Abstract
Background:
Endometrial cancer (EC) is a common gynecological malignancy, and the prognosis of advanced EC is unsatisfactory. The deregulated expression of RNA-binding proteins (RBPs) is closely associated with the occurrence and development of cancer. However, the role of RBPs in EC remains unclear. The aim of this study was to validate the prognostic values of RBPs combined with clinical factors.
Methods:
We downloaded the RNA sequencing and clinical data for EC from The Cancer Genome Atlas (TCGA) database. R software was used to identify the differentially expressed RBPs. Univariate and multivariate Cox proportional hazards regression analyses were performed to predict the 4 overall survival (OS)-related RBPs. We then constructed a nomogram combining the 4-RBP signature with clinical risk factors to assess the prognostic power. Furthermore, we validated the expression of 4 RBPs in our patient samples using quantitative real-time polymerase chain reaction (qRT-PCR) and explored the effect of cold-inducible RNA-binding protein (CIRBP) on EC tumor growth using cell proliferation experiments.
Results:
It is found that Shwachman-Bodian-Diamond syndrome (SBDS), CIRBP, MRPL15, and CELF4 were significantly related to the prognosis of EC patients. In addition, the nomogram showed better performance in OS predictions than the International Federation of Gynecology and Obstetrics (FIGO) stage. The qRT-PCR results showed that low CIRBP expression was associated with cell proliferation.
Conclusions:
In our study, we constructed a 4-RBP signature-based nomogram combined with clinical factors in EC that could effectively predict the prognosis of EC patients. The results provide novel insights into the development of treatment targets and prognostic molecular markers in EC.
Introduction
Endometrial cancer (EC) is a common malignancy in females. 1 The incidence and mortality have been rapidly increasing over time in several countries.2-4 Despite the rapid progress in surgery, radiotherapy, and chemotherapy, the results vary greatly in patients at different EC stages. 5 At present, risk stratification system that includes age, myometrial invasion, histological tumor grade, and the International Federation of Gynecology and Obstetrics (FIGO) stage is used to evaluate the prognosis of patients with EC. However, the evaluation of 5 common risk stratification systems shows that they are not very accurate for EC. 6 To improve the treatment effect, it is of great significance to understand the molecular mechanism of EC and further formulate effective early screening and diagnosis methods.
RNA-binding proteins (RBPs) contain RNA-binding domains (RBDs), which can interact with RNA and regulate post-transcriptional processes, such as transcriptional control, intracellular localization, RNA transport, sequence editing, and RNA editing. 7 To date, 1542 RBP genes have been identified in the human genome through genome-wide screening. 8 These RBPs perform various functions to maintain cell physiological homeostasis, especially in development and stress responses. 9 Most RBPs can compete or cooperate with the same mRNA to control mRNA turnover. 10 Moreover, some RBPs can control mRNA translation and pre-mRNA splicing.11,12 In addition to RNAs that participate in the translation machinery and its regulation (rRNAs, tRNAs, small interfering RNAs, and miRNAs), a growing knowledge of RBPs targets is shifting the focus toward noncoding RNAs, such as the long noncoding RNAs (lncRNAs) and circular RNAs (circRNAs).13,14
Many studies have found dysregulated expression of RBPs in a variety of human diseases, especially cancer. 15 In addition, recent genome-wide analysis has confirmed that many RBPs play important roles in the occurrence and development of cancers, such as colorectal, gastric, and lung cancers.16-18 A few RBPs have also been found to play critical roles in gynecological cancers. 19 For example, Musashi-1 inhibits the malignant characteristics of ovarian cancer and reverses paclitaxel resistance 20 ; ESRP1 is upregulated in ovarian cancer and promotes the transformation of ovarian cancer cells from an interstitial phenotype to an epithelial phenotype 21 ; and the stemness of EC cells is inhibited by the RBP RNPC1 by stabilizing MST1/2 mRNA. 22 However, the role of the most RBPs in EC remains unclear. A systematic functional analysis of RBPs will help to further study their role in EC.
In this study, we aimed to construct a nomogram based on the RBP signature and clinical parameters to predict overall survival (OS) in EC patients, which can help clinical decision-making and individualized treatment.
Materials and Methods
Date resources and processing
The RNA sequencing data of 35 normal endometrial tissue samples and 542 EC samples with corresponding clinical data were downloaded from The Cancer Genome Atlas (TCGA). We used the DESeq2 package to preprocess the raw data and exclude genes with an average count value less than 1. The DESeq2 package was also used to choose the differentially expressed RBPs according to|log2Fold Change| (log2FC) ⩾ 1 and an adjusted P < .05. The R software packages ggplot2 and pheatmap were used to visualize the volcano map and heatmap. The research design is illustrated with a flow chart (see Supplemental Figure S1 for details).
