Abstract
The objective of this research is to identify major subject areas of medical informatics and explore the time-variant changes therein. As such it can inform the field about where medical informatics research has been and where it is heading. Furthermore, by identifying subject areas, this study identifies the development trends and the boundaries of medical informatics as an academic field. To conduct the study, first we identified 26,307 articles in PubMed archives which were published in the top medical informatics journals within the timeframe of 2002 to 2013. And then, employing a text mining -based semi-automated analytic approach, we clustered major research topics by analyzing the most frequently appearing subject terms extracted from the abstracts of these articles. The results indicated that some subject areas, such as biomedical, are declining, while other research areas such as health information technology (HIT), Internet-enabled research, and electronic medical/health records (EMR/EHR), are growing. The changes within the research subject areas can largely be attributed to the increasing capabilities and use of HIT. The Internet, for example, has changed the way medical research is conducted in the health care field. While discovering new medical knowledge through clinical and biological experiments is important, the utilization of EMR/EHR enabled the researchers to discover novel medical insight buried deep inside massive data sets, and hence, data analytics research has become a common complement in the medical field, rapidly growing in popularity.
Introduction
Medical informatics is an interdisciplinary research field that applies information technology (IT) to the medical field for creating and analyzing data, information, and knowledge to improve healthcare and medicine.1–3 Despite the recently heightened interest, medical informatics is not a new field. Rather, it has been a long-running endeavor of applying IT to medical research for creating and processing data, information, and knowledge. According to Masic, 4 the bulk of the related research activity within this field began in the 1950s due to the rise of information communication technology (ICT). Masic noted that the first medical informatics article was authored by Ledley and Lusted in 1959, and it was the medical application of the electronic computer in diagnostics and therapy which established medical decision-making. Masic further claimed that the falling price of computers in the 1980s opened the door for an extensive development of health information technology (HIT) at all levels of the healthcare systems. He states that in the 1990s, medical/health research on the improvement of methods and techniques of artificial intelligence was actively conducted. The development and application of expert systems in medical diagnostics and therapy was also widely researched during this period. As such, the discussion of medical informatics cannot be separated from the development of technology.
Recently, the development of technology which enabled the healthcare industry to collect massive data and analyze for quality care and cost-effectiveness has prompted a big stream of research, which is data mining using electronic medical/health records (EMR/EHR).5,6 Because of the hope of what data mining can offer to the healthcare industry, President Bush called for nationwide use of EMR by 2014, and the Department of Health and Human Services (HHS) is involved in various aspects of achieving this goal. 7 Accordingly, the Administration and Congress have both been requiring the adoption, connectivity, and interoperability of HIT. 7 This effort will increase the adoption of EMR/EHR and is expected to increase research using the collected data. As such, it is not surprising to observe that this field grew very fast and became a mature independent field of study. 1 In fact, medical informatics is now the fastest growing field based on the number of publications in PubMed. 8
Couple with the policy and the advancement of technology, the academia observed the rapidly growing research interests, and scholars have attempted to understand a boundary for the field, but it is limited to a specific topic area 9 or a specific journal. 10 Although existing review studies assist in the understanding of the field of medical informatics, they were written before the wide spread of the EMR/EHR deployment or narrowly focused.1,11 Because EMR-related medical informatics articles have recently seen a rapid increase, and because those articles are the most cited in the field,10,12 it is important to investigate how these recent governmental efforts to adopt EMR/EHR and emerging technological capabilities to store and analyze big data have changed the field, and how they impact medical informatics as an academic discipline. As such, the purpose of this article is to comprehensively investigate the major subject areas over the last decade, as well as how those subject areas have changed and where they are heading.
In order to achieve the research objective, we identified the top 23 journal publications in the field of Medical Informatics within the past 12 years. Journals are identified based on the Institute for Scientific Information’s (ISI) Web of Knowledge Journal Citation Report (JCR) 2012. These journals are considered to be leading and shaping the field of study because of their impacts. The total number of articles amounts to 26,307 articles. With such a large number of articles, if one attempted to discover the trends of the field by manual means, it would be an overwhelming, if not impossible, task to process and categorize this vast quantity of articles in order to identify the trends of major subject areas in general and in a time-series by year. Even if it is possible, the outcome could be inaccurate or incomplete. The recent advances in data mining technologies and related software development efforts enable researchers to discover underlying patterns and trends from massive quantities of documents, as is the case in this study. Text mining is the automated or partially automated processing of text. It involves imposing meaningful structure upon text so that relevant information can be extracted from it.13–15 In this study, following the general guidelines presented in Delen and Crossland’s article from 2008, we used a hybrid methodology (text mining followed by data mining) to apply semi-automated text categorization to organize available research abstracts into logical categories (or topic clusters) by published years in order to observe the trends of each subject area over time.
The rest of the article is organized as follows: in section “Materials and methods,” we will discuss research methods that include how we identified the articles, the analytical techniques used, and the processes of those techniques. Section “Results” reports the findings followed by an interpretation of the findings in section “Discussion.” We conclude this article with some recommendations and future research directions.
Materials and methods
This section offers explanations on how sample articles are identified, searched, and finalized. The second part of this section deals with text mining techniques that were used for this article. We offered detailed explanations on the text mining process that we used for this manuscript in order to introduce the emerging technique to the field.
Data acquisition and preprocessing
The sample journals are identified using the ISI’s Web of Knowledge JCR 2012 in the field of medical informatics. This method is chosen because it is commonly used in academia as an indicator to assess journal rankings across disciplines. Scholars and publishers perceive highly ranked JCRs as prestigious or of such an esteemed quality that they commonly use them as their own research outlet. 16 As such, those journals shape and guide the directions of the field. Following the practice, we used all journals listed in JCR 2012 in the medical informatics area as our journal sample, and the criteria produced 23 journals shown in Table 1.
List of journals included in the study.
