Abstract
Collaborative and multicenter studies permit a large number of patients to be enrolled within a reasonable time and providing the opportunity to collect different data. Informatics platforms play an important role in management, storage, and exchange of data between the participants involved in the study. In this article, we describe a modular informatics platform designed and developed to support collaborative and multicenter studies in cardiology. In each developed module, data management is implemented following local defined protocols. The modular characteristic of the developed platform allows independent transfer of different kinds of data, such as biological samples, imaging raw data, and patients’ digital information. Moreover, it offers safe central storage of the data collected during the study. The developed platform was successfully tested during a European collaborative and multicenter study, focused on evaluating multimodal non-invasive imaging to diagnose and characterize ischemic heart disease.
Keywords
Introduction
Collaborative studies in cardiology are a major source of information for establishing and evaluating clinical practice, since they permit a large number of patients to be enrolled within a reasonable time and provide the opportunity to collect a large amount of data. Multicenter studies are based on defining specific clinical protocols to guarantee the application of the same procedures for collection of comparable data. 1 Usually, the organization of a multicenter study involves the definition of the following three entities:
Each institution involved in the study may play one or more roles in the network, leading to a complicated exchange of several data between the centers. In general, informatics platforms are recognized to be important in management, sharing, and exchange of data collected during cooperative and multicenter studies.2–4 For this reason, the use of an informatics platform is considered essential for cardiac multicenter studies, currently based on a multimodal evaluation approach, including the acquisition of clinical information, biohumoral data, and multimodal imaging datasets. In particular, recent improvements in non-invasive modalities for coronary anatomy and functional correlate evaluations lead to a widespread use of these techniques. Indeed, coronary abnormalities can be evaluated using multislice computed tomography (CT), which also offers the possibility of evaluating and quantifying coronary calcium (calcium score (CS)) and detecting the non-calcified plaque components (computed tomography artery (CTA)),5,6 while the functional effects, assessed in terms of hemodynamic (myocardial perfusion) or mechanical (wall motion and myocardial contractility) consequences of coronary stenosis, can be evaluated using radionuclide imaging (positron emission tomography–single photon emission computed tomography (PET-SPECT)) 7 and magnetic resonance imaging–echocardiography (MRI-ECHO),8,9 respectively. Additionally, anatomical and functional non-invasive modalities can be integrated in order to provide a hybrid and three-dimensional visualization of coronary disease with the corresponding qualitative and quantitative functional information.10–14
The purpose of this article is to describe an informatics platform designed and developed to support collaborative and multicenter studies in cardiology. The performance of the developed platform was evaluated by applying it to the collection, sharing, storage, and management of data acquired during a European study (EVINCI—Evaluation of INtegrated Cardiac Imaging for the Detection, Characterization and Monitoring of Ischemic Heart Disease—grant agreement: 222915—ClinicalTrials.gov Identifier: NCT00979199).
Methods
Informatics network structure
A hub-and-spokes network was designed and developed to support data management during a collaborative and multicenter study (Figure 1). The hub, representing the coordination center, allows data sharing among all centers and is responsible for data management.
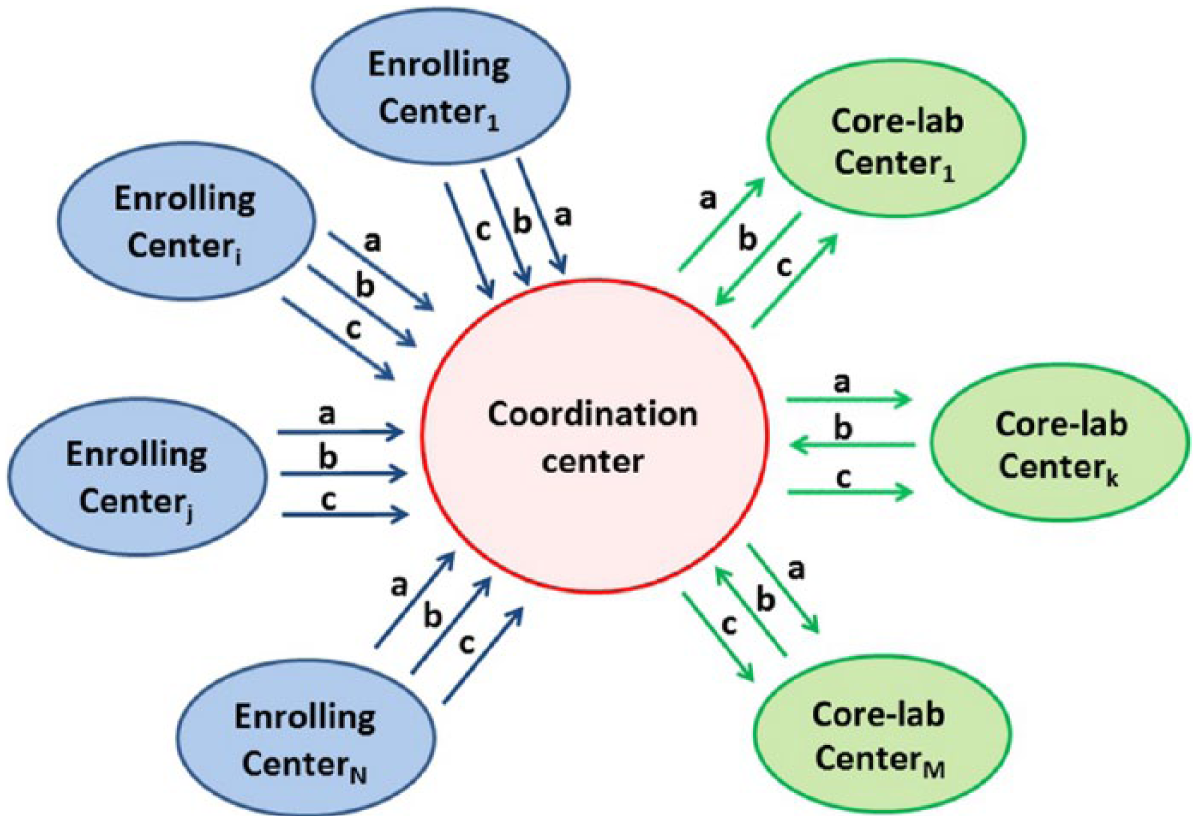
Developed hub-and-spoke informatics platform. The hub is the coordination center, and the various spokes represent the different channels used to transfer (a) biological data, (b) patient’s digital reports, and (c) raw image data.
Data transfer and communication include three channels: physical transfer of biological samples (i.e. blood samples; Figure 1, channel a), transfer of patient’s digital reports (Figure 1, channel b), and transfer of raw image data (Figure 1, channel c). A detailed description of the data flow through the network is given in Figure 2.
