Abstract
There is a growing body of literature on process mining in healthcare. Process mining of electronic health record systems could give benefit into better understanding of the actual processes happened in the patient treatment, from the event log of the hospital information system. Researchers report issues of data access approval, anonymisation constraints, and data quality. One solution to progress methodology development is to use a high-quality, freely available research dataset such as Medical Information Mart for Intensive Care III, a critical care database which contains the records of 46,520 intensive care unit patients over 12 years. Our article aims to (1) explore data quality issues for healthcare process mining using Medical Information Mart for Intensive Care III, (2) provide a structured assessment of Medical Information Mart for Intensive Care III data quality and challenge for process mining, and (3) provide a worked example of cancer treatment as a case study of process mining using Medical Information Mart for Intensive Care III to illustrate an approach and solution to data quality challenges. The electronic health record software was upgraded partway through the period over which data was collected and we use this event to explore the link between electronic health record system design and resulting process models.
Introduction
Process mining 1 is an emerging approach for discovering and analysing business processes based on event logs extracted from information systems. An event log in an electronic health record (EHR) contains records of timestamped events that have taken place to the patients during their treatment. The approach has been applied to the analysis of healthcare processes 2 with the aims of improving quality of care, patient safety, and optimisation of resources. 3 Healthcare, with its emphasis on the quality (as well as the cost) of outcomes, is arguably more complex than many other industries. Many healthcare organisations have been slow to adopt large-scale information systems to manage and record their clinical data and activities but such EHR systems are becoming more common 4 and have the potential to provide a rich source of insights for process improvement. 5 The data are, however, sensitive and confidential 6 creating unique challenges.
Reviews of process mining in the general healthcare literature 7 and more specifically in cancer care 8 have reported common issues gaining approval for access to data, difficulty linking data from multiple sources, constraints due to anonymisation, challenges accessing domain experts to help understand business processes, and concerns over data quality. Healthcare processes are recognised as complex because the steps are nonlinear, unpredictable, and dependent on each individual’s medical and personal circumstances – such processes often not follow standard sequences. 9 Data quality issues have direct implications for the quality of healthcare provision. 10 For researchers, detailed understanding of data quality in health records is essential if we are to use them to draw conclusion about healthcare provision. 11 Data quality issues may be explained in terms of underlying bias in the way that users interact with the system. 12 Several studies have explored data quality issues related to process mining in healthcare,2,13–15 and one study has even used process mining as an approach to better understand healthcare data quality, 16 but none of them address the Medical Information Mart for Intensive Care III (MIMIC-III) data quality issues for healthcare process mining. The relationship between healthcare processes and data quality or bias was examined by Agniel et al. 12 using only lab test orders. In our article, we address the data quality issues across the full range of data types from a freely available EHR data so that other researchers can also explore this complex issue.
Weiskopf and Weng 17 developed a framework for assessing data quality in electronic health records using five dimensions linked to seven methods used to address quality issues. This framework has informed our approach. Our own experience of health data quality was shaped by work on process mining chemotherapy care data from a large UK cancer centre which took eight iterations surfacing data quality issues for domain expert review over 9 months with each iteration revealing unanticipated further complexity. 18 We developed a healthy respect for the data quality challenges in healthcare. Our approach applying the Weiskof and Weng framework has been to keep careful track of extract criteria and transform data manipulations before loading to standard analytics tools so that strategies to deal with data quality are documented and can be refined through multiple iterations.
One solution to progress methodology development is for process analytics researchers to try out their methods using a freely available research dataset so that they can compare results. In this article, we describe the potential to use the MIMIC-III dataset for process mining in healthcare. The dataset used in this research is MIMIC-III v1.3 released on 10 December 2015 (available at https://mimic.physionet.org). MIMIC-III is a freely available critical care database which contains the de-identified records of 46,520 intensive care unit (ICU) patients at the Beth Israel Deaconess Medical Centre (BIDMC) in Boston, USA, covering 58,976 admissions between June 2001 and October 2012. The data were largely sourced from the hospital’s Philips CareVue (CV) EHR until that system was replaced in 2008 by a new EHR called iMDSoft MetaVision (MV). Our article aims to (1) highlight the opportunities for using MIMIC-III for healthcare process mining, (2) provide a structured assessment of MIMIC-III using the Weiskopf and Weng 17 EHR data quality framework, and (3) explore the potential impact of the change in the hospital EHR software in 2008 on data quality and our confidence in resulting process models. The method for de-identification of the data presents several challenges for process mining which we discuss. We provide a worked example of cancer treatment as a case study in process mining to illustrate an approach to data quality challenges.
This article is structured as follows: section ‘Using the MIMIC-III data for process mining’ describes the description of the MIMIC-III used for process mining, Section ‘Methodology’ presents the followed methodology and the addition of a new stage. Section ‘Results’ illustrates the application of the proposed method, the quality assessment and the analysis of the potential impact of the system changes. Finally, section ‘Discussion’ and section ‘Conclusion’ describe the discussion and conclusions.
Using the MIMIC-III data for process mining
The MIMIC-III data are available as a set of downloadable files which can be used to create a relational database with 26 tables. 19 In our previous works,20–22 examples of process mining using MIMIC-III have been published, some details of the data structures relevant to process mining were given, and methods to use MIMIC-III database for process mining were implemented. There were problems with data quality mentioned, but there was no comprehensive discussion on data quality assessment. In this article, we examine the methods for using MIMIC-III and provide a structured data quality assessment. The MIMIC-III dataset contains medical event data such as vital signs, medications, laboratory results from hospitals and clinics, charted observations during a patient’s stay in the ICU, and other clinical notes including nursing notes and discharge summaries. 19 Patients were included in MIMIC-III if they had at least one period of care in an ICU but the data available cover all of their hospital data for all episodes, it therefore provides a comprehensive example of EHR data from a large and busy hospital.