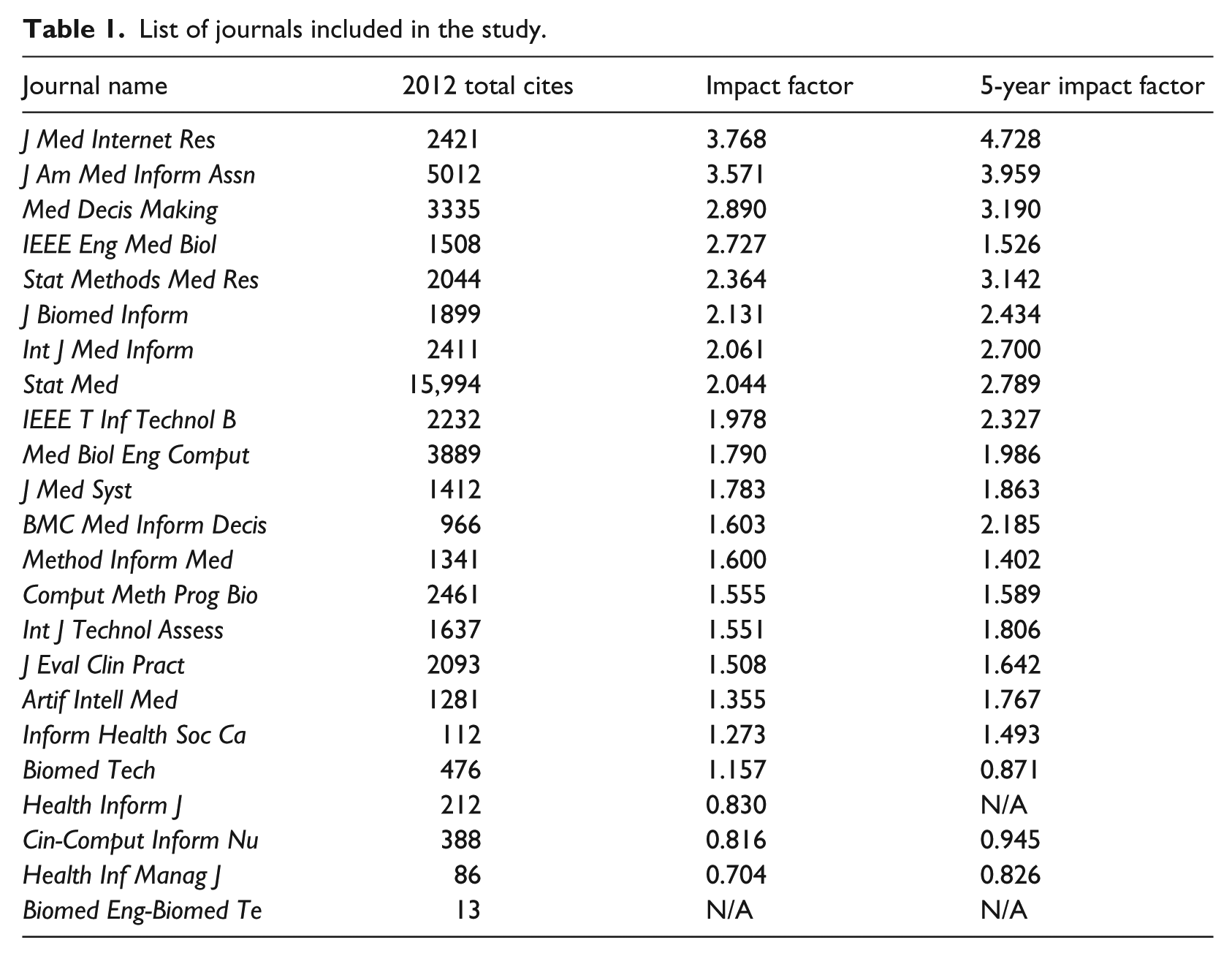
The dates of the included journals range between 2002 and 2013. The beginning year, 2002, was chosen because the application of HIT to the medical field became widely utilized in the timeframe. This was due to the demands for decreasing paperwork through the adoption of HIT and preventing medical errors via evidence-based treatments.17–21 Despite the popular application of HIT to medical informatics, sparse research has been conducted after this timeframe. The ending timeframe, 2013, was set because many of the journals’ summaries for 2014 were unavailable when we searched the articles in March 2014 as PubMed does not make journal articles available until 1 year after publication.
Based on the 23 journals, we went through each journal in the PubMed website and retrieved the title, abstract, publication date, and journal name.
22
Two of the journals began included in PubMed after 2002. More specifically,
After going through each journal’s search process, 26,307 articles were identified. Among them, editor’s notes, book reviews, and commentaries were excluded from the dataset as the purpose of this project was to cluster the words in the abstract of research papers. Also, the above items are not research papers, and thus they are not peer reviewed. Furthermore, they do not have abstracts. After removing those items, the data sample was reduced to 21,464. All searched articles were directly transferred from the PubMed website to EndNote and then to an Excel file in order to analyze clusters in SAS Enterprise.
Text mining methodology
Text mining is the semi-automated process of extracting patterns to discover knowledge from large amounts of unstructured data sources. 23 Text mining is closely related to data mining in that it has the same purpose and uses the same processes, but with text mining, the input to the process is a collection of unstructured text files such as Word documents, PDF files, text excerpts, and XML files. The benefits of text mining are in areas where large amounts of textual data are being generated, such as academic research literature (the one that is used in this study), finance (quarterly reports, media commentaries), medicine (discharge summaries, doctor notes), law (court orders), biology (molecular interactions), technology (patent files), and marketing (customer comments).
Text mining process
In order to succeed, text mining studies should follow a sound methodology based on lessons learned and best practices. A standardized process, such as CRoss Industry Standard Process for Data Mining (CRISP-DM), is also needed for text mining. At a very high level, the text mining process can be represented with a context diagram where the inputs, output, controls (i.e. constraints), and mechanisms (i.e. enablers) are captured as directed arrows as in Figure 1.
