Abstract
BACKGROUND:
Tissue biopsy remains the conventional technique for tumor genotyping. The main limitations are it is invasive and provides only partial snapshot during disease progression. Liquid biopsy approaches via plasma and urine are possible alternatives, and the current study aims to provide comparative analyses for plasma and urine derived disease genotyping.
METHODS:
Blood and urine specimens were collected from 150 individuals with metastatic colorectal cancer (mCRC). Patients had multiple metastases and advanced stages of cancer. Common genetic mutations including KRAS and BRAF genetic abnormalities were evaluated. Patients were also serially monitored and compared.
RESULTS:
In all cases, plasma and urine cell free DNA were successfully recovered and were of good quality for genetic analysis. Median recovered DNA from both urine and plasma samples were higher in mCRC patients than healthy volunteers indicating disease associations. Among the identified mutations, matched tumor tissue profiles compared to results from plasma ctDNA was 92%. For urine cell free DNA, the concordance among the identified mutations was 91%. Both sample types were closely matched to reference standards of tissue biopsy and indicated good clinical utility. Serial measurements indicated trends within each patient group that was linked with disease outcome.
CONCLUSIONS:
In the current study, our data indicated that both plasma and urine cell free DNA can be utilized to address possible disease progression in colorectal cancer patients. More importantly, this also provide risk stratifications that correlated to disease outcome. This can potentially aid in early clinical intervention for patients with possibly worse outcomes.
Introduction
Advances in technology have permitted detailed molecular detection of metastatic colorectal cancer (mCRC) crucial for targeted therapies [1, 2]. mCRC is one of the leading causes of mortality worldwide [3]. Challenges in mCRC remain for early detection as the disease may not be symptomatic at initial stages [4]. There are also difficulties in patients’ follow ups to determine disease progression. Current techniques rely on conventional tissue biopsies to gain access to tumor materials [5] and radiographic imaging to understand the extend of the disease [6]. Cancer is a complex disease and may morph constantly as a result of clinical intervention to adapt to the new environment [7]. Patients will benefit from regular genetic monitoring to understand changes at the molecular levels that are critical for treatments such as cetuximab and panitumumab [8]. Tissue profiling may not also comprehensively address genetic heterogeneity in the disease as sampling are usually localized. It does not also cover secondary tumors as a result of metastasis.
Recent advances in oncology research showed that molecular diagnosis is possible with tumor materials from peripheral blood [9] and urine [10]. The methods are attractive as these are less invasive than tumor biopsy and may cover the disease heterogeneity better [11]. This approach mainly relies on cell free DNA released from necrotic cells present in the blood circulatory system [12]. The blood circulatory system is one convenient route for the dissemination and movement of tumor cells to secondary sites during the process of metastasis [13]. A good portion of this material is then removed by the body’s renal system and present in the urine [14]. The term liquid biopsy has most often been used to associate with disease detection using these means [15]. The clear advantage is ease of sample collection, which makes this an ideal technique for long term serial monitoring of disease changes.
Current clinical studies have uncovered various potential uses of these alternative tumor materials. Plasma DNA has been extensively covered in various studies but urine testing for cell free DNA is limited. In several longitudinal analyses, the concentration of plasma DNA in disease patients were clearly associated with disease prognosis [16]. Several groups had also demonstrated good agreement with tumor tissues for various tumor markers [17, 18]. In the current study, we aimed at testing the utility of plasma and urine cell free DNA in late stage mCRC patients and provide a direct comparison of both parameters. This fills the gap to better understand the kinetics of urine cell free DNA. In the post-treatment phase, patients were monitored and their trends in cell free DNA tracked to capture mCRC dynamics.
Baseline clinicopathological factors
Baseline clinicopathological factors
Study design and patient population
A total of 150 patients recruited were examined. All patients provided informed consent to be part of the study. The institutional review board (IRB) approved all procedures and ensured ethical compliance. Blood and urine were collected on the same day to prevent discrepancy. Patients were selected randomly before mutational profiling. The inclusion criteria were patients with advanced stage of colorectal cancer and all had multiple metastases with intact primary tumor in the colon. Details of patients’ characteristics are provided in Table 1. As part of the study, serial samplings were taken at different time points from each participant. A total of 30 healthy volunteers participated to provide healthy reference measurements. All healthy volunteers were of certain disease free and had no history of cancer. Serial sampling was performed on both patients and healthy donors at monthly intervals.
