Abstract
Colorectal cancer incidences are on a rise in India. In this study, we have analyzed the mutation frequencies of six potential biomarkers, their coexistence, association with clinicopathological characteristics, and tumor location in Indian colorectal cancer patients. Next-generation sequencing was performed to identify mutations in the six potential biomarker genes using formalin-fixed paraffin-embedded tissue blocks of 112 colorectal cancer patients. The mutation frequency observed in KRAS, BRAF, PIK3CA, NRAS, TP53, and APC was 35.7%, 7.1%, 16.1%, 6.3%, 39.3%, and 29.5%, respectively. The significant associations of mutations were KRAS with age less than 60 years (p = 0.041), PIK3CA with males (p = 0.032), tumor stage I–II (p = 0.013), lack of metastasis in lymph nodes (p = 0.040), NRAS with rectum (p = 0.002), and APC with T2 stage of tumor growth (p = 0.013). No single patient harbored mutations in these six genes or any five genes simultaneously. Significance was noted in coexistence of KRAS with APC (p = 0.024) and mutual exclusion of KRAS with BRAF (p = 0.029). PIK3CA exon 9 was observed to be more frequently associated with KRAS mutations than PIK3CA exon 20 (p = 0.072). NRAS mutations were mutually exclusive with BRAF and PIK3CA mutations. As per our knowledge, this is the first next-generation sequencing–based biomarker study in Indian colorectal cancer patients. Frequent coexistence of gene mutations in pairs and triplets suggests that synergistic effect of overlapping mutations might further trigger the disease. In addition, infrequent coexistence of multiple gene mutations hints toward different signaling pathways for colorectal cancer tumorigenesis.
Introduction
Colorectal cancer (CRC) has been identified as the third most common cancer worldwide, which accounted for 1.4 million cases in 2012. 1 In Asia, CRC incidences have been rapidly increasing in most of the countries, specifically China, Japan, Singapore, and South Korea. 2 India is among the lowest risk of CRC in Asia where colon cancer ranks eighth in men and ninth in women, whereas rectal cancer ranks ninth among men and does not figure in the top 10 cancers in women. 3 Multiple signaling pathways are associated with transition of CRC from adenomatous polyp into adenocarcinoma, which includes Wnt-β catenin pathway, MAPK/ERK pathway, PI3K/Akt pathway, and p53 pathway. The cumulative effect of activation and inactivation of numerous oncogenes and tumor suppressor genes, respectively, deregulates these signaling pathways and leads to CRC. This study includes six frequently mutated genes in CRC: KRAS, BRAF, NRAS (MAPK/ERK pathway), PIK3CA (PI3K/Akt pathway), TP53 (p53 pathway), and APC (Wnt-β catenin pathway). Mutations in these six genes have been extensively studied worldwide due to the high frequency found in CRC patients, and this study investigates these mutations with respect to Indian population. Several CRC-targeted therapies and prognosis testing are based on mutations in the signaling genes KRAS, BRAF, NRAS, and PIK3CA. In addition, TP53 and APC mutations have been associated with most human cancers and play a crucial role in CRC tumorigenesis.