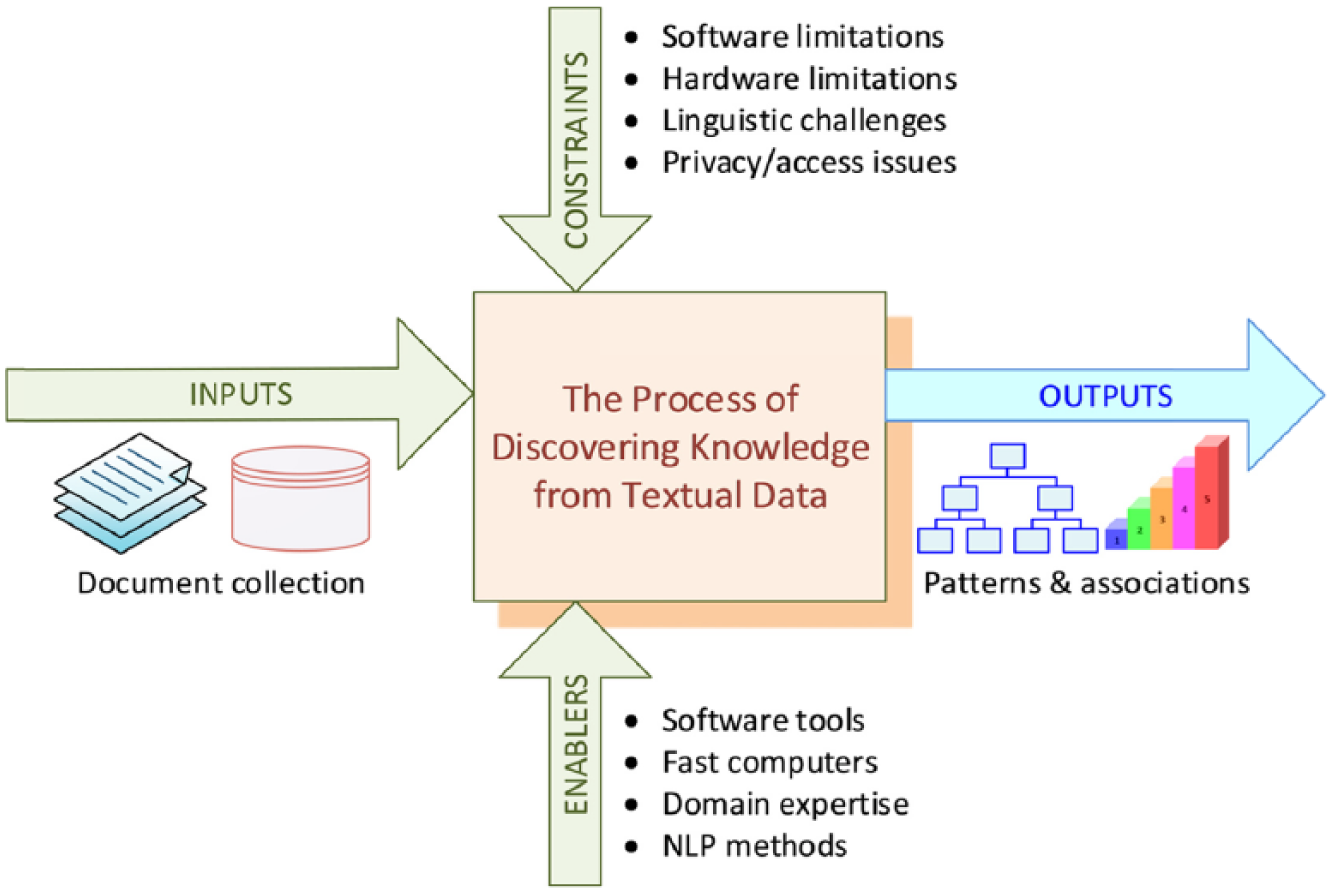
High-level context diagram for text mining of published literature.
The context diagram can be decomposed into three consecutive activities/phases, each of which having specific inputs to generate certain outputs (see Figure 2). If, for some reason, the output of a task is not that which is expected, a backward redirection to the previous task execution is necessary. Figure 1 provides a graphical presentation.
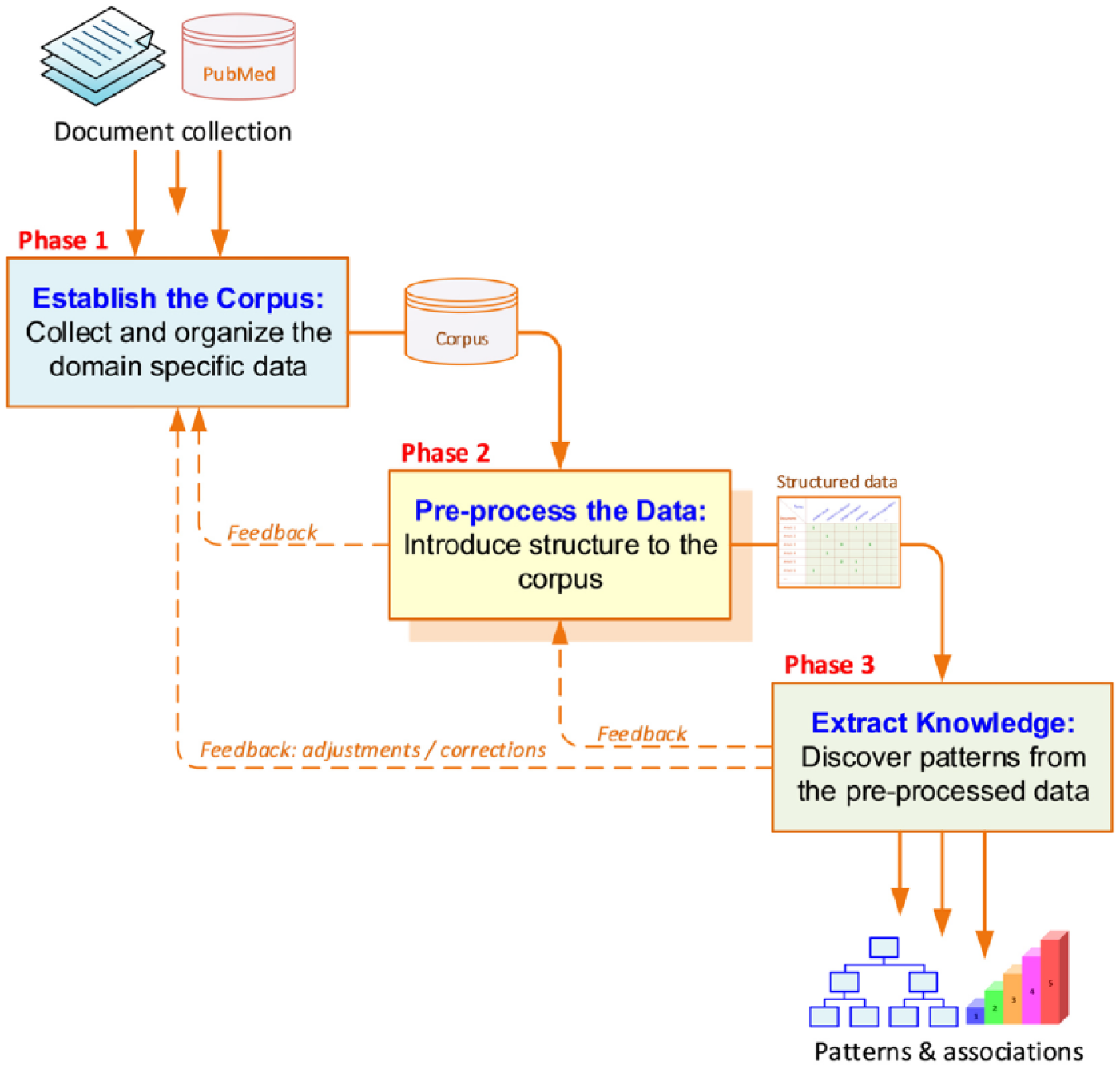
Process for text mining and knowledge discovery.
Phase 1: establish the corpus
The main purpose of the first task activity is to collect all of the documents related to the context (domain of interest) being studied. In this specific study, we retrieved article summaries of the selected sample from the PubMed website. The collected documents were then transformed and organized in a manner in which they are all in the same representational form (e.g. ASCII text files) for computer processing.
