Abstract
Adalat (Nifedipine) is a calcium-channel blocker that is also used as an antihypertensive drug. The drug was approved by the US Food and Drug Administration in 1985 but was discontinued in 1996 on account, among other things, of interactions with other medications. Nonetheless, Adalat is still used in other countries to treat congestive heart failure. We examine all the congestive heart failure electronic health records of the largest medical center in Israel to discover whether, possibly, taking Adalat with other medications is associated with patient death. This study examines a semantic space built by running latent semantic analysis on the entire corpus of congestive heart failure electronic health records of that medical center, encompassing 8 years of data on almost 12,000 patients. Through this semantic space, the most highly correlated medications and medical conditions that co-occurred with Adalat were identified. This was done separately for men and women. The results show that Adalat is correlated with different medications and conditions across genders. The data also suggest that taking Adalat with Captopril (angiotensin-converting enzyme inhibitor) or Rulid (antibiotic) might be dangerous in both genders. The study thus demonstrates the potential of applying latent semantic analysis to identify potentially dangerous drug interactions that may have otherwise gone under the radar.
Introduction
This case study describes the discovery of an unexpected potential drug interaction through text analysis using latent semantic analysis (LSA) of medical records concerning congestive heart failure (CHF) as recorded in the electronic health records (EHR) system of a large medical center. CHF is one of the central causes of hospitalization in the United States. Advances in big data solutions allow for storage, management, and mining of large volumes of structured and semi-structured data, such as compound healthcare data. 1 That applies to healthcare too where there is a wealth of data ready for use within the healthcare systems. Nonetheless, there is a lack of effective analysis tools to find hidden relationships and trends in those data. Medical data mining has a huge potential for exploring the hidden patterns in the datasets of the medical domain. These patterns can be utilized for clinical diagnosis. This case study tells the story of how we used a standard text analysis method to identify rarely occurring dangerous drug interactions with the medication Adalat, interactions that warrant special medical attention. This was the first attempt in this large medical center to apply text analysis to identify such cases. Putting this case study into context, while medical data mining presents methodological and technical solutions to deal with the analysis of medical data and construction of prediction models, 2 according to Srinivas et al., 3 there is an absence of productive analysis tools to discover hidden relationships and trends in such data. This case study describes a successful attempt at that based on analyzing archival healthcare data.
Text analytics is a way to unlock the meaning from unstructured text. Broadly speaking, what the text mining method applied in this case study does is to find relationships between pairs of terms across documents in a corpus based on creating and then analyzing a text-to-document matrix (TDM). Perhaps unlike a human reader, text analysis does this on a superficial level of frequencies of the two terms appearing together. Superficial in this context means that prior schemata, such as prior knowledge of the human reader, do not intervene in the analysis, which could give text analysis methods a relative edge in some cases. Indeed, LSA may highlight relationships that might otherwise contradict “known” relationships. Moreover, a human reader while seeking novel new insight might ignore trivial known relationships, and hence not notice when they occur in unusual contexts that might shed new unexpected light on old problems. Moreover, text analysis allows the assigning of distance measures (e.g. cosine or correlation) to each relationship among pairs of terms, and in doing so allows identifying the relatively potentially more important relationships. In addition, text analytics can reduce confirmation bias, where researchers find what they are looking for and ignore that which contradicts their expectations. This was rather evident in this case study as the medical teams in both the medical center and the university did not expect the drug interaction results. Closely related, a human reader might be susceptible to cognitive overload when analyzing thousands of terms, but not so an automated technique. Moreover, the method described in this article allows also the discovery of indirect relationships, that is, when two terms are related but primarily or even only through their relationship to a third term. We wish to emphasize that the objective of the article was not so much to develop a new method as to show the applicability of an existing method in a new context.
The need to identify what might otherwise be ignored by a human reader and especially indirect relationships was pertinent in the context of this case study that analyzed a very infrequently used drug. As a drug rarely used, its drug interactions and other issues of potential danger might have gone undetected by a learned medical doctor who might just not associate them with those conditions, especially when the interaction is related to a third term. The potential of applying algorithms to create medical insight is well established (e.g. Tison et al. 4 ). We add to that line of research a demonstration of the potential of LSA and its application to specifically highlight less frequently occurring combinations of terms that reveal potentially dangerous interactions in rarely used drugs. In the analyses that we ran, our objective was to seek infrequent medical condition and medications, because, this being a world-class hospital, we did not expect to detect irregularities in typical treatment regimens. We chose LSA specifically out of the many machine learning languages because it was used previously for an equivalent study on health insurance data. 5
We hasten to add that these objectives should be put into proportion. The results reported here are analyses of correlations in archival data. Causality cannot be inferred, and there is no doubt that more is happening than the correlations and our subsequent reading of some of the related medical records reveal. The results suggest that this method could be brought in to support phase IV clinical trials, that is, testing side effects of drugs after they have been approved and are in circulation with the public at large. 6 Specifically, that may be achieved by indicating individual medical records that medical examiners may wish to review.