Baseline cell free DNA from plasma and urine specimens was also recovered from these volunteers. Mean age of patients and healthy cohorts were 53 and 52 years old respectively. For all 180 subjects of the study, basic clinicopathological factors are shown in Table 1. Including all serial measurements, a total of 1,080 plasma and urine samples were processed. Molecular profiling was performed for both KRAS and BRAF mutations. Patient survivals were subsequently followed up and correlated to cell free DNA measurements from urine and plasma. At baseline profiles, tumor tissue biopsies indicated 72 KRAS and 38 BRAF positive colorectal cancer patients. The rest were of wildtype characteristics. We performed baseline comparisons with healthy volunteer’s blood and urine specimens, assuming all were of wildtype characteristics.
Liquid biopsy sample collections and initial purifications
Peripheral blood was collected in a 10 ml K2 EDTA blood tube. For urine, the first morning urine was collected. 40–50 ml of urine specimen was taken and transferred to a 50 ml sterile centrifuge tube. Both specimens were processed within 3 hrs. For blood sample pre-processing, plasma was extracted via direct centrifugation at 4
Profiling of plasma and urine specimens
Cell free DNA was immediately purified fromplasma and urine specimens as it may degrade with prolong storage in the media. To purify nucleic acid, the Qiagen’s QIAmp Circulating Nucleic Acid Kit (Qiagen Inc, USA) was used. The processes used spin columns coupled with multiple centrifugation steps and followed manufacturer’s instructions. Sample was eluted in TE buffer (20 ul) and measured on the Nanodrop 2000 (ThermoScientific Inc., USA). All purified DNA specimens were then stored at 20
95 94 98
Post PCR, the reaction plates were analyzed on the QX200 droplet reader (Bio-Rad Inc., USA) using the QuantaSoft software included.
Normality testing using the Shapiro-Wilk test was performed on recovered DNA concentrations from both urine and plasma specimens. Agreement between matched tumor genetic results were compiled to address assay sensitivity and specificity. Correlations between healthy volunteers at reference measurements were performed using Student t test. All data presented