Phase 2: pre-process the data (create the term-by-document matrix)
Upon the establishment of the corpus, the term-by-document matrix (TDM) is created using the corpus. In the TDM, rows represent the documents and columns represent the terms (Figure 3). The relationships between the terms and documents are characterized by indices (i.e. a relational measure that can be as simple as the number of occurrences of the term in respective documents). As can be seen in Figure 4, there are several consecutive tasks that need to be carried out to create the clusters.
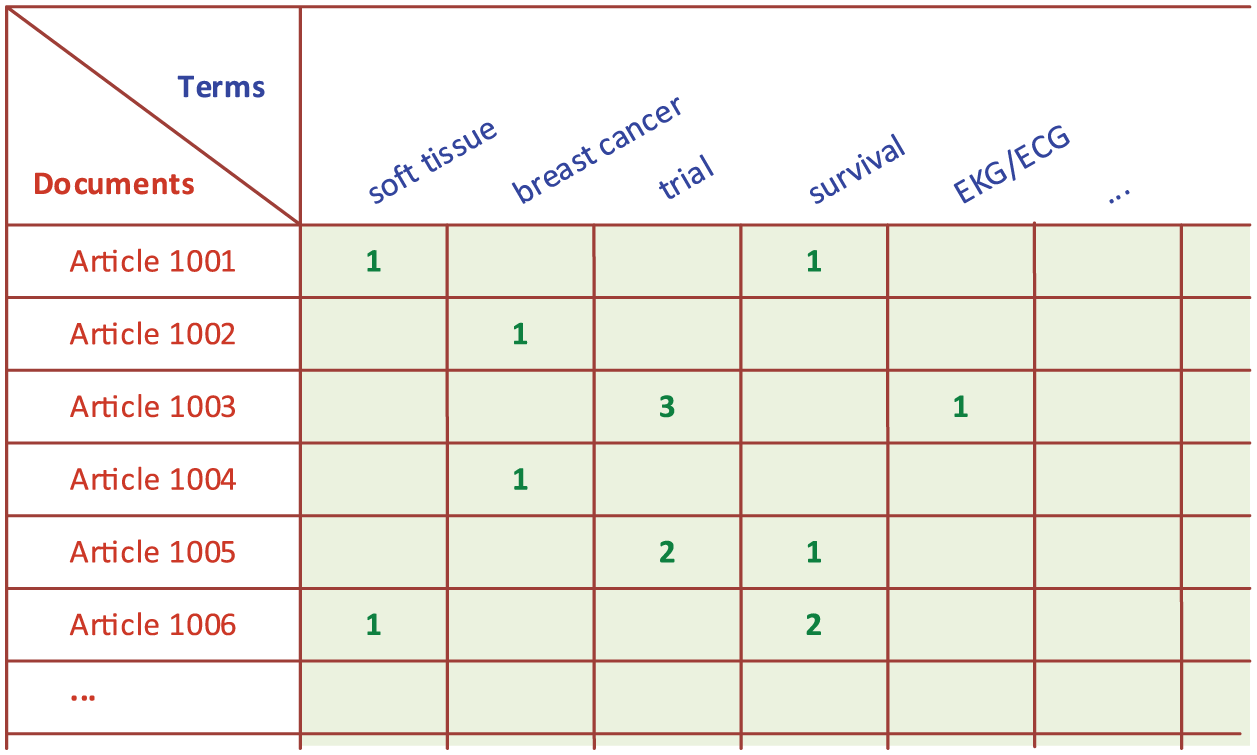
Sample term-by-document matrix.
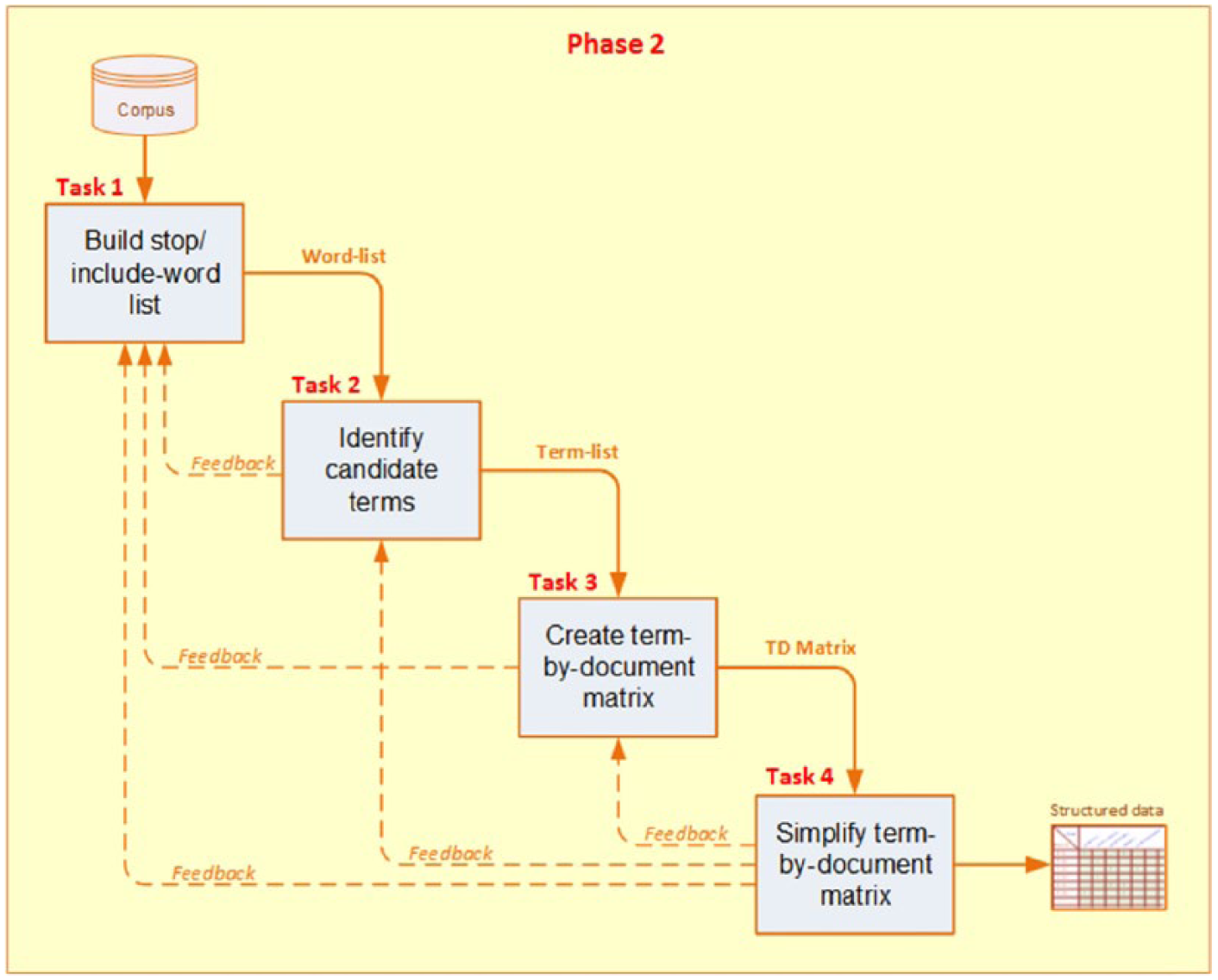
Decomposition of “pre-processing the data” phase.
