Abstract
Background:
Lung adenocarcinoma (LUAD) is a common type of malignant tumor with therapeutic challenges. Cancer-associated fibroblasts (CAFs) promote LUAD growth and metastasis, regulate the tumor immune response, and influence tumor treatment responses and drug resistance. However, the molecular mechanisms through which CAFs control LUAD progression are largely unknown. In this study, we aimed to determine the correlations between CAF-related genes and overall survival (OS) in patients with LUAD.
Methods:
We acquired the gene expression data and clinical information of 522 patients with LUAD patients from The Cancer Genome Atlas (TCGA) and 442 patients with LUAD from the Gene Expression Omnibus (GEO) databases. CAF infiltration levels were assessed using the Microenvironment Cell Population (MCP) counter, the Estimating the Proportions of Immune and Cancer cells (EPIC) algorithm, and Tumor Immune Dysfunction and Exclusion (TIDE) scores. A CAF-related gene network was constructed using the Weighted gene co-expression network analysis (WGCNA). Based on the CAF-related genes, univariate Cox regression and Least Absolute Shrinkage and Selection Operator (LASSO) Cox regression analyses were performed to identify prognostic genes. Gene expression levels within the prognostic model were validated using the Cancer Cell Line Encyclopedia (CCLE) databases and Western blotting.
Results:
Our results demonstrated that high CAF scores were associated with lower survival rates in patients with LUAD. Gene modules that were highly correlated with high CAF scores were closely associated with tissue characteristics and extracellular matrix structures in LUAD. In addition, correlations between CAF scores and responses to immunotherapy and chemotherapy were observed. Finally, we found that SNAI2 expression was higher in lung cancer tissues than in normal tissues.
Conclusion:
Deepening our understanding of the influence of CAFs on tumor progression and treatment response at the molecular level can aid the development of more effective therapeutic strategies. This study provides important insights into the functional mechanisms of action of CAFs in LUAD and highlights their clinical implications.
Introduction
Lung cancer is among the most common malignancies worldwide and is the leading cause of cancer-related deaths. 1 The incidence of lung cancer has declined in the last century; however, mortality rates remain high. Non-small-cell lung cancer (NSCLC) accounts for approximately 80% to 85% of all lung cancer cases. 2 Lung adenocarcinoma (LUAD) is the predominant subtype of NSCLC worldwide and is a highly heterogeneous tumor. 3 Many patients with LUAD have similar pathologies, gene mutations, and disease stages; however, their prognoses differ considerably depending on the location and time of the gene mutation and the tumor microenvironment (TME).4,5
The TME negatively influences the efficacy of immunotherapy because it suppresses immune cell function and promotes cancer cell survival, local invasion, and metastatic dissemination.6-8 The TME of LUAD is mainly composed of various highly heterogeneous cell types, including tumor cells, fibroblasts, endothelial cells, immune cells, vasculature, and the extracellular matrix (ECM).8-10 Fibroblasts found in tumor tissues, termed cancer-associated fibroblasts (CAFs), are one of the major types of stromal cells that influence homeostasis. 11 CAFs promote tumor progression by secreting factors that stimulate angiogenesis, invasion, and cancer cell proliferation. 12 Moreover, they enhance the formation of an immunosuppressive network during antitumor therapy by secreting growth factors and cytokines, paracrine signaling, and ECM remodeling.13-15 Therefore, the use of CAFs as candidate targets for targeted molecular therapy is an appealing and emerging strategy.
Snail family transcriptional repressor 2 (SNAI2) is a classical epithelial-mesenchymal transition (EMT)-related transcriptional inhibitor with an N-terminal SNAG domain and a C-terminal DNA-binding domain. 16 SNAI2 plays an important role in the progression of several cancers. Reprogramming of mesenchymal fibroblasts by SNAI2 contributes to tumor fibrous tissue proliferation and promotes progression and invasive metastasis of ovarian cancer by regulating EMT through the lncRNA AC005224.4/miR-140-3p/SNAI2 axis.17,18 FBXO28 can inhibit hepatocellular carcinoma invasion and metastasis by promoting protein kinase A (PKA)-dependent phosphorylation-mediated degradation of SNAI2. 19 SNAI2 can significantly influence the TME by reactivating the tumor stroma and generating an immunosuppressive microenvironment in prostate cancer. 20 In breast cancer, ASB13 inhibits metastasis by promoting SNAI2 degradation. 21 However, whether SNAI2 is a CAF-related gene requires further investigation.
Furthermore, the clinical relevance of and molecular mechanisms by which CAFs regulate tumor progression remain unclear. In this study, we aimed to construct a model to determine the correlations between CAF-related genes and overall survival (OS) in patients with LUAD. CAF infiltration scores were calculated using the Estimating the Proportions of Immune and Cancer cells (EPIC) method, Microenvironment Cell Population (MCP) counter, and Tumor Immune Dysfunction and Exclusion (TIDE) scoring to determine the relationship between the CAF scores and OS in patients with LUAD based on data from the publicly available Gene Expression Omnibus (GEO) and The Cancer Genome Atlas (TCGA) databases. Weighted gene co-expression network analysis (WGCNA) was used to identify the hub modules that were most strongly correlated with CAF infiltration.