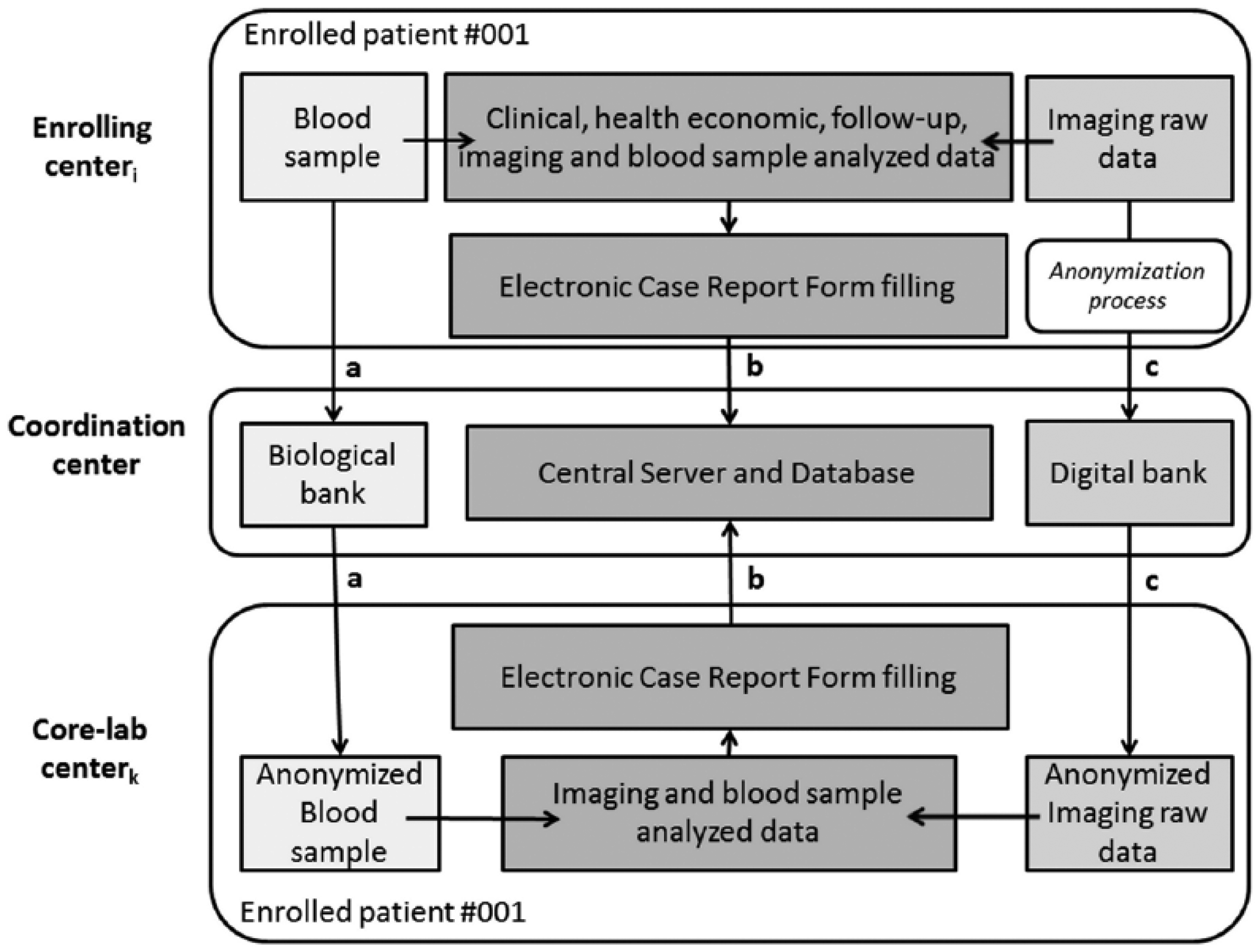
Detailed representation of data flow through the network.
The designed platform guarantees that for each involved center, data transfer is accomplished following local defined protocols, which can be different among involved centers. A detailed description of the activities of each center is given in the following.
Coordination center activities
The coordination center (Figure 2—central level) is responsible for storing the data arriving from enrolling centers (channel a and channel c) and to host the central server for the patient’s database management (channel b). Database information is used for the statistical analysis at the end of the study. To achieve this, a biological bank, a central server and database, and a digital bank are available, for storage of biological samples, electronic case report forms (eCRFs), and raw image data, respectively. The coordination center is also responsible for sending the biological samples (channel a) and anonymized raw image data (channel c) to the core-lab centers, following the same protocols adopted for reception from enrolling centers. The developed informatics platform also allows the coordination center to receive the eCRF data from core-labs (channel b), resulting from the blinded analysis. The three modules developed for coordination center activities (i.e. biological bank, central server and database, and digital bank) will be described in the following sections.
Biological bank
Biological samples are expected to be collected for each enrolled patient. A standardized procedure for handling the biological samples, containing detailed instructions for drawing blood samples, pre-analytical treatment, storage, and shipment, was defined in the study protocol and sent by email to the enrolling centers by the coordination center.
The developed network guarantees saving the results of blood sample analysis and also the possibility of shipping the collected blood samples by special international delivery mail to the coordination center for storage in the Biological Bank (BioBank). The BioBank, located at the coordination center, is an operational facility where standardized procedures are used for the collection, conservation, and management of biological samples. After receiving confirmation by the coordination center, the designed procedure sends the blood samples to the core-lab centers for blinded analysis.
Digital bank—Clinical Trial Picture Archiving and Communication System
The informatics platform was designed and developed in order to share and distribute the image raw data within the network. A digital bank was implemented as a Clinical Trial Picture Archiving and Communication System (PACS) in order to store and distribute raw image data from enrolling to core-lab centers for blinded analysis. Hence, the clinical trial server is able to receive digital imaging and communications in medicine (DICOM) images from the enrolling centers and to redistribute them automatically to the core-lab centers. Figure 3 shows the structure of the system.
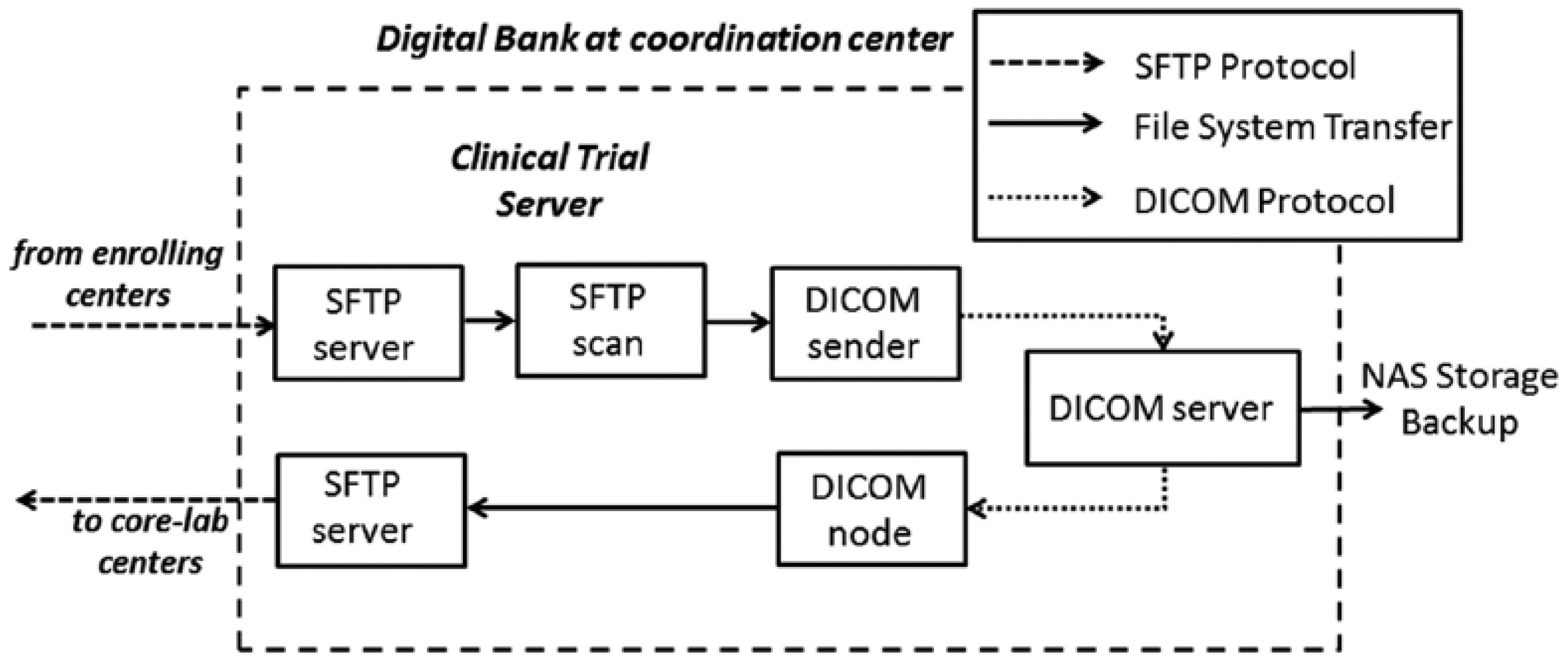
General schema of the method used to store and share the imaging raw data between the centers involved in the multicenter study.