were expressed as mean and standard deviation unless otherwise specified. Serial measurements were tested using analysis of variance (ANOVA). Analyses of survival outcomes were performed using Kaplan Meier (KM) estimate. The hazard ratio (HR) between different group of patients was computed to assess the clinical utility of cell free DNA. All statistical analyses were deemed significant if
Comparison of results at baseline with genotyping from primary tumor tissue. (A) Concordance rates among different study groups. Healthy volunteers and patients with wildtype KRAS/BRAF were 100% concordant. (B) Sensitivity and specificity among urine based cell free DNA test. (C) Sensitivity and specificity among plasma based cell free DNA test.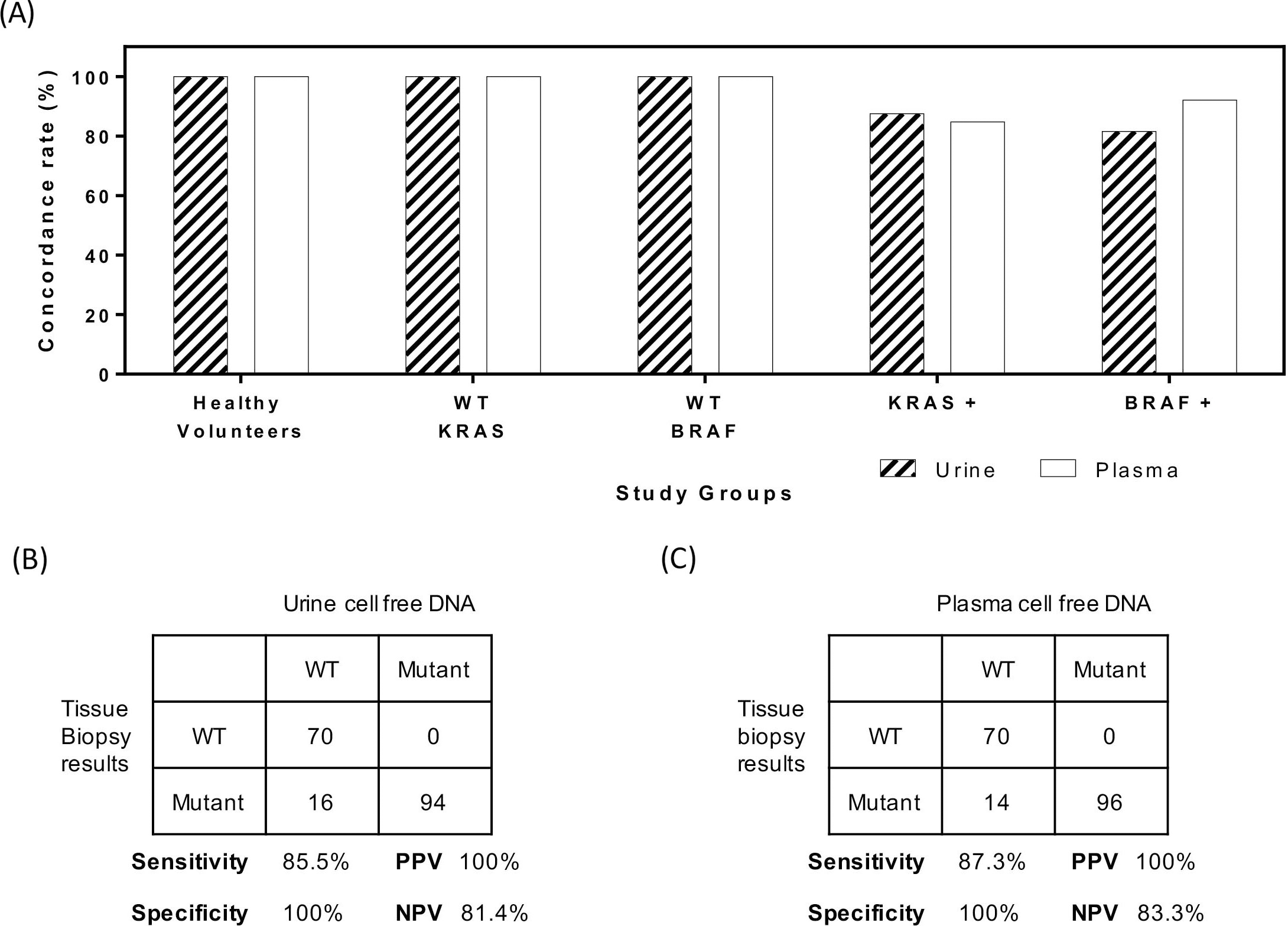
Study design and matched tumor correlation at baseline
A total of 150 patients with primary colorectal cancer were recruited and subsequently followed through in a serial measurement study. Patient recruitment was performed between 2015 to 2017. In addition, 30 healthy volunteers were used as wildtype controls in the genetic profiling of KRAS andBRAF genes. Figure 1a collates the results from genetic profiling of urine and plasma specimens. For healthy volunteers, 100% were wildtype for KRAS andBRAF identified from cell free DNA of plasma and urine specimens. This was similarly observed in patient groups with wildtype KRAS and BRAF detected from tumor biopsies. In the patient’s group with KRAS mutations, we observed a good agreement between tumor tissue and cell free DNA. For urine and plasma cell free DNA, the concordance of patients with mutations were 85% and 87% respectively. To address the sensitivity and specificity for using plasma or urine cell free DNA, we compiled individually the results as shown in Fig. 1b and c. For plasma DNA, the overall agreement with primary tissue biopsy samples was 92% with an established sensitivity and specificity of 87.3% and 100% respectively. For urine cell free DNA, the overall agreement was 91% and corresponding sensitivity and specificity were 85.5% and 100% respectively. The results indicated a high percentage of mutant DNA present in cancer patients were readily detectable in both plasma and urine specimens.