KRAS (Kirsten rat sarcoma viral oncogene homolog) gene is the most commonly altered gene in the RAS family, which is found mutated in 17%–25% of all cancers and 35%–45% of CRCs. 4 Mutations in KRAS are highly frequent in the codons 12, 13, 59, 61, and 146.5,6 Recently, KRAS mutation has been elucidated as a negative predictive biomarker used to detect resistance to the anti-epidermal growth factor receptor (EGFR) monoclonal antibody therapy. 7 BRAF (v-raf murine sarcoma viral oncogene homolog B1) gene mutations are persistent in 15% of all cancers, and the V600E mutation is present in 10% of all CRCs.8,9 A significantly higher frequency of BRAF mutations has been reported in females and in the proximal colon.10–13 Furthermore, KRAS and BRAF mutations are observed to be nearly mutually exclusive of each other.11,12,14,15
PIK3CA (Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit alpha) gene is involved in PI3K/Akt pathway and is found mutated in approximately 10%–20% CRCs where nearly 80% mutations are found in two hotspot regions: exons 9 and 20. 16 PIK3CA mutations are more frequent in the proximal colon.17,18 NRAS (neuroblastoma RAS viral oncogene homolog) is the second most important RAS protein involved in CRC, after KRAS. 19 NRAS is generally found mutated in 15% of all cancers 20 and 5% of all CRCs. 19 NRAS mutations have been previously reported to have a strong association with KRAS wild type (p = 0.001) and are generally found to be mutually exclusive with KRAS mutations.21,22
The tumor suppressor (TP53) gene regulates DNA repair mechanism and apoptosis. The TP53 mutation is observed in nearly 50% CRCs, with the most frequently observed mutations between exons 5 and 8 at codons 175, 213, 245, 248, 273, and 282.23,24 In Asia, the p53 mutation frequency varies between 23% and 52.6%.25–28 TP53 was observed to be frequently mutated with APC.29,30 Adenomatous polyposis coli (APC), a tumor suppressor gene involved in Wnt-β catenin signaling pathway, is found mutated in nearly 80% sporadic CRCs. 31 The mutation cluster region (MCR) is present in exon 15 between codons 1286 and 1513. Higher frequency of APC mutations has been reported in distal colon than proximal colon.32,33
Materials and methods
Patients and specimens
This study was performed on Indian patients undergoing surgery for CRC or diagnosed with metastatic CRC at Sir Ganga Ram Hospital, Delhi, India, between March 2013 and May 2015. Tumor samples were obtained from 146 patients; however, 34 tumor samples were filtered out during tumor content estimation, DNA extraction, and next-generation sequencing (NGS). The remaining 112 CRC patients with confirmed histopathology or biopsy of CRC were included in this study. Clinical characteristics of all patients were recorded including age, gender, tumor location, tumor size, Tumor, Lymph-node, Metastasis (TNM) stage, distant metastasis in liver, or other parts. Follow-up end point was December 2015.
For all these patients, formalin-fixed paraffin-embedded (FFPE) tissue was used for DNA isolation. All the FFPE tissue blocks were taken from the Department of Histopathology, Sir Ganga Ram Hospital, and were checked thoroughly for tumor content. FFPE tissue blocks with more than 30% tumor content were taken for the study. Five to eight sections of 10 µm were used for DNA extraction using microtome. The DNA was isolated using Qiagen AllPrep DNA FFPE Kit (Cat No./ID: 80234, Qiagen) as per manufacturer instructions.
Library preparation and sequencing
DNA was quantified for double stranded DNA (dsDNA) contents using QuBit (Thermo Fisher, USA), and protein contamination was checked by measuring the 260:280 ratio using Nanovue (Agilent, USA). Samples with 260:280 ratio within the range 1.7–2.0 were qualified. The DNA was also further qualified by running the Illumia Infinium assay. Samples with delCT value ⩽2 were qualified for further processing. As recommended by the manufacturer, 200–300 ng of dsDNA was used for generating sequencing libraries using TruSeq Amplicon cancer panel (Cat.No.FC-130-1008, Illumina, USA). The panel includes 212 amplicons of 170–190 bp length, targeting hotspot mutation in 48 cancer-related genes. DNA libraries were denatured and diluted as per Illumina protocol. Samples were run on MiSeq machine using MiSeq Reagent Kit V2 (Cat.No.MS-102-2002, Illumina).
