Abstract
Background:
Colorectal cancer (CRC) remains a global health issue with high morbidity and mortality. According to molecular profiling, genetic mutations such as KRAS, NRAS, and BRAF play a significant role in tumor biology, treatment effectiveness, and patient prognosis. The exact effect of these alterations on overall survival (OS) is, however, unclear. This systematic review and meta-analysis estimated the combined effect of KRAS, NRAS, and BRAF mutations on CRC survival relative to wild-type tumors.
Methods:
This study focused on research conducted in the last decade and was registered on PROSPERO (Reg No: CRD420251003000). Search was done using PubMed, EMBASE, MEDLINE, and Google Scholar. Cohort, case control, and randomized controlled trials reporting Overall Survival Hazard Ratios (HRs) for CRC patients with and without KRAS, NRAS, or BRAF mutations were considered. Multiple reviewers independently extracted and assessed data quality using the Newcastle–Ottawa Scale. For heterogeneity, pooled HRs were computed using a random-effects model. Publication bias was determined by funnel plot asymmetry.
Results:
Nine trials with 3096 participants qualified. Compared to wild-type CRC, KRAS, NRAS, and BRAF mutations were substantially linked with lower overall survival (pooled HR > 1, P < .05). KRAS mutations were most common and consistently associated with poor survival, while BRAF mutations had the largest impact. Mild funnel plot asymmetry suggested publication bias, but risk-of-bias assessment showed moderate to high methodological quality across trials.
Conclusions:
This meta-analysis shows that KRAS, NRAS, and BRAF mutations are important for adverse prognostic markers in colorectal cancer, linked to reduced overall survival. Routine molecular testing for these mutations is vital to enable personalized treatment, improve patient stratification, and enhance clinical outcomes in colorectal cancer management.
Keywords
Introduction
Colorectal cancer (CRC) is a common cancer that affects the colon and rectum, ultimately leading to significant morbidity and mortality around the world.1-4 It ranks as the third most diagnosed cancer worldwide after lung cancer and breast cancer (which comprise 17% and 23% of the total new cancer cases and 23% and 14% of the total cancer deaths, respectively).2,5,6 CRC continues to present an increasing challenge to public health as 1.9 million new cases per year are reported.2,4,6,7
The knowledge of colorectal cancer has been profoundly altered by developments in molecular genetics, which have shown a complex landscape of genetic alterations that are crucial to tumor behavior and patient outcomes.8-10 KRAS, NRAS, and BRAF mutations play critical roles in colorectal cancer (CRC) pathogenesis through constitutive activation of the MAPK signaling pathway, driving tumor growth and progression. 11 These mutations are primarily somatic, with rare germline cases, and thus do not follow classical inheritance patterns in sporadic CRC.11,12 Risk factors associated with these mutations include right-sided tumor location, advanced age, smoking, and metabolic factors such as obesity. 10 Clinically, KRAS and NRAS mutations predict resistance to anti-EGFR therapies, and standard treatment involves chemotherapy-based regimens and anti-VEGF agents. BRAF mutations confer poor prognosis and are managed with combination chemotherapy (FOLFOXIRI + bevacizumab) or targeted therapy using BRAF inhibitors in combination with anti-EGFR agents in previously treated patients. 13 KRAS mutations are particularly noteworthy, as they have been observed in approximately 27% of colorectal cancer cases, marking them as the most mutated gene in colorectal malignancies.1,14-17 Conversely, BRAF-mutated patients account for almost 10% of the patients and are known to experience a very negative prognosis, often associated with a median survival rate of under 12 months.18-21 NRAS mutations occur with lower frequency; however, they play a significant role in the genetic complexity of colorectal tumors.22-25
New research has revealed various molecular patterns of the disease state, which has expanded information on their different roles in the disease process and can provide better prognostication and targeted therapies that can improve patient outcomes and further give insight into understanding the negative predictive factors of the response to monoclonal antibody targeting EGFR.26-29 In addition, the use of these mutations as prognostic markers that will guide clinical practice, particularly in metastatic disease settings, has received considerable attention.30-33 The aim of this study is to evaluate the prognostic significance of KRAS, NRAS, and BRAF mutations in colorectal cancer (CRC). By analyzing the impact of these genetic alterations on overall patient survival, the study seeks to determine their role as predictive biomarkers.