Total cell free DNA recovered from each participant and comparison between plasma and urine cell free DNA. (A) Plasma cell free DNA concentration among different study groups. (B) Urine cell free DNA among different study groups. (C) Spearman correlation coefficient among colorectal patients with wildtype KRAS/BRAF. (D) Spearman correlation coefficient among colorectal cancer patients with positive KRAS or BRAF mutations.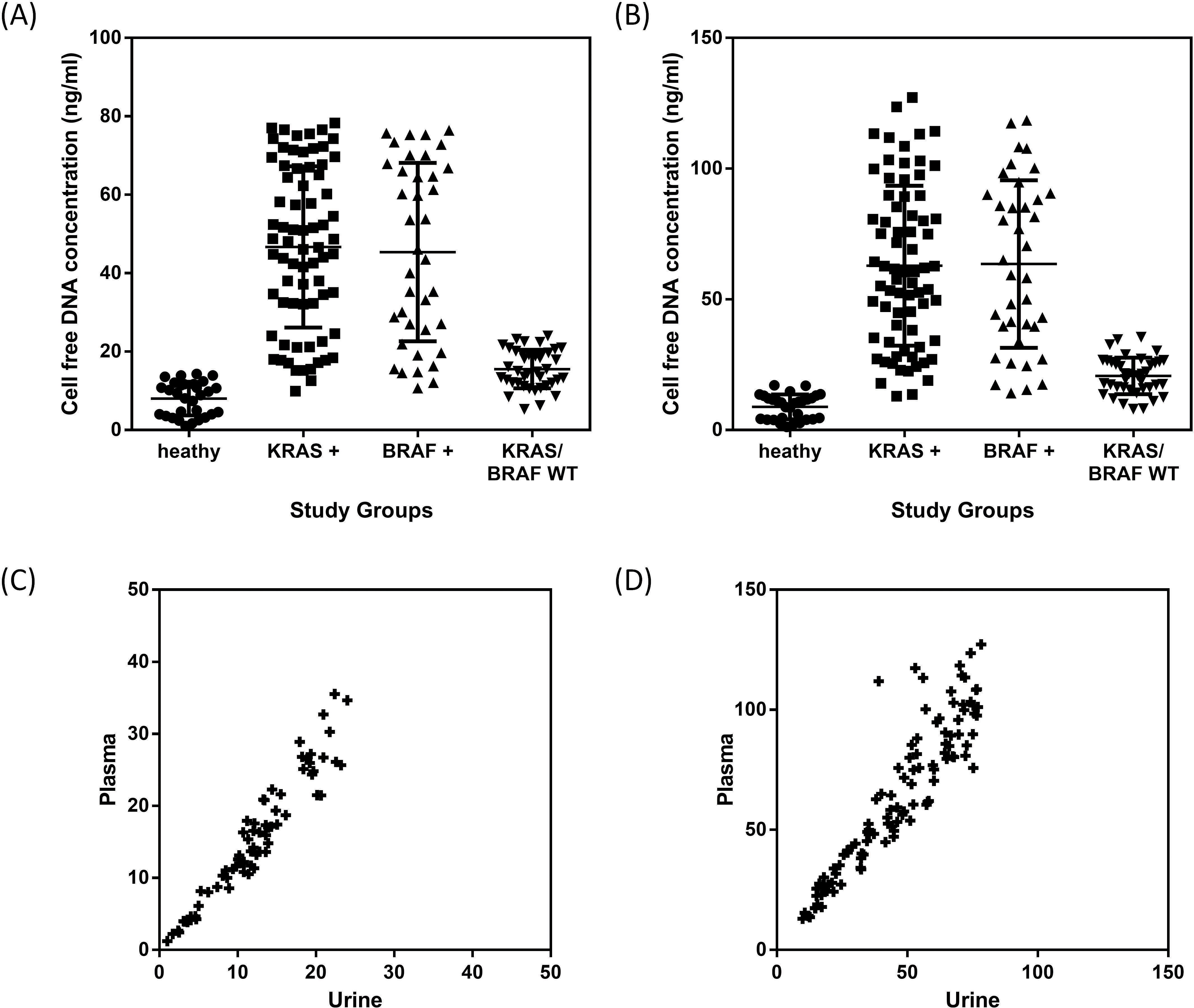
Serial monitoring of study participants using plasma and urine cell free DNA. (A) Measurement of concentration changes in colorectal cancer patients using urine cell free DNA. Healthy volunteers observed no significant variations based on a repeat measure ANOVA. (B) Measurement of concentration changes in colorectal cancer patients using plasma cell free DNA. Healthy volunteers were observed to have stable mean levels.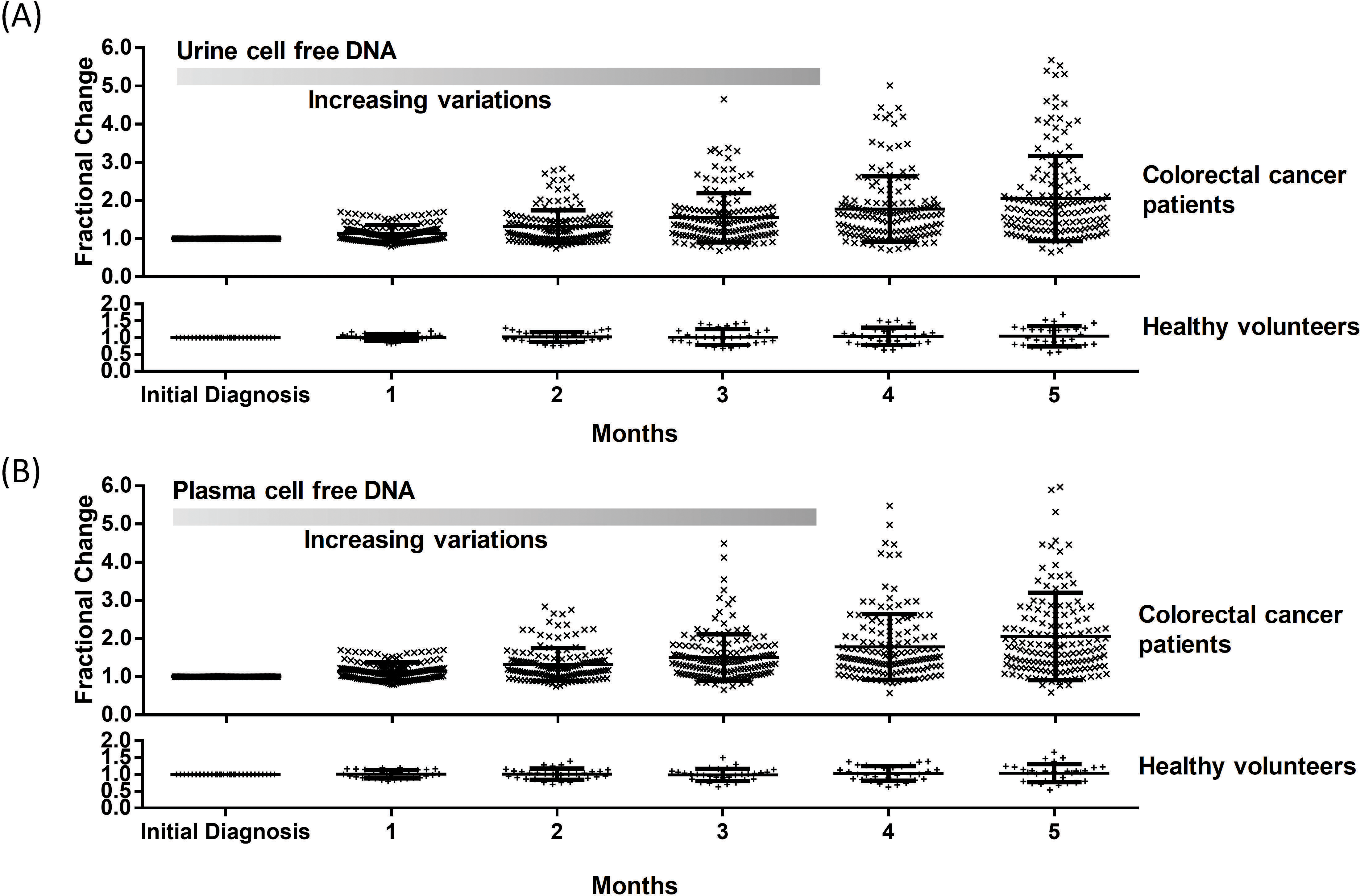
Survival analysis of colorectal cancer patients based on maximum recovered cell free DNA. (A) Plasma DNA separation based on median split of recovered cell free DNA. (B) Urine cell free DNA stratification of colorectal cancer patients demonstrating that concentrations of cell free DNA is useful to identify patients with poor outcome.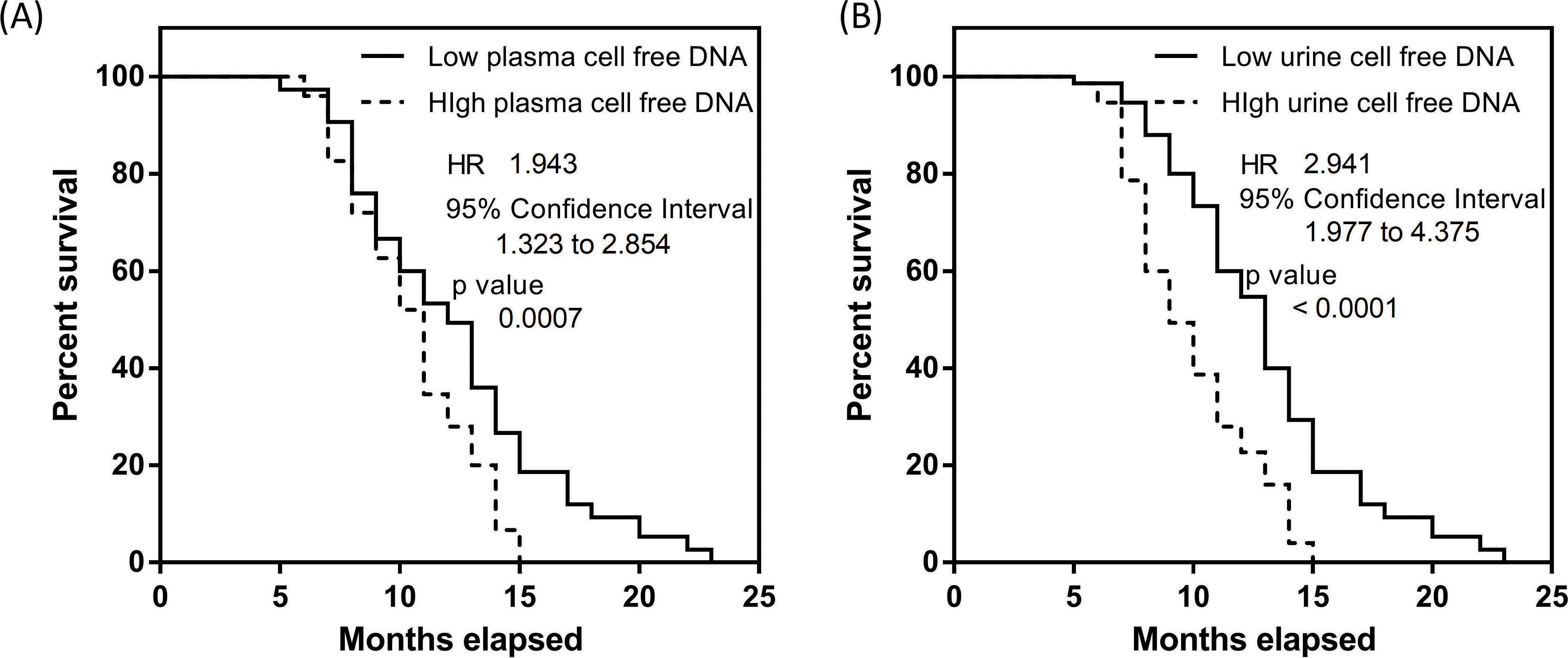
Levels of cell free DNA derived from both plasma and urine were compared among the different groups. This ascertains if different groups of study participants have different profiles. Figure 2a shows the results of plasma cell free DNA among healthy volunteers (mean