Data analysis
The trimmed FASTQ files were generated using MiSeq Reporter (v2.6.2) from Illumina. The reads were aligned against the whole genome build hg19 in MiSeq Reporter. Subsequent analysis was done using Strand NGS v2.1.6. Reads that failed vendor QC, reads with average quality less than 20, reads with alignment score below 90, and reads with ambiguous characters were all filtered out. The Strand® NGS variant caller was used to detect variants at locations in the target regions covered by a minimum of 10 reads with at least two variant reads. Variants with a decibel score of at least 50 were reported. Substitution variants with a StrandBias more than 100% and InDel variants in homopolymer stretches more than 4 with supporting reads less than 10% were filtered out. The variants were further analyzed for their effect on the protein using the “SNP Effect Analysis” functionality in Strand NGS. This provides the effect with respect to each valid transcript. The variants were further annotated with COSMIC (Catalogue of Somatic Mutations in Cancer) database (v71) to provide information on whether that variant has been observed before in the context of any tumor type. Samples were randomly chosen and validated through Sanger sequencing, which were found to be 100% concordant with NGS analysis.
Statistical analysis
The data were analyzed using the SPSS 17.0 statistical software. Individual information and baseline characteristics were summarized using descriptive characteristics. Continuous variables were presented as mean ± standard deviation (SD). Categorical variables were expressed as frequency and percentage. The Pearson’s chi-square test or the chi-square test of association was used to determine the relationship between two categories, which included comparison of different clinicopathological characteristics with gene mutations and coexistence of two gene mutations. The p-value less than 0.05 was considered to be statistically significant.
Results
In this study, 112 CRC samples were examined for clinicopathological characteristics related to the patients and the tumors (Table 1). CRC was more frequently observed in males (60.7%) than females (39.3%). In addition, a higher proportion of patients were observed to be aged less than 60 years (65.2%), with the median age of 53 years ranging from 23 to 86 years. Primary tumors were significantly higher in colon (82.1%) than rectum (17.9%), where the tumors were equally distributed in proximal (right) colon and distal (left) colon, accounting for 41.1% each. Cecum was included within the proximal colon while sigmoid colon was included in the distal colon. The size of tumor at the time of surgery was reported to be more than or equal to 5 cm in 77 (68.8%) patients. A higher proportion of patients were observed to have tumor stage III (40.2%), followed by stage IV (36.6%) and stage I-II combined (23.2%). The N0, N1, and N2 stages of lymph node invasion were observed to be similar. However, a noteworthy variation was observed in the depth of invasion of tumor in the layers of colon/rectum, where T3 represented 67% of all cases, followed by 25.9% cases in T4, 7.1% cases in T2, and no case observed in T1. Finally, presence and absence of distant metastasis were similar (51.8% vs 48.2%) in the study population.
Clinicopathological data of 112 Indian CRC patients.

CRC: colorectal cancer.
The most frequently mutated gene was TP53 (48 mutations), followed by KRAS (43 mutations), APC (39 mutations), and PIK3CA (19 mutations). BRAF and NRAS show the lowest case of gene mutations, with eight mutations each. Tumor profiling of CRC patients was performed in parallel to genetic alterations (Table 2). KRAS mutation was significantly linked to age, where age less than 60 years was more prevalent (p = 0.041). PIK3CA mutations were predominantly harbored by male patients (p = 0.032) and were likely to be associated with tumor stage I-II (p = 0.013) and lack of metastasis in lymph nodes (p = 0.040). NRAS mutations were frequently observed in rectum over colon (p = 0.002). Finally, occurrence of APC mutations was associated with T2 stage of tumor growth in the walls of large intestine (p = 0.013), which refers to spread of tumor into the muscle layer of colon/rectum.
Correlation of KRAS, BRAF, PIK3CA, NRAS, TP53, and APC gene mutations with respect to clinicopathological data of the patient cohort.

Significant values.