The developed system guarantees that each enrolling and core-lab center receives its own login and password in order to upload and download, respectively, the DICOM images, using secure file transfer protocol (SFTP) facility on a SFTP server at the coordination center. All images should be anonymized prior to transfer within the network. The developed platform is based on an automatized procedure developed as a shell script that periodically scans specific receiving folders inside the SFTP server in order to find new uploaded files. When new files are detected, a DICOM sender is activated. It sends images to a DICOM server that works as a centralized DICOM image manager built on top of the Open Source software dcm4che. 15 All images are also saved on an external repository network attached storage (NAS) used as a backup system for image data.
Raw images of each enrolled patient are received and certified by the certification team of the coordination center, which is also in charge of sending the checked raw data images to a DICOM Node present on the SFTP server. This procedure is necessary in order to redistribute the raw images to the core-lab centers by an automatic procedure that recognizes the image acquisition modality and distributes them to the right core-lab SFTP download folder. The certification step (quality control) by the coordination center is necessary to assure the control and standardization of the anonymization before sending data from coordination to the core-lab centers.
The DICOM system, and the server that processed and distributed images to the core-lab centers, is based on open source software. We used dcm4che 15 and DCMTK 16 tools in order to create the DICOM image processing nodes. The main PACS architecture was based on dcm4che, a widely used Open Source PACS solution, with continuous updates and valuable support from the developers’ community. As dcm4che is developed in Java, some parts of the described system, where the processing time is critical, were implemented using DCMTK, a collection of libraries and applications implementing large parts of the DICOM standard written in a mixture of ANSI C and C++. In particular, DCMTK was used to create channels to compress and decompress DICOM images.
OpenIndiana 17 was used as a storage machine operative system, including an SFTP server. OpenIndiana is the best option when dealing with storage systems due to the robustness of the illumos kernel (a Solaris kernel derivative) and the native presence of several tools not available in common LINUX distributions. OpenIndiana includes (a) ZFS as filesystem supporting 16 TB of storage: in the project, we used the snapshot feature extensively in order to do fast backup with the system running, and also the RAID-Z feature in order to maintain consistency of the system even with the failure of one or more hard drives; (b) DTrace, a tracing framework that can be used to obtain an overview of a running production system and troubleshooting running applications.
The use of a “full” DICOM protocol was discouraged due to several considerations. First, several image modalities (such as MRI) do not support multislice/multiframe DICOM, so the number of DICOM files to transfer for each patient might be very high (several thousand files). It would make raw image transfer by DICOM protocol impractical. Second, security policies at several network centers did not allow direct access from local PACS to the Internet network, as PACS store sensitive patient’s data. The adopted solution allows a local center to anonymize patients’ data on the local PACS and to move them on a local workstation connected to the external network, respecting the local security protocols since only anonymized data are involved in the procedure. Data can be moved using physical supports as CDs or DVD or by network, depending on skills and informatics technology available at the center. Data can be packed and compressed by an archiving utility and transferred to the coordination center by SFTP protocol.
Central server and database
The central server and database located at the coordination center were developed using an open source platform called Bio-Medical Framework (BMF 2.13.4, http://www.ftgm.it/bmf). The system is Open Source and is released under GNU Lesser General Public License. The BMF has been developed in Java since 2003 and it benefits from all the language features. The system was equipped with a web user interface and complied with current regulations on the handling of sensitive data. HL7 standard was used in the design and in the development of the described framework.
The web interface was developed to allow centers to enroll patients, to receive an identification code for each enrolled patient, and to enter the patient’s data. Entered information are then stored in a central database. The same web system also allows the core-lab centers to enter results from their blinded analysis. To standardize the procedure for collection of cardiac data, dedicated eCRFs have been designed and developed, including all possible parameters required by a multicenter cardiac study. An example of provided eCRF is shown in Table 1. The eCRF structure includes general fields including identifiers of the patient and center performing the exam and information about the clinical condition of the patient. Several structures were defined to collect information from each specific acquisition protocol, including cardiac MRI, PET, SPECT, ECHO, CTA, and invasive coronary angiography (ICA). Each specific structure includes acquisition information (e.g. stressor information), global clinical indexes, such as ejection fraction and stroke volume, and segmental clinical indexes following the American Heart Association’s (AHA) standard segmentation. 18 To improve data consistency, the eCRF fields corresponding to segmental indexes can be filled only using predefined values to be selected by drop-down lists. Table 2 shows an example of predefined lists used for MRI imaging modality. Figure 4 shows graphical examples of imaging eCRFs. In particular, the functional (MRI) eCRFs are represented.
Example of MRI eCRF.
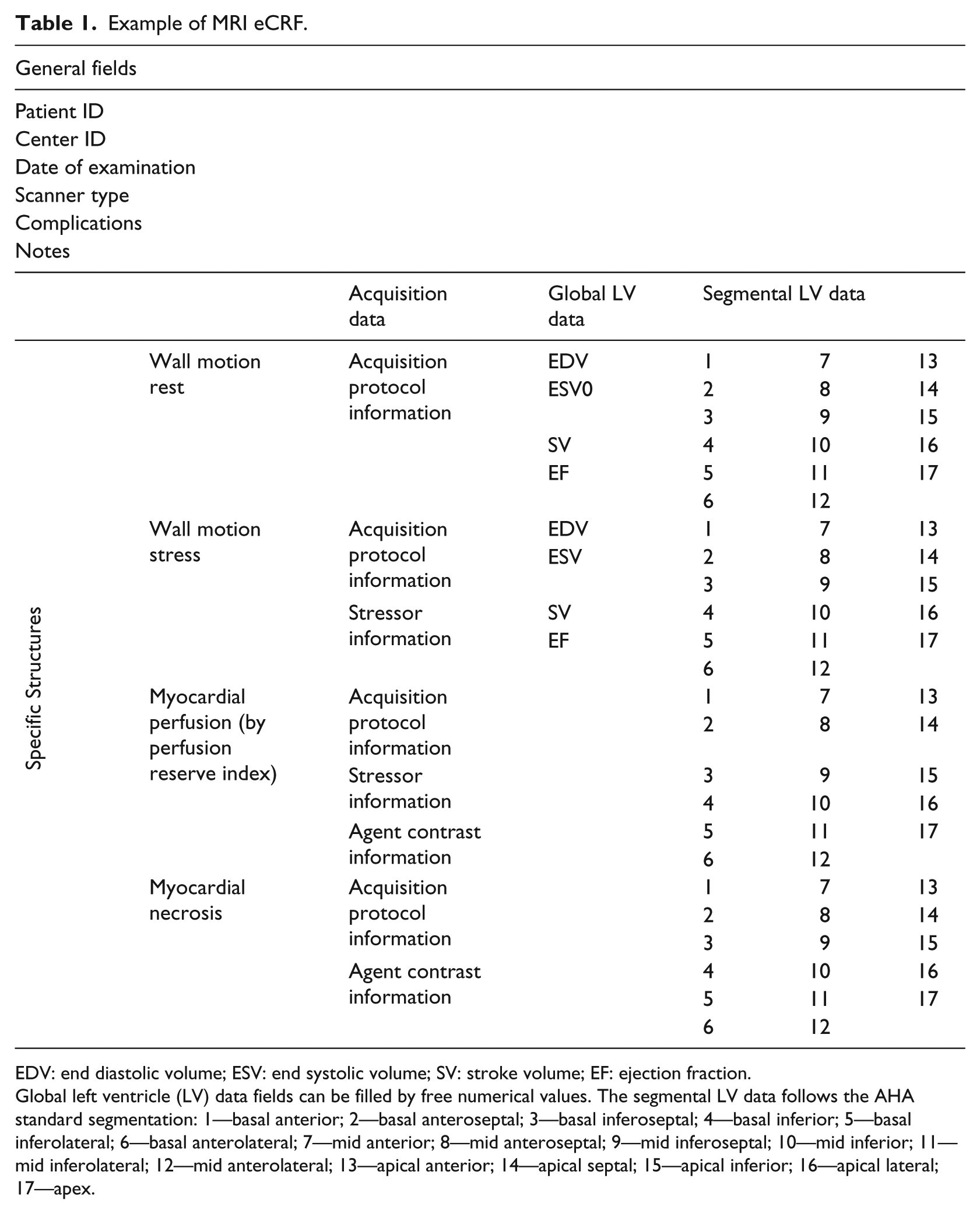
EDV: end diastolic volume; ESV: end systolic volume; SV: stroke volume; EF: ejection fraction.