Material and Methods
Data of patients with LUAD
The gene expression profiles of 522 patients with LUAD and corresponding patient clinical characteristics, including outcome, age, and survival time, were downloaded from TCGA (https://gdc.nci.nih.gov/). The batch effects were revised using the sva R package. The fragments per kilobase of transcript per million mapped reads (FPKM) format was used for the RNA-seq data from TCGA datasets. The normalized FPKM values were converted to transcripts per million (TPM) and log2(TPM + 1) transformed. In addition, we included the normalized gene expression profiles of 442 patients with LUAD from the GSE72094 dataset from the GEO database (https://www.ncbi.nlm.nih.gov/geo/), with complete information regarding OS-related transcriptomics, in the validation dataset.
Assessment of CAF scores
CAF infiltration scores were evaluated using three methods—estimation of the marker gene expression–based MCP counter, 22 the EPIC algorithm, 23 and TIDE scores downloaded from the TIDE database (http://tide.dfci.harvard.edu). Tumor sample data obtained from TCGA and the GEO databases were divided into high- and low-CAF-score groups according to the cutoff value of the CAF score. Then, we analyzed the relationship between CAF scores and survival using the Survival R package, and samples were screened based on P-values < .05.
CAF co-expression network construction and hub gene selection
Gene co-expression network analysis and the selection of hub genes that target CAF infiltration were performed using the WGCNA R package, 24 and the genes with the top 5000 median absolute deviations (MADs) in both the TCGA and GSE72094 datasets were chosen for WGCNA analysis. Thereafter, the appropriate soft-thresholding power β for adjacency computation was graphically confirmed. Subsequently, the adjacency matrix was clustered using a topological overlap matrix (TOM) and the corresponding dissimilarity matrix (1–TOM); a hierarchical clustering dendrogram was constructed, and genes with similar expressions were divided into different modules. The first principal component of each module expression was summarized by the module eigengene (ME). Pearson’s correlations between the ME and clinical features were calculated, and the module with the strongest correlation was selected for subsequent analyses. Next, we evaluated module membership (MM) and gene significance (GS) for the traits of individual genes in the hub module, and hub genes with GS > 0.4 and MM > 0.6 were selected. Finally, the overlapping hub genes between the TCGA and GSE72094 datasets constituted the final set of hub genes.
Establishment and validation of the CAF-related prognostic signature
CAF-related hub genes from TCGA were subjected to univariate Cox regression analysis to screen for prognosis-related hallmark genes using the Survival R package, with P < .05. To construct a prognostic correlation model, we performed Least Absolute Shrinkage and Selection Operator (LASSO) 25 Cox regression analysis. CAF-related prognostic genes from TCGA were used as the training set, and CAF-related hub genes from the GEO dataset were used as the testing set. Patients in the two datasets were divided into high- and low-risk groups based on their median risk score. Risk scores were calculated using the following formula:
Gene set enrichment analysis and single-sample gene set enrichment analysis of TCGA cohort
To explore unique Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways and hallmark gene sets in the high- and low-risk groups, gene set enrichment analysis (GSEA) was performed based on the hallmark gene sets, using the clusterProfiler package in R, and pathways with P < .05 were considered to be significantly enriched. Single-sample GSEA (ssGSEA) was performed on several of the enriched pathways using the GSVA R package.
Cancer cell line encyclopedia and HPA datasets
To verify the expression levels of genes involved in the construction of prognostic models, we downloaded mRNA expression data from the Cancer Cell Line Encyclopedia (CCLE) database (https://portals.broadinstitute.org/ccle) 26 and compared the expression levels of these genes between fibroblasts and lung cell lines using the plyr R package and the Wilcoxon test. In addition, the protein expression of CAF-related genes in lung tissues was determined using immunohistochemical (IHC) images from the HPA online database (https://www.proteinatlas.org/). 27
Chemotherapeutic sensitivity predictions
To predict chemosensitivity in the high- and low-risk groups, we downloaded drug sensitivity and data representation files from the largest publicly available pharmacogenomics database, Genomics of Drug Sensitivity in Cancer (GDSC; https://www.Cancerrxgene.org/), 28 and performed chemosensitivity analysis using the oncoPredict R package.
