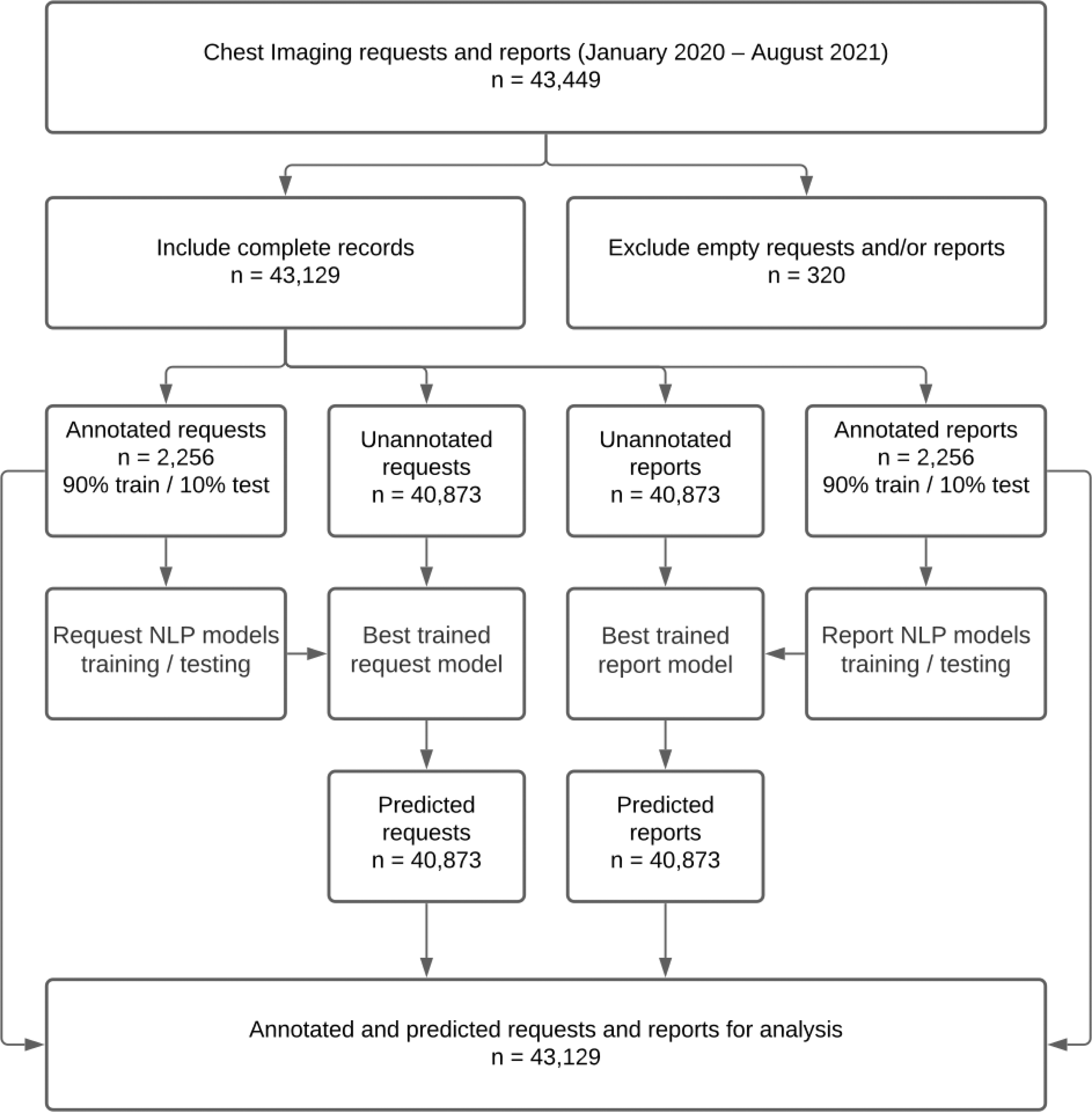
Ground Truth
In multilabel classification, each text item has multiple binary labels. For example, a chest imaging request that says, “Patient with fever and coughing. Infiltrate? Pleural fluid? Signs of COVID-19?” has positive labels for “Infiltrate”, “Pleural fluid”, and “COVID-19”, but negative labels for other items such as “Tumor”. In this manner, a model can be trained to classify texts using multiple labels simultaneously. For model training and testing, the requests (n = 2256) and reports (n = 2256) from the first 2 weeks of March 2020 and the first 2 weeks of April 2020 were annotated by a board-certified radiologist with 13 years of chest imaging experience. The two time periods were considered representative of cases without and with COVID-19, respectively, because the local rise of COVID-19 cases occurred towards the end of March 2020. The annotation process, performed in Microsoft Excel, consisted of assigning eight nonexclusive categories to the requests and 10 nonexclusive categories to the reports. The categories were based on the most frequent items found in the requests and the most frequent findings in the reports. Items had to be explicitly mentioned to be labelled as positive. For example, the COVID-19 label was given to requests that explicitly mentioned COVID-19 or coronavirus. For the reports, the “Normal” category was used only for those devoid of any abnormal findings.
After 5 months, the same radiologist reviewed all annotations to ensure consistency before the final model training was performed. This quality check resulted in the correction of 86 errors with the request annotations (4%) and 59 errors with the report annotations (3%).
Data Partitions
The annotated data was split into two groups, training data (90%) and testing data (10%), by using iterative_train_test_split from the skmultilearn library and used for model development.34,35 The remaining unannotated data (n = 40,873) from the original dataset was sent through the best-performing request and report models to receive classification predictions. The combined data (i.e. the combination of the annotated and predicted data) was used for the proof-of-concept study.
Models
Transformer model characteristics. Transformer models are language models that have been pretrained on text corpora in one or more languages. Listed here are the characteristics of the study’s five models. Fine-tuning was performed on Dutch radiology requests and reports.