Global left ventricle (LV) data fields can be filled by free numerical values. The segmental LV data follows the AHA standard segmentation: 1—basal anterior; 2—basal anteroseptal; 3—basal inferoseptal; 4—basal inferior; 5—basal inferolateral; 6—basal anterolateral; 7—mid anterior; 8—mid anteroseptal; 9—mid inferoseptal; 10—mid inferior; 11—mid inferolateral; 12—mid anterolateral; 13—apical anterior; 14—apical septal; 15—apical inferior; 16—apical lateral; 17—apex.
eCRFS values defined for segmental indexes of MRI modality.
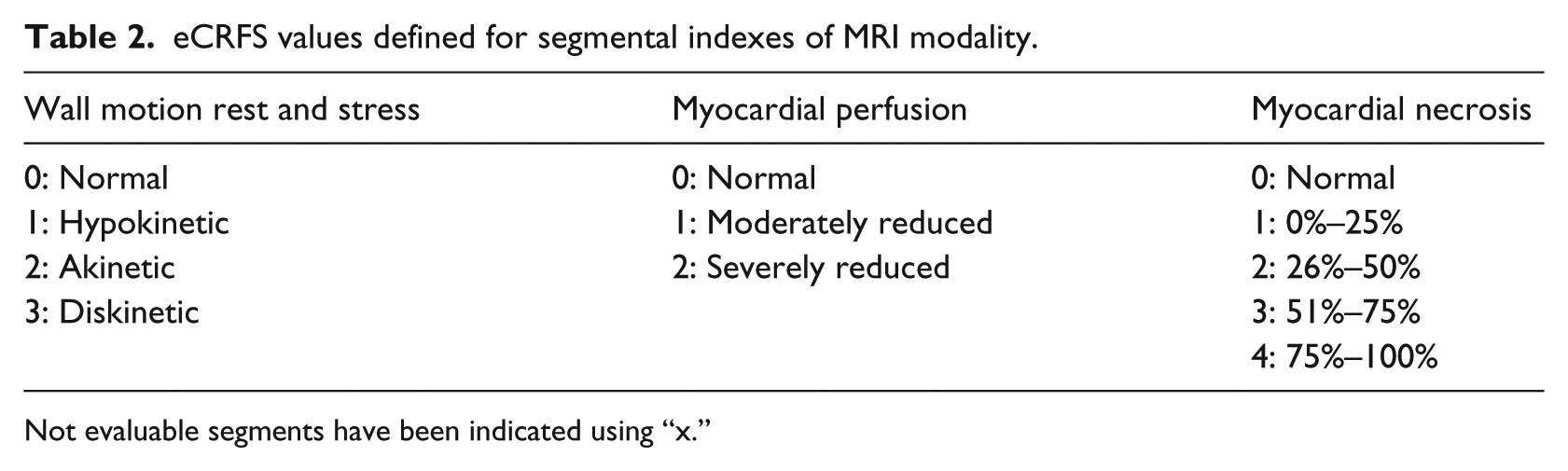

Not evaluable segments have been indicated using “x.”
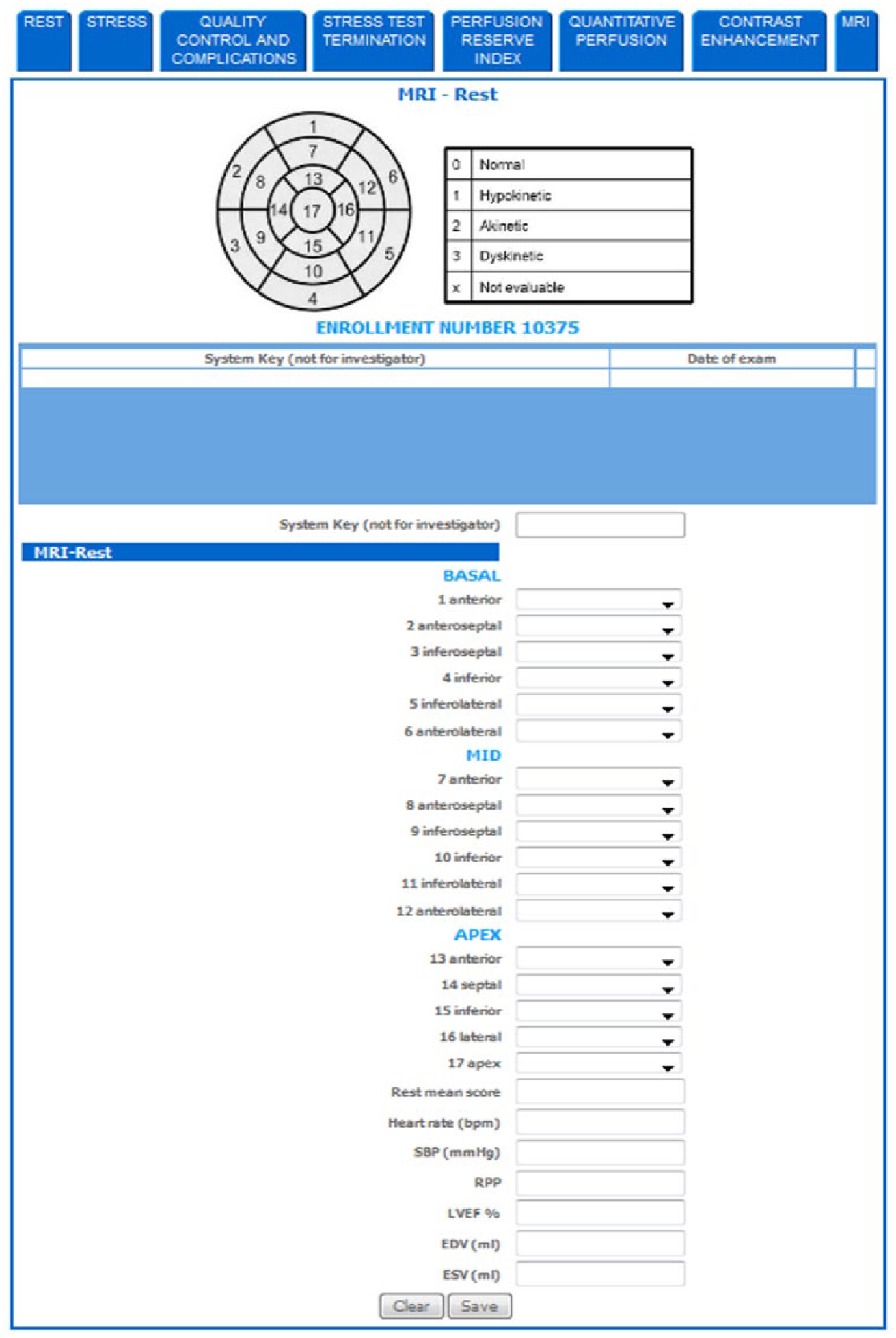
Example of MRI electronic case report forms (eCRFs).
The eCRFs were provided in English. The eCRFs have been designed to be modular with respect to fields to be enabled for different users. Indeed, the developed informatics platform allows the creation of different account profiles, for both enrolling and core-lab centers, in order to set rules to access and manage data. Generally, as core-lab centers replicate for validation of the analysis done at the enrolling centers, the same eCRF formats have been used by enrolling and core-lab centers. In some cases, as in blood sample analysis, the centralization of analysis is required in order to avoid inconsistency of results due to the methodological variability. In this case, enabled eCRF fields could be different for enrolling and core-lab centers.
