Abstract
Background:
Next-generation sequencing (NGS) has been widely used to identify targetable variants for patients with solid tumors, especially lung cancer. Circulating tumor DNA (ctDNA) has emerged as an alternative approach for tumor biopsy. However, the feasibility of ctDNA in detecting molecular variants remains debatable.
Methods:
Herein, we performed NGS on matched tissue and plasma samples from 146 Chinese patients with lung cancer. The concordance of variants between tissue and plasma samples was explored at patient and variant levels.
Results:
More than 80% of patients harbored at least one concordant variant in tissue and plasma samples. A total of 506 variants were shared between tissue and plasma samples, and 432 variants were identified in tissue only and 92 variants were identified in plasma only. The sensitivity and positive predictive value (PPV) of all variants detected in plasma were 53.9% and 84.6%, respectively. High concordance was observed in several driver genes. In details, epidermal growth factor receptor exon 19 deletion (EGFR 19del), EGFR p.S768I, anaplastic lymphoma kinase (ALK) fusion, rearranged during transfection (RET) fusion, and kirsten rat sarcoma viral oncogene homolog (KRAS) p.G12C achieved a sensitivity of 90%, 100%, 85.7%, 100%, and 85.7%, respectively. Four EGFR-altered lung adenocarcinoma patients who underwent ctDNA-based NGS at initial diagnosis benefited from first-line gefitinib/icotinib with a median progression-free survival of 379.5 days.
Conclusions:
Our work provided the clinical evidence of feasibility of ctDNA-based NGS in guiding decision-making in treatment. ctDNA-based NGA could be a reliable alternative approach for tissue biopsy in patients with lung cancer.
Introduction
Lung cancer is the most common malignant tumor and the leading cause of cancer-related death.1,2 The identification of cancer driver genes provides growing number of novel treatment options for lung cancer, in particular, for lung adenocarcinoma (LUAD). 3 Patients harboring actionable variants could benefit from personalized targeted therapies such as tyrosine kinase inhibitors (TKIs). 4 To date, an array of predictive biomarkers in lung cancer have been approved for clinical use, including epidermal growth factor receptor (EGFR) exon 19 deletion, EGFR exon 21 p.L858R, EGFR p.S768I, EGFR p.L861Q, EGFR p.G719X, EGFR exon 20 insertion, kirsten rat sarcoma viral oncogene homolog (KRAS) p.G12C, hepatocyte growth factor receptor gene (MET) exon14 skipping mutation, B-Raf proto-oncogene, serine/threonine kinase (BRAF) p.V600E, anaplastic lymphoma kinase (ALK) rearrangement, ROS proto-oncogene 1 receptor tyrosine kinase (ROS1) rearrangement, rearranged during transfection (RET) rearrangement, neurotrophic receptor tyrosine kinase 1/2/3 (NTRK1/2/3) fusion, programmed cell death ligand-1 expression, and tumor mutation burden.4-6
High-throughput next-generation sequencing (NGS) has been widely used to detect oncogenic driver variants and provides treatment suggestions. 7 In recent years, liquid biopsy with plasma emerged as an alternative approach for tumor biopsy. Cell-free DNA (cfDNA) consists of DNA fragments derived from necrotic and apoptotic eukaryotic cells, which has emerged as a promising noninvasive diagnostic approach in oncology. 8 Tumor-specific, circulating cfDNA (also called circulating tumor DNA [ctDNA]) is released by necrotic or apoptotic tumor cells. 9 The advantages of ctDNA sequencing includes non-invasiveness, short turn-around time, and overcoming tumor heterogeneity. 10 Therefore, it has been used to monitor the efficacy of targeted therapies, evaluate minimal residual disease, and identify drug-resistant mechanisms. 11 However, the concordance of genomic profiling between tissue and plasma remains to be a major concern during clinical practice.
Herein, we performed a retrospective study to describe genomic profiles of lung cancer with matched tissue and plasma samples. We aimed to investigate the feasibility of ctDNA in detecting driver alterations in Chinese lung cancer patients, and to analyze the correlation of concordance of oncogenic alterations between tissue and plasma with histological subtype.
Materials and methods
Patients and samples
Between December 13, 2017 and July 14, 2021, tumor tissue and matched plasma samples derived from 146 patients who were diagnosed with lung cancer at Quanzhou First Hospital Affiliated to Fujian Medical University were retrospectively enrolled. Paired formalin-fixed paraffin-embedded (FFPE) tissue and plasma samples of the patients were collected. The demographic, clinical, and genetic profiling information of the patients were retrospectively collected from a de-identified database. Progression-free survival (PFS) was defined as the date of receiving treatment to the date of progression. The study was approved by the Institutional Review Board of Quanzhou First Hospital Affiliated to Fujian Medical University on August 13, 2021 ([2017] 163). Written informed consent was obtained from each patient.
DNA extraction
Ten microliters of peripheral blood were collected and stored at 4°C, and then centrifuged at 2000 g for 10 min at 4°C. The supernatant was transferred to a new centrifuge tube and centrifuged again at 16 000 g for 10 min at 4°C. The supernatant was stored at −80°C for further analysis. cfDNA was extracted using the QIAamp Circulating Nucleic Acid Kit (Qiagen, Hilden, Germany). QIAamp DNA FFPE Tissue Kit (Qiagen, Hilden, Germany) was used to extract tumor tissue DNA from FFPE samples according to the manufacturer’s instructions. The Qubit 2.0 fluorometer and the Qubit dsDNA HS Assay Kit (Life Technologies, Carlsbad, USA) were used to measure DNA concentration. Tissue-specific variants are defined as those variants detected only in tissue not in plasma samples.