KRAS mutations
KRAS mutations were observed in 40/112 (35.7%) CRC patients, giving rise to a total of 43 mutations. One patient harbored three KRAS mutations (G12D, G10R, and E49K), one patient harbored two KRAS mutations (G12D and E63K), while the rest harbored one KRAS mutation each. A higher frequency of KRAS mutations was observed in male patients (25/40, 62.5%) than female patients (15/40, 37.5%) and in proximal colon (19/40, 47.5%) when compared to distal colon (16/40, 40%). The detailed distribution of KRAS mutations has been given in Table 3. Codons 12 and 13 together constituted 35/43 (81.4%) of all KRAS mutations. All KRAS mutations comprised of missense substitution. The three most frequent mutations were substitution of glycine with aspartate or valine at codon 12 and substitution of glycine with aspartate at codon 13. These three mutations,
Mutation spectrum of KRAS, BRAF, PIK3CA, NRAS, TP53, and APC genes.
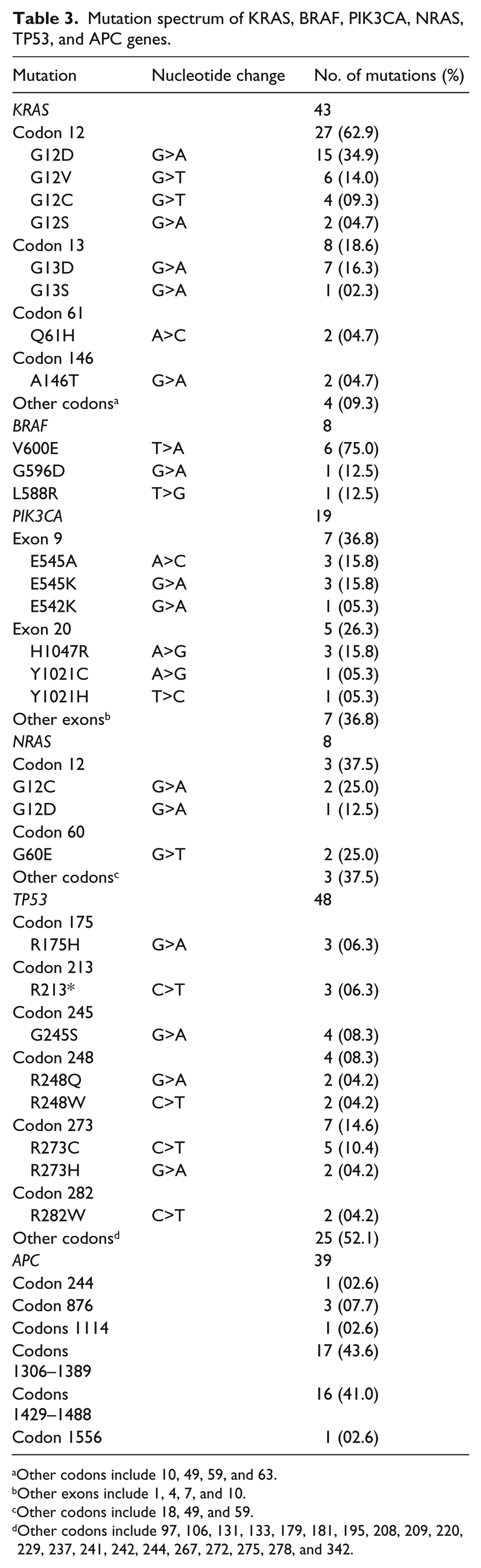
Other codons include 10, 49, 59, and 63.
Other exons include 1, 4, 7, and 10.
Other codons include 18, 49, and 59.
Other codons include 97, 106, 131, 133, 179, 181, 195, 208, 209, 220, 229, 237, 241, 242, 244, 267, 272, 275, 278, and 342.