The developed system uses the Boyce Codd Normal Form (BCNF) 19 for internal data storing and also allows exporting entered data to a format (.cvs format) compatible with software tools used for statistical analysis.
The central server includes a graphical user interface for monitoring patients’ enrollment and data consistency. The adherence of enrolling centers to the recruitment protocol is displayed, as well as the level of completeness of entered data. The number of patients enrolled by each center was constantly checked, and a reminder email was sent in order to improve adherence to the recruitment protocol. Reliability control of entered data was the responsibility of both enrolling and core-lab centers that performed the local and blinded analysis, respectively. Additionally, quality control of entered data, in terms of missing data or evident wrongly entered values, was periodically performed by the coordination center, improving the reliability and consistency of the central database. In these cases, a warning email was sent by the coordination center to the center involved.
Enrolling center activities
Each enrolling center (Figure 2—top level) is responsible for recruiting patients for inclusion in the study and for local analysis. In order to ensure the respect of patients’ privacy, the developed platform allows, after validation (respect of enrolling protocol), automatic identification of each enrolled patient in the Central Server, by receiving a progressive ID number (Center Code + Progressive enrollment code). This code identifies the information and data regarding each enrolled patient. Each enrolling center is also responsible for collecting data (such as clinical information, blood samples, and image raw datasets) from enrolled patients. Acquired data are generally stored at the enrolling center site following local defined procedures such as the Hospital Information Systems (HIS) and PACS adopted at the specific enrolling center. To ensure the transfer of collected data from the enrolling to the coordination center, various solutions have been designed and developed. In particular, the developed platform allows the delivery of biological samples, labeled with the patient ID assigned by the Central Server, via a special international delivery mail service (channel a). If the study protocol involves the acquisition of multiple biological samples, the platform generates a progressive exam ID to allow identification and tracking of multiple biosamples from a given individual that are captured at different times. Moreover, the transfer of raw image data to a digital bank (channel c), located at the coordination center, is guaranteed after anonymization by the enrolling center, using the patient ID assigned by the Central Server. To assure correct application of the anonymization procedure, including a masking step in cases where the patient’s data were “printed” on the image, the coordination center emails the enrolling center the instructions for the required anonymization and for the correct transfer of data, also suggesting the software tools to be used. Examples of software tools are “Dicom Cleaner” 20 and “FileZilla” 21 to use for anonymization and SFTP transfer, respectively.
The patient’s data, such as the clinical, health economics (including exam specific reimbursement, socio-demographics, and other patient information essential for calculating direct and indirect costs associated with examination and quality of life data), and follow-up information, as well as the image analysis results and specific analysis of blood samples, are transferred to the coordination center (channel b) using the eCRFs filled out manually by dedicated personnel.
Core-lab center activities
The developed platform allows each core-lab center (Figure 2—bottom level) to take responsibility for the blinded analysis of anonymized blood samples and imaging raw data sent by the coordination center (channel a and channel c, respectively), and for data quality control. Image quality control represents a key issue in large multicenter studies, as even using standardized acquisition protocols, image quality could be affected by several unpredictable factors as different scanner calibration and differences in acquisition procedure among different vendors. As the association between patients and identification codes is done at the coordination center, the core-lab center is not aware about the enrolling center that acquired the patient’s data. This procedure avoids a possible bias in data quality control. Indeed, it guarantees the transfer to the coordination center of the results of imaging and blood sample analysis (channel b) using the eCRFs filled out manually by dedicated personnel.
The methods used to transfer data between enrolling and coordination centers were also used to exchange data between coordination and core-lab centers.
Results
The proposed platform was used and evaluated during the EVINCI study which was designed to test the impact of a combined non-invasive anatomical and functional cardiac imaging strategy for the early detection and management of ischemic heart disease (IHD). 22 The EVINCI study enrolled patients with chronic chest pain and intermediate probability of coronary artery disease (CAD) based on Diamond and Forrester scale (20%–90%) modified after exercise electrocardiogram (ECG). 23 In the EVINCI protocol (Figure 5), non-invasive diagnostic tests including both anatomical and functional imaging were performed in all patients before invasive coronary angiography.
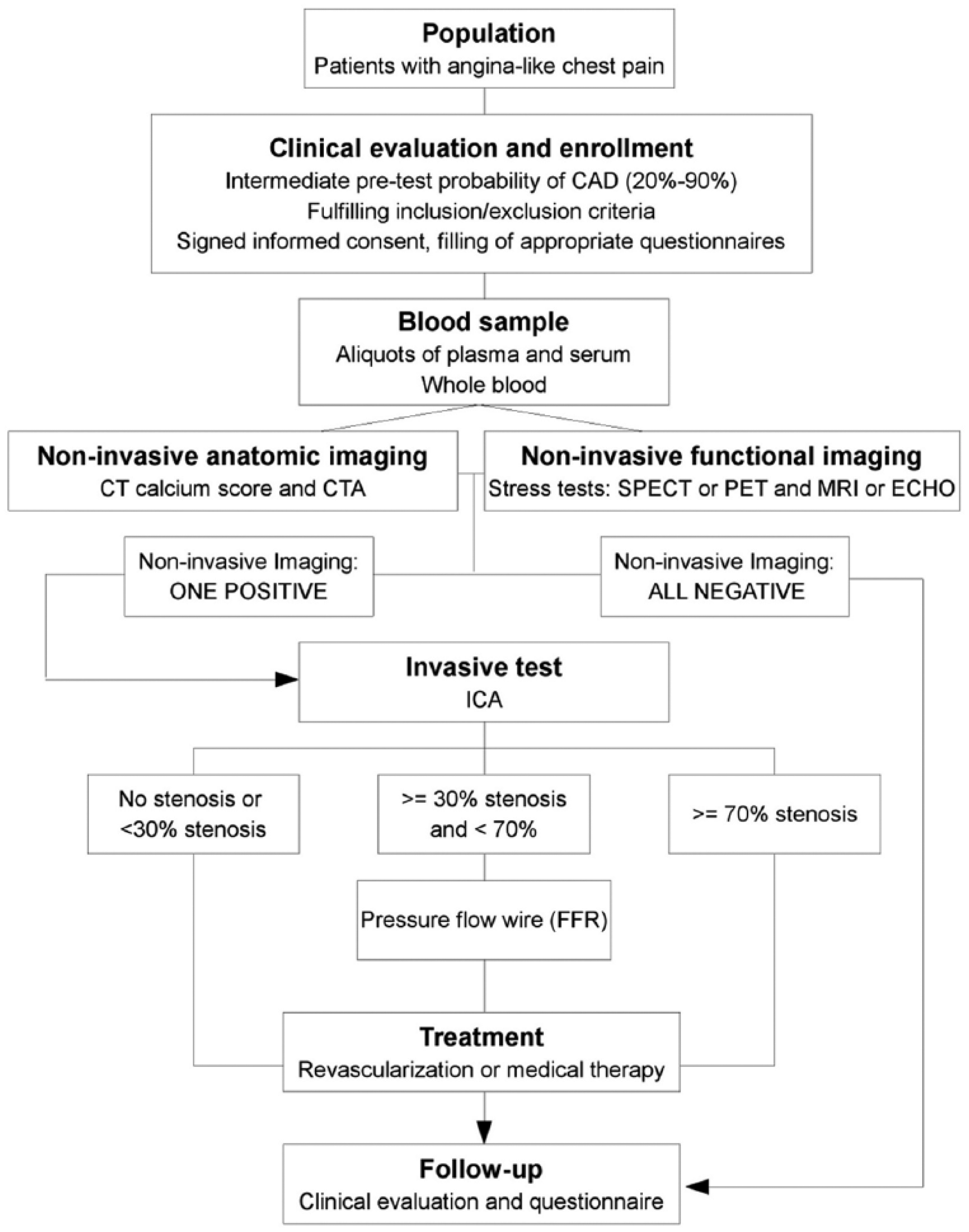
Flowchart of the EVINCI diagnostic work-up and follow-up.