Background
Literature review and our contribution to the literature
Studies have shown the feasibility of text mining in creating meaningful structured information from clinical content such as those found in pathology or radiology reports 7 and medical insurance records 5 as well as for predicting disease. 8 Those studies show the potential of text mining for efficient prediction of disease status and relationships across diagnoses from relatively small datasets of clinical discharge summaries and other reports.
This study looks at a larger body of medical reports: the complete set of EHR of a large medical center. Such EHR provide large amounts of data that if appropriately analyzed could lead to earlier identification of disease states such as heart failure 9 as well as rare drug interactions that might go undetected in a smaller dataset. However, clinical text is unstructured data, and fundamental questions that are easy to state in plain language are often difficult to reduce to practice with such data. 10 EHR also have the advantage of incorporating medical text reports/narratives in conjunction with structured data such as test result numbers. This allows the analysis of the text in the context of and in comparison with standard test results, such as echocardiogram (ECHO) test narratives. Including those structured test results could add a new dimension, as witnessed specifically for CHF patients. 11
The type of natural language processing (NLP) of EHR data applied here through LSA can automatically, and at least partly without resorting to a human expert, produce vast volumes of complete and structured coded data that may be available for many diverse medical applications, and so contribute to considerably improve healthcare. 12 In this case study, we show its application to pharmacovigilance (the collection, detection, assessment, monitoring, and prevention of adverse drug effects). 13 Automatic discovery of adverse drug reaction (ADR) from textual data has received attention in pharmacovigilance research. 14 Such an automatic method combined with text analytics’ methods can enable an automatic coding and parsing of medical events in real time, something that if produced by manually inspecting medical documents with clinical terminologies may be close to impossible as the number of ADR reports grows substantially every day. 15 In addition, machine learning of EHR may decrease the time needed for the validation of ADR reports compared to using manual coding and review. 16 The type of structured terminology analyses provided by machine learning of EHR may thus assist clinicians as well as enable the construction of knowledge bases to assist clinical points of care. 17
The detailed objective of this study, adding to that literature, is to present a case study that demonstrates the potential of applying a specific type of text analysis to EHR medical records to discover possible adverse drug combinations—demonstrated specifically with an investigation about Adalat (Nifedipine in the United States) (“Nifedipine is used to treat high blood pressure and to control angina (chest pain). Nifedipine is in a class of medications called calcium-channel blockers. It lowers blood pressure by relaxing the blood vessels, so the heart does not have to pump as hard. It controls chest pain by increasing the supply of blood and oxygen to the heart” (USA National Library of Medicine https://medlineplus.gov/druginfo/meds/a684028.html).) Adalat used to be a medicine of choice for dealing with CHF. Adalat/Nifedipine was approved by the US Food and Drug Administration (FDA) in 1985 (NDA 019478) under the proprietary name “ADALAT.” It is currently “Discontinued,” in part because of serious adverse effects (https://www.accessdata.fda.gov/scripts/cder/ob/results_product.cfm?Appl_Type=N&Appl_No=019478). Nifedipine was discontinued in part due to a study by Furberg et al. 18 that associated its use with increased mortality compared to control groups in high dosages, possibly related to an increase in angina symptoms, and therefore they recommended that “the use of moderate to high doses of short-acting Nifedipine in survivors of an acute myocardial infarction and patients with stable or unstable angina should be avoided.” Their observation does not appear in our data, hopefully because that recommendation was heeded to. Adalat is still used in Israel, albeit sparingly, and appears in our data in the records of 196 men and 236 women, including a relatively recent case of a man who had hematuria (blood in urine) and was given Adalat in 2013.