The data available to researchers have been curated by the MIMIC-III team at the MIT Lab for computational physiology to (1) address known data quality issues, (2) reconcile differences in the data formats of the Philips CareVue (CV) EHR and iMDSoft MetaVision (MV), and (3) de-identify the data to make it suitable for external research use. 19 The methodology for anonymisation was in accordance with US Health Insurance Portability and Accountability Act (HIPAA) recommendations to remove all personally identifiable data (names, addresses, phone numbers, etc.) with one important exception. HIPAA recommends the removal of event dates and times on the basis that these present unique patterns that could aid re-identification – such an approach would have prevented process mining. The approach to anonymisation used in MIMIC-III was date shifting. All dates were shifted into the future (between 2100 and 2200) by a random offset generated for each patient. Time of the day, day of the week, and approximate seasonality were conserved during date shifting. Time in the MIMIC-III database is stored with one of two suffixes with different resolutions: time (down to the minute) and date (down to the day). Most data are recorded with a time indicating when the observation was made (charttime) and when it was validated (storetime).
The MIMIC-III dataset does not contain an event log but 16 out of 26 tables do contain timestamped information which can be used to construct the sequence of events for a patient. Tables are linked by identifiers: subject_id refers to a unique patient, hadm_id to a unique admission, and icustay_id to a unique ICU stay. Process mining focuses on event data but other tables provide supplementary data. For example, when the chartevents table is used, we need to refer to d_item table to get the corresponding label of item_id. Diseases and procedures are encoded using the International Classification of Diseases Version 9 (ICD-9) codes, and the mapping for these can be found in diagnoses_icd and procedures_icd tables.
One de-identification step was to obstruct identifiable times through random date shifting and this creates a challenge and limitations for process mining. Specifically if dates have been shifted by a random amount, workflow analysis looking at busy days is not meaningful. 2 In the MIMIC-III, dates have been shifted so that they are internally consistent for the same patient, but randomly distributed in the future. Similarly, the impact of bottlenecks, for example, of patients waiting for care on a busy day, cannot be deduced. Despite the date shifting, the MIMIC-III dataset still contains detailed information of real healthcare processes for individual patient during their time in the hospital including comprehensive details on administrative activities (admission, discharge, transfer to a ward, etc.) and clinical activities (triage, test and scans, diagnosis, etc.). 23 These data can reveal much about the care processes as they are experienced by each individual patient.
Methodology
Our goal to understanding suitability of the MIMIC-III dataset for process mining in healthcare is focused on the assessment of data quality. The process mining methodology in this research follows the L* lifecycle model, suggested by Van der Aalst et al. 1 but with some adaptation. The standard L* model consists of five stages: plan and justify (Stage 0), extract (Stage 1), create a control-flow model and connect it to the event log (Stage 2), create an integrated process model (Stage 3), and provide operational support (Stage 4). Stage 4 of the L* lifecycle model is only relevant for researchers who can influence the operation of the organisation and was therefore beyond the scope of our study.
We needed to introduce an additional Stage between Stage 0 and Stage 1, to prepare the MIMIC-III data for process mining. This new Stage involved the preparation steps to reconstruct the MIMIC-III data files into a relational database. This exercise helped us understand the underlying data model before Stage 1 and informed further iterations (see Figure 1). In earlier work, we develop a question-driven methodology for process mining in healthcare 24 and we used this approach to motivate our investigation.
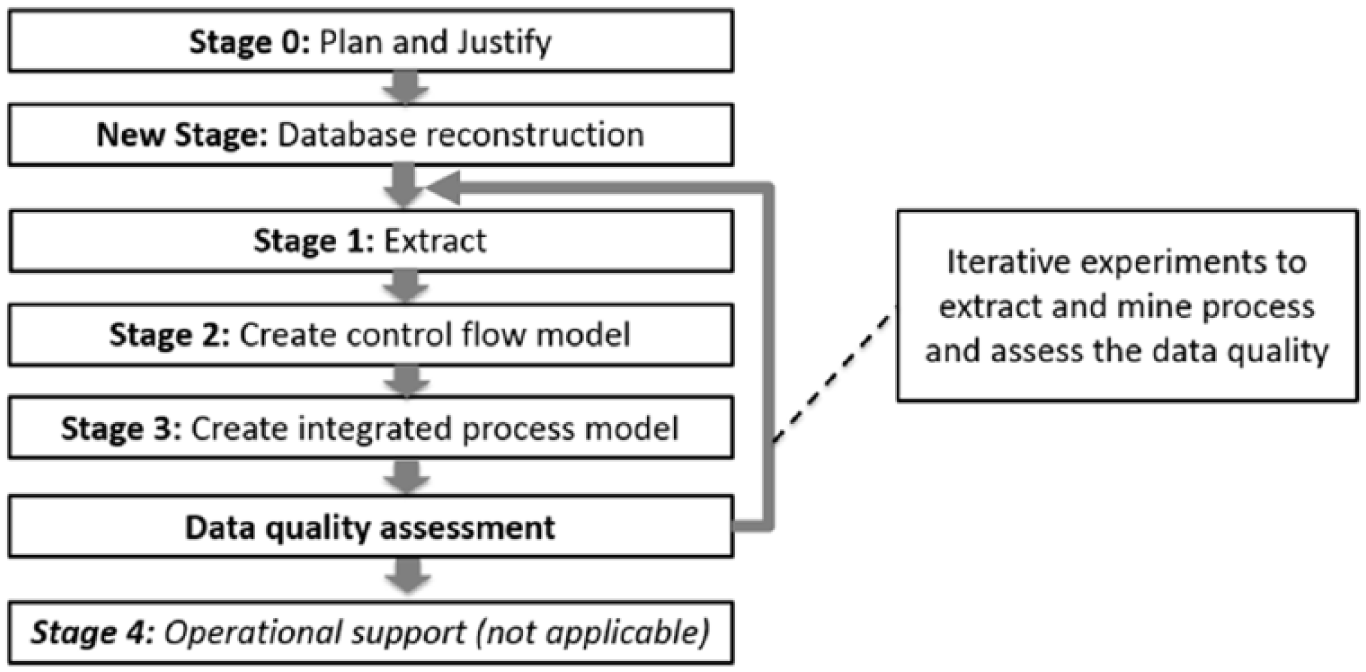
Adaptation of the L* life-cycle model to include data quality assessment.