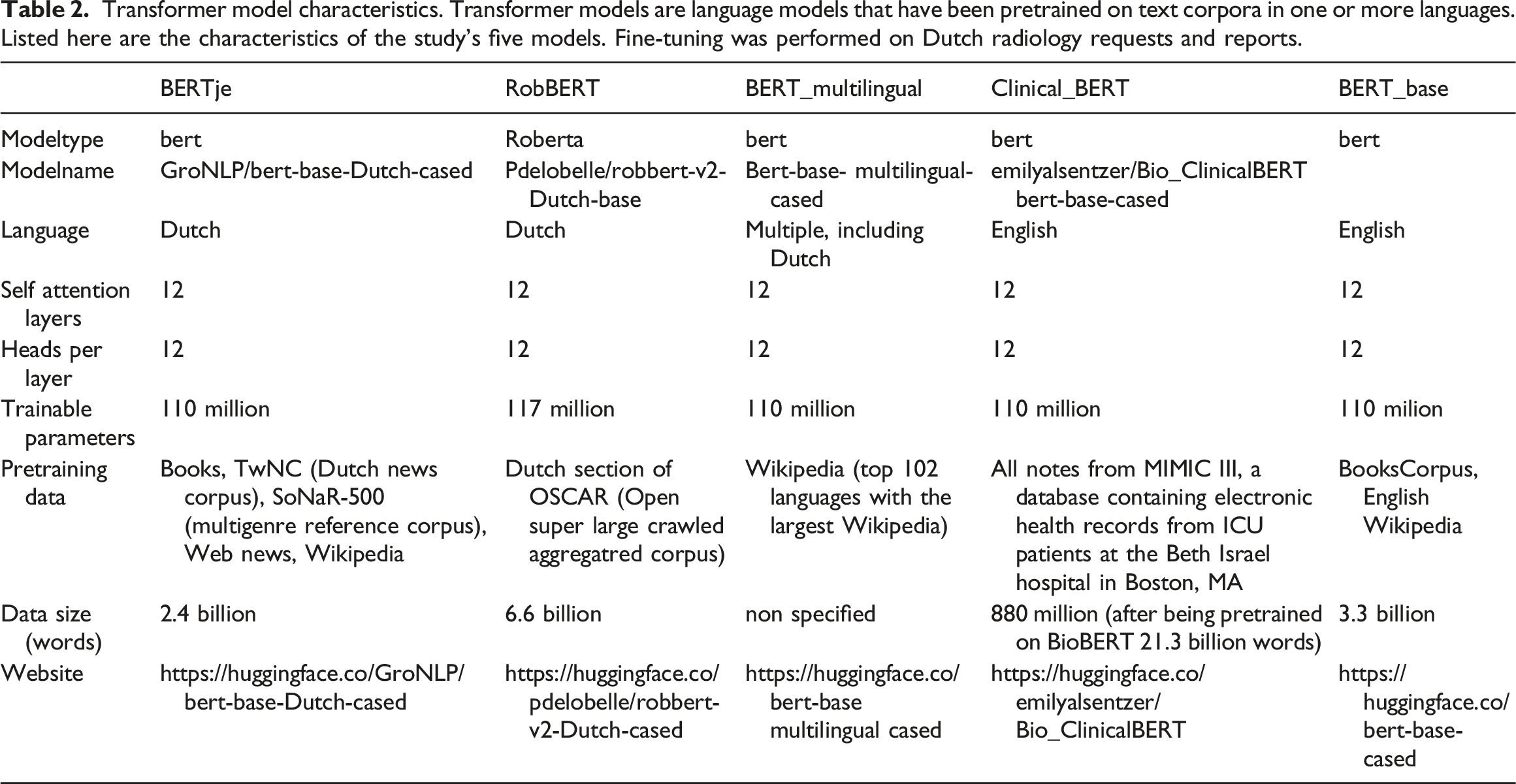
Acting as a baseline, an LSTM model was trained according to the methodology described in our previous work 29 except for the eight request and 10 report classes used for multilabel (instead of single-label) classification of the last layer.
Training
Model training parameters of the MultiLabelClassificationModel from the simpletransformers library. Parameters can be changed to optimise the training process for a specific situation.

Evaluation
For each of the five model types, three trained models—the models with the best accuracy according to the evaluation during training, the trained model after 2000 iterations, and the trained model after 4000 iterations—were stored and used for evaluation. This approach was implemented to identify the impact of training duration on performance. For the 15 trained transformer models and the baseline LSTM models using the testing sets, model performance was evaluated by calculating sensitivity, specificity, positive predictive value (PPV), negative predictive value (NPV), area under the curve (AUC), and F1-score. This resulted in 128 and 160 sets of performance metrics for request and report data, respectively. Confidence intervals were calculated. Using box plots, the following factors were compared to evaluate their impact on performance: 1. Model type (LSTM vs. Bertje vs. RobBERT vs. BERT-clinical vs. BERT-multilingual vs. BERT-base), 2. Data type (short requests vs. long reports), 3. Data prevalence (separate analysis of the request and report items), and 4. Training strategy (overall best vs. 2000 iterations vs. 4000 iterations).
Statistical significance was assessed using Python and the SciPy Library. An uncorrected p-value of <0.05 was considered statistically significant.
Proof-of-concept Study
To identify patterns on a large scale, a proof-of-concept study was performed with automated annotations of the unlabelled data. Thus, all the predicted request and report data (n = 40,873) was entered into the best-performing trained request and report models, respectively, to obtain predicted labels.
The variable number of items for each request and report equalled the sum of the positive labels per request or report. For all request categories, the likelihood of positive findings in the corresponding reports was calculated using contingency tables of each combination of requests and report categories. A heatmap was created that showed the diagnostic yield of the reported item for all single-item requests.
The variation of request items per week was visualised and combined with hospital admission data (retrieved from a publicly available dataset 37 ) originating from the same region as the radiology data.