Specifying all the documents as rows in the matrix;
Identifying all of the unique terms in the corpus (as its columns), excluding the ones in the stop term list;
Calculating the occurrence count of each term for each document (as its cell values).
Commonly, the corpus includes a rather large number of documents, which is time-consuming and, more importantly, it might lead to the extraction of inaccurate patterns. These dangers of large matrices and time-consuming operations pose two questions. 24 The first question asks how does a researcher identify the best representation of the indices for optimal processing by text mining algorithms. The most commonly used method for this question is to transform the term frequencies. For this method, the input documents are indexed and the initial word/term frequencies (by document) are computed in order to summarize and aggregate the extracted information. Specifically, terms that occur with greater frequency in a document may be the best descriptors of the contents of that document. It is not, however, reasonable to assume that the term counts themselves are proportional to their importance as descriptors of the documents. For example, even though a term occurs three times more often in document A than in document B, it is not necessarily reasonable to conclude that this term is three times more important as a descriptor for document A as it would be for document B.
In order to have a more consistent TDM for further analysis, these raw indices should be normalized. In a statistical analysis, normalization consists of dividing multiple sets of data by a common value in order to eliminate differing effects of different scales among the data elements to be compared. Raw frequency values can be normalized using a number of alternative methods that include log frequency, binary frequency, and term frequency (TF) into inverse document frequency (IDF).
The most commonly used representation is TF/IDF, which is the one used in this study (Table 2). It works as follows: a term such as
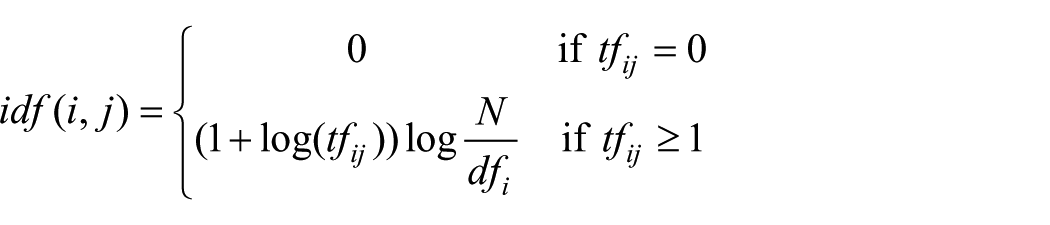
Simple example of TF/IDF.
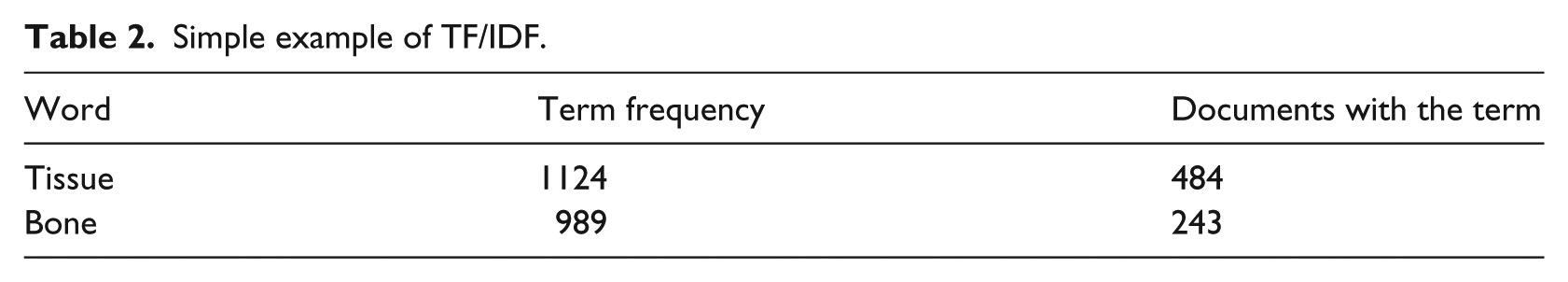
where
Following the formula above, TF for tissue is 1+ (log(1124/484)) = 1.37 and the TF for bone is 1+ (log(989/243)) = 1.61. IDF for tissue is log(2253/484) = 0.67 and for bone is log(2253/243) = 0.97. The TF/IDF for tissue is 1.37*0.67 = 0.92, and for bone, it is 1.61*0.97 = 1.56. As shown in this calculation, although the word tissue appears more frequent than that of bone, because tissue appears more frequently in other documents in Cluster 1, its weight is lower (0.67) than that of bone (0.97), and thus the relative importance of the term bone is higher in this example.
The second question asks how would a researcher reduce the dimensionality of TDM. Because, the TDM is often very large and rather sparse (most of the cells are filled with zeros), this answer is more tractable to handle. While several options are available for reducing such matrices to a manageable size, singular value decomposition (SVD) is a method of representing a matrix as a series of linear approximations that expose the underlying meaning–structure of the matrix. The goal of SVD is to find the optimal set of factors that best predict the outcome. This article adopted the SVD method.
Phase 3: extract the knowledge
Using the well-structured TDM, we then extracted patterns, which are represented in clusters. Clustering is an unsupervised process, whereby objects or events are placed into “natural” groupings. An unsupervised process is one that uses no pattern or prior knowledge to guide the clustering process. The unsupervised clustering process groups an unlabeled collection of objects (e.g. documents, customer comments, and web pages) into meaningful clusters without any prior knowledge. The basic underlying assumption is that relevant documents tend to be more similar to each other than to irrelevant ones. If this assumption holds, the clustering of documents based on the similarity of their content improves search effectiveness. 26
Finding the “optimal” number of clusters is not an easy task, since there is not a mathematical formula (a closed-form algorithm) developed for it. It is still an experimental heuristic process where the number of clusters are gradually increased from a small number to a large number (or decreased the other way around) to reach a point where the number of clusters are the “optimal” representation of the underlying multi-dimensional dataset. The optimality is determined by measuring the balance between in-cluster similarities and intro-cluster dissimilarities using the Euclidean distance. The Euclidean method is popularly used because of its capability to extract distinct and yet meaningful clusters. 11 This is the heuristic experimental process that we followed in determining a six-cluster representation of the dataset.