Materials and method
Study setting and design
The data consist of all the Cardiology Department EHR at Chaim Sheba Medical Centre in Israel. Sheba is the largest medical center in Israel, serving a population of over 1.5 million people in the Tel-Aviv Region and serving about 200,000 emergency room visits a year. The data contain all the medical tests and written reports of the 11,949 patients who visited the department between 1 January 2008 and 31 December 2016. There was one record for each patient. That record contained also all the test results and written reports of all the visits that each patient made, including test results from other departments in the hospital. The data were de-identified, replacing the patient ID with an artificial code and removing other obvious identifiers. Average patient age was 77.26 years (standard deviation (SD) = 12.42). The oldest person was 109 years old. Only 634 patients were younger than 55.
Data preparation
The data contained 1729 variables (structured and unstructured), including demographics and every possible CHF test result as applicable to that patient. Only the textual part of the data was analyzed. The results of the medical tests were included as is. So, if, for instance, the result of an ECHO test was “minimal aortic stenosis,” then “minimal aortic stenosis” was included in the data. The written assessments were mostly in Hebrew, others were in English. The Hebrew text was translated as is by running an application program interface (API) to Google Translate, using R, a statistical software package. A sample of 30 translated texts, chosen in random, were verified manually for their translation accuracy by a dual native speaker of Hebrew and English. This verification was about the correct translation of the medication names, medical conditions, and physiology terms. Comparing the original with the translated text, no errors were detected in the translations. To put this into context, the medical texts being analyzed were quite repetitive, as one might expect, and contained a limited number of physiological terms, medical conditions, and medications described in straightforward, short, unambiguous sentences. Moreover, in some cases, the term as used in English was just written in Hebrew letters, such as “hypertrophy,” “septum,” “tricosphalopathy,” “anaphroplasia,” and “systolic,” so the translation and verification were unequivocal. We also translated the English text back into Hebrew to verify that the translations were accurate. The medical keywords of interest to this study (i.e. medication names, medical conditions, and physiology) were translated back correctly, although the grammar, which is not the topic of this study, occasionally became somewhat inaccurate and sometimes synonyms replaced the original words. We did not limit the inclusion of terms. That was a deliberate research decision. Had we introduced a lexicon then it might have introduced a bias, such as external information that is not part of the original data, into the analysis. We wanted to avoid such a case so that there could be no doubt that the results are derived from the data and method alone, not from any external source. Text was not limited by their length either. The average number of words in a medical record of a man was 307.20 (SD = 211.35, minimum = 14, maximum = 1483), which is significantly larger (t = 18.75, p < .0001), than that of an average number of words in a medical record of a woman at 242.36 (SD = 166.23, minimum = 18, maximum = 1341).