PPI network and functional enrichment analysis
In this study, we used the STRING database to identify protein-protein interaction (PPI) information of the differentially expressed RBPs. 23 Then, we imported the PPI pairs identified by the STRING database into Cytoscape 3.7.0 software and established a PPI network. Moreover, we used the Database for Annotation, Visualization, and Integrated Discovery (DAVID) online tool and cluster profiler to annotate the filtered RBPs. Then, we used the cluster profiler package to analyze Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways, Gene Ontology (GO), and Disease Ontology (DO). 24 A P value of less than .05 was considered statistically significant.
Selection and verification of prognosis-related RBPs
Univariate Cox regression analysis was performed on the differentially expressed RBPs using the survival software package. Then, we further used the R software package glmnet to construct a multigene signature using least absolute shrinkage and selection operator (LASSO) regression to predict the prognosis of EC. 25 Next, we used the UALCAN database to compare the differential expression of common cancer types, stages, and their normal adjacent tissues. 26 The expression profiles of 4 central RBPs were detected at the translation level using the Human Protein Atlas (HPA) online database. 27 We constructed receiver operating characteristic (ROC) curves to evaluate the ability to distinguish between normal and cancerous tissues in GraphPad Prism 7.0 software. The prognostic value of the 4 hub RBPs in EC was verified using the Kaplan-Meier (KM) plotter and OncoLnc online tools.
Establishment and evaluation of the 4-RBP prognostic signature
The regression coefficients of the 4 hub RBPs were obtained from a multivariate Cox proportional hazard regression model. Then, we used the linear combination method to combine the expression levels and coefficients of each hub RBP to obtain the risk score. We divided the patients into the high- and low-risk groups (median as cut-off point) on the basis of the risk score. Time-dependent ROC curves were drawn, and the area under the ROC curve (AUC) values at 1, 3, and 5 years were calculated. To further evaluate the prognostic performance of the 4-RBP signature, we used the KM method and the log-rank test to evaluate the survival difference between the high- and low-risk groups. Then, we performed univariate and multivariate Cox proportional hazards regression analyses to evaluate whether the risk score and some clinical parameters could be used as independent prognostic factors. Stratified analysis was performed to assess the diagnostic ability of the 4-RBP signature for other clinical prognostic parameters at different levels.
Construction and validation of the RBP signature-based prognostic nomogram
Based on independent prognostic parameters screened by univariate and multivariate Cox proportional hazards regression analysis, a composite nomogram was constructed using the R software package rms to predict the 1-, 3-, and 5-year OS probabilities. The ability of the nomogram to compare the model-predicted OS probability with the actual OS probability was evaluated by a concordance index (C-index) and calibration curve. We used the R software package survivalROC to draw the tROC curve to evaluate the predictive accuracy of independent prognostic parameters. We also compared the discrimination ability of prognostic parameters by calculating the AUC.
Preparation for human EC samples
The study was approved by the medical research ethics committee of the International Peace Maternal and Child Health Hospital (the ethical approval ID is [2015] No. 2, and the date of approval is April 19, 2016). From October 2019 to October 2020, we collected 36 EC tissue samples and 36 normal uterine tissue samples in the Department of Gynecologic Oncology, Shanghai International Peace Maternity and Child Health Hospital. All patients were diagnosed according to the histopathology reports from biopsy after surgery, and none of them received chemotherapy, radiotherapy, or hormone therapy before surgery. The tumor stages and histological grades were determined according to the criteria of FIGO 2018 staging system. Normal uterine tissue samples were collected from participants who had undergone hysterectomy for hysteromyoma. We stored all tissue samples at −80°C until use.
Quantitative real-time polymerase chain reaction analysis
Total RNA was isolated from samples using TRIzol reagent (Takara, Dalian, China), and the RNA concentration was determined by a NanoDrop ND-2000 (NanoDrop, USA). Then, we used PrimeScript RT Master Mix (Takara, Dalian, China) to reverse-transcribe 500 ng of RNA and amplified cDNA with Hieff® qPCR SYBR Green Master Mix (Yeasen, Shanghai, China). In accordance with the manufacturer’s instructions, we conducted quantitative real-time PCR (qRT-PCR) using the QuantStudio 7 Flex system (Life Technologies, USA). For the detection of mRNA expression levels, we used actin as the control. The primer sequences used for qRT-PCR are listed in Supplemental Table S1. The ΔΔCt method was used for quantification.