Genetic somatic mutation analysis
Genetic somatic mutation data for the LUAD samples were downloaded from TCGA. The maftools R package was used for the recognition and visualization of somatic variants in both the low- and high-risk groups. The tumor mutational burden (TMB), a sum of the nonsynonymous mutations per million bases in coding regions, has been proposed as a biomarker of immunotherapy efficacy. 29
Cell culture
The human LUAD cell lines H1573, H1975, HCC827, A549, and SPC-A-1 were purchased from the Kunming Institute of Zoology. These cell lines were cultured in RPMI 1640 medium supplemented with 10% v/v fetal bovine serum (FBS), 10 000 units of penicillin, and 10 mg/mL streptomycin, at 37°C in an incubator with 5% CO2. The human bronchial epithelial cells BESA-2b, obtained from iCell, China, were cultured in Dulbecco’s modified Eagle’s medium (DMEM) supplemented with 10% v/v FBS, 10 000 units of penicillin, and 10 mg/mL of streptomycin at 37°C in an incubator with 5% CO2.
Western blot analysis
A total of 1 × 106 cells were collected and lysed in radioimmunoprecipitation assay (RIPA) lysis buffer (Beyotime, Shanghai, China). The cell lysate was centrifuged to isolate proteins (30 or 35 μg), which were collected, resolved using Sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) with a 10% resolving gel, and electroblotted onto Polyvinylidene fluoride (PVDF) membranes (Millipore, Carrigtwohill, Germany). The membranes were blocked with 5% (w/v) dry skimmed milk in Tris buffered saline (TBS) and incubated overnight at 4°C with primary antibodies against SNAI2 (Proteintech, Cat#12129-1-AP, USA). The membranes were then washed with TBS containing 0.1% (v/v) Tween-20 and incubated with a horseradish peroxidase–conjugated secondary antibody (Proteintech, USA).
Statistical analysis
All statistical analyses were performed using the R software. The Wilcoxon test was used for pairwise comparisons. The Survival and Survminer R packages were used for OS analysis using the Kaplan–Meier curve with the log-rank test. P < .05 was considered statistically significant.
Results
Survival analysis of CAF scores in patients with LUAD
We calculated CAF infiltration scores using the EPIC, MCP counter, and TIDE methods. OS analysis using the Kaplan–Meier curve with the log-rank test demonstrated that the high-CAF-score group exhibited worse OS than the low-CAF-score group in both the GSE72094 (Figure 1A) and TCGA (Figure 1B) cohorts of patients with LUAD. This highlights the importance of further research on CAF in LUAD.

Survival analysis of patients with lung adenocarcinoma (LUAD) using cancer-associated fibroblast (CAF) scores. (A) Patients with LUAD and increased CAF infiltration had worse overall survival (OS), as revealed by Kaplan–Meier analysis of The Cancer Genome Atlas (TCGA) data. (B) Patients with LUAD and increased CAF infiltration had worse OS, as revealed by Kaplan–Meier analysis of the GSE72094 data.
Construction of CAF-related modules and selection of hub genes
To construct a co-expression network of CAF scores, we performed WGCNA in the GSE72094 and TCGA cohorts. Regarding the TCGA cohort, the CAF scores had the strongest positive correlation with the green module (Figure 2A, R = 0.76, P < .001). For the GSE72094 cohort, the CAF scores had the strongest positive correlation with the yellow module (Figure 2B, R = 0.84, P < .001). Accordingly, we further explored the genes in these two modules and screened 91 genes of the 204 genes in the green module of TCGA dataset and 159 genes in the yellow module of GSE72094 as hub genes, which were visualized using a Venn diagram (Figure 2C).

Cancer-associated fibroblast (CAF)-related modules constructed using weighted gene co-expression network analysis (WGCNA). (A, B) Module–trait relationships showing correlations between each gene module eigengene and phenotype in The Cancer Genome Atlas [TCGA] (A) and GSE72094 cohorts (B). (C) Venn diagram presenting overlapping TCGA and GSE72094 module genes. (D) Gene ontology (GO) analysis of 91 genes and the enriched biological process (BP), cellular component (CC), and molecular function (MF) terms. (E) Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis of the enriched pathways of the 91 genes. (F) Forest diagram showing overall survival (OS)-associated genes, as revealed by univariate Cox analysis of TCGA data. (G) Least Absolute Shrinkage and Selection Operator (LASSO) Cox regression was conducted to construct the prognostic model.
Then, we performed gene ontology (GO) and KEGG analyses of the 91 hub genes. ECM and extracellular structure organization in the biological process (BP) category, collagen-containing ECM in the cellular component (CC) category, and ECM structural constituents in the molecular function (MF) category were the main significantly enriched GO terms. These are key components of ECM processes in the TME (Figure 2D). The enriched KEGG pathways were mainly involved in desmoplastic and proliferation processes, such as focal adhesion, ECM–receptor interaction, the PI3 K-Akt signaling pathway, and the TGF-beta signaling pathway (Figure 2E). These enrichment terms are associated with the prognosis of patients with LUAD.