After that initial transformation, a semantic space was created out of those records. Using standard R modules, uppercase letters were cast into lowercase, standard stop-words were removed (words such as “and” and “or” that add little semantic value when running text analysis but that appear many times in typical text), and special characters and numbers were removed. The records were entered with no other changes. A TDM was then created out of those data, depicting the text to document frequency matrix. Although it is sometimes regarded as standard to run stemming in text analysis, we did not do it in this case because carefully reading the translated medical records revealed that doing so would not add value. Our objective there was to make as few changes to the original data as possible. Nonetheless, running the same analysis with stemming produced the same results as shown in Table 1.
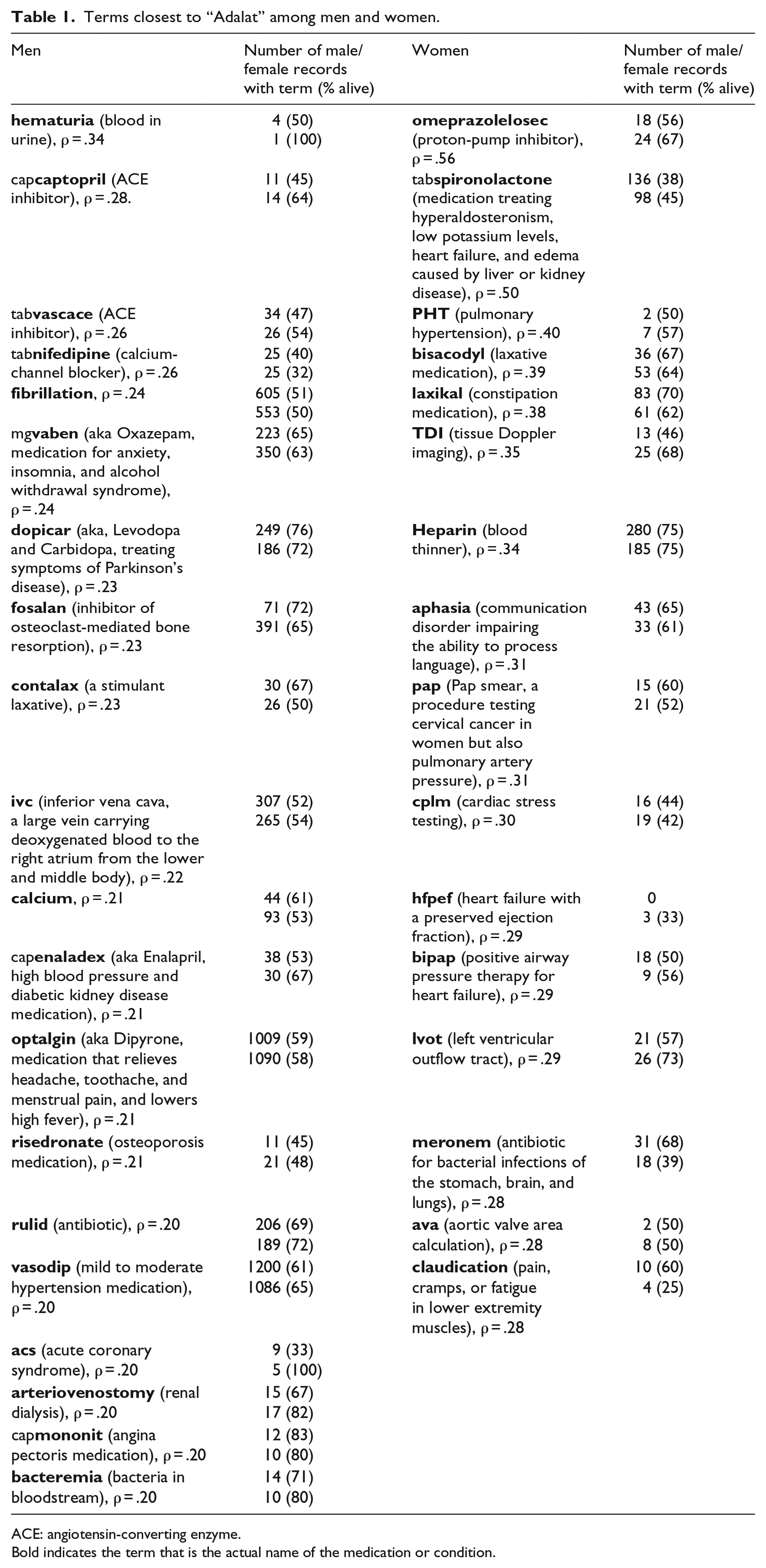
Terms closest to “Adalat” among men and women.
ACE: angiotensin-converting enzyme.
Bold indicates the term that is the actual name of the medication or condition.
The creation of the TDM was done separately for men and women, resulting in two matrices. The separation between men and women was done because it is known that CHF symptoms and treatments differ by gender. There were 6372 medical records of men and 5577 of women. The two resulting TDMs show the frequency of every retained term (word) in each document (medical record). As it is standard in LSA, semantic spaces were created by log-entropy weighting each TDM and running a singular-value decomposition (SVD) transformation. TDM weighting decreases the influence of high-frequency terms, to increase the model’s sensitivity to other semantic information.
5
This additional weighting is done by analyzing the product of the local weight of each term within the document (calculated as

Analysis process.
Running an SVD on a TDM produces a semantic space composed of a matrix of terms to dimensions, a matrix of documents to dimensions, and a singular-value matrix that by multiplying the previous two matrices by it can reconstruct the original data, providing enough dimensions are allowed. The terms to dimensions matrix factors the terms into dimensions so that the terms that load high on the same dimension share a lot of variance - that is, they tend to co-occur highly, emphasizing the similarity of the terms. On each of the two term-to-term matrices (men and women separately), we ran Pearson’s correlations of each term with all other terms. As LSA gives more weight to infrequent terms and to infrequent co-occurrences, this approach amounts to looking for the proverbial needle in the haystack in the less taken paths where others might miss it.
Results
After the correlation matrices were created out of the two semantic spaces, the 50 closest neighbors (“neighbors” is an R function that produces the n closest terms to a specified term in a vector) to the term “adalat” (all the terms were in lowercase at this stage) were identified for men and women. These terms are shown in Table 1, including the correlation coefficients. The results shown in the table are limited to the names of medications and medical conditions only. As the accepted shorthand practice in the dataset is to add “tab” and “cap” as a prefix to the medication name, representing tablet and capsule, respectively, the values in Table 1 show in bold the part of the term that is the actual name of the medication or condition. Showing the ability of LSA to also identify indirect relationships in the data, while the terms “pap” and “adalat” do not appear together in any of the records, LSA identifies that they are connected. Indeed, “adalat” and “cervical” appear together in three records, and “pap” and “cervical” in two.