BRAF mutations
BRAF mutations were comparatively uncommon in CRC patients (8/112, 7.14%). A single BRAF mutation was observed in every patient. All BRAF mutations were observed to be mutually exclusive with KRAS mutations. BRAF mutations were equally distributed in proximal and distal colons (3/8, 37.5% each) but were observed to be more frequent in male patients (7/8, 87.5%) than female patients (1/8, 12.5%). All eight mutations comprised of missense substitution. The most frequent BRAF mutation in our study was
PIK3CA mutations
Mutated PIK3CA was identified in 18/112 (16.1%) CRC patients. Each patient harbored one PIK3CA mutation except for one patient with two PIK3CA mutations (R88Q, H1047R). A higher frequency of PIK3CA mutations was observed in male patients (15/18, 83%) than female patients (3/18, 17%) and in proximal colon (10/18, 56%) than distal colon (6/18, 33%). Majority PIK3CA mutations were observed in exon 9 (7/19, 36.8%), exon 20 (5/19, 26.3%), and exon 1 (4/19, 21%). The remaining three exons (exon 4, exon 7, and exon 10) harbored only one PIK3CA mutation each (1/19, 5.3%; Table 3). Every PIK3CA mutation was identified to be missense substitution type. The three most frequent mutations observed were
NRAS mutations
The incidence rate of NRAS mutations was 7/112 (6.25%) in CRC patients. Only one patient harbored two NRAS mutations (G60E, T58I) simultaneously. NRAS mutations were frequent in female patients (5/7, 71.4%) than male patients (2/7, 28.6%) and in rectum (5/7, 71.4%) than colon (2/7, 28.6%). The two colon mutations were situated in the distal side. Coexistence of NRAS with KRAS was observed in one case. NRAS mutations mainly belonged to codon 12 (37.5%) and codon 60 (25%) which together comprised 55.6% of all NRAS mutations. One NRAS mutation was observed each in codons 18, 49, and 59 (Table 3). No mutations were observed in codon 13. All eight NRAS mutations were categorized under missense substitution. The two most frequent mutations observed were
TP53 mutations
The frequency of TP53 mutations was 44/112 (39.3%) in CRC patients, which harbored a total of 48 TP53 mutations. One patient harbored three TP53 mutations (C275Y, V97I, R175H), two patients harbored two TP53 mutations each (R175H, N131K; C229R, R213*), while the rest harbored one TP53 mutation each. The TP53 mutations were observed to be higher in distal colon (20/44, 45.5%) over proximal colon (13/44, 29.5%) and in male patients (25/44, 56.8%) over female patients (19/44, 43.2%). The six hotspots for TP53 mutations—codons 175, 213, 245, 248, 273, and 282—individually accounted for 3, 3, 4, 4, 7, and 2 TP53 mutations, respectively (Table 3). These codons together constituted 23/48 (47.9%) of all TP53 mutations. Missense substitution mutation (45/48, 93.8%) were most common, followed by nonsense substitution mutation (2/48, 4.2%) and frameshift deletion mutation (1/48, 2.1%). The most frequent TP53 mutations were
APC mutations
APC mutations were persistent in 33/112 (29.5%) CRC patients, contributing toward 39 mutations. One patient harbored three APC mutations (S1389FS*26, L1488FS*19, and S1465FS*3), four patients harbored two mutations each (R1432*, R1450*; P1373S, R1450*; K1308*, R1450*; and S1355P, A1366V), while rest harbored one mutation each. Majority of the APC mutations were identified between the codons 1306 and 1556 (Table 3). APC mutations were highly frequent in male patients (20/33, 60.6%) over female patients (13/33, 39.4%) and a bit high in proximal colon (13/24, 54.2%) over distal colon (11/24, 45.8%). The APC mutations largely comprised of substitution mutations (27/39, 69.2%) followed by frameshift mutations (12/39, 30.8%). In substitution mutations, the most common mutation was nonsense (22/27, 81.5%) followed by missense (5/27, 18.5%) whereas deletion (11/12, 91.6%) and insertion (1/13, 8.3%) were observed in frameshift mutations. The most common APC mutation,
Coexistence of two mutations
The coexistence of two gene mutations is highly fluctuating (Figure 1). The significant coexistence of KRAS and APC is reported in 18 cases (p = 0.024). Also, a significance is noted in mutual exclusion of KRAS with BRAF mutation (p = 0.029). Other frequently coexisting mutations include KRAS and TP53 (13 cases), TP53 and APC (10 cases), KRAS and PIK3CA (9 cases), PIK3CA and TP53 (6 cases), and PIK3CA and APC (6 cases). NRAS mutations were mutually exclusive with BRAF and PIK3CA mutations. A nearly significant result was found in association of KRAS and PIK3CA mutations where PIK3CA exon 9 was highly associated with KRAS mutations while PIK3CA exon 20 was not (p = 0.072).