Results
Data
Figure 2 displays the number of requests and reports included in the study, as well as each file’s word count, which varied among modality type (CR or CT) and document type (request or report). For example, the requests’ word count was the lowest. CR requests were the smallest texts, and the largest texts were CT reports. Number of and word count for radiography (CR) and computed tomography (CT) requests and reports.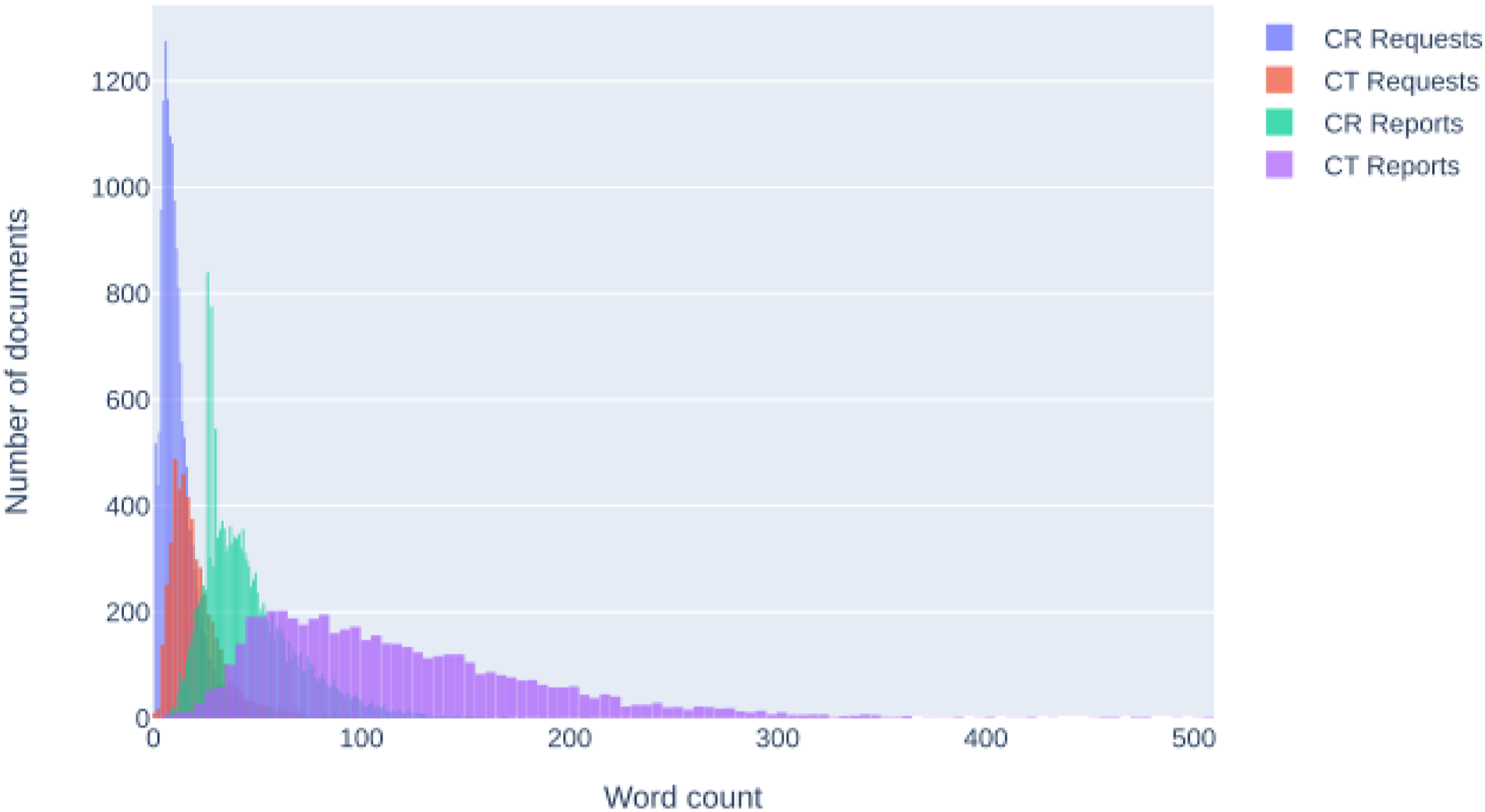
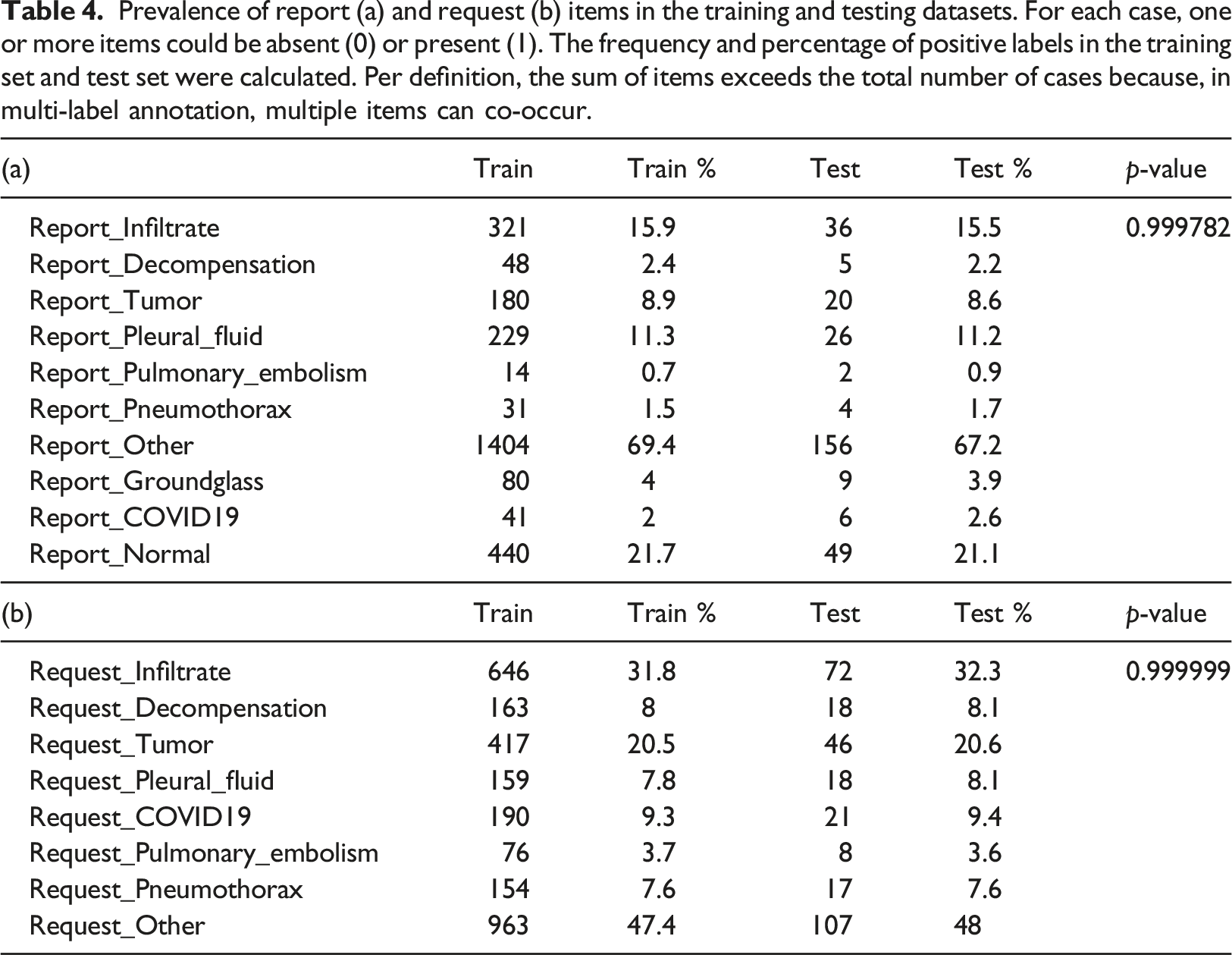
Prevalence of report (a) and request (b) items in the training and testing datasets. For each case, one or more items could be absent (0) or present (1). The frequency and percentage of positive labels in the training set and test set were calculated. Per definition, the sum of items exceeds the total number of cases because, in multi-label annotation, multiple items can co-occur.