Cell culture and transfection
Human endometrial epithelial cells (EECs) and EC cell lines (Ishikawa, KLE, AN3CA, HEC-1A, and HEC-1B) were purchased from the American Type Culture Collection (ATCC, VA, USA). The EECs were cultured in epithelial cell medium (MingZhoubio CO., Ltd.). The EC cells were cultured in Dulbecco’s modified Eagle medium (DMEM)/F12 (Gibco, Auckland, New Zealand) containing 10% fetal bovine serum (Gibco, Carlsbad, CA, USA), 100 μg/mL penicillin, and 100 U/mL streptomycin (Gibco) at 37°C in a 5% CO2 humidified atmosphere. According to the manufacturer’s protocol, we used Lipofectamine 3000 (Invitrogen, NY, USA) to transfect cells. However, 2 cold-inducible RNA-binding protein (CIRBP) overexpression plasmids were constructed by cloning the coding region sequence (CDS) of human CIRBP into the pLX304 vector purchased from Shanghai Jiao Tong university, School of Medicine (http://dnacore.shsmu.edu.cn).
Cell proliferation assay
Cell proliferation was measured using a Cell Counting Kit-8 (CCK8) and a 5-ethynyl-20-deoxyuridine (EdU) assay kit following the manufacturer’s directions (Yeasen, Shanghai, China). Then, we used a SpectraMax 190 microplate reader (Bio-Rad Model 680) to calculate the absorbance values at 450 nm. The EdU cell lines were photographed and counted in 5 fields/well under a microscope at x200 magnification.
Statistical analysis
All experiments were performed in triplicate. Data were analyzed with SPSS software (version 19.0) (SPSS, Inc., Chicago, IL, USA) and are presented as the mean ± SD. Then, we used an unpaired Student’s t-test or one-way analysis of variance (ANOVA) to calculate the statistical significance of the results. A P value of less than .05 was considered statistically significant.
Results
Identification of differentially expressed RBPs in EC patients
In total, 1542 RBPs were included in our study and 176 RBPs were identified as differentially expressed genes (|log2FC| ⩾ 1, adjusted P < .05), which contained 112 upregulated and 64 downregulated RBPs. The results are shown in Figure 1A and B.

Heatmap and volcano plot of 176 RBPs in EC: (A) heatmap and (B) volcano plot.
PPI network and functional analysis
To investigate the potential relationship among differentially expressed RBPs in EC, we constructed a PPI network. This PPI network consisted of 159 nodes, including 103 upregulated RBPs and 56 downregulated RBPs, and 761 edges (Figure 2A). Then, we used GO, KEGG, and DO enrichment analyses to elucidate the functions of these key RBPs. The top enriched results are shown in Figure 2B to D. Functional annotation revealed that “RNA splicing,” “RNA splicing, via transesterification reactions with bulged adenosine as nucleophile,” and “mRNA splicing, via spliceosome” were the most significantly enriched GO terms (Figure 2B). In the KEGG analysis, spliceosomes and ribosomes were the most enriched (Figure 2C). Meanwhile, DO analysis results showed that the key RBPs were associated with urinary system cancer, germ cell cancer, and embryonal cancer (Figure 2D).