Methods
This systematic review was done with the guidance of the Preferred Reporting Items for Systematic Review and Meta-Analysis (PRISMA) guidelines.34,35 The study was registered on PROSPERO (Reg No: CRD420251003000). Each author contributed to formulating the study protocol.
PICO
Using the PICO framework the aspect the authors wished to address were P: patient with colorectal cancer, I: operative and nonoperative intervention (chemotherapy, immunotherapy, radiotherapy), C: Patient with KRAS, BRAF and NRAS mutations compared to patient with sporadic/wild type colon cancer, O: overall survival rate.
Search strategy
An electronic search was performed on PubMed, EMBASE, and Medline databases for relevant studies. Google Scholars was used for the gray literature search. This search was performed by two independent reviewers (AAO and MO) using a predetermined search strategy designed by (MO). The search included the following terms and Boolean operators (“KRAS” OR “NRAS” OR “BRAF”) AND (“colorectal cancer” OR “colon cancer” OR “rectal cancer”) AND (“prognosis” OR “survival” OR “outcome”); KRAS OR NRAS OR BRAF AND “colorectal cancer” OR “colon cancer” OR “rectal cancer” AND prognosis OR survival OR outcome; KRAS OR NRAS OR BRAF OR AND “colorectal cancer” OR “colon cancer” OR “rectal cancer” AND prognosis OR survival OR outcome. Included studies were limited to English language. The search was restricted to studies within the last 10 years. All duplicates were electronically removed before screening, and studies considered appropriate were reviewed. Final search for the review was performed on May 21, 2025.
Inclusion and exclusion criteria
The studies included were based on study design (cohort studies, case control studies, RCT). All peer-reviewed journals of 10 years involving studies of mutation and non-mutations of KRAS, NRAS, and BRAF in CRC were included. The outcome of interest was overall survival (reported in hazard ratio) of patients who had confirmed mutant genes and non-mutant genes, regardless of treatment received, initial stage of disease, or presence of multiple genetic alterations. Studies that did not specifically analyze KRAS, NRAS, or BRAF mutation status or where mutation is not clearly defined or assessed using validated methods were excluded. Review articles, abstracts from conferences, case reports, case series with less than 10 patients, narrative reviews, systematic reviews, meta-analyses, and studies involving only sample blocks (gross specimen) and secondary data were excluded.
Data extraction and assessment
Each reviewer (MO, AAO, MO, AE, and EM) read the final studies included to ensure all inclusion criteria were met before extracting the following data: Authors name, Year of publication, Study design, Study title, Number of patients included, Molecular analysis, and Overall Survival outcome. The risk-of-bias and quality-of-methodology assessment was performed in accordance with the Newcastle-Ottawa scale. 36 In cases of discrepancies in the views of reviewers, a third reviewer served as an arbiter.
Statistical analysis
Hazard ratios were log-back transformed, and respective standard errors were calculated from confidence intervals with the following formula: (ln [upper CI] − ln [lower CI]) / 3.919928. Estimation was done using the Inverse Variance method and the Log of HR. Statistical heterogeneity was determined using Chi2. A random-effects model with the restricted maximum likelihood method was used to obtain the pooled hazard ratio estimates and respective confidence intervals. Results were graphically presented with a forest plot. The random-effects model was preferred over the fixed-effects model because heterogeneity was anticipated, mainly because of the variety in the adjuvant treatment administered and in the proportion of stage II vs III tumors across the studies. 37 The test was based on P < .050 indicating statistical significance. The risk-of-bias and quality-of-methodology assessment was done on the Newcastle-Ottawa scale. Meta-analysis was performed using Review Manager. 38
Result
Literature search
Overall, 21 814 studies were identified, and 6369 duplicates were removed. The remaining 14 457 were found ineligible by the automation tool. Of the 988 remaining, 952 were excluded following title and abstract screening, and 27 were eliminated following full text screening. Nine full texts were reviewed and all included. The study identification is summarized in the PRISMA flow diagram (Figure 1).