Results
Figure 5 shows the most frequently surfaced words/terms in each of the six clusters using a word cloud representation. The counts and percentages of each cluster over time are provided in Figures 6 and 7.
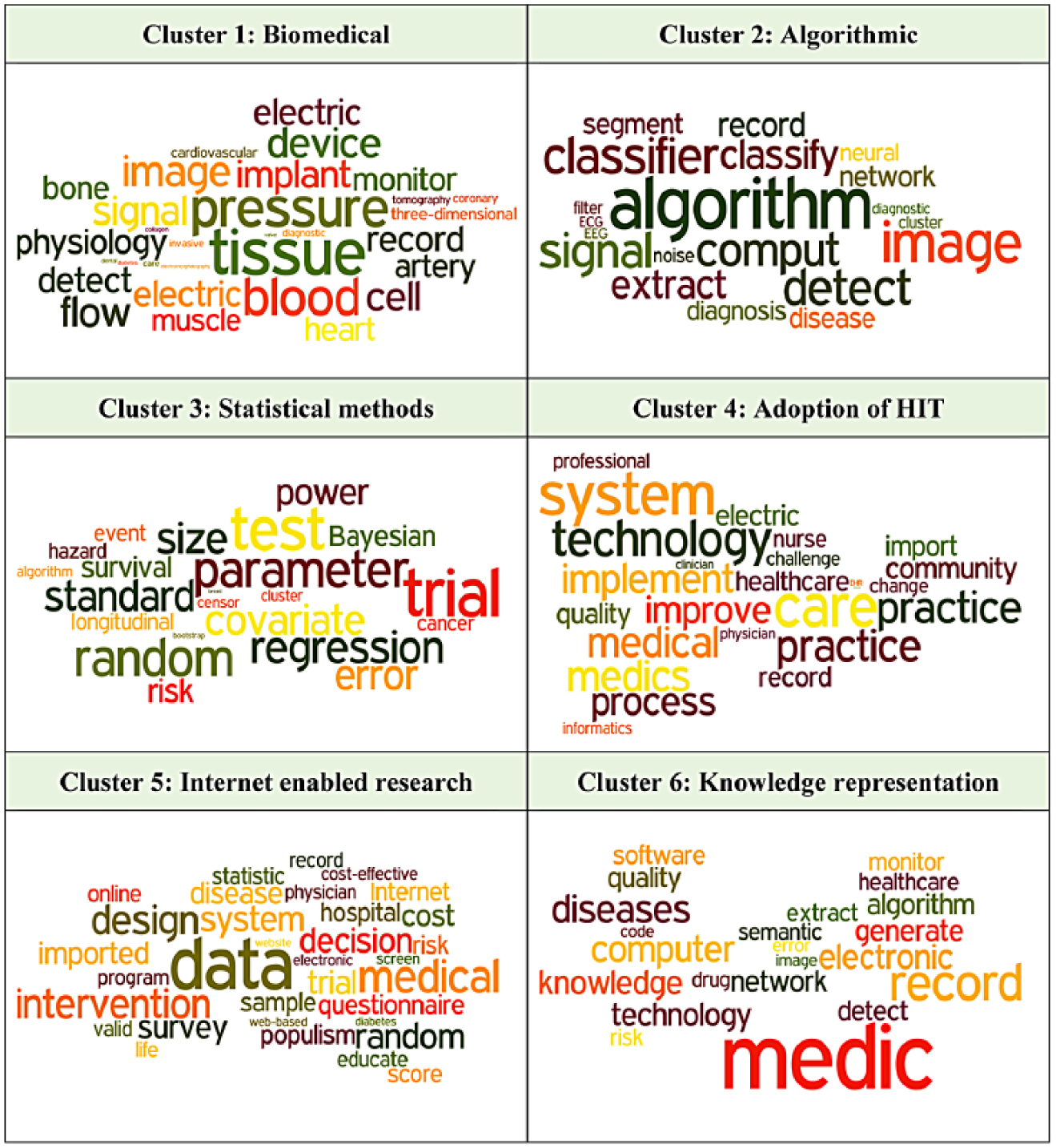
Frequently surfacing words in cluster.
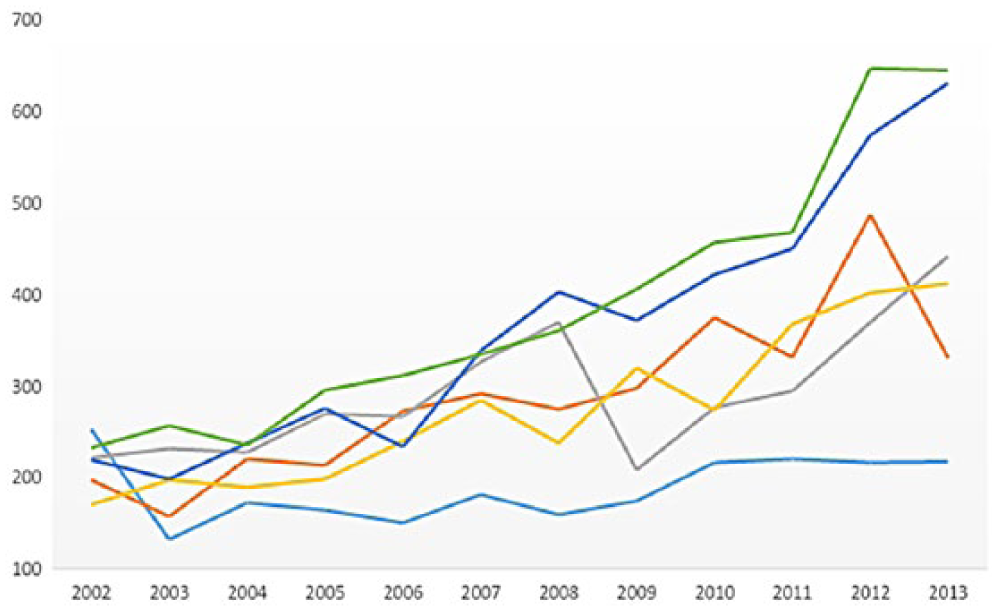
Research trends by count.