In serial monitoring of cancer patients, changes in cell free DNA were monitored for all participants over an extended period of 5 months. Interestingly, results were seen to be relatively stable comparing plasma to urine specimens for healthy volunteers (Fig. 3a). A repeated measure ANOVA showed no significant differences (
Trends in recovered cell free DNA relate to survival outcome
Interestingly, as we addressed each patient separately, we observed that in both cases for urine and plasma cell free DNA, not all patients experienced an increase in DNA levels when compared to baseline measurements. Approximately 10% of the entire cohort measured using both plasma and urine specimens experienced a decrease in levels. We postulated that these differences in trends may be disease related and potentially have prognostic utility. Following up with patients over and extended 2 years, we performed the KM analysis for both plasma and urine testing as shown in Fig. 4a and b. To highlight if changes in cell free DNA comparing to baseline are significant, we split the patient cohort into two equal sized groups. The first group had larger variations in cell free DNA among the 5 measurements taken post baseline and the remaining had lower cell free DNA changes. In plasma DNA analysis, we observed a significant split of the cohort with a determined HR of 1.94 (95% CI 1.32 to 2.85) as shown in Fig. 4a. Median survival rates of the patient group with poorer outcome was 11 months. Similarly, for urine cell free DNA (Fig. 4b), HR was 2.94 (95% CI 1.98 to 4.38). Median survival of patients with higher urine cell free DNA had median survival of 9 months compared with the alternative group with 13 months. The results indicated that cell free DNA may be prognostic and highlights the value of continual monitoring for cancer patients.
Discussion
In the current comparative analysis of paired plasma and urine samples, we observed several interesting trends and correlations for colorectal cancer patients. The need for alternative tumor material sampling other than tumor tissue biopsy arises as continual monitoring of cancer patients will better aid in their clinical management. Plasma and urine are attractive as specimen collections are relatively straightforward and more acceptable to patients. In the current study addressing advanced stage colorectal cancer, where access to tumor tissue is challenging, these assays provide alternative strategies to profile the disease.
To ascertain relevance of cell free DNA in plasma and urine to the primary tumor, molecular characterization using KRAS and BRAF mutations was investigated. These mutations typically reflect poor prognosis for colorectal cancer patients [19] and are critical for clinical management. For healthy volunteers, we confirmed all were wildtype for KRAS and BRAF using cell free DNA derived from either plasma or urine. This is similarly true for colorectal cancer patients with wildtype KRAS or BRAF detected by tumor biopsies. These demonstrated good specificities in both urine and plasma tests. Majority of patients with either KRAS or BRAF mutations were correctly identified and no significant differences were observed between the concordance rates in KRAS or BRAF detection using cell free DNA. Collectively, plasma cell free DNA had a marginally higher concordance rates with matched tumor biopsy results than urine cell free DNA as shown in Fig. 1. Our results are largely coherent with other studies that investigated the use of plasma cell free DNA [20, 21, 22]. Our results demonstrate much better concordance than the results shown by Kuo et al. [20], who observed than the agreement rate was 78.8% for KRAS mutations. This is likely due to the differences in patient cohort where in our case all patients were in advanced stages. In newer technologies presented by Powell et al. [22], better clinical sensitivities were shown and with the advent of new assays, we expect the efficacy of the assay to improve. In additional, our results demonstrate that urine is also a viable source for accurate detection of key driver mutations of the disease. In direct comparison of both assays in this study, we detected a good correlation among both assays. Using a Spearman’s correlation, we noted that the levels of recovered nucleic acid were similar in both cases. Our analyses also revealed that this is irrespective of the presence of mutations as the correlation coefficients in the wildtype study group was similar to the patient group that had KRAS or BRAF mutations.