PRISMA flow diagram.
Study characteristics
The studies included in this systematic review (Table 1) are Schirripa et al, 20 Margonis et al, 39 Shindoh et al, 40 Amikura et al, 41 Brunsell et al, 42 Kim et al, 43 Hatta et al, 44 Sakai et al, 45 and Saadat et al. 46
Summary of studies included (the table below shows the individual study characteristics, including the number of participants, study design, year the study was published, and the comparative number of mutant genes versus wild-type genes).
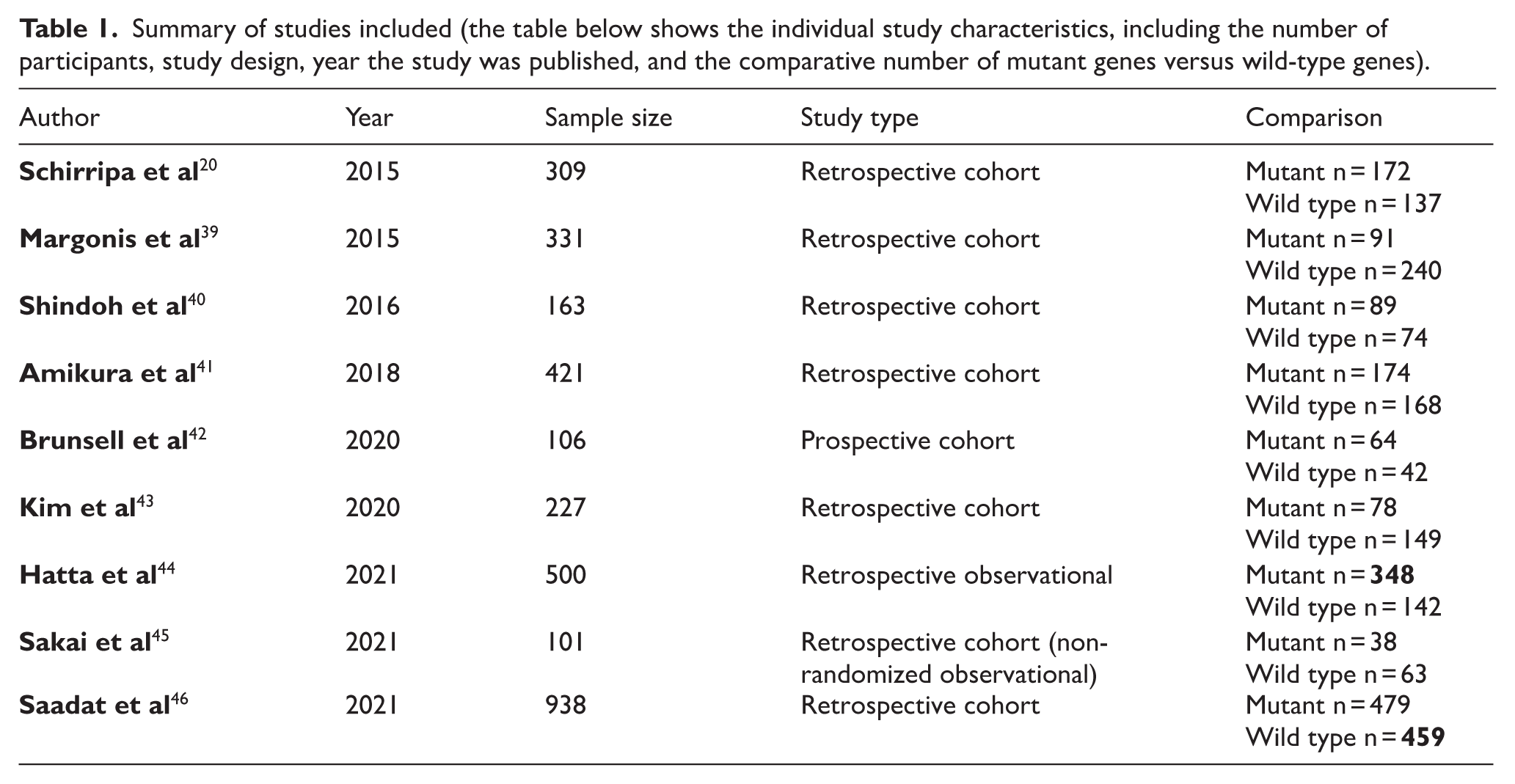
Schirripa et al 20 discusses the prognostic impact of BRAF and RAS mutations in metastatic colorectal cancer patients undergoing liver resection. The primary endpoint was relapse-free survival (RFS). The final study population included 309 patients divided into three groups of RAS, BRAF, and wild-type CRC. Margonis et al 39 on the other hand, assess how specific KRAS mutations influence outcomes in patients undergoing liver resection for colorectal liver metastases (CRLM) in a single-center retrospective cohort study done between 2003 and 2013 with 331 patients.