Our assessment of data quality follows the Weiskopf and Weng 17 framework with a specific focus on the impact of data quality for process miners. The data quality assessment would mark various issues in the process characteristics and the quality of an event log for process mining research.2,13 In term of process characteristics, EHR databases including MIMIC-III face common challenges due to the voluminous data – large number of cases and events, case heterogeneity – large number of distinct traces, event granularity – large number of distinct activities, process flexibility, and concept drifts. Those problems would be identified by showing a few basic metrics such as the number of cases in the event log, the number of unique activities, and the number of distinct traces. While the quality of the event log would be discussed by identifying four broad categories of problems, which are missing data, incorrect data, imprecise data, and irrelevant data. Those four categories would be applied to check problems in case, event, activity name, and timestamp. Methods and dimensions in the Weiskopf and Weng data quality framework could be viewed as the first level quality assessment with the details being the process mining – specific metrics.
The implementation of the L* lifecycle is presented in section ‘Process mining with the MIMIC-III dataset’, with a worked example of process mining using MIMIC-III. Data quality assessment is presented in section ‘Data quality assessment of MIMIC-III for process mining’. The effects of the EHR system changes are presented in section ‘How does the 2008 change in the EHR system affect data quality?’. The current tools used by our group are (1) relational database: PostgreSQL version 9.5 using PgAdminIII for SQL-based data extract, (2) data transformation: Python programming language, (3) process mining: the ProM toolkit version 6.4.1 as a standard process mining toolset, (4) experiment documentation: standard templates in Microsoft Word, (5) version management: GitHub repository to support rapid and iterations (this project is at https://github.com/apkurniati/mimic3qualityforpromin).
Results
Process mining with the MIMIC-III dataset
Stage 0: plan and justify
The justification for this research is based on recognising a need for a high-quality, freely available research dataset that can be used internationally to progress process mining methodology development. Planning for this study was based on three research questions:
Q1 was addressed by presenting a cancer-specific worked example in this article to illustrate our approach to data quality in sections ‘New stage: database reconstruction’ to ‘Stage 2: create control flow model and connect event log’. Q2 will be addresses by applying Weiskopf and Weng’s 17 EHR data quality assessment framework in section ‘Data quality assessment of MIMIC-III for process mining’. Q3 will be addressed by investigating the differences in processes and data quality of the CareVue (CV) and MetaVision (MV) systems in the MIMIC-III dataset in section ‘How does the 2008 change in the EHR system affect data quality?’.
New stage: database reconstruction
This stage included the reconstruction of the MIMIC-III dataset to create a relational database in PostgresSQL. This included downloading 26 csv files (more than 6 GB in total) along with scripts to import the data into the PostgreSQL database following the guidelines at https://mimic.physionet.org/gettingstarted/dbsetup. The concept-level entity-relationship (E-R) diagram for the resulting database is presented in Figure 2. The interpretation of events and their significance was based on an understanding of the entities and their relations described in the data descriptors paper
14
. It is possible to extract specific table(s) in the MIMIC-III database for process mining, but our approach was to reconstruct and explore the fullest possible set of data and judge relevance in later stages. This was done to maintain the data quality for the next stages, which are the extract, create control-flow model, create integrated process model, and data quality assessment stages. The entities in bold contain
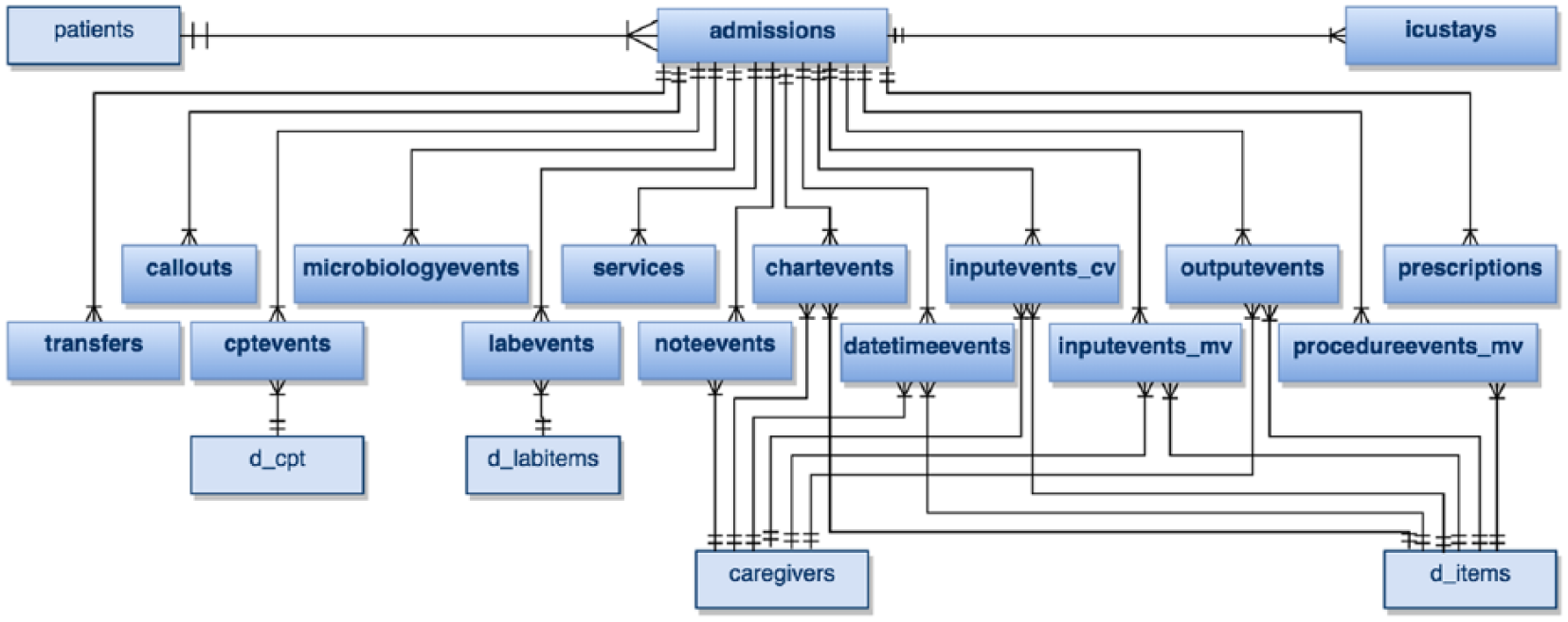
Concept-level entity-relationship (E-R) diagram of the MIMIC-III database.