Prognostic risk model based on CAF-related genes
We screened 28 OS-related genes (ITGA5, MN1, SNAI2, COL6A1, FRMD6, NID2, GFPT2, COL5A3, COL6A2, MMP14, LOX, CLMP, TMEM158, TNFAIP6, CPXM1, LOXL2, PXDN, COL5A1, COL5A2, GPC6, MXRA5, CTHRC1, VCAN, ADAM12, SPOCK1, COL12A1, GREM1, and COL11A1) with P < .05 from the 91 common hub genes to establish a prognostic risk model based on CAF-related genes in patients with LUAD using LASSO-penalized Cox analysis (Figure 2F). Initially, 522 samples from TCGA were used as the training group, and 442 samples from GSE72094 were used as the testing group; a cross-validation method was used to optimize the prognostic model (Figure 2G). Subsequently, patients with LUAD were divided into low- and high-risk groups according to the median cutoff values. Kaplan–Meier curves revealed that the OS of patients with LUAD in the low-risk group was longer than that of the patients in the high-risk group in both the TCGA and GSE72094 datasets (Figure 3A and B).
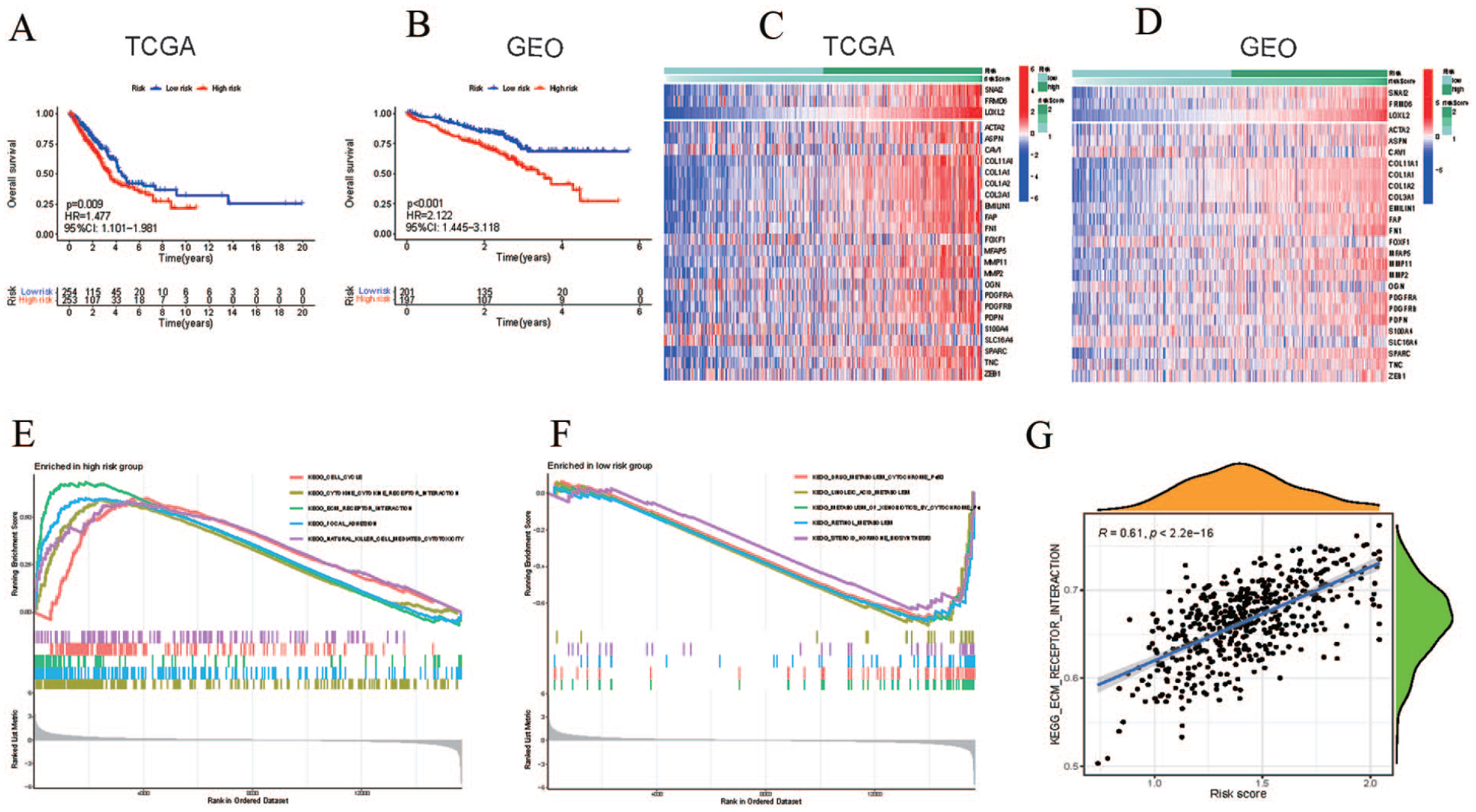
Differences in pathways between the two groups. (A, B) Kaplan–Meier survival curves for patients with lung adenocarcinoma (LUAD) in the high- and low-risk groups from The Cancer Genome Atlas (TCGA) and GSE72094 cohorts. (C, D) Heat map revealing the expression patterns of the three identified cancer-associated fibroblast (CAF) genes and the CAF risk scores in TCGA and GSE72094. (E, F) GSEA of KEGG gene sets in the high- and low-risk groups. (G) Single sample gene set enrichment analysis (ssGSEA) results and the positive correlation between CAF risk scores and extracellular matrix (ECM)–receptor interaction-related genes in the TCGA cohort.