According to the study protocol, the CTA test for coronary anatomy and CS evaluations, and non-invasive functional stress tests for myocardial perfusion assessment (chosen among PET and SPECT) and/or for global and regional myocardial wall motion assessment (chosen among ECHO and MRI) is mandatory. Moreover, blood samples were collected for each patient. After non-invasive screening, and when appropriate, patients with positive tests underwent invasive heart catheterization as a reference method to define the presence and extent of functionally significant obstructive coronary disease (i.e. coronary disease potentially causing ischemia and indicating revascularization). The consortium of the EVINCI study comprised 19 European centers distributed among 9 European Countries, including centers for patient’s enrollment (14 enrolling centers located in Italy, Finland, Switzerland, Germany, Poland, Spain, United Kingdom, and France) and for blinded analysis of biohumoral and imaging data (9 core-lab centers located in Italy, Netherlands, Finland, Spain, United Kingdom, and Germany). The coordination center was located in Pisa (Italy). Each European center could have more roles in the Consortium.
The study was approved by the Local Ethics Committees and followed the principles outlined in the Declaration of Helsinki. The EVINCI research program has been conducted in accordance with any relevant local or national rules and regulations of the country where the research has been carried out.
Data collection started in March 2009 and was concluded in June 2012. The EVINCI study enrolled 697 patients (Figure 6).
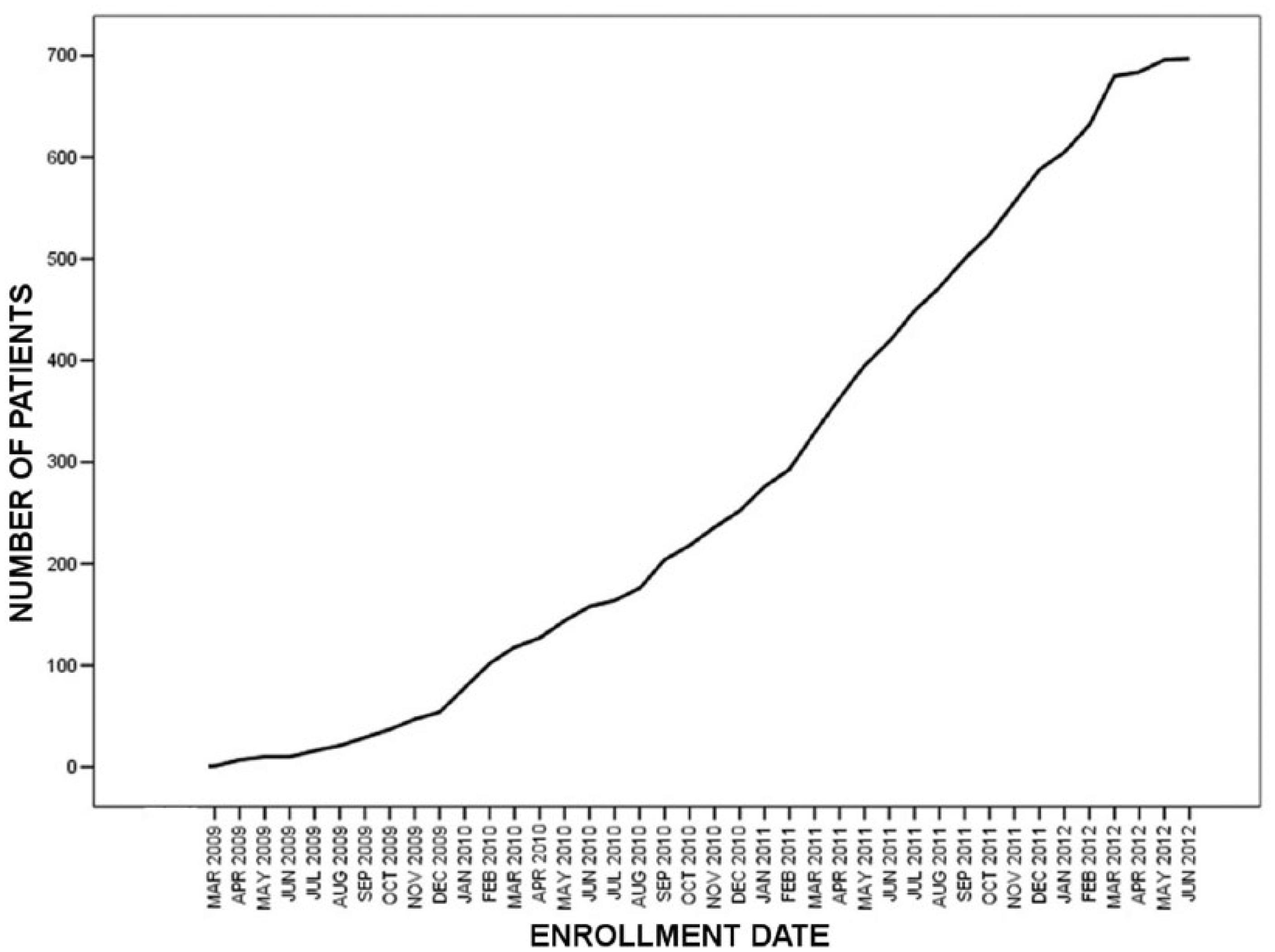
Cumulative curve representing enrollment progress from March 2009 to June 2012.
For each modality, the following total number of examinations is performed by the enrolling centers: 599 CTA (86% of enrolled patients), 408 ICA (59% of enrolled patients), 135 PET (19% of enrolled patients), 396 SPECT (57% of enrolled patients), 353 ECHO (51% of enrolled patients), and 166 MRI (24% of enrolled patients). Figure 7 shows a summary of the collection of biohumoral and imaging information. A detailed description is provided in the following.
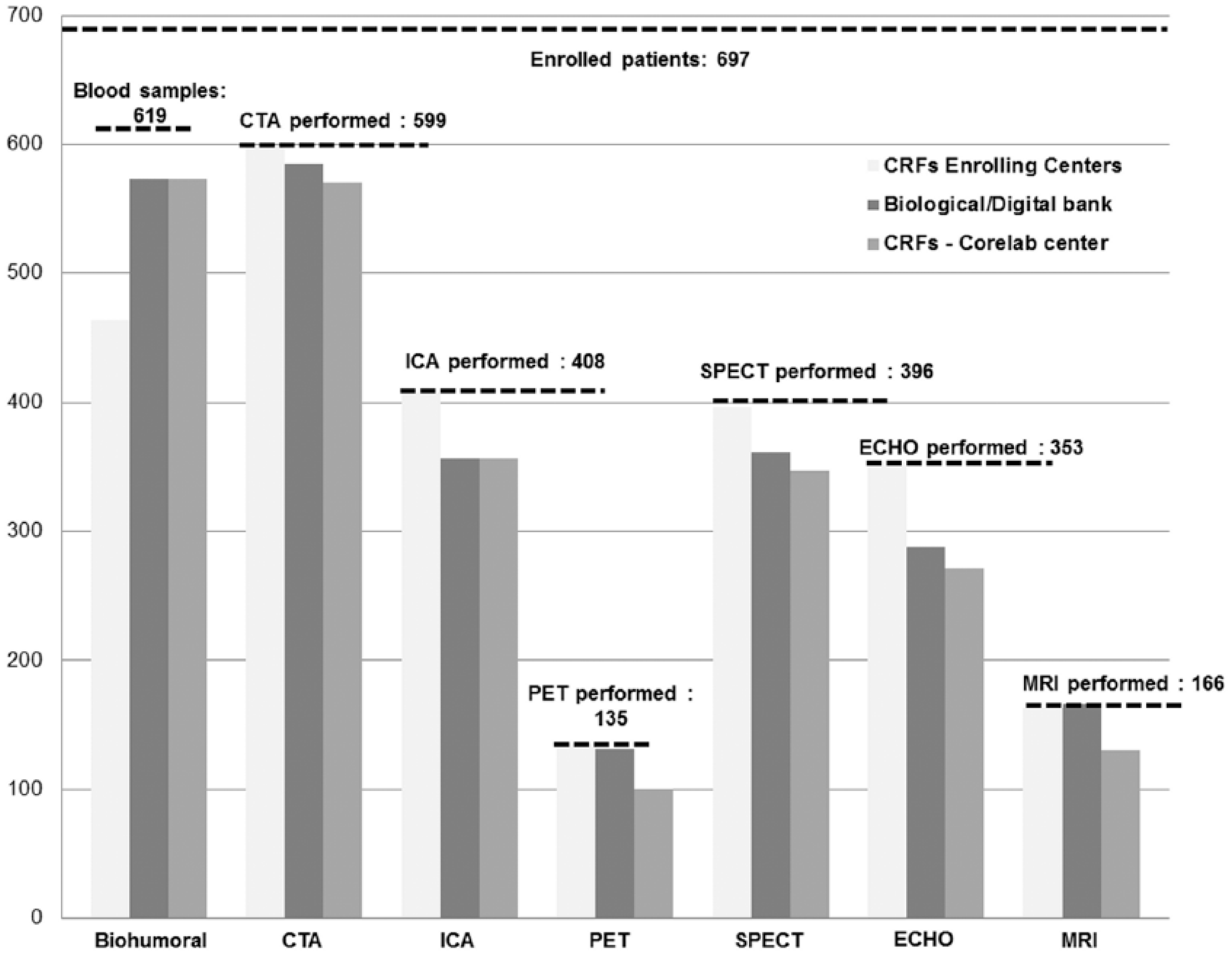
Total number of collected EVINCI data.