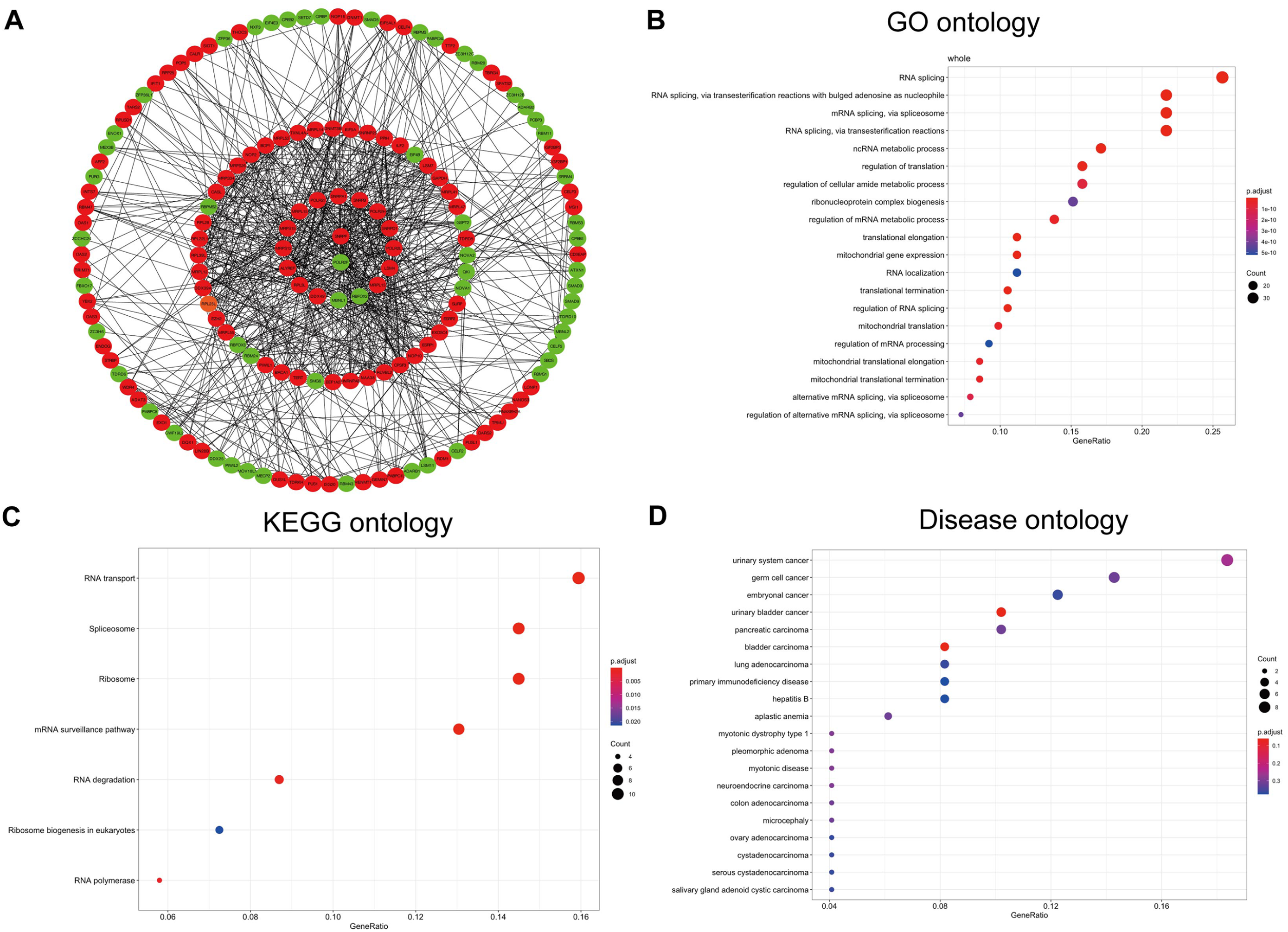
PPI network and functional analysis of key RBPs: (A) PPI network of proteins encoded by the RBPs, including 159 nodes and 761 edges. Red nodes represent upregulated RBPs, while green nodes represent downregulated RBPs. (B) GO analysis identified top 20 most significant GO terms. (C) KEGG enrichment analysis of key RBPs. (D) DO analysis identified top 20 most significant DO terms.
Selection of prognosis-related RBPs
To investigate the prognostic significance of 159 RBPs that were identified from the PPI network, we used univariate Cox regression analysis and the multiple stepwise Cox regression. However, 4 RBPs, namely, Shwachman-Bodian-Diamond syndrome (SBDS), MRPL15, CIRBP, and CELF4, were the most valuable prognostic RBPs. Table 1 shows the univariate and multivariate Cox analysis results for these RBPs. MRPL15 and CELF4 were upregulated, while SBDS and CIRBP were downregulated. Moreover, MRPL15, CELF4, and SBDS with a hazard ratio of >1 were considered as risky prognostic RBPs, while CIRBP with a hazard ratio of <1 was regarded as a protective prognostic RBP (see Supplemental Figure S2 for details).
Univariate and multivariate Cox analysis of the association of the 4 hub RBPs with EC patient’s OS in the TCGA cohort (n = 539).

Abbreviations: CI, confidence interval; HR, hazard ratio; RBP, RNA-binding proteins; TCGA, The Cancer Genome Atlas.