Moreover, three CAF-related genes (SNAI2, FRMD6, and LOXL2) were highly expressed in the high-risk group in both the TCGA and GSE72094 datasets, which is consistent with the results obtained using various CAF markers (Figure 3C and D). GSEA was performed on the TCGA dataset between the high- and low-risk groups, and the essential KEGG signaling pathways were involved in the cell cycle, cytokine–cytokine receptor interactions, and ECM–receptor interactions in the high-risk group (Figure 3E). Genes in the low-risk group were mainly enriched in drug metabolism, cytochrome P450, linoleic acid metabolism, and the metabolism of xenobiotics by cytochromes (Figure 3F). The ssGSEA results also showed that CAF risk scores were positively correlated with ECM–receptor interaction (Figure 3G).
Comparison of the TMB and the efficacy of immunotherapy and chemotherapy between the high- and low-risk groups
Immunotherapy has become a powerful clinical treatment strategy for patients with cancer. 30 Using the TIDE method, we investigated the potential clinical efficacy of immunotherapy in patients with LUAD. We found that the low-risk group displayed significantly lower TIDE scores than the high-risk group in both the TCGA and GSE72094 datasets (Figure 4A and D), which implied that patients in the low-risk group had lower potential for immune evasion and were more likely to benefit from immune checkpoint inhibitor (ICI) therapy than those in the high-risk group. In the TCGA dataset, the low-risk group had similar proportions of responders (56%) and non-responders (44%); however, the proportion of non-responders was significantly higher than that of responders (81% vs 19%) in the high-risk group (Figure 4B). Similar results were obtained for the GSE72094 cohort (Figure 4E). These findings suggested that the low-risk group exhibited higher sensitivity to immunotherapy. Moreover, the Area under the curve (AUC) values were 0.77 and 0.794 in the TCGA (Figure 4C) and GSE72094 cohorts, respectively (Figure 4F), indicating the excellent sensitivity and specificity of our CAF prognostic model for immunotherapy response prediction.

(A-F) Tumor Immune Dysfunction and Exclusion (TIDE) immunotherapy prediction analyses. (A, D) TIDE cancer-associated fibroblast (CAF) risk scores were higher in the high-risk groups than those in the low-risk groups in the GSE72094 and The Cancer Genome Atlas (TCGA) cohorts. (B, E) Proportions of responders and non-responders in the high- and low-risk groups in the GSE72094 and TCGA cohorts. (C, F) Receiver operating characteristic (ROC) curves of the risk signatures in the GSE72094 and TCGA cohorts. (G, H) Oncoplots showing the top 20 mutational genes in the high- and low-CAF-risk groups of the TCGA cohort. (I) Boxplot displaying the tumor mutation burden (TMB) values of the high- and low-risk groups. (J) CAF risk scores were significantly positively correlated with TMB values.
The top 20 genes with the highest mutational frequencies were identified in the low- and high-risk subgroups. Surprisingly, the genes (TP53, TTN, MUC16, CSMD3, RYR2, LRP1B, ZFHX4, USH2A, KRAS, XIRP2, FLG, SPTA1, NAV3, ZNF536, COL11A1, FAT3, ANK2, PCLO, CSMD1, and APOB; Figure 4G) and frequencies of mutations were highly similar between the two groups. In addition, we found that the high-risk group had higher TMB values than the low-risk group (Figure 4H), and the CAF risk score was significantly positively correlated with the TMB value (Figure 4I).
Chemotherapy is the standard approach for treating LUAD. Low-risk patients with LUAD exhibited increased sensitivity to cisplatin, crizotinib, docetaxel, gemcitabine, savolitinib, trametinib, and paclitaxel. In contrast, the high-risk group was estimated to be more sensitive to vinorelbine in the TCGA cohort (Figure 5).