Brunsell et al 42 investigates the genetic heterogeneity of driver gene mutations (KRAS/MRAS/BRAF) in colorectal liver metastases (CRLMs) among 106 patients with CRLMs. Amikura et al 41 also evaluated the effect of all-RAS mutations on survival outcomes and recurrence patterns in patients undergoing liver resection for colorectal liver metastases. They all showed that RAS mutations were independently associated with worse primary outcomes of overall survival, and furthermore, lung metastases occurred significantly more often in RAS mutant patients. Moreso, Saadat et al 46 investigates the impact of RAS mutation location on survival outcomes in patients with colorectal liver metastases in a single-center retrospective study from 1992 to 2016, including patients with CRLM associated with worse overall survival (OS) (Time from diagnosis or treatment initiation to death). Overall, findings were kept with Amikura et al 41 and Brunsell et al. 42
Shindoh et al, 40 Sakai et al, 45 and Kim et al 43 studied the significant impact of survival outcomes (survival rate at specific time points) and recurrence patterns in patients with KRAS mutation status undergoing resection for colorectal liver metastases and the prognostic impact of primary tumor location (right-sided vs. left-sided) and KRAS mutational status. Shindoh et al 40 and Sakai et al 45 concluded that mtKRAS is associated with poor survival outcomes due to higher lung recurrence and shorter TSF. However, Kim et al 43 concluded that KRAS status alone did not significantly impact survival. Furthermore, Hatta et al 44 evaluated how resection margin status and KRAS mutation affect long-term outcomes in patients undergoing curative liver resection for colorectal liver metastases in a retrospective study of 500 patients and concluded that although KRAS status impacted survival, it had no impact on the margin status of the tumor.
Patient characteristics
The patient characteristics (Table 2) from the reviewed studies focus on sex distribution and median age across the different genetic mutation groups: BRAF mutant, RAS mutant (KRAS and NRAS), and wild-type (wt) colorectal cancer. Males generally outnumbered females in most studies, except in Schirripa et al, where the distribution was more balanced. Shindoh et al reported a significant male predominance in the wild-type group (61 males vs. 28 females, P = .021). Brunsell et al had no females in the BRAF-mutant group (P = .006). Saadat et al showed a large discrepancy in the RAS mutant group (407 males vs. 72 females). The median age varied across studies but typically ranged between 58 and 68 years. Shindoh et al reported a higher median age in the RAS mutant group (65 vs. 61 years, P = .028). Margonis et al noted a trend toward younger age in the RAS mutant group (51.1 vs. 48.5 years, P = .07). Kim et al and Amikura et al reported older median ages in RAS mutant groups (68 years). BRAF mutants are shown to be rare, with a total of 16 patients,20,42 with some studies reporting no cases, like Margonis, Shindoh, and Hatta. The RAS mutant, however, had a larger cohort with a total of 1306.
Summary of studies that evaluated patient characteristics as predictors for OS.