Model Performance
Training and testing
The models’ performance metrics from the training dataset are summarised in Figures 3 and 4. Raw data of the model performance metrics with 95% confidence intervals is provided in the Supplemental appendix (Table A1). Summarised performance metrics of five transformer models and one LSTM model. For each model type, three models with different training durations were evaluated. Colours indicate training duration: two or three epochs (according to evaluation during training), 7.8 and 15.7 epochs for the transformer-based models, and 256 and 128 epochs for the LSTM models (best performance empirically). The sensitivity (a), specificity (b), positive predictive value (c), negative predictive value (d), AUC (e), and F1-score (f) were calculated for all labels and displayed as box plots per model type. Summarised performance metrics for all request and report items, as well as all models and training schedules. Request and report items are sorted in descending order, from left to right, according to prevalence.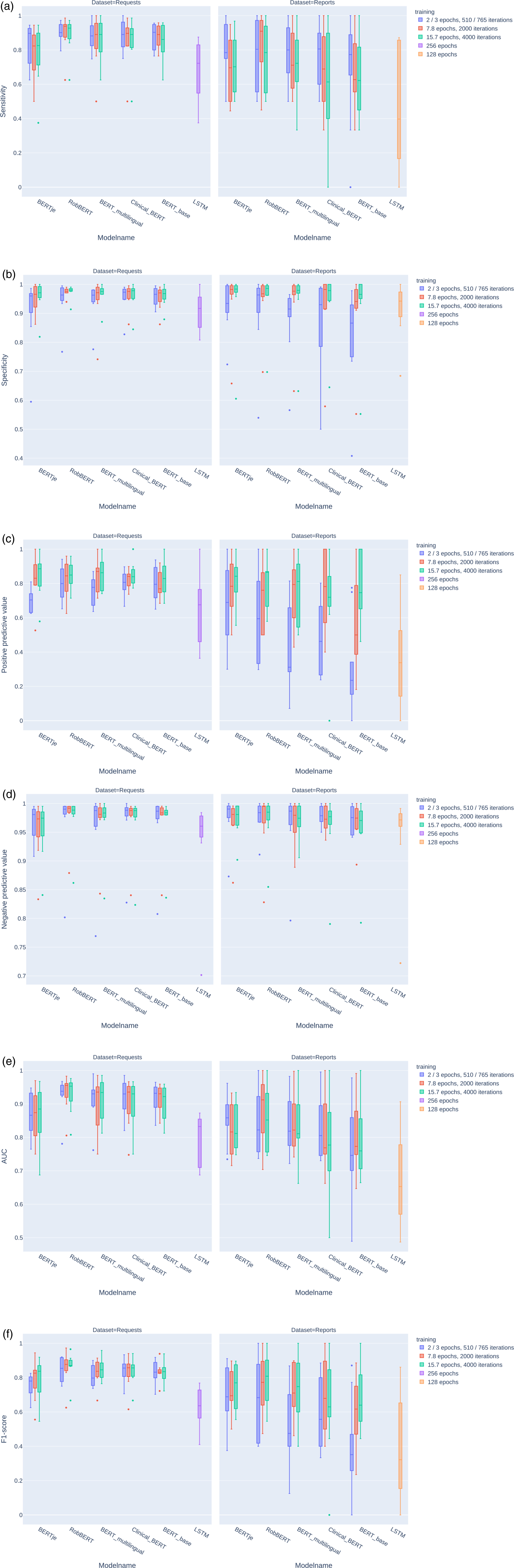
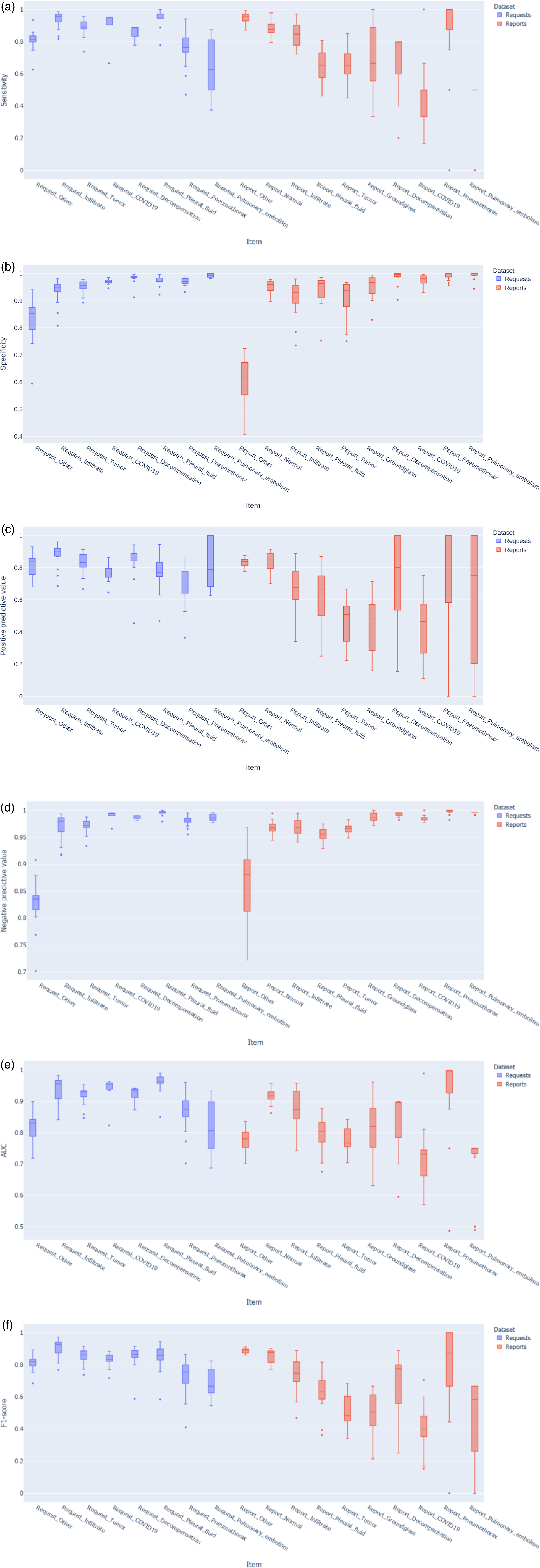
The metrics for the models trained on the request dataset were better than the report dataset models. In addition, the latter showed considerably more variation in performance. This variation occurred between the models and the training strategies. Especially true for the reports, the transformer models that were pretrained with Dutch data performed better, needing only 2–3 training epochs and improving only modestly after additional training. The specificity of the multilingual and English pretrained models improved substantially with additional training, but the AUC did not change significantly when training strategies were compared because the sensitivities decreased after prolonged training.
For both requests and reports, each model demonstrated a high specificity (> 0.90) and negative predictive value (> 0.95). Also, the sensitivities and positive predictive values were more variable and greater for the request and report items of higher prevalence in the datasets.