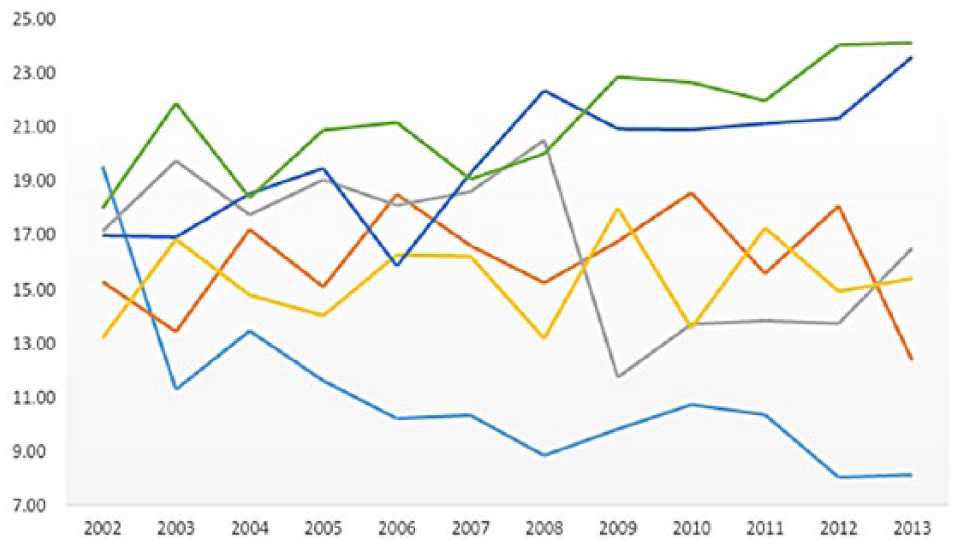
Research trends by percentage.
The words in Figure 5 are captured using the entire collection of abstracts of each cluster. As with other cluster analyses, each word in each cluster is not totally independent. For example, the two words of “blood” and “pressure” in Cluster 1 can be two independent words or a combined term. Therefore, the multi-word selections are usually identified as a single unit and are called
Cluster 1: biomedical
The most frequently surfaced words in Cluster 1 include tissue, pressure, blood, image, flow, device, and signal. Those words propose that medical informatics research in this cluster directly deals with utilizing the capability of technology to experiment in order to improve patient treatments. More specifically, the term “tissue” is used in the context of tissue differentiation in fractured vertebra with and without fixation devices, the numerical optimization of gene electrotransfer into muscle tissue for minimizing muscle tissue damage, the simulation of tissue differentiation and bone regeneration, and a patient-specific tumor and normal tissue for prediction of the response to radiotherapy.27–30
The major context of “pressure” is used in conjunction with blood and artery. Included examples are blood pressure long-term regulation, oscillometric measurement of systolic and diastolic blood pressures, a procedure for the evaluation of non-invasive blood pressure simulators, neural set points for the control of arterial pressure, the estimation of mean arterial pressure from the oscillometric cuff pressure, and the feasibility of measuring coronary blood flow.31–36
The term “image” is mainly used in the context of guiding treatments. Some examples include image-guided oral implantology, design of the image-guided biopsy marking system for gastroscopy, extraction of three-dimensional (3D) information on bone remodeling in the proximity of titanium implants in SRmuCT image volumes, 3D patient-specific geometry of the muscles involved in knee motion from selected magnetic resonance imaging (MRI) images, and jaw tissues segmentation in dental 3D computed tomography (CT) images using fuzzy-connectedness and morphological processing.37–41
Other frequently appearing words are “electric,” “device,” and “implant.” This category of research includes electromagnetic interference with active implantable devices, electromagnetic interference of cardiac rhythmic monitoring devices to radio frequency identification, implantable devices for long-term electrical stimulation of denervated muscles, and computer methods for automating preoperative dental implant planning.42–45
These results reveal that medical informatics research in this category mainly deals with discovering knowledge and improving the ability to treat patients through experimentation. The research by count in this category receives a stable academic interest over time in Figure 5. However, Figure 7 shows that this type of research had the highest percentage of interest (19.52) in 2002, yet its academic interest has decreased since that time. This was especially the case when it abruptly dropped in 2003. In fact, since 2003, the number of publications within this discipline has shown to be the lowest among the six categories. This indicates that the recent application of technology in the medical field is used for much more than just understanding a patients’ physical body.
Cluster 2: algorithmic
Cluster 2 captures many computer-aided technical languages such as algorithm, image, signal, comput [compute, computer], classifier, classify, extract, detect, disease, clinic, record, segment, and network. Those clustered words propose that scholars in this cluster use algorithms and neural networks to extract images and detect diseases, and the collected images and recorded data are further classified in order to better diagnose diseases. For example, an algorithm is used to classify images, genes, micro-array data of ovarian cancer, epilepsy diseases, and an effective cardiac arrhythmia.46–51 Algorithms are also frequently used to analyze signals. 52
Computerized images are also used to diagnose tumors, cancer, bone disease, and dental treatments. Those collected images are further analyzed and classified. Examples are bone disease classification using collected 3D image analysis and the artificial intelligence diagnosis of dental deformities in cephalometry images using a support vector machine.53,54 Neural networks or computers are used to record and categorize patterns using the collected and recorded information, which in turn is the basis for data mining analysis and anomaly detections. Examples are electroencephalogram (EEG) recordings for a better description of sleep, automatic classification of long-term ambulatory electrocardiogram (ECG) records according to type of ischemic heart disease, automated detection of neonate EEG sleep stages, epileptic seizure detection in EEGs using time-frequency analysis, central sleep apnea detection from ECG-derived respiratory signals, a detection of Alzheimer’s disease using independent component analysis (ICA)-enhanced EEG measurements, and epileptic EEG signal detection using time-frequency distributions.55–61
The main subject matter in this cluster is the utilization of algorithms, neural networks, and computational technology to group or categorize diagnoses or symptoms so that one can discover patterns of symptoms and identify anomalies. Figure 5 shows that the number of publications has increased; however, in terms of relative research interests compared to the other clusters in Figure 6, it shows a stable research interest over time.
Cluster 3: statistical methods
Frequently surfaced words in Cluster 3 are trial, test, random, parameter, regression, standard, error, covariate, size, Bayesian, power, risk, and longitudinal. Those clustered words suggest that the medical informatics research in this group mainly deals with various statistical methods applied to medical trials. This may be the reason that “trial” and “test” appear most frequently in this cluster. Also, terms such as standard, error, covariate, size, and power are all closely related to statistical analyses. A few example articles derived from Cluster 3 are further discussed below.
Statistical methods applied to medical informatics research are Bayesian strategies for monitoring clinical trial data, Bayesian analysis of multicenter trial outcomes, Bayesian approach to phase I cancer trials, techniques for incorporating longitudinal measurements into analyses of survival data from clinical trials, estimation and testing based on data subject to measurement errors, regression analysis for multiple-disease group testing data, power and sample size calculation for log-rank testing with a time lag in treatment effect, joint modeling of multivariate longitudinal measurements and survival data with applications to Parkinson’s disease, and regression analysis for the examination of the benefits of group testing.62–69
The trend of this research group shows an interesting pattern. It steadily increased from around 12 percent in 2002 to 21 percent in 2008 and then rapidly dropped to about 12 percent in 2009; however, the research interest of this group steadily increased since then.