Frequency of six gene mutations with combination of double mutations.
Coexistence of three and more mutations
The correlation between different gene mutations is given in Figure 2. The six genes included in this study were not found to be mutated simultaneously in a single patient. Also, no patient harbored a mutation in any five genes. The four mutations, KRAS, PIK3CA, TP53, and APC, coexisted in two cases while KRAS, NRAS, TP53, APC and BRAF, PIK3CA, TP53, APC coexisted in one case. In addition, coinciding KRAS, TP53, and APC mutations were found in four cases. Three cases were observed for co-mutation of three genes—KRAS, PIK3CA, and APC—two cases were observed for co-mutation of each of the three genes—KRAS, PIK3CA, TP53 and PIK3CA, TP53, APC. One case was observed for co-mutation of each of the three genes—KRAS, NRAS, TP53; KRAS, NRAS, APC; BRAF, PIK3CA, APC; BRAF, TP53, APC; and NRAS, TP53, APC. No case was observed with co-mutation of BRAF, PIK3CA, and TP53.

The interrelationship between occurrence of gene mutations: (a) KRAS, PIK3CA, TP53, APC; (b) BRAF, PIK3CA, TP53, APC; and (c) KRAS, NRAS, TP53, APC.
Discussion
In this study, CRC was observed to be higher in male patients (60.7%) than female patients (39.3%) which corresponds to the results reported by studies worldwide.1,34 The cancer was observed predominantly in colon (82.1%) over rectum (17.9%) where colon cancer was equally distributed between proximal and distal colon. Previously, some studies have reported a higher rate of CRC incidence in proximal colon35–37 whereas others report a higher rate in distal colon.33,38 We also observed KRAS mutations to be significantly associated with younger population, with age below 60 years (p = 0.041). Previous literature reports either no significance of KRAS mutations with respect to age or higher rate in patients above 50 years.14,39,41 Other significant values were observed in PIK3CA mutations, which were highly frequent in males (p = 0.032), tumor stage I-II (p = 0.013), and not associated with lymph nodes involvement (p = 0.040). Earlier, no significant association was observed between PIK3CA mutations and gender or tumor stage.18,39,41 NRAS mutations were significantly linked to rectum than colon (p = 0.002); however, no such observance was made in the past. 21 Also, APC was observed to be associated with T2 stage of tumor growth in the walls of large intestine (p = 0.013).