Validation of the expression of hub RBPs
To further determine the expression of these hub RBPs in EC, we used expression profiles from the UALCAN database to show that SBDS and CIRBP were expressed at lower levels in tumor tissues than in normal tissues, while MRPL15 and CELF4 were expressed at higher levels (Figure 3A) (P < .001). In addition, the protein expression of hub RBPs was verified using the HPA database. Apart from CIRBP missing in the HPA database, immunohistochemistry staining showed that compared with normal samples, the expression of SBDS was lower and the expression of MRPL15 was higher in the tumor samples. However, no difference was found for CELF4 protein expression (Figure 3B). Finally, ROC curve analysis was performed to assess the ability of SBDS, CIRBP, MRPL15, and CELF4 to distinguish EC from normal tissues (Figure 3C).

Validation of 4 significantly prognostic RBPs in EC: (A) Expression pattern of the 4 RBPs between normal tissues and tumor tissues. (B) Immunohistochemistry of SBDS, MRPL15, and CELF4 in normal tissues and tumor tissues based on the HPA. (C) The ability of SBDS, CIRBP, MRPL15, and CELF4 to distinguish EC samples from normal samples was evaluated by ROC curve analysis and AUC statistics.
In addition, we explored the association between the expression levels of the 4 RBPs and histopathological information, such as histological subtype and individual cancer stage. SBDS, CIRBP, and MRPL15 were significantly associated with histological subtype (see Supplemental Figure S3A for details; P < .05). SBDS, CIRBP, MRPL15, and CELF4 were significantly associated with individual cancer stage (see Supplemental Figure S3B for details; P < .05).
Survival analysis of hub RBPs
To further elucidate the prognostic value of the hub RBPs in EC, the KM plotter was used to determine the relationship between hub RBPs and OS, and the patients were split by selecting the best cut-off. Low expression of CIRBP and high expression of SBDS, MRPL15, and CELF4 were related to the poor prognosis of EC patients (Figure 4A; P < .05). The OS of each key RBP was analyzed through the online KM survival analysis tool (OncoLnc). Taking the median expression of 4-RBP as the dividing point, the samples were divided into a high expression group and a low expression group. The results showed that SBDS, MRPL15, and CELF4 were negatively correlated with OS in EC patients, while CIRBP was positively correlated with OS in EC patients (Figure 4B). These results suggested that SBDS, CIRBP, MRPL15, and CELF4 could be prognostic markers for EC.

Prognostic gene characteristics of 4 hub RBPs in EC patients: (A) Relationship between the expression level of SBDS, CIRBP, MRPL15, and CELF4 and OS was performed using KM plotter. (B) Association between SBDS, CIRBP, MRPL15, and CELF4 expression and OS was performed with the online KM survival analysis tool (OncoLnc).
Construction and analysis of the prognosis-related genetic risk score model
According to the Cox coefficients of the 4 genes, we constructed a 4-gene-based risk score. Furthermore, the risk score of each EC patient was calculated, and the patients were divided into a low-risk group and a high-risk group using the median risk score as the cut-off value (Figure 5A). Figure 5B shows the survival status of all patients. The heatmap revealed that the expression levels of SBDS, MRPL15, and CELF4 in the high-risk group were higher than those in the low-risk group, while CIRBP expression was higher in the low-risk group (Figure 5C). Based on the risk score model, the time-dependent ROC analysis showed that it performed well in survival prediction, and the AUC was 0.68, 0.70, and 0.72 at 1, 3, and 5 years, respectively (Figure 5D). Moreover, the KM survival curve analysis demonstrated that the OS of the high-risk group was worse than that of the low-risk group (Figure 5E).

Risk score analysis of 4-RBP prognostic model in TCGA EC cohort: (A) Risk score distribution of EC patients based on the risk score model. (B) Survival status distribution of EC patients in low- and high-risk groups. (C) Heatmap of the 4 prognostic RBPs expression profiles in low- and high-risk groups. (D) Time-dependent ROC analysis of 4-RBP signature. (E) KM survival analysis of 4-RBP signature.
Next, the relationship between the conventional clinical risk factors and risk score was analyzed. We found that FIGO stage, tumor grade, and the 4-RBP signature could independently predict the prognoses of EC patients using univariate and multivariate Cox proportional hazards regression (Table 2).
Univariate and multivariate Cox analysis of the association of the 4-RBP signature and clinical risk factors with EC patient’s OS in the TCGA cohort (n = 539).

Abbreviations: CI, confidence interval; FIGO, Federation International of Gynecology and Obstetrics; HR, hazard ratio; RBP, RNA-binding proteins; TCGA, The Cancer Genome Atlas.