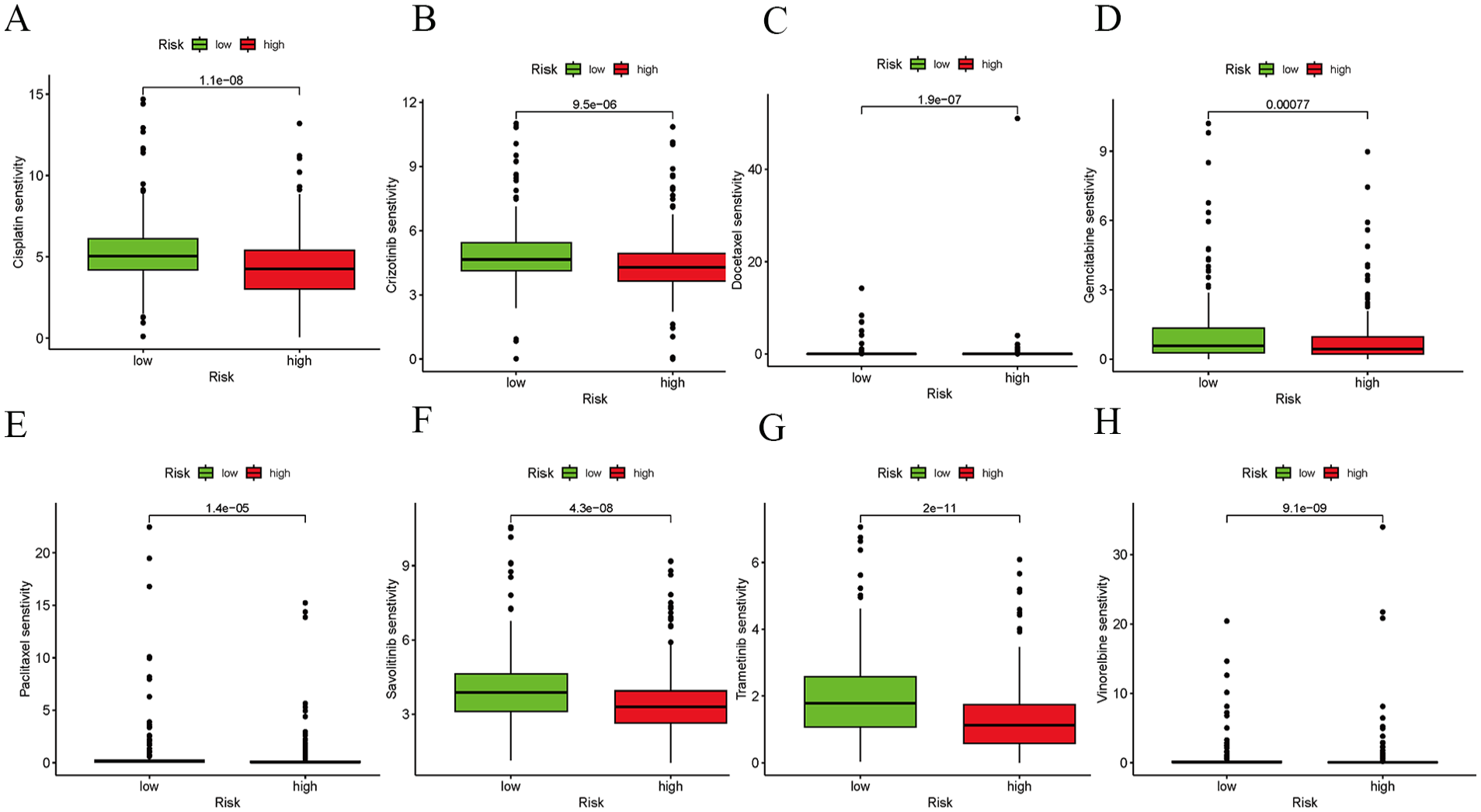
(A-H) Boxplot showing a comparison of the sensitivities of the high- and low-risk groups to several chemotherapy drugs.
Validation of CAF-related genes
Through analysis of the CCLE database, we found that the mRNA expression of SNAI2, FRMD6, and LOXL2 was high in fibroblast cell lines (Figure 6A). In addition, IHC data confirmed that SNAI2 expression was high in both cancerous and normal lung tissues (Figure 6B). However, we found that SNAI2 was not expressed in CAF cells (Supplemental Figure 1). The Western blot results showed that SNAI2 expression was higher in lung cancer cells (H1975, HCC827, A549, SPC-A-1, and H1573) than that in Beas-2b pulmonary epithelial cells (Figure 6B). Moreover, we found that the percentage of cells in the S phase significantly increased, while the proportion of those in the G2/M phase and G1 phase significantly decreased after knocking down SNAI2 in LUAD cells, indicating that the cell cycle of LUAD cells was halted in the S phase. Thus, these results suggest that knocking down SNAI2 may significantly inhibit the proliferation of LUAD cells via cell cycle arrest in the S phase (Supplemental Figure 2). These results suggest that SNAI2 is a CAF-specific marker.

Analysis of the expression of cancer-associated fibroblast (CAF)-related genes. (A) mRNA expression levels of the three CAF-related genes in CAFs and lung adenocarcinoma (LUAD) cell lines. (B) Protein expression of SNAI2 in LUAD tissues from the Human Protein Atlas database. (C) Western blotting demonstrated the protein expression of SNAI2 in LUAD cell lines.