Stage 1: extract
The database reconstruction supports multiple possible extractions so that Stage 1 can be repeated many times. In all experiments, we followed a standard documentation template. In the example below (Figure 3), we select records of cancer patients in the admissions table based on the ICD-9 codes for cancer diagnoses which are 140x-239x 25 creating an event log of [case_id, activity, timestamp]. The result table was then saved in .csv format and loaded into ProM or other available process mining tools.
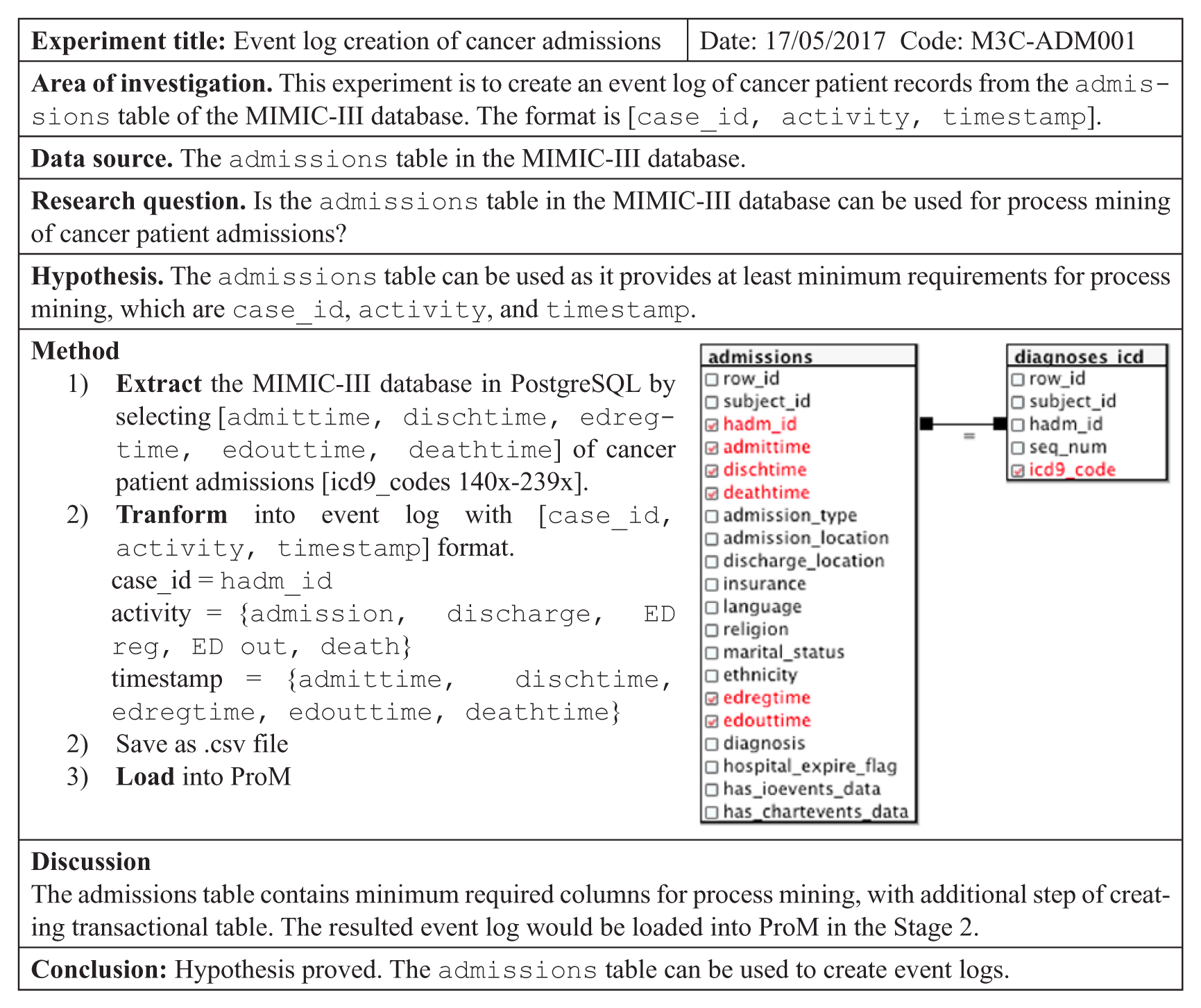

Example of experiment documentation for a simple extract.