Biological bank
Blood samples were sent to the coordination center to populate the biological bank: 619 blood samples (89% of enrolled patients) were sent to the coordination center. Due to drawbacks occurring during the shipment procedures (mainly thawing of blood samples), 7% of shipped samples were unavailable for analysis. Thus, 573 samples (93% of shipped samples) were stored in the BioBank.
Blood samples stored in the biological bank were also sent to the core-lab centers for blinded analysis. All samples were correctly received by the core-lab centers.
Digital bank
Raw data of each imaging modality were sent to the coordination center to populate the digital bank. At the end of the study, the digital bank comprised: 585 CTA datasets (98% of performed examinations), 356 ICA datasets (87% of performed examinations), 131 PET datasets (97% of performed examinations), 361 SPECT datasets (91% of performed examinations), 288 ECHO datasets (82% of performed examinations), and 166 MRI datasets (100% of performed examinations).
Anonymized raw imaging data stored in the digital bank were also sent to the core-lab centers for blinded analysis. All image data were correctly received by the core-lab centers.
Central server and database
For each enrolled patient, information from enrolling and core-lab centers was collected. A total number of 697 clinical eCRFs (100% of enrolled patients), 464 biohumoral eCRFs (67% of enrolled patients), 597 health economic eCRFs (86% of enrolled patients), and 465 follow-up eCRFs (67% of enrolled patients) were filled-up by the enrolling centers.
For each diagnostic imaging modality, the corresponding eCRFs were entered by the enrolling centers to populate the dedicated web application. In particular, 599 CTA eCRFs (100% of performed examinations), 408 ICA eCRFs (100% of performed examinations), 135 PET eCRFs (100% of performed examinations), 396 SPECT eCRFs (100% of performed examinations), 353 ECHO eCRFs (100% of performed examinations), and 163 MRI eCRFs (98% of performed examinations) were adequately filled and considered consistent for following statistical analysis.
After blinded analysis, the eCRFs were entered and completed by the corresponding core-lab centers to populate the dedicated database. In particular, 570 CTA eCRFs (97% of datasets stored in the digital bank), 356 ICA eCRFs (100% of datasets stored in the digital bank), 100 PET eCRFs (76% of datasets stored in the digital bank), 347 SPECT eCRFs (96% of datasets stored in the digital bank), 271 ECHO eCRFs (94% of datasets stored in the digital bank), 130 MRI eCRFs (78% of datasets stored in the digital bank), and 573 biohumoral eCRFs (100% of samples stored in the biological bank) were adequately filled by core-lab centers and considered consistent for subsequent statistical analysis.
Following the study protocol, several centers submitted formal request to the Steering/Writing Committee for sub-studies. After approval, relevant data were successfully extracted from the database by the coordination center and sent to the sub-study responsible.
Discussion
Collaborative and multicenter cardiac studies may involve a large number of centers located in different countries, and with different roles in the study network. As the multimodal approach becomes predominant in large studies, several types of patient information, ranging from biological samples to multimodal image data, are expected to be acquired, stored, and shared during the study’s progress. This aspect requires the development of a reliable network and informatics systems able to support the management, transfer, storing, and standardization of all collected data.
In this article, we showed the reliability of an informatics platform designed to support the activity of each center involved in the study and tested during an EU-funded project. The study was designed with the main purpose of assessing the medical and economic impact of a combined non-invasive anatomical and functional cardiac imaging strategy for the early detection and management of IHD. Considering the crucial objective of the study, a large network of centers in different European countries was created for data collection. The proposed informatics platform effectively supported sharing, transfer, management, storage, and standardization of the large amount of acquired and collected data.
In the EVINCI study, the biological samples involved were blood samples acquired from all patients at enrolling centers. All samples were correctly marked with the patient ID assigned by the central server. An acceptable percentage (89% of enrolled patients) of blood samples were transferred from enrolling centers to the coordination center. Missing data were correctly marked by the informatics platform, but they were not sent by the enrolling center to the coordination center. Some delays during the shipment of samples, resulting in the thawing of samples, led to the unavailability of 7% of shipped samples.
A relevant problem successfully addressed during the study was the process of sending and receiving multimodal raw datasets using the digital bank module (channel c). The developed PACS architecture fulfilled the needs of the project, thanks to the robustness and usability of the dcm4che package. The use of DCMTK to implement several critical procedures allowed us to satisfy processing time constraints. The PACS and SFTP services were well supported by the OpenIndiana OS. The only drawback of the system was the lack of a graphical user interface, so the system administration required highly skilled personnel.
Concerning the transfer of imaging datasets, 92% of images acquired at enrolling centers were successfully transferred to the coordination center. All images received by the coordination center were successfully transferred to core-lab centers.
The anonymization procedure worked in all datasets, even in the case of ECHO and secondary capture CT images, where the patient’s data were not only coded in the DICOM header but also “printed” on the image data itself and masking software was needed.
A lower transfer rate from enrolling centers was observed for ICA and ECHO raw datasets. This finding could be explained by the fact that in some enrolling centers, the digital storage of ICA and ECHO images is not performed routinely as these images are used “on-line” for clinical evaluation and reporting. Moreover, in the case that problems occurred at enrolling centers for using SFTP protocol, due to technical issues such as the local limitation of a firewall, the CD/DVD shipment of raw data has been also permitted. This specific request was required by only one clinical center having problems in setting SFTP connection. The use of a manual procedure to anonymize DICOM images induced a work overhead at enrolling centers. Centralized anonymization would be impractical as it would imply transfer of sensitive patient’s data and for the technical reasons previously discussed in subsection “Digital bank—Clinical Trial Picture Archiving and Communication System,” The use of a local procedure allowed enrolling centers to use local, established procedures if available or to setting one with the collaboration of the coordination center. Integration of the anonymization procedure in the DICOM image transfer protocol would be desirable in the further development of the platform.
Both enrolling and core-lab centers adequately used the web application for eCRFs filling in the Central Server and Database. The data insertion procedure driven by the web-application interface was generally well accepted by the personnel at remote centers and the learning curve was considered satisfactory. The eCRFs filling percentage was totally satisfactory in the case of imaging eCRFs (around 100% and 94% for enrolling and core-lab centers, respectively) and acceptable (at least mandatory fields for final statistical analysis were filled-up) in the other cases. Although the careful design of the web-application interface was expected to reduce data insertion errors, data quality would depend also on the correctness of original medical record at enrolling centers. The quality control procedure done by core-lab centers mitigated the problem allowing the individuation of enrolling centers providing inconsistent data.