Cluster 4: adoption of HIT
The frequently surfaced words in medical informatics research for Cluster 4 are system, care, technology, implement, improve, practice, process, medics, and medical. As the frequently surfaced words suggest, the research in this cluster mainly deals with the adoption and effectiveness of HIT including electronic patient record systems, electronic prescriptions, data sharing, and electronic reminders in a healthcare setting. A few examples are the determinants of primary care nurses’ intention to adopt an electronic health record in their clinical practice, introduction of eHealth to nursing homes, implementation of health information exchange for public health reporting, HIT implementation in critical access hospitals, hospital implementation of HIT and quality of care, improvement of HIT adoption and implementation through the identification of appropriate benefits, integration of a nationally procured electronic health record system into user work practices, and prediction of the influence of the electronic health record on clinical coding practice in hospitals.70–77
Another group of frequently surfaced terms are “process” and “improve.” A major context of those words is commonly captured with the adoption of HIT: the process of developing technical reports for healthcare decision makers, the role of methodologies in improving the efficiency and effectiveness of care delivery processes, the measurement of healthcare process quality, and a process for the consolidation of redundant documentation forms.78–81 Also examined is the effectiveness of HIT for patient care delivery, as well as the communications between nurses, physicians, and patients.82,83
In sum, this group of research deals with the effective adoption and use of HIT and EMR, and the question of whether or not HIT improves quality. Although some fluctuations are observed within the sample period, they are within the range of 13–18 percent. As such, this topic shows a relatively stable research interest during the research period.
Cluster 5: Internet-enabled research
Cluster 5 is composed of data, medical, intervention, design, decision, system, random, survey, cost, trial, Internet, and online. Based on the frequently surfacing words, this group of medical informatics research utilizes the Internet or online services to improve quality care, treat patients, and use online resources for medical research. More specifically, design and intervention are often used in conjunction with research design; however, unlike Cluster 3, this group of research utilizes the Internet. A few examples are the design of a website on nutrition and physical activity for adolescents, an Internet-based intervention to promote mental fitness for mildly depressed adults using a randomized controlled trial, evaluation of a community-based intervention to enhance breast cancer screening practices, and technology-based interventions for mental health in tertiary students.84–87
The Internet is also used for interventions to promote seeking treatment for mental health, online alcohol intervention, and online intervention for a health behavior change campaign.88,89 Further examined using the Internet in this cluster is sample randomization. Included examples are the accuracy of geographically targeted Internet advertisements on Google AdWords for recruitment in a randomized trial, recruitment to online therapies for depression using a pilot cluster randomized controlled trial, comparison of questionnaires via the internet to pen-and-paper in patients with heart failure, and reach, engagement, and retention in an Internet-based weight loss program in a multi-site randomized controlled trial.90–93
Based on the research in this cluster, the Internet and the web revolutionized the ways in which medical informatics’ research is conducted. This group of research concerns how the healthcare industry can better leverage the Internet and the web in order to provide better treatments for patients. This group of research consistently increased from 2002 until 2013. Since 2009, it has become one of the two clusters which contain the most prominent topics. This may be attributed to the general public’s accessibility of the Internet, their skill sets, the demands from the public to deliver information online, and cost-saving pressures.
Cluster 6: knowledge representation
Cluster 6 includes medic, record, diseases, computer, electronic, knowledge, technology, and network. As the frequently surfacing words propose, this cluster of research mainly deals with strategic use of collected medical data such as EMR/EHR to improve quality and reduce errors. A few commonly appearing subjects are the use of EMR to enhance detection and reporting of vaccine adverse events, the use of EMR data for quality improvement in schizophrenia treatment, and the discovery of notifiable diseases using EMR.94–97
Text mining is actively utilized to categorize various diseases. Examples are medical text classification for the Vaccine Adverse Event Reporting System, semantic classification of diseases in discharge summaries using a context-aware rule-based classifier, and automated evaluation of electronic discharge notes to assess quality of care for cardiovascular diseases using Medical Language Extraction and Encoding System (MedLEE).98–100 Also, using EHR clinical notes on heart-related symptoms (the Framingham heart failure diagnostic criteria), 101 used a text mining technique to detect early heart failure and improved the understanding of the early detection of heart failure. Data mining is also used for clinical oral health documents to analyze the longevity of different restorative materials. 102 Some of the articles explicitly use the term knowledge discovery. Examples are knowledge-based biomedical word sense disambiguation using document classification, the role of domain knowledge in automating medical text report classification, Mayo clinical text analysis and knowledge extraction system, a knowledge discovery and reuse pipeline for information extraction in clinical notes, and providing concept-oriented views for clinical data using a knowledge-based system.103–107
In sum, this cluster of research deals with utilizing EMR/EHR for categorizing or discovering treatment-related knowledge. Also, data and text mining is used in order to discover knowledge buried within text and data. Like Cluster 1, this group of research strives to find better treatments, but this research group utilizes EMR/EHR. This group of research also adopts text and data mining techniques to categorize diseases. The academic interest of this cluster is consistently increasing as each year goes by and has become one of the two most researched topics since 2008.
Discussion
Interestingly enough, the counts and the percentages of each cluster in Figures 5 and 6 between 2002 and 2004 did not show notable differences across clusters; however, distinctively different patterns among clusters began to emerge in 2005. The most notable publication increases are in the clusters,
Conclusion and future research
This study explored the trends of medical informatics research using articles published between 2002 and 2013. The main objective of this study was to understand where the field of medical informatics has been, where it is heading, and identify a boundary of the field as its subject area has matured as an academic field. According to the findings, as with any other field that goes through different academic interests in different times, an increase or decrease in medical informatics publications corresponds to the needs of the field and the opportunities enabled by the capabilities of HIT. Despite the advantages, the utilization of EMR/EHR data is still in a nascent stage that is necessary for the support of evidence-based medicine, 7 and thus we anticipate continual and rapid growth of Internet-based and data-driven, evidence-based research.
The purpose of this study was to identify the scope and the trend within this field. In order to achieve these research objectives, we focused on big streams of research. As such, for future research, it is recommended to investigate smaller clusters, which can provide insights on emerging fields of study.
The sample of this study is drawn from the major medical informatics journals provided by the ISI’s Web of Knowledge JCR 2012. Because smaller journals often publish innovative or non-traditional ideas about the field, in future studies, it is recommended to include smaller journals in the field and investigate how those journals integrate new emerging ideas, and by doing so, define/redefine the field.
This study did not include the statistical significance of the changes in publications over the study period in order to focus on the identification of the scope and boundary of the field. It will be, however, an interesting idea to calculate statistical significances of the publication changes over the time period and discover what drives such changes.
This article has some limitations. Because we chose 23 journals as a representative sample, the research findings do not collectively represent all medical informatics journals. Therefore, readers need to be cautious when they apply these research findings to a bigger sample or a different sample drawing.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