In this study, KRAS was found mutated in 35.7% CRC patients. This is in accordance with the average level of KRAS mutations (35%–45%) found in CRCs 4 and the mutation rates observed in Serbia, Turkey, and Saudi Arabia (32%–34.7%); lower than the mutation rates observed in Iraq, Spain, and Slovenia (46.2%–48%); and higher than the mutation rates observed in Greece and Morocco, and previously reported rate in India (23.5%–29.3%).14,15,38 We observed KRAS mutations to be more frequent in male patients than female patients (62.5% vs 37.5%). Earlier, a even distribution of KRAS mutations has been reported in both genders.18,39,41 Also, KRAS mutants were frequent proximal colon (47.5%) than distal colon (40%), which is similar to previous studies that reported a high frequency in proximal colon than distal colon (60%, 40%, 10 respectively, and 38%, 30%, 8 respectively). The frequency of KRAS mutations at codons 12 and 13 were observed to be 24% and 7.1%, respectively, which is lower than the previous Indian study (47% and 19%, respectively). 42 KRAS was chiefly observed to be mutated in codons 12 and 13, which together constituted 81.4% of all KRAS mutations. Previous studies have reported similar inference (81% and 90%, respectively).11,43 The three most frequent mutations observed in this study, Gly12Asp, Gly12Val, and Gly13Asp (32.6%, 11.6%, and 16.3% of all KRAS mutations, respectively), are similar to the earlier reported mutation frequencies in Germany (36%, 21.8%, and 18.8% of all KRAS mutations, respectively). 44
We identified BRAF mutations in 7.14% CRC patients, which is similar to Asian reports (5.6%) 37 and other countries including Spain, Turkey, Morocco, Slovenia, and China (5.1%–6.2%) but lower than the mutation rates observed in Serbia (17.8%). 15 Earlier, BRAF has been reported to be mutated in 9.8% CRC patients in India. 14 BRAF mutations were proportional in proximal and distal colons (6.5% each) and more frequent in male patients (10.3%) over female patients (2.3%), concordant to the previous data.6,13 However, some studies have reported higher BRAF mutations in proximal colon and in females.10–12,14 We observed only eight cases of BRAF mutations, which might be the reason behind this disparity. The V600E mutation alone constituted 75% of all BRAF mutations. This is comparable to generally reported results (i.e. 80%). 8 In Turkey, Spain, Slovenia, Serbia, and past Indian study, all BRAF mutations were reported to be V600E.14,15
The mutation frequency of PIK3CA was observed to be 16.1% in this study which falls within the range reported in CRCs (10%–20%). 16 This PIK3CA mutation rate is comparable to the mutation rates observed in Italy, France, Japan, and Australia (14%–17.8%), lower than the mutation rates in south Italy (28%) and higher than the mutation rates observed in Singapore, France, China, Poland, Switzerland, and past Indian study (2.2%–10.1%).14,17,39,41,45–47 PIK3CA mutations were higher in male patients (22.1%) than female patients (6.8%). Previous studies have either reported more PIK3CA mutations in females 41 or a even distribution between males and females.18,55 Also, PIK3CA mutations have been reported to be evenly distributed within tumor site; 18 however, we observed higher mutation frequency in proximal colon (21.7%) over distal colon (13%). Together, exon 9 and exon 20 were observed to account for 63.1% of all PIK3CA mutations, which is less than earlier reports (i.e. nearly 80%). 16 The PIK3CA mutation frequency at exon 9 (6.25%) and exon 20 (4.5%) is comparable to the Poland study (4.5% and 5.1%, respectively). 47
In this study, NRAS was mutated in 6.25% CRC tumors, similar to the mutation rates observed in Asia (i.e. 7.2%) and worldwide (i.e. 5%).19,20 A higher mutation rate has been reported in Greek and Romania (9.75%) 48 and a lower rate in Japan, China, Belgium, and America (2.2%–4.2%).18,41,45,46 Previous literature reports a even distribution of NRAS mutations with respect to gender and tumor location.18,22 However, we observed NRAS mutations to be more frequent in female patients (11.4% vs 2.9%) and all the three NRAS mutations identified in the colon were located distally. Previously, NRAS mutations were reported to be predominant in codon 61 over codons 12 and 13. 49 In this study, codon 12 represented 33.3% of all NRAS mutations; however, no mutations were observed in codons 61 or 13.