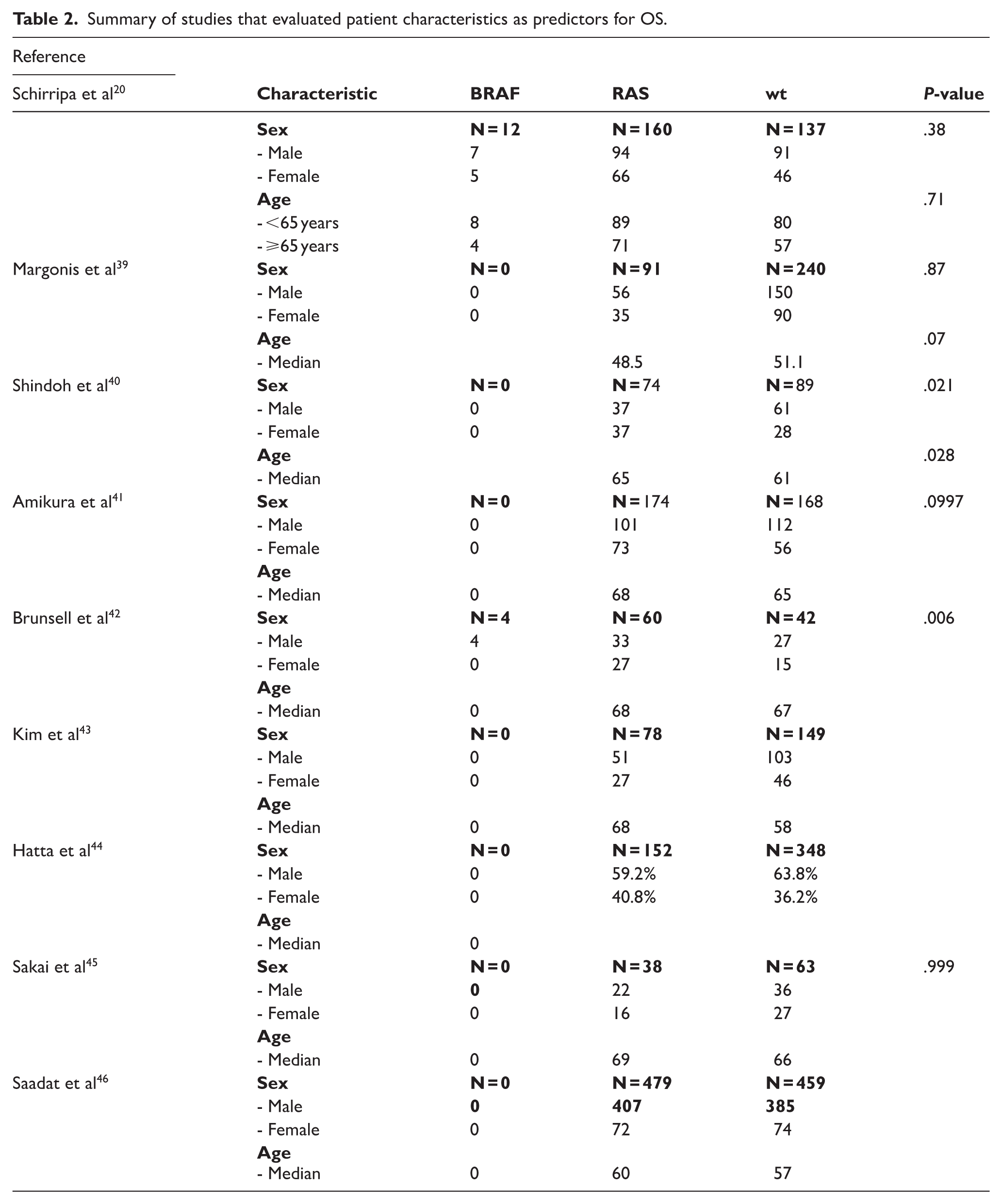
This summary highlights consistent trends (e.g., age, sex distribution) while noting outliers and statistical significance where applicable.
Molecular analysis
Multivariate analysis (Table 3) across the nine studies consistently indicated that KRAS mutation correlated with reduced overall survival. Hazard ratios varied between 1.21 and 3.3, with confidence intervals excluding 1.0 in 90% of the studies, demonstrating statistical significance. The highest risk was noted in Brunsell et al 42 (HR: 3.3, 95% CI: 1.6–6.5), whereas the lowest significant association was found in Kim et al 43 1.420 (0.90–2.25); P = .042.