Stage 2: create control flow model and connect event log
The next stage in the L* model is to create control flow models and link these to the event log. For this example, three main plugins were used: (1) ‘
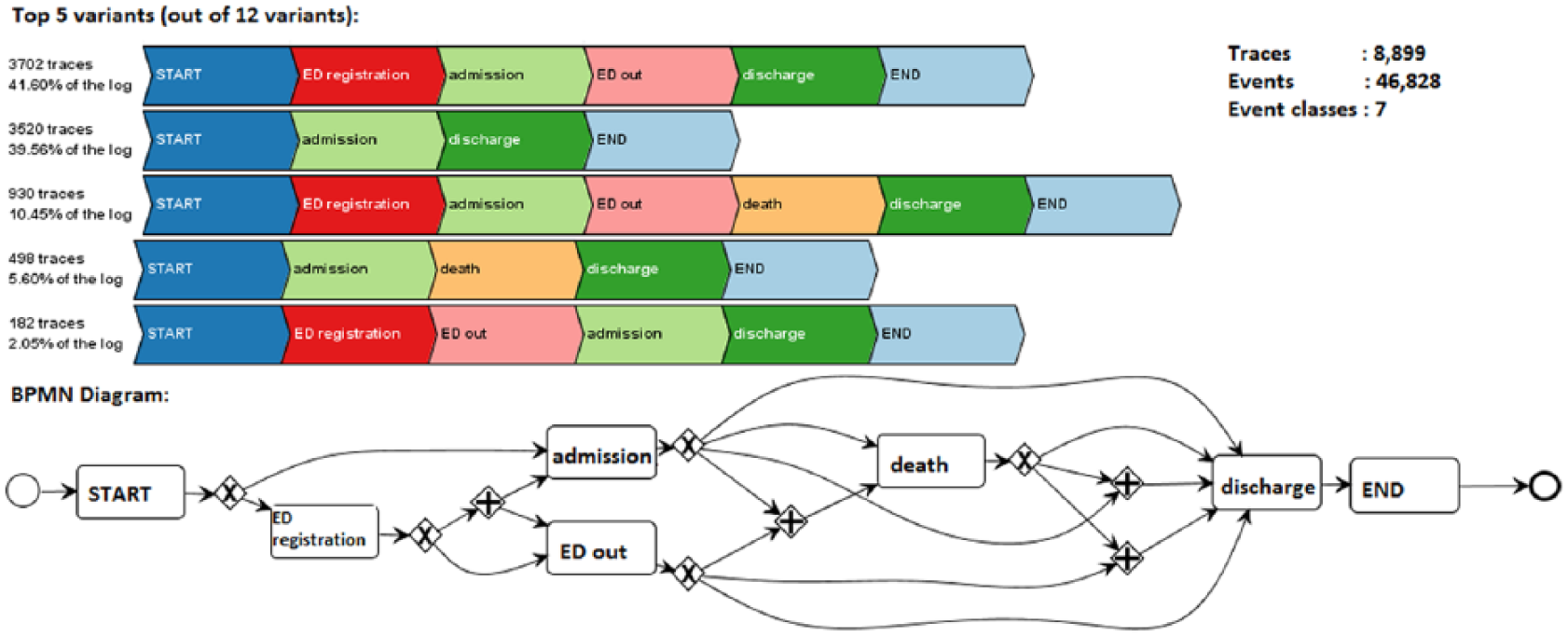
Five most common variants and BMN process model of admissions.
The top part of Figure 4 represents the five most common variants out of 12 variants. The process model in the bottom part of Figure 4 was in Business Process Modelling Notation (BPMN). This process model fitness was 100 per cent, precision was 81.713 per cent and generalisation 85.275 per cent. We then reviewed the resultant model with a UK-based oncologist for sense checking. The apparent data quality issue is that discharge took place after death was investigated and was found to reflect the hospital’s standard administrative process. There are also variations in administrative steps. For example, when patients were being admitted (admission) after discharged from the emergency department (ED out), they might be admitted before ED out (in variant 1) or ED out before admitted (see variant 5). In variant 1, admission – ED out durations are 118 min in average with 89 min median. In variant 5, ED out – admission durations are 132 min in average with 46 min median.
Stage 3: create integrated process models
In Stage 3, the models are extended with other perspectives (e.g. time and resources). Our analysis studies the effect of the change of EHR and this is discussed in section ‘How does the 2008 change in the EHR system affect data quality?’.
Data quality assessment of MIMIC-III for process mining
Element presence
The first data quality assessment method is
Element presence checking: the event tables.
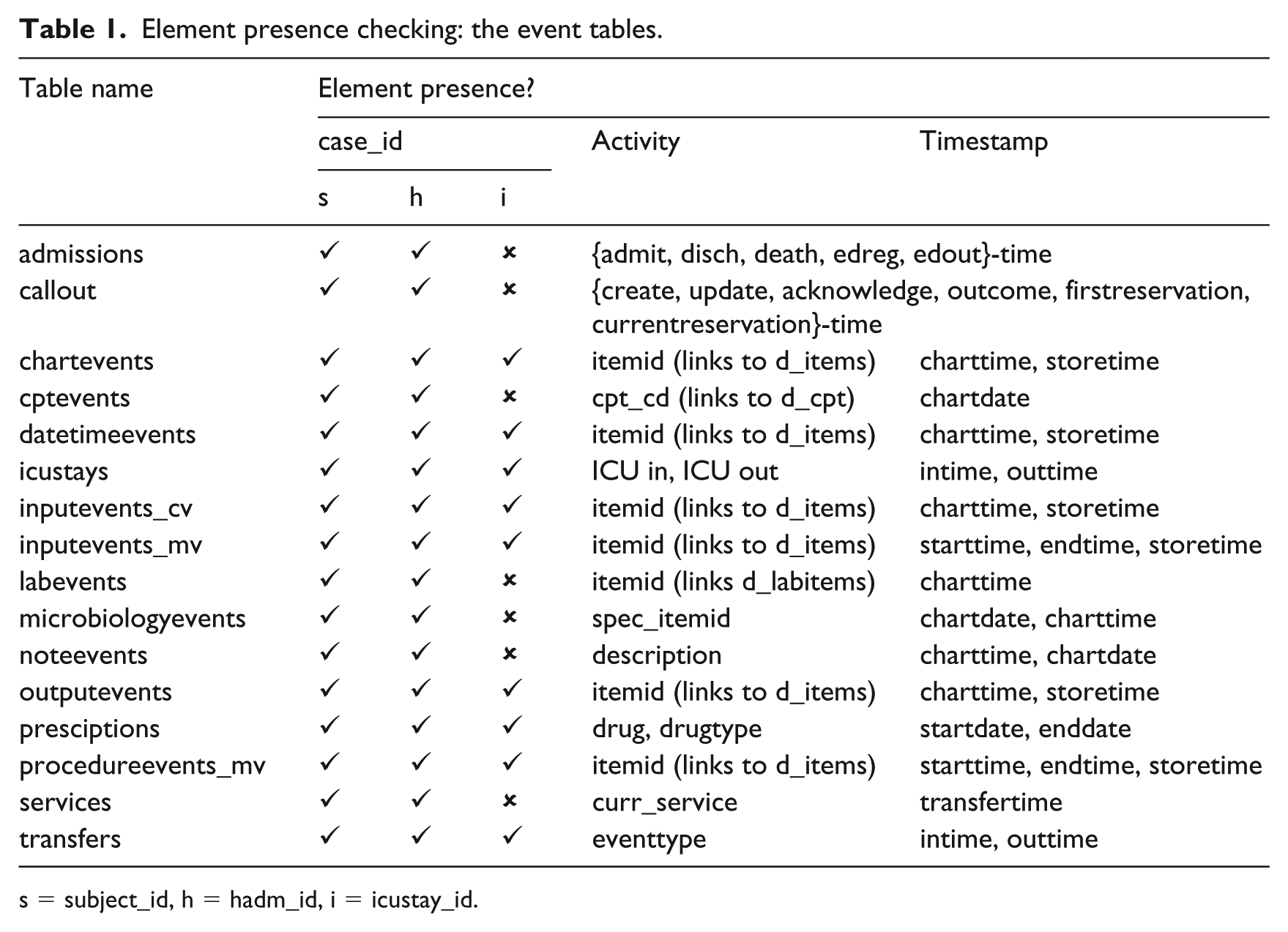
s = subject_id, h = hadm_id, i = icustay_id.