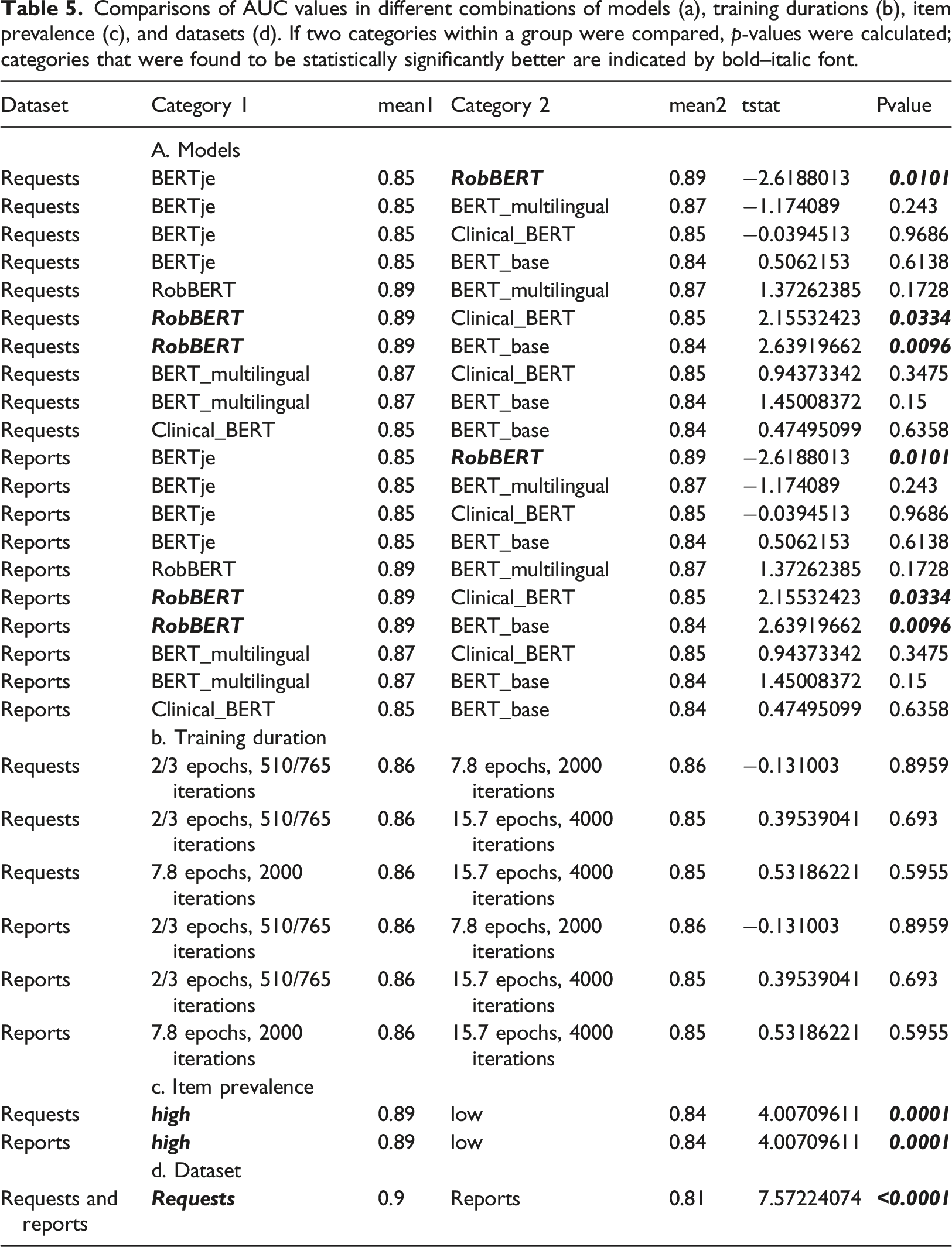
Comparisons of AUC values in different combinations of models (a), training durations (b), item prevalence (c), and datasets (d). If two categories within a group were compared, p-values were calculated; categories that were found to be statistically significantly better are indicated by bold–italic font.
Comparisons of F1 scores in different combinations of models (a), training durations (b), item prevalence (c), and datasets (d). If two categories within a group were compared, p-values were calculated; categories that were found to be statistically significantly better are indicated by bold–italic font.
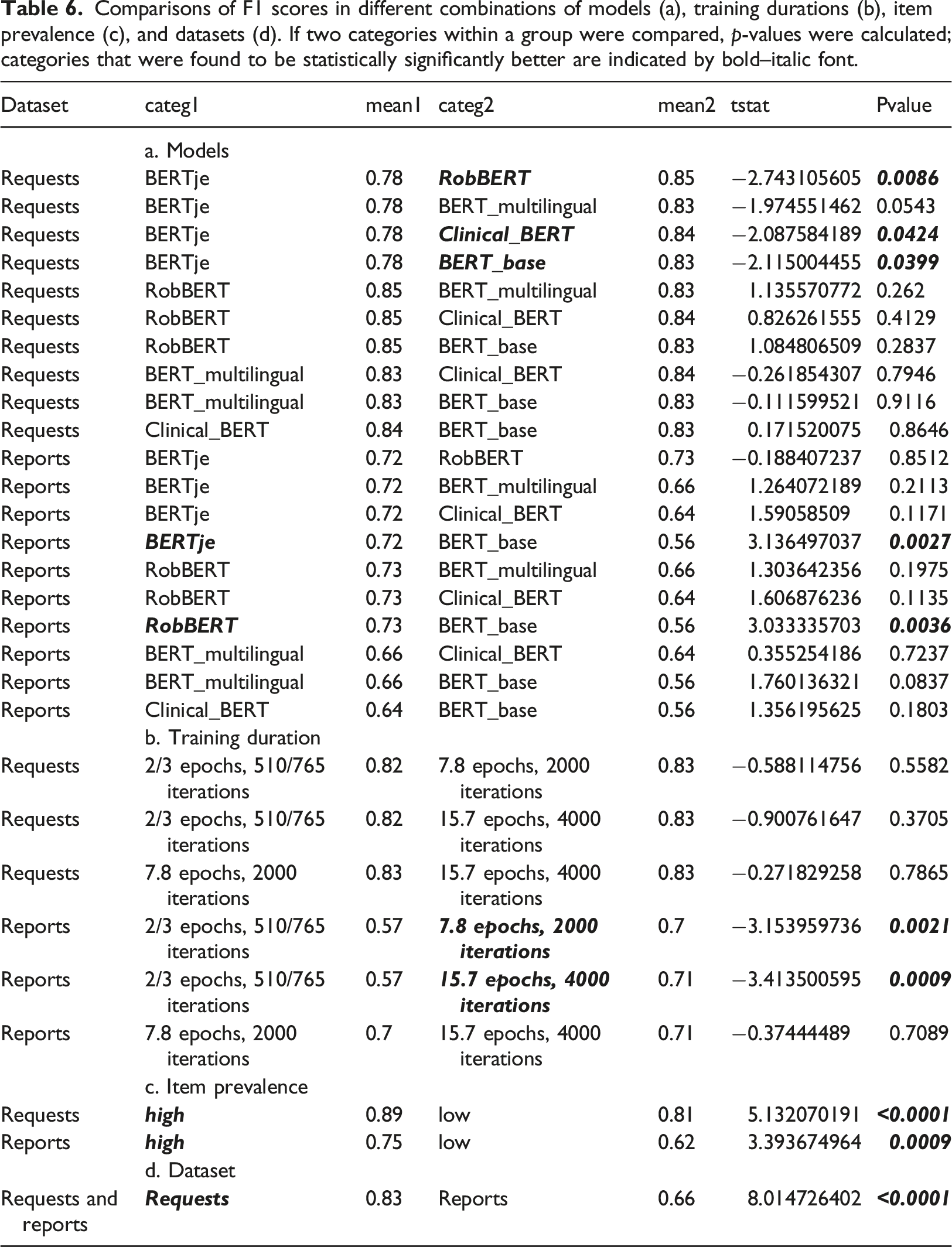
The transformer models outperformed the baseline LSTM models, but the RobBERT model surpassed all the others. The RobBERT model that trained for 4000 iterations was chosen as the best model overall. For the requests, the AUC values varied from 0.808 [95% CI (0.757–0.859)] for the item Request_Pulmonary_embolism to 0.976 [95% CI (0.956–0.996)] for the item Request_infiltrate. For the reports, the AUC values varied from 0.746 [95% CI (0.689–0.802)] for the item Report_COVID-19 to 1.0 [95% CI (1.0–1.0)] for the (infrequent) item Report_Pneumothorax. The AUC for the classification of normal reports was 0.95 [95% CI (0.922–0.979)]. In the comparison of F1-scores the differences between models were less pronounced, but did show significantly better results for longer training duration. Both the comparisons of AUC and F1-score demonstrated larger differences between results of data prevalence and text size compared to differences between models.