Summary of studies that evaluated molecular mutation as a predictor of OS.

The primary outcome of interest was overall survival (OS). However, several studies reported alternative but closely related time-to-event outcomes, including disease-specific survival (DSS) or relapse-free survival (RFS). To maximize data utilization, hazard ratios for these survival metrics were harmonized on the logarithmic scale and treated as estimates of OS, in line with established meta-analytic conventions. Sensitivity analyses excluding studies reporting RFS demonstrated no significant change in pooled effect estimates.
Forest plot of pooled studies
The forest plot (Figure 2) below combines results from all nine studies to measure the overall effect of KRAS, NRAS, and BRAF mutations on overall survival (OS) in patients with colorectal cancer (CRC). All the studies showed a hazard ratio (HR) > 1, signifying a higher risk of death compared to patients with wild-type (non-mutated) tumors. The pooled HR (represented by the diamond at the bottom) is to the right of the line of no effect (HR = 1), thus indicating statistical significance. The heterogeneity of the analysis using chi and Tau was chi 2 8.94 and I 2 of 11% indicating a moderate heterogeneity, and the random-effects model was used appropriately to account for this variation.

Forest plot showing overall effect of KRAS, NRAS, and BRAF mutations on overall survival (OS) in patients with colorectal cancer (CRC).
Funnel plot of pooled studies
The funnel plot (Figure 3) for this meta-analysis of KRAS, NRAS, and BRAF mutations vs. wild-type CRC represents each study’s hazard ratio (HR) plotted (X-axis) against its standard error (Y-axis). The funnel plot demonstrated mild asymmetry, with a slight underrepresentation of small studies reporting non-significant or negative associations. This shape suggests potential publication bias, where studies finding no prognostic effect of KRAS, NRAS, or BRAF mutations may be less likely to be published. However, the symmetry among larger studies, which clustered near the pooled estimate, indicates that the overall finding of poorer survival in mutation-positive colorectal cancer is robust and unlikely to be solely due to bias.”

Funnel plot of comparison: meta-analysis of KRAS, NRAS, and BRAF mutation vs WT colon cancer, outcome.
Risk-of-bias assessment
The risk-of-bias assessment, summarized in the risk-of-bias table and graph (Figures 4 and Figure 5), demonstrated that the included studies were generally of moderate to high methodological quality based on the Newcastle–Ottawa Scale. 36 Selection bias was low overall, with all studies employing validated genetic testing methods to confirm mutation status. The main source of potential bias arose from limited adjustment for key confounding variables such as the clinical parameters in some studies, which may influence hazard ratio estimates. However, the consistent direction of effect across studies, together with the use of a random-effects model, mitigates this limitation. Overall, the risk-of-bias findings support the validity of the pooled estimate while underscoring the importance of comprehensive adjustment in future prognostic research.

Risk-of-bias summary: review authors’ judgments about each risk-of-bias item for each included study.

Risk-of-bias graph: review authors’ judgments about each risk of bias presented as percentages across all included studies.
Discussion
This systematic review and meta-analysis offer a thorough look at how mutations in KRAS, NRAS, and BRAF impact overall survival (OS) for patients with colorectal cancer (CRC). By examining data from nine different studies, we see a clear pattern: tumors with these mutations tend to lead to poorer survival outcomes compared to those without these genetic changes. The aggregated results suggest that patients with these mutations face a 40%–60% increased risk of mortality, emphasizing the serious implications of these molecular alterations.
Among the mutations studied, KRAS is the most common, consistently showing detrimental effects on OS. The hazard ratios for KRAS mutations vary between 1.4 and 3.3 across different studies, reinforcing the already established connection between these mutations and resistance to anti-EGFR therapies, as well as a more aggressive form of cancer. NRAS mutations, although less frequent, also indicate a worse prognosis, with a notable risk of increased mortality, further highlighting their significance despite having fewer studies to reference. BRAF mutations stand out with the most severe impact, often leading to a doubling of mortality risk. These mutations are more commonly found in right-sided tumors, which tend to exhibit high-grade histology, microsatellite instability, and aggressive tumor characteristics with poor treatment responses. Thus, assessing KRAS, NRAS, and BRAF mutations aids in creating a comprehensive approach to risk stratification for CRC.