Patients were risk-stratified according to FIGO stage, tumor grade, and age. We found that the classification efficiency was limited when the 4-RBP signature was applied to some subgroups. The KM curves showed that the survival rate of patients in the high-risk group was significantly lower than that in the low-risk group for the Stage I and III subgroups. However, for the Stage II and IV subgroups, the difference in the 4-RBP signature was not statistically significant (Figure 6A). When stratified by tumor grade, only the 4-RBP signature in the grade 3/4 subgroup divided patients into high- and low-risk groups with significantly different survival rates (Figure 6B). We also observed that the OS of patients with high-risk scores was worse than that of patients with low-risk scores in both ⩽60 and >60 years of age groups (Figure 6C). Therefore, although the 4-RBP signature could be viewed as an independent prognostic predictor for EC patients, its performance was limited to specific subgroups.

Risk-stratified analysis of the 4-RBP signature in patients with EC. KM survival analysis of patients in different subgroups: (A) Stages I/II/III/IV, (B) Grade 1/2, grade 3/4, and (C) Age ⩽ 60 and >60 years.
Construction and validation of a nomogram based on the 4-RBP signature and clinical risk factors
Clinical risk factors, such as the FIGO stage and tumor grade, are vital predictors of OS in EC patients. Thus, we constructed a compound nomogram based on the 4-RBP signature, FIGO stage, and tumor grade to predict different-year OS of EC patients (Figure 7A). By drawing a vertical line between the total score axis and each prognosis axis, we calculated the possible 1-, 3-, and 5-year survival rates of EC patients. Then, we used the C-index and calibration plots to evaluate the discrimination and calibration ability of the prognostic nomogram. The C-index was 0.751 for the entire dataset. The predicted results of the 4-RBP prognostic nomograms were consistent with the actual results according to the calibration chart of patient survival prediction (Figure 7B to D).

A 4-RBP signature-based nomogram and calibration plots. (A) Nomogram integrated 4-RBP-based risk score, FIGO stage, and tumor grade. (B to D) Calibration plots of the nomogram were used to predict OS at 1, 3, and 5 years.
Time-dependent ROC curves were used to further compare the prognostic performance of the nomogram with different prognostic factors. The AUC value of the nomogram was greater than that of FIGO stage and the 4-RBP signature, which indicated that the nomogram may perform best in predicting OS (see Supplemental Figure S4A to C for details). Thus, the newly developed prognostic nomogram concentrated the advantages of the 4-RBP signature and 2 clinical risk factors, which improved their prognostic predictive efficiency for EC patients.
Validation of the expression profiles of 4 RBPs expression in our patient samples
The expression profiles of 4 RBPs were verified in clinical tissues by qRT-PCR. The clinical characteristics are described in Supplemental Table S2. As Figure 8B demonstrates, the expression of CIRBP in tumor tissues was obviously lower than that in normal tissues (Figure 8B). The relative expression levels of MRPL15 and CELF4 in tumor tissues were markedly elevated compared with those in normal tissues (Figure 8C and D). However, there was no significant difference in the expression of SBDS between tumor tissues and normal tissues (Figure 8A). More samples may need to be collected to validate the expression of RBPs.

Relative expression level of 4 RBPs in clinical samples: (A) SBDS, (B) CIRBP, (C) MRPL15, (D) CELF4, (E) qRT-PCR showing the expression of CIRBP in normal EECs and 5 EC cell lines, and (F) analysis of CIRBP mRNA in HEC-1A cell stably overexpressing CIRBP. (G to H) CCK-8 assay and EdU assay were performed to determine the ability of proliferation in HEC-1A cells transfected with oe-CIRBP or vector (magnification, ×200; scale bars = 50 μm).
It has been reported that CIRBP is decreased in the EC by Western blot and immunohistochemical analysis. Moreover, this decreased expression may lead to the uncontrolled proliferation of EC cells, but this possibility has not been verified. 28 Therefore, we chose CIRBP for further study. We found that HEC-1A cells showed the lowest expression of CIRBP and that Ishikawa (ISK) cells showed the highest expression of CIRBP among EC cell lines (Figure 8E).= Thus, we selected the HEC-1A cell line to upregulate the expression of CIRBP by transfecting plasmids of CIRBP1 and CIRBP2 plasmids, which have high transfection efficiency for subsequent experiments (Figure 8F). CCK-8 assays demonstrated that the upregulation of CIRBP significantly inhibited proliferation and viability (Figure 8G). Similarly, EdU assays revealed that the upregulation of CIRBP greatly decreased the percentages of EdU-positive cells (Figure 8H). These experiments suggested that CIRBP suppressed the proliferation of EC cells.