Predictions and proof-of-concept study
Diagnostic yield per report item. The request items represent the presence of items, irrespective of the number of other items that are also present in the same request. For example, a request with a positive label for “Infiltrate”, with or without other categories in the same request, has a 28.1% likelihood of receiving a corresponding report with a positive label for “Infiltrate”.
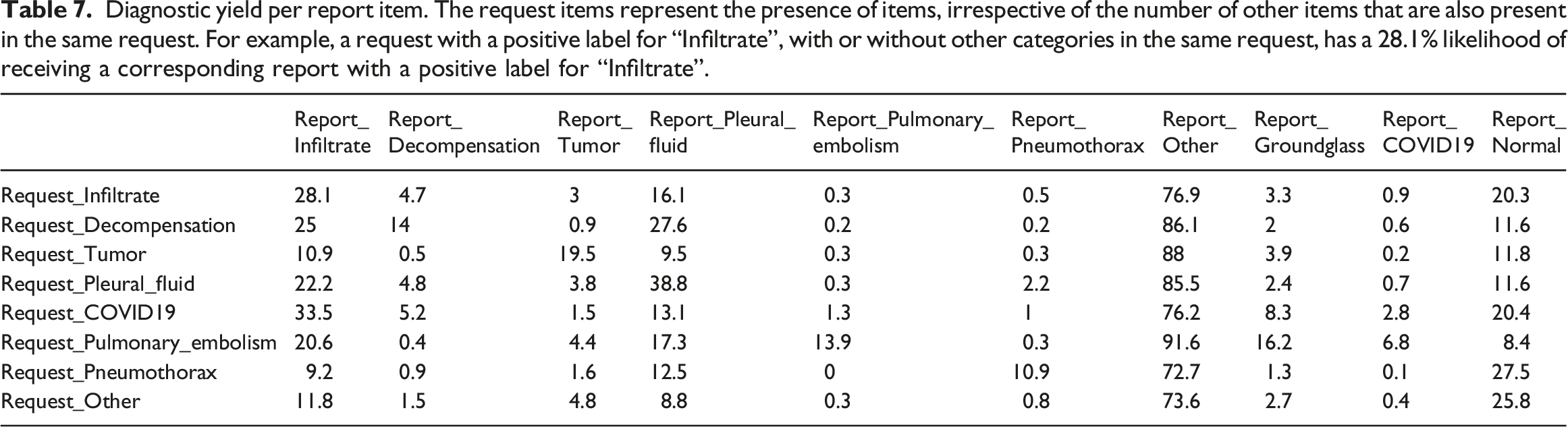
Figure 5(a) and Figure 5(b) illustrates the variability of the diagnostic yield per request category. For example, infiltrates are found when they are specifically mentioned in requests, but they can also be found even if requests specify other categories. (a) Heatmap of the diagnostic yield of single-request items for different report categories. The colour indicates the percentage of positive findings for a report item. For example, a request mentioning pleural fluid has a 70% chance of receiving a report with a positive result for pleural fluid. (b) Diagnostic yield of different report findings for all single-request items. Some report findings are found only on corresponding requests. For example, pulmonary embolism is only found in examinations with pulmonary embolism on the request. Other imaging findings, like infiltrate, are found in various examinations independent of the request.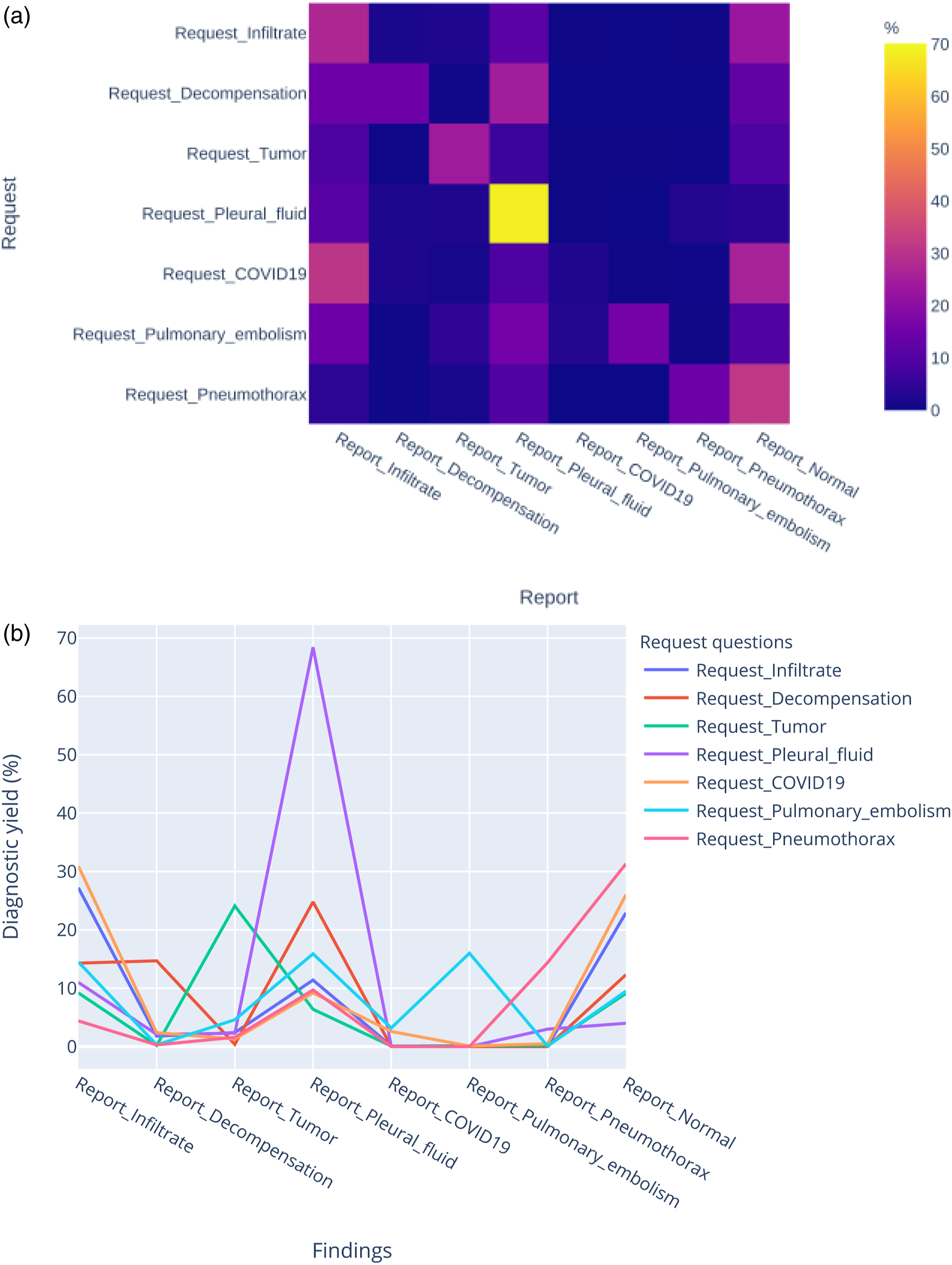