As can be seen in Table 1, there is practically no overlap between the terms of medicines or medical conditions that are most highly correlated with “adalat” among men and women. To add context, we then looked at the raw data in the medical records. In Table 1, the highest correlated term with Adalat among men is hematuria. Only one woman in the dataset had the term “hematuria” appear in her medical record, as compared to seven men, and no woman had both terms “hematuria” and “adalat” appear together. That specific woman died at 63, suggesting the condition is more rare among women. (And, there was little in the medical record to suggest otherwise why she died. Her ECHO average regurgitation was “Trivial” and her ECHO hemodynamics was “normal” with also normal sinus rhythm, but her latest ECHO hemodynamics was “severely increased.”) Another noticeable difference between men and women is Vascace (Cilazapril). Vascace is an angiotensin-converting enzyme (ACE) inhibitor that might be dangerous for pregnant women (http://www.medicinesinpregnancy.org/Medicine–pregnancy/ACE-Inhibitors/). Indeed, Vascace is correlated highly with Adalat only among men in Table 1, while in the medical records themselves among the 11 women in whose medical record it appears together with Adalat all are at least 64 years old. The above would seem to confirm that there is no obvious reason to suspect that the medical center used Adalat in any inappropriate manner.
Looking at the potential of this method to identify adverse drug interactions, interesting results are the cases of Rulid and Captopril. Among the nine men in whose medical reports appear both Rulid (antibiotic) and Adalat, seven are reported dead. Likewise, among the 11 women who took Rulid and Adalat, 9 are reported dead. Obviously, we cannot say that the combination of Rulid and Adalat is the cause of death, but it may be an indication that these factors warrant closer examination, as might be the case with phase IV testing. Another case of a possible adverse drug combination with Adalat is Captopril. This drug is in high correlation with Adalat among men in the SVD analysis (ρ = .28), but not so among women (ρ = .13). Among the five men in whose record both drugs appear, all are dead. Among the 11 such women, 9 are dead. To put those numbers into perspective, in the entire dataset, 56.71 per cent of records pertaining to men indicate that the patient is dead, and 59.38 per cent indicate so for women.
Interestingly, comparing those results with the database at the Medical School of the authors, a database that encompasses medical data from six hospitals in the Mid-Atlantic region of United States with 424,100 patient records showed that there was no combination on record of Nifedipine (the US equivalent of Adalat) ever appearing together with Rulid, nor with its alternative spelling Rulid nor its US brand name Roxithromycin. Likewise, Nifedipine never appeared with Captopril. We caution to add, however, that this US database has data starting only in 2007. We did not have access to other datasets.
Discussion and conclusion
Adalat has been discontinued in the United States by order of the FDA since 1996, including all its brand names. The results concerning Captopril and Rulid mentioned in the medical reports together with Adalat might be interpreted as another reason in support of that decision as well as suggesting the possibility of using this method in phase IV testing as a way of pinpointing potential lethal drug interactions. This does not come to suggest that LSA can definitely identify causal relationships, as there is much more at play than only the correlations among terms across the medical documents, especially as the method we applied highlights the low co-occurrence cases. That said, these results confirm expected patterns, such as an evident reluctance to give women Vascace (known in the United States as Cilazapril), and, as such, are reassuring. The results also show that Adalat was given for different conditions among men than women, and suggest that the medical center did indeed perform medical profiling by gender. Just how important such profiling might be as a preventive measure is possibly implied in that the one woman who suffered from hematuria died at the age of 63 without any obvious reason showing in her ECHO tests. We hope these results encourage medical text analysis research as a possible additional tool in phase IV testing. As a case study, the results show that LSA can, with relative ease, be applied to discover adverse drug interactions and to verify that medication is used correctly in consideration of gender and age issues.
Although our study is limited to CHF patients and to the specific medication in probe (Adalat), there is a potential for similar studies in other chronic diagnoses and other medications. Future research can study connections, relationships, and correlations between other types of medications using text analysis as done in our study. This research shows the potential of such studies to discover unexpected and even dangerous correlations between medication that can be avoided or at least known and controlled henceforth.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