The MIMIC-III database provides three types of identifiers which can be used as
Data element agreement
(1) When we use subject_id or hadm_id as the case_id, the admissions and transfers tables are complete. But when we use icustay_id, only icustays and transfers tables are complete. All other tables can be used as needed, but the completeness can be checked by reference to those tables. (2) Time in the MIMIC-III database is stored as datestamps in three tables (cptevents, noteevents, and prescriptions) and as timestamps in the other tables (admissions, icustays, etc.). This might encounter issues in process mining when combining tables with different time/date stamps resulting in incorrect temporal order throughout a patient’s encounter. For example, all prescriptions are recorded with time 00:00 in the prescriptions table. Combining it with the admissions table which does have detailed timestamps would result in prescription appearing to precede the admission. (3) The MIMIC-III data descriptor said that the dates have been shifted into the future to the years 2100 to 2200 but we found some tables had records of events before and after those dates. This happens because some tables also contain historical patient data, for example test results from chemistry and haematology in labevents table. This presents no immediate data quality issues but could lead to errors if dates were used in selection.
Distribution comparison
Completeness checking.
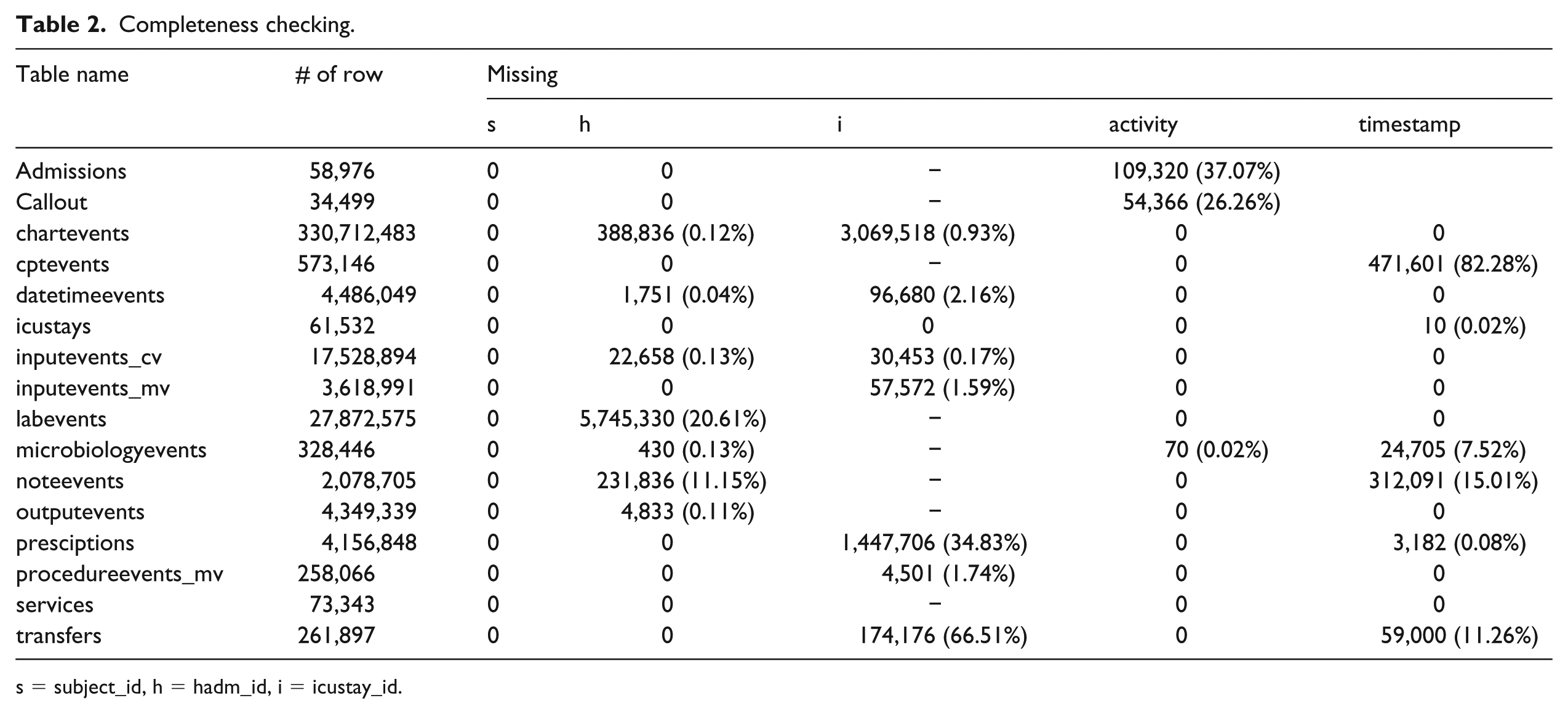
s = subject_id, h = hadm_id, i = icustay_id.
Table 2 shows that subject_id is complete in all event tables, but hadm_id and icustay_id are not. There are 70 missing spec_itemid in microbiologyevents table, but those can be replaced completely by spec_type_desc. In the admissions table, there are missing timestamps which represent patients who are not death or not admitted in the ED. This was illustrated in Figure 4. The callout table also has some missing timestamps (see Table 2). The MIMIC-III documentation mentions that the collection of callout data only began part way through the MIMIC-III database and with date shifting this missing data has been spread at random. This incompleteness should also be considered when using this table. Incomplete timestamps in the charttime of microbiologyevents and noteevents tables can be derived by linking to chartdate, but consider that the granularity level would be different. This is also happened in cptevents, prescriptions, and transfers tables so process mining would be unreliable.