Levels of recovered cell free DNA for different study groups highlighted a distinctive pattern for patients with KRAS or BRAF mutations. Both urine and plasma cell free DNA analyses showed similar trends. Mean recovered concentrations of cell free DNA were much higher than patients with wildtype characteristics or healthy volunteers. Given that patients with KRAS or BRAF mutations tended to have poor prognosis, we postulate that the levels of cell free DNA may be related to clinical outcome as well. Overall, we observed that mean recovered cell free DNA from urine and plasma were much higher in colorectal cancer patients, coherent with prior studies [23, 24] performed on plasma samples. A clear advantage of liquid biopsy via urine and plasma is the ability for serial monitoring of patients as shown in Fig. 3. Normalizing by the baseline reference levels obtained with either the plasma or urine specimen, we observed in a span of 5 months a monotonic increasing level of cell free DNA in the patient cohort. Conversely, healthy volunteers’ measurements showed no significant changes as established using a repeated measure ANOVA test. To address if the changes in cell free DNA significantly impact the prognosis of these patients, we split the patient group equally based on an ascending ranking of recovered cell free DNA and performed a KM analysis. This was tracked over 2 years with regular follow up calls or visits. In both cases using either plasma or urine cell free DNA, the results favored patients with lower cell free DNA during the monitoring period. A direct comparison between urine and plasma cell free DNA however showed that the HR was higher in the case for patient stratifications using urine cell free DNA. Potentially, both plasma and urine can be good candidates to aid in long term monitoring of colorectal cancer patients.
Our study showed highly feasible uses of cell free DNA derived from urine and plasma samples. This has clear advantage over classical tumor biopsy. A major limitation of the assay is the small numbers of false negative results in the patient group with mutations. This is likely due to low concentrations of mutant DNA masked in background normal DNA [25, 26, 27] as observed in other studies. It may not be entirely feasible to replace tumor biopsy at this juncture until more sensitive molecular techniques can be applied but plasma and urine cell free DNA serves well to complement existing disease management strategies. The study by Thomsen et al. [27] also showed insignificant correlative analysis to treatment outcome, which demonstrates that these assays require much more validation studies before clinical adoption. We noted that serial monitoring of patients can be beneficial for better risk stratifications and possibly other forms of cancer detection such as circulating tumor cells (CTCs) can further aid to refine the process. CTCs from peripheral blood may work better as background blood cells can be effectively removed [28]. Future studies may focus on implementing other companion tests to enhance the detection of these key driver mutations for colorectal cancer. In the current study, we addressed common mutations in both KRAS and BRAF that are linked to treatment efficacy. Given the wide spectrum of mutations present in these genes, it is of value for a large scale genomic analysis to address disease heterogeneity. This would be valuable to better utilize cell free DNA in colorectal cancer management. Additionally, in earlier concordance analysis (Fig. 1A), we observed different sensitivities associated with KRAS and BRAF testing. Further studies to compare the sensitivities with larger sample sizes or using both assays in tandem will be useful before the adoption into wider clinical practice.
Conclusions
Our data established clear and effective use of cell free DNA from either plasma or urine specimens for colorectal cancer patients. The results showed highly concordant data for key driver mutations found on tumor tissues and cell free DNA form urine and plasma. A direct comparison of urine and plasma cell free DNA showed relatively similar trends in serial monitoring of patients. This potentially has clinical utility to address current shortfalls for sensitive monitoring of colorectal cancer patients during disease progression.
Footnotes
Acknowledgments
This work was supported by research grants provided by Xiangyang Central Hospital.
Conflict of interest
All authors declare no conflict of interest.
Supplementary data
ddPCR assay information for KRAS
Mutation
Locus
Assay ID (Mutation: WT)
Amplicon size (bp)
c.38G
A
chr12:25398281
dHsaCP2500598: dHsaCP2500599
57
c.35G
T
Chr12:25398284
dHsaCP2500592: dHsaCP2500593
57
c.35G
A
Chr12:25398284
dHsaCP2500596: dHsaCP2500597
57
c.35G
C
chr12:25398284
dHsaCP2500586: dHsaCP2500587
57
c.34G
A
Chr12:25398285
dHsaCP2500588: dHsaCP2500589
57
c.34G
T
Chr12:25398285
dHsaCP2500584: dHsaCP2500585
57