Figure 6 shows the variability in request volume over time. Radiology data can be combined with data from other sources; in this case, the rise of COVID-19 requests corresponds to an increase in hospital admissions. The first wave of COVID-19 led to an overall reduction of imaging, including requests in the “Infiltrate” category. Radiology CR (a.) and CT (b.) requests per week with positive labels in the categories “Infiltrate” or “COVID-19” and hospital admissions per week for COVID-19 patients. The request categories are nonexclusive: each request can have labels in one or more categories.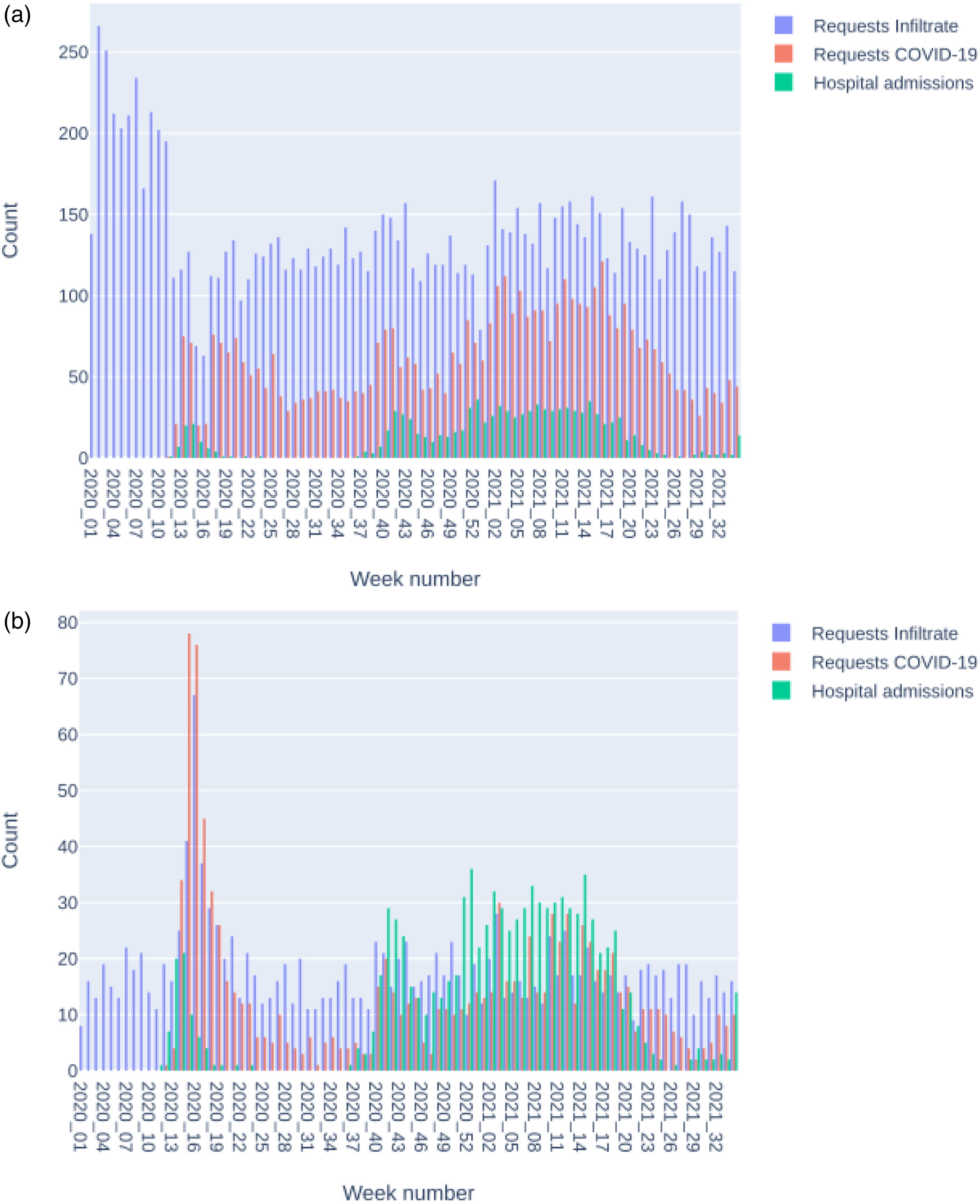
Discussion
In this study, we developed a pipeline for deep learning NLP in the context of radiology and compared five transformer models and one LSTM model. Distinctive characteristics of our work are the number of different models that we compared, as well as the multilabel (instead of single-label) classification of both radiology reports and requests. We now discuss (1) the possible reasons for the obtained results, (2) this work’s contribution and its comparison with published research, and (3) the challenges and limitations of this study.