Validity checking
The first finding was about the ICD-9 codes for diagnoses and procedures. The MIMIC-III dataset provides reference tables, which are d_icd_diagnoses and d_icd_procedures. But, there are 144 out of 14,711 (0.98%) missing codes in d_icd_diagnoses table, and there are 16 out of 258,082 (0.01%) missing codes in d_icd_procedures table.
The second finding was that there are duplicates between different tables, such as datetimeevents and admissions table, labevents and chartevents tables. For example, there is a hospital admit date in the datetimeevents table which is duplicated with admittime in the admissions table. The admissions table is specifically providing information of patient admissions in the hospital and has a complete set of admission and discharge times. Meanwhile, the datetimeevents table contained all date measurements about a patient in the ICU including hospital admission dates and it was found incomplete. Our investigation compared all records and found that there were 24,549 admissions recorded both in the admissions and datetimeevents tables with 1696 records are matched to the duplicate on the other table, but the others are not. In the 24,354 of 24,549 duplicated admissions (99.2%), admission dates in the admissions table were recorded earlier than those in the datetimeevents. Our suggestion to handle this issue was to use only the valid records and ignore the duplicated records in the other table(s). We suspect there are probably more issues like this with other tables.
How does the 2008 change in the EHR system affect data quality?
Our third research question specified in the Plan and Justify Stage in section ‘Stage 0: plan and justify’, aimed to identify the effects of the change in EHR system in 2008 from CareVue (CV) to MetaVision (MV). Specifically any impact on data quality. Because the MIMIC-III anonymisation used date shifting a simple comparison of dates was not possible. However, when MIMIC-III was constructed it was not possible to reconcile inputevents so there are separate inputevents_cv and inputevents_mv tables (the only separated tables for the two systems). Our approach was to work backwards from these two tables to identify which hospital admissions had been recorded on which EHR. We used this logic to insert a field in each table that indicated which of the two systems had been used and then used these new fields to extract event logs before and after the new EHR. The checking on itemid column in d_items table is presented in Table 3.
Details of itemid in d_item table.
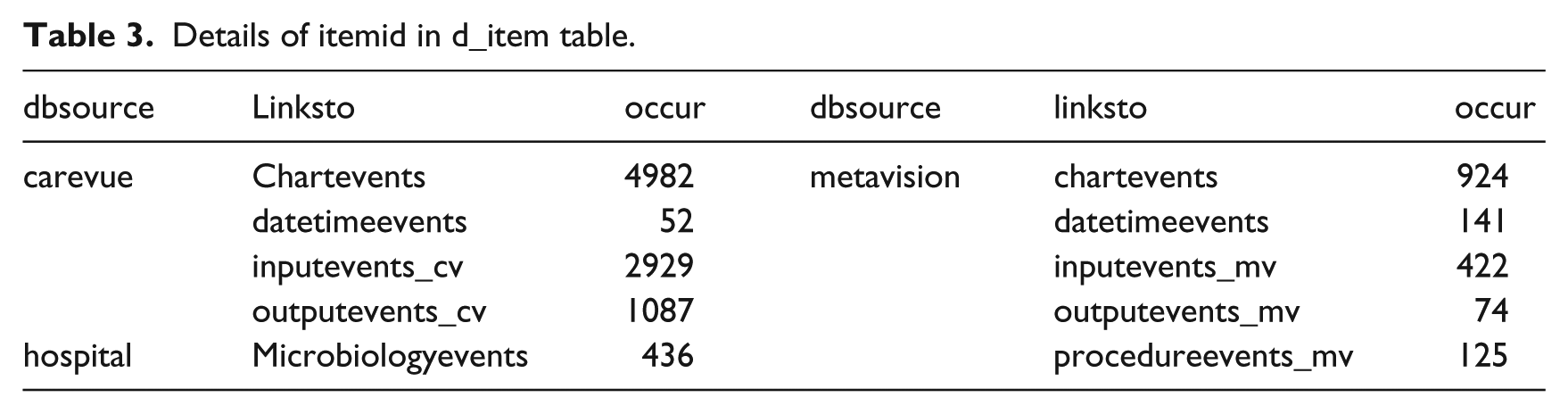
Table 3 shows that d_items table has a column to identify the database source (dbsource) and which tables link to those sources (linksto). This means that we could identify the differences between the EHR systems through four tables: chartevents, datetimeevents, inputevents, and outputevents. We could also identify the sources of each record in the other tables by referring to those four tables. The dbsource hospital would be ignored at this point because our focus was on the CV and MV systems used. Identifying the original EHR is possible because we marked each admission of cancer patients in the admissions table with CV, MV, or both (if the admission is recorded both in CV and MV) and created separate event logs from each data sources. We then compared the models discovered from CV and MV event logs using DifferenceGraph plugin in ProM. This plugin supports the identification of differences and commonalities between two process models. The result is presented in Figure 5.
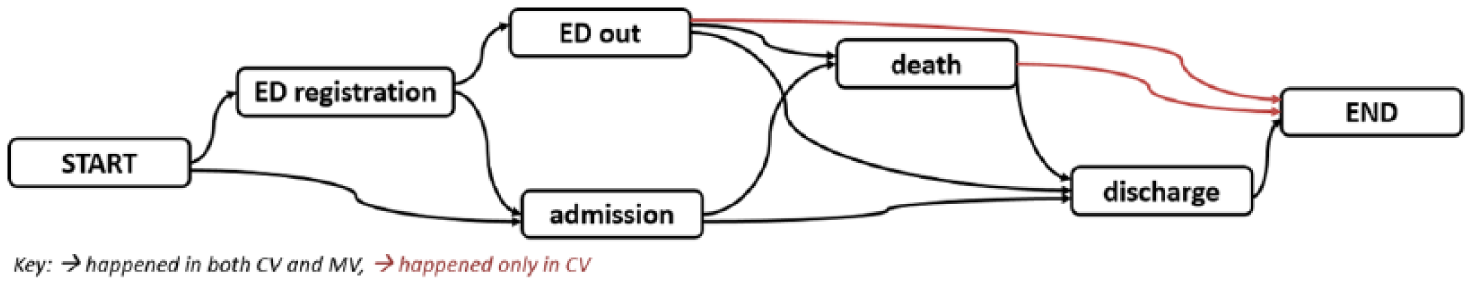
DifferenceGraph of admissions in CV and MV. It is evidence that in the new system, all patients have discharge event as the last event in the case.