Discussion
CAFs have long been considered to play a crucial role in the development and prognosis of LUAD. Our results revealed that CAF infiltration was associated with poor prognosis in patients with LUAD in two independent cohorts (GSE72094 and TCGA). Patients with high CAF scores had lower survival rates than those with low CAF scores, which highlights the significant role of CAF in the TME.
CAF-related genes in multiple cancer types have been an important focus of research to identify critical molecular pathways in CAF subtypes that may be associated with clinical outcomes, disease progression, and immunotherapy resistance.31-34 Recently, there has been a decline in the incidence of lung cancer owing to improved cancer screening and treatment in clinical practice. 35 This implies that targeted cancer control, interventions, early detection, and treatment can help reduce cancer-related mortality. 36 Some studies have demonstrated several roles of CAFs in LUAD; however, the molecular markers of CAF need to be elucidated to better classify tumor subtypes and facilitate the establishment of CAF-specific targeted therapies. Furthermore, the Heterogeneous nuclear ribonucleoprotein K (HNRNPK)/Chloride channel 3 (CLCN3) axis promotes the progression of LUAD through CAF–tumor interactions, 37 and CAF activation can facilitate the invasion of breast cancer cells by CAFs. 38
Cancer progression, recurrence, and metastasis are strongly associated with the TME, of which the ECM is a key component. 39 The ECM regulates cell differentiation, proliferation, migration, death, and survival and plays a crucial role in maintaining cellular homeostasis. 40 Disruption of the ECM composition and structure is linked to carcinogenesis and cancer progression. 41 By constructing a co-expression network of CAF-related genes and screening for hub genes, we identified gene modules that were highly correlated with CAF scores, as well as functionally enriched GO and KEGG pathways. Specifically, GO analysis revealed that CAF-related genes were mainly associated with the ECM structure and tissues, which is consistent with the findings of previous research, indicating that CAF-related genes play important roles in ECM remodeling, tumor development, and metastasis. 38 In addition, KEGG analysis revealed that the enriched pathways of CAF-related genes, such as ECM–receptor interactions, were related to cancer development. The mechanical remodeling of the ECM by CAF is crucial for tumor cell migration and invasion. 42 Changes in these enrichment pathways have been implicated in the development and metastasis of LUAD, which further supports the association between CAF and LUAD. A total of 28 genes that were significantly associated with the survival of patients with LUAD were identified using univariate Cox regression analysis, and a prognostic risk model based on CAF-related genes was established using the LASSO analysis. The training and testing groups in the TCGA and GSE72094 cohorts were divided into low- and high-risk subgroups based on the median. Kaplan–Meier curves showed that the OS of the low-risk group was better than that of the high-risk group, confirming that CAF-related genes may serve as predictive biomarkers for LUAD. Furthermore, our results showed that three CAF-related genes (SNAI2, FRMD6, and LOXL2) were highly expressed in the high-risk group, which is consistent with the panel of CAF markers collected from the TCGA and GSE72094 datasets. Additional experiments confirmed the high mRNA expression of SNAI2, FRMD6, and LOXL2 in fibroblast cell lines in the CCLE database. In addition, Western blotting validated the higher expression of SNAI2 in lung cancer cell lines compared with that in normal tissues. Moreover, we found that knocking down SNAI2 can significantly inhibit the proliferation of LUAD cells.
SNAI2 (SLUG) is a C2H2 zinc-finger transcriptional repressor belonging to a three-member family of SNAIL proteins (SNAIL, SNAI2, and SMUC). 43 SNAI2 is known for its role in orchestrating EMT, during which tumor cells lose their epithelial characteristics and acquire a fibroblast-like cell phenotype and enhanced migratory abilities. 18 Recent studies have revealed additional roles of SNAI2 in cancer progression, including in the activation of tumor-initiating cells, and cell cycle regulation, invasion, and metastasis.44-46 In addition, SNAI2 is a key transcription factor in the regulation of squamous cell differentiation, which is a hallmark histopathological feature of lung squamous cell carcinoma. 31 Our findings suggest that SNAI2 may serve as a specific marker for CAFs.
CAFs can regulate the formation of an immunosuppressive network to weaken antitumor immunity.47,48 Our study also explored the differences between the low- and high-CAF groups in terms of TMB and response to immunotherapy and chemotherapy. The results showed that in both the TCGA and GSE72094 cohorts, the low-risk group had significantly lower TIDE scores than the high-risk group, suggesting that patients in the low-risk group had a lower potential for immune escape and were more likely to benefit from ICI therapy than those in the high-risk group. Furthermore, our findings revealed a significant positive correlation between CAF risk scores and TMB values. Regarding chemotherapy, the low-risk group exhibited higher sensitivity to drugs such as cisplatin, crizotinib, docetaxel, gemcitabine, sunitinib, and paclitaxel, whereas the high-risk group exhibited higher sensitivity to etoposide. Thus, further exploration of correlations between CAF-related genes and OS in patients with LUAD is needed, including more basic experiments and clinical samples, which we will address in our future studies.