The slight difference between examinations sent to core-lab centers and eCRFs entered by core-lab centers can be explained by the fact that some multimodal datasets were not considered usable for processing by core-lab centers, although they were considered clinically useful by enrolling centers. As previously noted (subsection “Core-lab center activities”), the exclusion of some data set from the project database is a common finding in large multicenter studies due to local changes in the acquisition procedures. For that some data were correctly received by the core-lab centers but not processed leading to unfilled or partially filled eCRFs.
In general, the BMF framework exploited in the study facilitated developing a friendly and robust interface for data insertion and management, as also previously shown in other multicenter-networked studies. 24 The experience acquired in using BMF guided the choice to use it during the multicenter European study. Many other commercial, proprietary, and Open Source platforms are used for clinical trial management. Recent survey showed the heterogeneous adoption of these systems across Europe. 25 Because the heterogeneity of systems may be a hindrance in data exchange, the adoption of standards such as Clinical Data Interchange Standard Consortium (CDISC; http://www.cdisc.org) represents an added value. Regarding this aspect, OpenClinica (http://www.openclinica.com) may be an interesting and valuable alternative to the proposed framework. Additionally, among other Open Source software tools, 26 an important example that may be considered for evaluation is the patient research system (Shared Health Research Information Network (SHRINE)) described by McMurry et al. 27 that permits to aggregate as many patient information as possible from a large number of centers. Currently, the BMF framework does not include a context-sensitive help system. Thus, the user support was provided by instruction sent by email and by the helpdesk service. The adoption of more sophisticated help systems may improve usability.
The information technology available at centers involved in the study was jeopardized going from the use of paper reports to sophisticated HIS system. For this reason, the use of eCRFs by web interface allowed standardization of data, avoiding also delay and costs associated with “paper mail shipment.” At coordination center, the use of full digitalized data collection procedure supported a reduction in the workload of coordination center personnel making faster the collection, control, and correction of the entered data. Conversely, the workload of enrolling and core-lab centers could have been increased due to the need to populate the web application. However, the reliability analysis to compare the content of local data with eCRFs would be the responsibility of enrolling and core-lab centers.
This study has a couple of additional limitations. First, the user evaluation provided in this article only aimed to prove the feasibility of the framework for an initial case study, although the described multicenter study covers nearly all cardiac imaging modalities and several European clinical centers. Further evaluative work, including different cases and a more rigorous validation, will be completed and published upon in the future. In particular, the case study is focused on patients scheduled for coronary angiography that perform all the examinations in a short time at the same center. Although the system should support shared acquisition of data of the same patient through the network, this scenario could be challenging and the robustness of the system on this aspect should be assessed.
Second, during the development, we did not consider the standardized eCRFs proposed by the CDISC through the Clinical Data Acquisition Standards Harmonization (CDASH) initiative. 28 Future objectives include the improvement in the development informatics system, such as compliance to CDASH standards, and exploitation in additional multicenter studies for further evaluations and analysis.
The modular characteristic of the proposed informatics platform makes it adaptable for different collaborative studies. If biological samples and/or raw image data transfer are not needed, the related channels (channel a and channel c) can be excluded without affecting the core of the network. If the study involves few or even just one acquisition modality, the related eCRFs alone can be enabled. Hence, our expectation is that the developed platform could be successfully used in other collaborative studies.
Moreover, the availability of this large amount of data made possible innovative applications in multimodal imaging integration and educational fields. This has been demonstrated on a subsample of the study population, leading to the development of a tool for high-resolution 14 integration of complementary information coming from different imaging modalities. Moreover, the established image and patients’ information databases have been exploited for educational purposes, as shown by the developed web-based educational tool (“eduCAD”) directed to young cardiologists for training in the appropriate use of multimodal imaging technology in patients with suspected IHD. 29 The web-based educational tool has been included in a dedicated website (http://www.educad.org). Finally, the described platform could also be considered at starting point for the development of an informatics network to be used for remote medical second opinion and telemedicine, making the exchange of information more reliable. 30
Conclusion
According to the described and discussed results, the developed hub-and-spokes informatics platform guarantees the safe and reliable sharing, collection, and storage of different kinds of data (e.g. clinical, biohumoral, and imaging) useful for collaborative and multicenter studies in cardiology and also for further technological development related to clinical reporting, integration of cardiac imaging information, and education.
Footnotes
Acknowledgements
The authors would like to thank all contributors for informatics platform design.
Informatics core members: Martina Marinelli, Institute of Clinical Physiology—CNR, Pisa, Coordinator of data management and software engineer in multimodal data integration; Vincenzo Positano, Fondazione G. Monasterio CNR-Regione Toscana, Pisa, Scientific consultant for multimodal data integration; Maurizio Mangione, Fondazione G. Monasterio CNR-Regione Toscana, Pisa, Manager of Central Server and Database; Paolo Marcheschi, Fondazione G. Monasterio CNR-Regione Toscana, Pisa, Manager of Clinical Trial PACS and digital bank; Stefano Puzzuoli, Institute of Clinical Physiology—CNR, Pisa, Programmer of Central Server and Database; Natalia Esposito, Institute of Clinical Physiology—CNR, Pisa, Collaborator in data management; Giuseppe Andrea L’Abbate, Fondazione G. Monasterio CNR-Regione Toscana, Pisa, Software developer in data integration and web design.
Core-lab centers: Juhani Knuuti, Turun yliopisto, Turku, ICA and PET core-lab center; Mikko Pietilä, Turun yliopisto, Turku, ICA core-lab center; Stephan G. Nekolla, Nuklearmedinische Klinik der TU, München, PET core-lab center; Richard Underwood, Royal Brompton and Harefield NHS Trust, London, SPECT core-lab center; Jose Luis Zamorano, Servicio Madrileno del la Salud, Madrid, ECHO core-lab center; Covadonga Fernández-Golfín, Servicio Madrileno del la Salud, Madrid, ECHO core-lab center; Luis Miguel Rincón Díaz, Servicio Madrileno del la Salud, Madrid, ECHO core-lab center; Rosa Sicari, Institute of Clinical Physiology—CNR, Pisa, ECHO core-lab center; Giancarlo Todiere, Fondazione G. Monasterio—CNR-Regione Toscana, Pisa, MRI core-lab center; Massimo Lombardi, Fondazione G. Monasterio: CNR-Regione Toscana, Pisa, MRI core-lab center; Artur JHA Scholte, Academisch Ziekenhuis Leiden: Leids Universitair Medisch Centrum, Leiden, CTA core-lab center; Michiel De Graaf, Academisch Ziekenhuis Leiden: Leids Universitair Medisch Centrum, Leiden, CTA core-lab center; Chiara Caselli, Institute of Clinical Physiology—CNR, Pisa, Biohumoral core-lab center; Daniela Giannessi, Institute of Clinical Physiology—CNR, Pisa, Biohumoral core-lab center; Daniele Rovai, Institute of Clinical Physiology—CNR, Pisa, Clinical core-lab; Alessia Gimelli, Fondazione G. Monasterio—CNR-Regione Toscana, Pisa, Clinical core-lab.
Statistical and health economics analysis: Fabio Mariani, Institute of Clinical Physiology—CNR, Pisa, Statistician for data analysis; Giuseppe Turchetti, Scuola Superiore Sant’Anna, Institute of Management, Pisa, responsible of the Health Economics Evaluation group; Stefania Bellelli, Scuola Superiore Sant’Anna, Institute of Management, Pisa, researcher in the Health Economic Evaluation group; Valentina Lorenzoni, Scuola Superiore Sant’Anna, Institute of Management, Pisa, researcher in the Health Economic Evaluation group; Francesca Pierotti, Scuola Superiore Sant’Anna, Institute of Management, Pisa, researcher in the Health Economic Evaluation group.
EVINCI coordinator: Danilo Neglia, EVINCI project coordinator.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The activity received funding from the European Community’s Seventh Framework Program (FP7/2007-2013) under grant agreement no. 222915 (EVINCI Study). Unrestricted grant and computer technology was made available by GEHC.