It’s worth noting that variability in confidence intervals and moderate differences in the included studies may stem from diverse mutation detection methods, variations in patient demographics like tumor location and stage, and different treatment protocols. That said, using a random-effects model helps address these variations and supports the reliability of our pooled results. The consistent effects observed and the lack of significant outliers further reinforce the conclusion that having these mutations is linked to worse survival outcomes.
From a clinical perspective, these findings underscore the essential role of precision oncology in managing CRC. Routine molecular testing for KRAS, NRAS, and BRAF mutations is crucial, not just for understanding prognosis but also for informing treatment plans. For example, knowing a patient’s mutation status can indicate whether they should avoid anti-EGFR monoclonal antibodies and consider BRAF inhibitors for BRAF-mutated tumors instead. Additionally, the association between mutation status and recurrence patterns, such as lung-dominant recurrence in KRAS mutant cases, suggests that tailored surveillance strategies, focusing on specific imaging techniques, could help detect relapses early. Integrating these genetic markers with standard clinical factors can enhance risk prediction models, allowing for better patient stratification and optimal treatment intensity and potentially preventing overtreatment in lower-risk patients while also identifying high-risk individuals who may benefit from clinical trials or innovative therapies.
Although only nine studies met the inclusion criteria, the restricted number reflects the rigorous eligibility requirements emphasizing high-quality, methodologically robust analyses reporting hazard ratios for overall survival stratified by KRAS, NRAS, or BRAF mutation status. Notably, several of the included studies were based on liver metastasis or hepatic resection cohorts. These populations are representative of advanced colorectal cancer stages in which molecular profiling is routinely undertaken. Given that RAS and BRAF mutations represent systemic molecular alterations rather than site-specific phenomena, their prognostic implications likely extend beyond hepatic disease. Nevertheless, this concentration of studies in metastatic settings should be acknowledged as a factor that may influence the generalizability of our findings to earlier-stage colorectal cancer.
However, it’s important to acknowledge some limitations. For instance, potential publication bias could mean that studies showing no significant associations are underrepresented, as indicated by mild asymmetry in the funnel plot. Differences in how confounding factors are accounted for and variability in patient eligibility criteria, like tumor stage, clinical parameters, and prior treatments, might influence the estimates of effect sizes. Future research should delve deeper into subgroup analyses and standardize adjustments for confounding factors to improve prognostic assessments. Furthermore, this meta-analysis was restricted to studies published within the past ten years to ensure methodological consistency with modern molecular diagnostic techniques and current management protocols. While this decision improves clinical relevance, it may have excluded earlier investigations with pertinent data, potentially introducing selection bias. Additionally, the limited number of studies available constrains generalizability and underscores the need for further large-scale, prospective analyses across diverse colorectal cancer populations.
Conclusion
This meta-analysis indicates that mutations in KRAS, NRAS, and BRAF are not only significant but also serve as adverse prognostic markers in colorectal cancer. Their prevalence is closely tied to a meaningful reduction in overall survival rates, making molecular testing essential for thorough risk assessment and personalized treatment strategies. Despite the limitations, this study provides a contemporary synthesis of the prognostic relevance of KRAS, NRAS, and BRAF mutations in colorectal cancer, reinforcing their value as biomarkers for risk stratification and personalized management. By incorporating these genetic markers into everyday clinical practice, we can enhance precision oncology efforts, improve patient stratification, optimize treatment selection, and ultimately boost outcomes for individuals diagnosed with CRC.
Footnotes
Ethical Considerations
Not applicable.
Informed Consent
Not applicable.
Patient Consent
Not applicable.
Author Contributions
All authors reviewed and approved the final manuscript and agreed to be accountable for all aspects of the work.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
All data available upon request via corresponding author
Trial Registration Number and Date
PROSPERO Reg No: CRD420251003000 and 10 June 2025. The protocol was submitted during the trial registration process.