Discussion
Despite the progress in diagnosis and treatment in recent decades, the incidence and mortality rates of EC are still rising. 29 It is necessary to explore the pathogenesis of EC and identify effective prognostic biomarkers to improve the survival rate of patients. At present, we often use the FIGO staging system to predict the prognosis of EC patients. 30 However, a single clinical parameter has poor predictive ability for prognosis. The data based on molecular biology bring new vision to tumor prognosis analysis. Therefore, in the era of precision medicine, a more reliable EC prognosis model combined with other biomarkers is urgently needed.
Many recent studies have shown that RBPs are very important in the occurrence and development of numerous tumors. 31 However, we know little about the expression patterns and roles of RBPs in EC. Thus, we identified the differentially expressed RBPs between cancer tissues and normal tissues based on published TCGA-EC RNA sequencing and clinical data and constructed a PPI network to systematically explore potential functional pathways. Then, univariate and multivariate Cox regression analyses were used to verify potential biomarkers related to the prognosis of EC, and a 4-RBP signature was developed to predict the prognosis of EC patients, including SBDS, CIRBP, MRPL15, and CELF4. The 4-RBP signature has a great predictive value for the OS of EC patients and their subgroups.
These results demonstrated that differentially expressed RBPs were greatly enriched in RNA splicing, ncRNA metabolic processes, regulation of translation, and regulation of cellular amide metabolic processes using functional pathway enrichment. DO analysis showed that EC was the most similar to urinary system cancer. KEGG pathway analysis showed that abnormal expression of RBPs regulated the occurrence and development of EC by affecting the RNA transport pathway, spliceosome, ribosome, mRNA surveillance pathway, and RNA degradation. Moreover, previous studies have shown that RBPs can interact with genetic mutations that are present in EC. For example, the RBP Piwil1 caused epigenetic alteration of the PTEN gene via upregulation of DNA methyltransferase in type I EC. 32
In this study, we found that SBDS, MRPL15, and CELF4 were positive prognostic RBPs, while CIRBP was a negative prognostic RBP. SBDS was named because its mutation is highly related to Shwachman-Bodian-Diamond syndrome, and SBDS is involved in rRNA processing and 80S ribosome assembly. 33 Several studies have revealed SBDS to have both oncogenic and tumor-suppressive functions in cancer through its dual regulator of the MDM2-p53 circuit. 34 Similar to our findings, SBDS was positively related to the overall survival in EC. 35 Moreover, SBDS is highly associated with tumor stage, grade, diabetes, and hypertension in patients with EC. 36 The outcome suggested that the higher expression of SBDS could result in EC progression and indicated that EC had a higher degree of malignancy. As a human mitochondrial ribosomal protein (MRP), MRPL15 provides energy for cell growth in the form of ATP. Moreover, high transcription levels can predict breast cancer recurrence and increased tamoxifen resistance. 37 MRPL15 can also be used as a target gene of miR-26b-5p and has carcinogenic properties in Burkitt lymphoma. 38 Our analysis suggested that the high expression of MRPL15 was related to poor prognosis. CELF4 (also known as BRUNOL4) encodes proteins with 3 domains that are used to bind RNA recognition motifs. CELF4 may participate in RNA selective splicing during specific cell development. 39 Some studies have identified the methylation genes in EC samples using the methylomics method and verified that the methylation level of CELF4 in cervical scraps is the highest, which can be used to detect EC. 40 In our study, the expression of CELF4 was upregulated and correlated with poor prognosis. These results showed that CELF4 played a complex role in cancer, though more research is needed to analyze the mechanism of CELF4 in EC. A member of the cold shock protein family, the CIRBP is also known as A18 hnRNP or CIRP. It performs its function by binding the RNA-binding activity of its target genes in cells or triggers inflammatory reactions extracellularly as a secreted damage-related molecular pattern. 41 It has been reported that CIRP may be involved in the regulation of the cell cycle in normal endometrium, and the loss of its expression may be related to endometrial carcinogenesis. 28 This may explain our results that CIRBP was expressed at lower levels in EC tissues than in normal samples, which indicated the importance of low CIRBP expression for the development of EC.
To further improve the prognostic ability of the 4-RBP signature, we combined clinical prognostic factors with the risk score to construct a highly accurate predictive nomogram to quantify the possibility of individual OS. Then, its tROC survival analysis over time in the 1-, 3- and 5-year groups showed that it has the best performance in OS prediction with the risk score and FIGO stage. We also used qRT-PCR to validate the expression of 4 RBPs in clinical samples, and the results agreed with bioinformatics analysis. Moreover, we found that overexpression of CIRBP led to the suppression of cell proliferation by CCK-8 and EdU experiments, suggesting that CIRBP may play a critical role in the progression of EC and may be a treatment and prognostic target.