Figure 5 shows the differences between admissions in CV and MV, which found changes in the last activities happened in those two systems. The admissions in CV (35833 admissions) ended with either discharge (35788/ 99.874%), death (26/ 0.073%), or ED out (19/ 0.053%). But all admissions in MV (19623 admissions) ended with discharge (100%), suggesting that MV has better administrative records than CV. The process model of CV fitness was 99.96 per cent, precision was 87.84 per cent, and generalisation was 90.06 per cent; while the process model in MV fitness was 99.99 per cent, precision was 93.82 per cent, and generalisation was 89.159 per cent. This suggested that both models are able to replay the observed behaviour (high fitness), describe the system generally (high generalisation), and not allow for too much behaviour (high precision). The combination of interpreting event frequencies, the DifferenceGraph and the conformance measures leads to the conclusion that the EHR system change did affect the process model and quality requiring further investigation.
Discussion
Process mining has been used in many case studies in healthcare data3,14,26 and MIMIC-III as a freely available de-identified healthcare database has been made available to researchers and used in many research projects. There are many opportunities for using process mining on MIMIC-III, but only three published examples so far.20–22 Agniel et al. 12 used lab test orders and linked data quality bias to evolving healthcare processes. In this article, we have explored the broader data quality issues that future process mining researchers will need to understand in order to use the full range of EHR data. We have provided a worked example of cancer treatment to illustrate this.
When using MIMIC-III for process mining, the L* lifecycle can be followed. However, an additional stage between Stage 0 and Stage 1 for database reconstruction is needed. Given that researchers using MIMIC-III are unlikely to have direct access to the Beth Israel Deaconess Medical Centre that provided the data, it should be clear that explicit operational support (Stage 4) is not applicable. In our worked example, we have demonstrated that the MIMIC-III dataset can be used for process mining with many interesting results. For example, detecting the hospital’s standard administrative process, investigating variations in the treatment steps, and visualising differences and commonalities between multiple process models. Such analysis can provide insights and learning opportunities for potential clinical improvements and help to progress methodology development in process mining. The ability to compare and publish against a freely available database (such as MIMIC-III) creates opportunities for international collaboration.
We assessed MIMIC-III using the completeness, plausibility, correctness, and concordance dimensions of the Weiskopf and Weng
17
EHR data quality framework. We did not assess the data by the
One important data quality issue in using MIMIC-III for process mining is the way that anonymisation has been achieved using a date shifting. All real dates have been shifted into the future by a random offset generated for each patient. For example, two patients seen on the same day may have admission dates of 2100-06-07 and 2185-04-08. This anonymisation approach makes it impossible to do some analysis in process mining. Public holidays such as Thanksgiving may well impact on processes but its effect will be distributed over a 100-year time period. Similarly, it is not possible to identify workflow looking at busy days, comparison of weekend versus weekday workloads, and bottleneck analysis of patients waiting time on a busy day.
A further challenge in the MIMIC-III database is the change of the EHR system that took place in 2008. The CV system was used in 2001–2008 and the MV system used in 2008–2012. The two systems have some data in different formats and the item ids differ. Our study found that the EHR system change did affect the process models and therefore quality of process mining. Our partial solution to this problem involves using the two different inputevents tables to mark the original EHR on the other tables on our copy of the database. This enabled us to create separate event logs from the two EHR and this made it possible to compare process models from the same process across the two different systems. Our comparison approach identified other differences and commonalities between process models in the two EHR systems in section ‘How does the 2008 change in the EHR system affect data quality?’. These differences and commonalities provide insights of the different process execution, specific activities, their order of executions, and those missing. These findings suggest that an awareness of EHR system changes is important to understand the root cause of some data quality issues.
Conclusion
This article has answered the three research questions developed in the initial stage of the study. First, we demonstrated the applicability of process mining using the MIMIC-III dataset and performed a data quality assessment. The additional stage in the L* lifecycle involved database reconstruction and made it possible to do Stage 1 to Stage 3 as many times as needed. We believe this iterative approach to identifying and understanding complex data quality concerns is essential. The use of a standard template for experiment documentation provides control over the iterative approach and helps document each data quality issue and how it can be mitigated in experiment design. We found that it was possible to mine complex hospital processes with existing techniques to analyse processes for different groups of patients. Second, we used the Weiskopf and Weng framework successfully to identify multiple data quality issues in section ‘Data quality assessment of MIMIC-III for process mining’ and ‘How does the 2008 change in the EHR system affect data quality?’. MIMIC-III has been used by EHR researchers to generate a range of insights and the data quality issues we have explored for process mining can therefore be assumed to be generally relevant to EHR research. Finally, we identified the effects of the change of the EHR system on the process and specifically on data quality, and we found evidence that the change of EHR did affect the process model and quality of results. Process mining researchers that have a concern about this may follow our technique to select records from either of the two EHR systems.
The most significant contribution of MIMIC-III to process mining is the opportunity to develop techniques which can be shared with and validated by other researchers and replicated against EHR dataset across the world. Future work will explore several aspects. To improve data quality using data cleaning strategies, including resolving missing data and aggregation of similar events. To apply different discovery algorithms to find process models with better conformance to the event log. To investigate the effect of EHR system changes on the healthcare processes. Finally, to focus on advanced analysis with more clinically based research questions from medical experts and comparison studies with real-life datasets.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship and/or publication of this article: This research is supported by the ClearPath Connected Health Cities Project and the Indonesia Endowment Fund for Education (LPDP).