A mutant form of TP53 was persistent in 39.3% CRC patients. This result is in accordance with the frequencies reported by Sweden, Italy, Ireland, and Netherlands (38%–41%); higher than mutation rates observed in Taiwan, Switzerland, South Korea, Australia, and previously reported rate in India (20%–36%); and lower than mutation rates observed in Austria, the United Kingdom, Scotland, Japan, the United States, China, France, Spain, Poland, and Norway (45%–84%).25,29,30,38,50–53 Similar to the reported literature, TP53 mutations were more frequent in male patients and in distal colon.33,54,55 The mutation frequency in codons 175, 213, 245, 248, 273, and 282, observed in our study (6.25%, 6.25%, 8.3%, 8.3%, 14.6%, and 4.2%, respectively), can be compared to previous reports (9%, 3%, 4%, 9%, 8%, and 5%, respectively). 53 TP53 mutation in these six hotspots codons together accounted for 47.9% of all TP53 mutations.
The frequency of APC mutations was observed to be 29.5% in this study. This mutation rate is comparable to Korea, Netherlands, the United States, and China (30%–37.3%); lower than the mutation rates observed in Norway, France, the United Kingdom, Germany, Japan, Scotland, and past Indian study (46.7%–66%); and higher than the mutation rates observed in Iran, Tunisia, and Hungary (20.8%–23.3%).29,38,56,57 APC mutations were observed to be evenly distributed between proximal and distal colons (28.3% vs 23.9%) as well as male and female patients (29.4% vs 29.5%). Similar results have been reported by previous literature.29,32,33 Majority APC mutations were located between the codons 1306 and 1556. This is in accordance with study held by Attard et al. 58 that concluded that Asians have a high frequency of APC mutations between codons 1099 and 1694. There was one case with mutation at codon 1309, which is the hotspot for sporadic CRC in Chinese population. 36
This study provides a comprehensive evaluation of six gene mutations in Indian CRC patients, upgrading the number of genes examined, methodology used, and gene fragments previously studied in Indian population.14,38,42 Mutual exclusion of simultaneous mutations in six or any five genes and rare combination of gene mutation in quadruplets suggests independent pathways for CRC tumorigenesis. Only two coexisting mutations were found to be significantly related. First, BRAF was significantly associated with KRAS wild type (p = 0.029). This mutual exclusion of KRAS and BRAF is supported by previous literature.11,12,14,15 Second, KRAS and APC were found to significantly coexist in 18 cases (p = 0.024). Such relevance has not been reported earlier to the best of our knowledge. Also, NRAS and KRAS mutations were observed to be coexisting in one case. Generally, NRAS and KRAS have been observed to be mutually exclusive 21 ; however, recent studies have reported rare cases of their coexistence.18,22,40,48 In our study, both the cases with coexistence of NRAS and KRAS mutations harbored a KRAS mutation at codon 12, which corresponds to the previous reports.48,60 In addition, three cases had coexisting NRAS and TP53 mutations, which have not been reported earlier to the best of our knowledge.
Conclusion
As per our knowledge, this is the first NGS-based biomarker study in Indian CRC patients. KRAS, BRAF, PIK3CA, NRAS, TP53, and APC are frequently altered in CRC and the mutation rates were comparable to worldwide reports. No patient harbored a mutation in six genes or five genes together, and only four patients harbored a mutation simultaneously in four genes. This suggests that CRC can be initiated through different signaling pathways. The six genes were frequently altered together in pairs and triplets suggesting that their synergistic effect might escalate CRC tumorigenesis. However, a mutual exclusion was noted between KRAS and BRAF, BRAF and NRAS, and NRAS and PIK3CA. Further studies involving a greater study population are required to validate the biomarkers for developing targeted therapies with better performance covering a wider range of therapeutic targets.
Footnotes
Acknowledgements
This work was mainly performed in the Department of Research, Sir Ganga Ram Hospital. The authors thank Strand Life Sciences for their assistance in data retrieval and preparation, and Mrs Parul Takkar for statistical analysis.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards. This article does not contain any studies with animals performed by any of the authors.
Funding
This study has been funded by Merck Serono India.
Informed consent
Informed consent was obtained from all individual participants included in this study.