Explanation of Results
The major difference between the LSTM models and the transformer models was that the former was trained only on the training data and the latter was pretrained on large corpora and took the context of words into account. This meant that, in our study, the transformer models needed less training with the training data to reach high performance levels. The Bertje and RobBERT models were pretrained with Dutch text, and not only was their better performance with short training duration expected, but it also confirmed one of our hypotheses. The multilingual and English models’ performance improved substantially after longer training, indicating the adaptability of transformer models. However, for pretraining, both language and type of text are relevant. For example, BERT-clinical was trained with English medical texts and, consequently, performed better than BERT-base. The results did not empirically explain the superior performance of the RobBERT model, but the model characteristics provide clues that can explain the differences. First, the RobBERT model was pre-trained on a corpus in the Dutch language, and second, the size of this corpus surpassed that of the other pre-trained Dutch model.
Prevalence impacts model performance, especially regarding sensitivity and positive predictive value. Accordingly, classification metrics change because of the differences in the prevalence of request and report labels. The datasets’ class imbalances were greater for the reports than for the requests; they were probably not fully compensated for by the application of class weighting. The word counts (and variations in word count) of the reports were greater than those of the requests, which explains the higher performance of the request models compared to the report models. This variability of performance, dependent on text characteristics and prevalence, confirmed our hypothesis about the comparable performance of multilabel classification compared to the single-label classification of our previous work. 29 The study results confirm previous results that transformer algorithms have higher performance in classification task on shorter texts and text with less class imbalance. 29
The proof-of-concept study illustrated and confirmed our hypothesis regarding diagnostic yield variations among different request categories. The degree of variation was difficult to estimate beforehand: just as one disease can cause several abnormalities, this was reflected in the radiology report findings. Therefore, multilabel classification provides a better reflection of the data than single-label classification.
Contributions and Comparison with the Literature
Natural language processing is increasingly applied to radiology reports, 15 but comparable studies of transformer-based models applied to chest imaging are scarce. Our work adds to the existing evidence that transformer models can reliably classify radiology reports and demonstrates the feasibility of combining both request and report classification models in the same pipeline. In addition, the study demonstrated the impact of dataset characteristics, such as item prevalence and text length. The ability of transformer models to classify both radiology requests and reports is important because the information provided by NLP models enables large-scale data retrieval for a myriad of other downstream tasks, such as analysing imaging results over extensive periods, as demonstrated in the proof-of-concept study. This work, therefore, contributes to the notion that NLP just as other applications of artificial intelligence can augment the ability of radiologists in patient care. 38
Wood et al. applied BioBERT to neuroradiology reports and achieved superior performance compared to our study. 39 It is important to remember that BioBERT combines language and medical context in the pretraining dataset; however, both studies saw differences in performance dependent on class labels.
Bressem et al. 40 compared four BERT models’ classifications of chest radiography reports. The best-performing model achieved the best pooled AUC of 0.98. The higher performance compared to our study can be explained by the inclusion of chest radiographs from intensive care patients, which would increase the prevalence of dataset findings, and the exclusion of reports that provided information about a single item without mentioning the absence or presence of other items. Our dataset is, therefore, less homogenous, but better reflects data in daily practice. Preselection can improve results, but this type of exclusion must be taken into account when applying the trained model to new data. The omission of preselection allowed us to use the trained models for predictive purposes on an unseen dataset.
Venturelli et al. assessed the appropriateness of referrals for imaging and other diagnostic procedures by analysing the requests’ content using a commercial software package. 41 Their study’s focus was on classification results (appropriate vs. not appropriate) and not the applied method’s performance, which impeded comparison with our study. However, the study demonstrated the feasibility of using requests for assessing appropriateness and is an excellent example of a future application for transformer-based NLP.
A systematic review of studies regarding the diagnostic yield of head CT scans of patients with syncope declared that a small sample size was a limitation of many studies and advocated large prospective research. 42 Our pipeline can be applied to such research in various subspecialties because of its applicability to large datasets. Pons et al. applied NLP to the long-term evaluation of the diagnostic yield of head CT for patients with minor head injuries. 43 Similar to our study was the use of information about the indications of imaging and information about the diagnostic results. The authors also used a small part of the data to train a model to extract information from a larger dataset.
Annarumma et al. 44 applied NLP to annotate chest radiographs to train a deep-learning image classification model for critical, urgent, nonurgent, and normal categories. The F1-scores for the extraction of the presence or absence of radiologic findings within the free-text reports ranged from 0.81–0.99 and were in the same range as our study. Annarumma et al. used a rule-based approach compared to deep learning in our study. The advantage of a deep learning pipeline is that it can be used for other annotated datasets; in contrast, a rule-based method requires manual feature engineering. Niehues reported a computer vision model for chest radiographs that was trained with labels derived from transformer-based NLP and demonstrated promising results comparable to those from expert radiologists. 27
The exceptional performance of the multilabel classification of radiology reports is not unique for transformer models. Short et al. reported a multilabel classification of mammography reports 45 and compared rule-based methods with a combination of convolutional and recurrent neural networks. Besides the methods applied, other differences between our studies included the relatively structured, homogeneous data used and the word-level (instead of document-level) classification performed.
Challenges and Limitations
No formal assessment of randomness of data or evaluation metrics was performed. Because all models were trained with the same data set, this is not supposed to have an impact on the results. Another limitation is that during training for all models all parameters were kept constant. Further performance optimization could be possible by hyperparameter tuning.
As already mentioned, model performance was impacted by class imbalances. This was relevant for both the training and testing datasets. Sensitivity values would have been more robust with a larger testing dataset size. Future research should incorporate additional methods to overcome low sensitivity due to class-imbalance. 46 Alternative performance measures can be considered in case of class-imbalance. 47 Furthermore, manual annotation was performed by a single person. Thus, it was not possible to assess inter-annotator agreement. To ensure the quality of the dataset, the same person performed an unblinded check for label consistency. External validation of a dataset annotated by multiple radiologists is needed to assess the generalisability of the trained models.
Conclusion
Transformer-based NLP is feasible for the multilabel classification of chest imaging request and report items, even after the fine-tuning needed by pretrained, language-specific models. The developed pipeline makes it possible to combine information from radiology requests and reports on a large scale to assess radiology utilization and diagnostic yield. Diagnostic yield in chest imaging varies with the information in the requests; therefore, the inclusion of NLP analyses of requests is recommended for quality control of and research into chest imaging.
Highlights
• Transformer-based natural language processing is feasible for the multilabel classification of radiology requests and reports. • Training and testing on a dataset containing 2256 requests and reports demonstrated good results. • Among five transformer models and one LSTM model, the RobBERT model surpassed the others and was used for the multilabel classification of 40,873 radiology requests and reports. • Diagnostic yield in chest imaging varies with the information in the requests.
Supplemental Material
Supplemental Material - The natural language processing of radiology requests and reports of chest imaging: Comparing five transformer models’ multilabel classification and a proof-of-concept study
Supplemental Material for The natural language processing of radiology requests and reports of chest imaging: Comparing five transformer models’ multilabel classification and a proof-of-concept study by Allard W. Olthof, Peter MA van Ooijen and Ludo J Cornelissen in Health Informatics Journal
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