Furthermore, there may have been sample bias in the public database used. In addition, confounding factors, such as patient clinical characteristics, may have affected the results. Future studies should focus on investigating the function of SNAI2 in LUAD and the underlying molecular mechanisms, as well as designing clinical trials to study the potential of CAFs as a therapeutic target.
Conclusions
Overall, our findings enhance our understanding of the role of CAFs in LUAD progression and provide a foundation for future research and clinical applications. We will further study the potential of the CAF marker SNAI2 in clinical trials in the future. SNAI2 has potential as a CAF therapeutic target and could be combined with existing treatment options, such as chemotherapy and immunotherapy, to reduce the development of drug resistance. In addition, the CAF marker SNAI2 has possible applications in prognosis assessment and treatment response monitoring of patients with LUAD to help clinicians better manage the disease and optimize treatment plans.
Supplemental Material
sj-tif-1-onc-10.1177_11795549241280506 – Supplemental material for SNAI2 as a Prognostic Biomarker Based on Cancer-Associated Fibroblasts in Patients With Lung Adenocarcinoma
Supplemental material, sj-tif-1-onc-10.1177_11795549241280506 for SNAI2 as a Prognostic Biomarker Based on Cancer-Associated Fibroblasts in Patients With Lung Adenocarcinoma by Tian-Tian Li, Qing-Gang Hao, Zhao-Wei Teng, Yuan Liu, Jia-Fan Wu, Jun Zhang and Li-Rong Yang in Clinical Medicine Insights: Oncology
Supplemental Material
sj-tif-2-onc-10.1177_11795549241280506 – Supplemental material for SNAI2 as a Prognostic Biomarker Based on Cancer-Associated Fibroblasts in Patients With Lung Adenocarcinoma
Supplemental material, sj-tif-2-onc-10.1177_11795549241280506 for SNAI2 as a Prognostic Biomarker Based on Cancer-Associated Fibroblasts in Patients With Lung Adenocarcinoma by Tian-Tian Li, Qing-Gang Hao, Zhao-Wei Teng, Yuan Liu, Jia-Fan Wu, Jun Zhang and Li-Rong Yang in Clinical Medicine Insights: Oncology
Supplemental Material
sj-tif-3-onc-10.1177_11795549241280506 – Supplemental material for SNAI2 as a Prognostic Biomarker Based on Cancer-Associated Fibroblasts in Patients With Lung Adenocarcinoma
Supplemental material, sj-tif-3-onc-10.1177_11795549241280506 for SNAI2 as a Prognostic Biomarker Based on Cancer-Associated Fibroblasts in Patients With Lung Adenocarcinoma by Tian-Tian Li, Qing-Gang Hao, Zhao-Wei Teng, Yuan Liu, Jia-Fan Wu, Jun Zhang and Li-Rong Yang in Clinical Medicine Insights: Oncology
Supplemental Material
sj-tif-4-onc-10.1177_11795549241280506 – Supplemental material for SNAI2 as a Prognostic Biomarker Based on Cancer-Associated Fibroblasts in Patients With Lung Adenocarcinoma
Supplemental material, sj-tif-4-onc-10.1177_11795549241280506 for SNAI2 as a Prognostic Biomarker Based on Cancer-Associated Fibroblasts in Patients With Lung Adenocarcinoma by Tian-Tian Li, Qing-Gang Hao, Zhao-Wei Teng, Yuan Liu, Jia-Fan Wu, Jun Zhang and Li-Rong Yang in Clinical Medicine Insights: Oncology
Footnotes
Author Contributions
L-RY and JZ: Conceptualization, Writing—review & editing. Z-WT: Data curation, Funding acquisition. Q-GH, J-FW, and YL: Formal analysis, Validation, Software. T-TL: Methodology, Resources, Project administration. T-TL and Q-GH: Writing—original draft.
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by Kunming University of Science and Technology and the First People’s Hospital of Yunnan Province Joint Special Project on Medical Research (grant number: KUST-KH2022002Z), the Joint Special Fund of Applied Fundamental Research of Kunming Medical University granted by the Science and Technology Office of Yunnan (grant number: 202301AY070001-299), the Yunnan Province Orthopedics and Sports Rehabilitation Clinical Medical Research Center Open Project (grant number: 2022YJZX-GK04), and the National Natural Science Foundation of China (grant number: 82060167). The funding agencies had no role in the design, conduct, write-up, or publication of the study.
Data Availability Statement
The datasets generated during the current study are available in the [TCGA] repository, [https://portal.gdc.cancer.gov/] and in the [GEO] repository,[![]() ].
].
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.