For patients with EC, the 2021 ESGO/ESTRO/ESP guidelines advocate molecular classification and propose a new prognostic risk stratification based on both pathologic and TCGA molecular features. Many studies have further investigated the prognostic value of TCGA molecular subgroups based on clinical features (such as myometrial invasion, histotype, or lymph vascular space invasion). Non-endometrioid carcinomas had a worse prognosis in all TCGA subgroups. 42 In both TCGA groups, deep myometrial invasion (DMI) appeared to affect recurrence independently, but further studies are needed to assess the prognostic impact of DMI separately. 43 The lymphovascular space invasion (LVSI) showed independent prognostic value, increasing the risk of death from any cause, EC-related deaths, and recurrent or progressive disease by 1.5- to 2-fold compared with TCGA signature. 44 In our study, we constructed a nomogram based on RBPs and clinical factors in EC to predict the prognosis of EC. Next, we may further explore the impact of the nomogram on the prognostic value of each TCGA molecular subgroup, which may be helpful for better risk stratification of patients in the future.
In general, our study provided novel insights into the role of RBPs in the occurrence and development of EC. We found that SBDS, CIRBP, MRPL15, and CELF4 are cancer-related RBPs and may have the potential to become biomarkers in EC therapy. In addition, rather than a single routine clinical parameter, our prognostic nomograms based on 4 RBPs could help clinicians predict the survival results of EC patients and provide reference for treatment guidance, which is more conducive to clinical application. However, this study has some limitations. First, our prognostic model was only based on the TCGA database and a limited clinical patient cohort and had not been validated in other databases. Second, we should conduct a prospective study to further verify our results because our study was only based on retrospective analysis. Last but not least, our findings were mostly based on public databases using bioinformatics analysis, so more functional experiments in vivo and in vitro are needed to verify these results. We only upregulated the expression of CIRBP in the HEC-1A cell line; we also need to knock out CIRBP in the ISK cell line to verify the influence of CIRBP on cell proliferation or any other properties of EC cells. In the future, we will design more mechanistic experiments, such as Western blotting and immunohistochemistry, to clarify the molecular mechanism based on the results of this study.
Conclusions
In conclusion, we systematically evaluated a large amount of bioinformatics data to investigate the expression mode, potential functions, and prognostic value of differentially expressed RBPs in TCGA-EC. The 4 RBPs may be potential biomarkers and new molecular therapeutic targets in EC. We have developed a nomograph based on the 4-RBP signature and clinical risk factors (TNM stage and tumor grade), which can accurately predict the OS of EC patients and aid in clinical decision-making and individualized treatments. Among the 4 RBPs, 1 hub gene, CIRBP, was downregulated in EC tissues and cells, and low CIRBP expression was related to cell proliferation. We will implement more research to elucidate its underlying mechanism in the pathogenesis of EC.
Supplemental Material
sj-docx-1-onc-10.1177_11795549221123620 – Supplemental material for Prognostic Values From Integrated Analysis of the Nomogram Based on RNA-Binding Proteins and Clinical Factors in Endometrial Cancer
Supplemental material, sj-docx-1-onc-10.1177_11795549221123620 for Prognostic Values From Integrated Analysis of the Nomogram Based on RNA-Binding Proteins and Clinical Factors in Endometrial Cancer by Shuang Yuan, Xiao Sun and Lihua Wang in Clinical Medicine Insights: Oncology
Footnotes
Acknowledgements
The authors thank all involvers for their support to our research.
Declaration of Conflicting Interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Natural Science Foundation of China (grant no. 81572547), Shanghai Municipal Key Clinical Specialty (grant no. shslczdzk06302), and the Clinical Research Fund of the International Peace Maternity and Child Health Hospital (grant no. GFY 1553). The results shown in the article are part based on data generated by the TCGA Research Network: ![]() .
.
Author Contributions
LW and SY designed the study. SY performed the experiments, analyzed the data and prepared the manuscript. XS and LW were responsible for project development and manuscript editing. All authors read and approved the final manuscript.
Ethics Approval and Consent
The study was approved by the medical research ethics committee of the International Peace Maternal and Child Health Hospital (the ethical approval ID is [2015] No.2 and date of approval is April 19, 2016). The patients/participants provided their written informed consent to participate in this study.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