Library construction and sequencing
A total of 20 to 80 ng of tissue DNA or cfDNA was fragmented by Covaris M220 focused ultrasonicator (Covaris, Inc., Woburn, MA, USA) followed by end repair, phosphorylation, dA addition, and adaptor ligation for library construction. The Agencourt AMPure XP beads (Beckman Coulter, Brea, CA, USA) were used to select DNA fragments with the range of 200 to 400 bp. Then, hybridization with capture probe baits, hybrid selection with magnetic beads, and PCR amplification were performed. Capture-based targeted sequencing was performed on tumor and plasma samples using a panel consisting of 168 cancer-related genes spanning 237 Kb of the human genome (Burning Rock Biotech, Guangzhou, China). The indexed samples were sequenced on Illumina NextSeq 500 with paired-end (Illumina, Inc., Hayward, CA, USA).
Sequence data analysis
The paired-end reads were mapped to the human genome (hg19) by Burrows-Wheeler aligner v.0.7.10. 12 Local alignment optimization, variant calling, and annotation were performed with the Genome Analysis Toolkit (GATK) v.3.2 13 and VarScan v.2.4.3. 14 DNA translocation analysis was performed with Factera v.1.4.3. 15 According to 1000 Genomes Project, Exome Aggregation Consortium, ESP6500SI-V2, and dbSNP databases, variants with over 0.1% population frequency were classified as single-nucleotide polymorphisms and excluded from further analysis. Remaining variants were annotated with the ANNOVAR software 16 and SnpEff v.3.6. 17 Allele frequency (AF) and copy number (CN) of genes were calculated. The threshold for single-nucleotide variants (SNVs)/insertions and deletions (Indels) detected from tissue and plasma was AF ⩾ 1% and AF ⩾ 0.1%, respectively. Genes with a CN more than 2.25 were considered with CN amplification, and those with a CN less than 1.5 were considered with CN deletion. Large genomic rearrangement (LGR) is defined as the deletion or duplication of single- or multi-exons at the DNA level. 18
Statistical analysis
The continuous variables were presented as mean or median. The categorical variables were presented as frequencies. Unpaired Wilcoxon signed-rank test was used to compare continuous variables, while two-sided Fisher’s exact tests were used to compare categorical variables, as appropriate. P < .05 was considered statistically significant. The sensitivity was defined as the ratio of number of true positive cases/variants to the sum of true positive and false negative cases/variants. The positive predictive value (PPV) was defined as the ratio of number of true positive cases/variants to the sum of true and false positive cases/variants. The sensitivity and PPV were calculated by using tumor biopsy samples as references. All bioinformatics analyses were performed with R (v.3.5.3, the R Foundation for Statistical Computing, Vienna, Austria).
Results
Study cohort
A total of 146 lung cancer patients were included in this study. There were 102 males (69.9%) and 44 females (30.1%) with a median age of 64 years. Most patients had LUAD (111 of 146, 76%). There were 15 patients with lung squamous cell carcinoma (LUSC, 10.3%), 4 patients with small cell lung cancer (SCLC, 2.7%), and 3 patients with large cell lung cancer (LCLC, 2.1%). The histological subtype of 13 patients was unknown. Most patients were diagnosed at stage IV (120/146, 82.2%). There were 24 patients at stage III (16.4%), 1 patient at stage II (0.7%), and 1 patient at stage I (0.7%). The median cfDNA concentration was 10.34 ng/mL with a range of 3.7 to 122.6 ng/mL. The median cfDNA amount extracted from plasma was 53 ng with a range of 16 to 515 ng. The clinicopathological characteristics of patients are summarized in Table 1.
Baseline characteristics of patients.

Abbreviations: LCLC, large cell lung cancer; LUAD, lung adenocarcinoma; LUSC, lung squamous cell carcinoma; SCLC, small cell lung cancer.
Genomic profiling in tissue and plasma
A total of 938 variants including SNVs/Indels, copy number variants (CNVs)/LGRs, and fusions were identified in 146 tissue samples. Meanwhile, there were 598 alterations identified in 127 of 146 plasma samples (87%). None of variants were identified in 19 plasma samples (13%). The genomic profiling of tissue and plasma indicated that TP53 (72% and 60%) and EGFR (47% and 40%) were the two most frequently altered genes (Figure 1) in tissue and plasma samples, followed by KRAS (16% and 12%), phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit alpha (PIK3CA) (11% and 8%), MET (10% and 5%), ALK (6% and 4%), and RET (5% and 5%) as shown in Figure 1.

Genomic profiling in tissue and plasma. PLA indicates plasma; TIS indicates tissue.
By-variant concordance between tissue and plasma samples
The concordance of alterations between tissue and plasma were further investigated. In total, 506 variants (SNVs/Indels/CNVs/LGRs/fusions) were identified both in tissue and plasma samples. Whereas, 432 variants were identified in tissue only and 92 variants were identified in plasma only (Figure 2A). The sensitivity of variants identified in plasma were 53.9% (506/938) with a PPV of 84.6%, when variants identified in tissue samples as a reference. Next, the sensitivity of plasma in detecting SNVs/Indels or CNVs was subsequently explored. There were 387 shared SNVs/Indels with 282 tissue-specific and 60 plasma-specific SNVs/Indels (Figure 2B). SNVs/Indels achieved a sensitivity of 57.8% (387/669) with a PPV of 86.6%. There were 61 CNVs identified in both tissue and plasma, 125 CNVs in tissue only, and 24 CNVs in plasma only (Figure 2C). CNVs achieved a 32.8% (61/186) sensitivity with a 71.8% PPV.
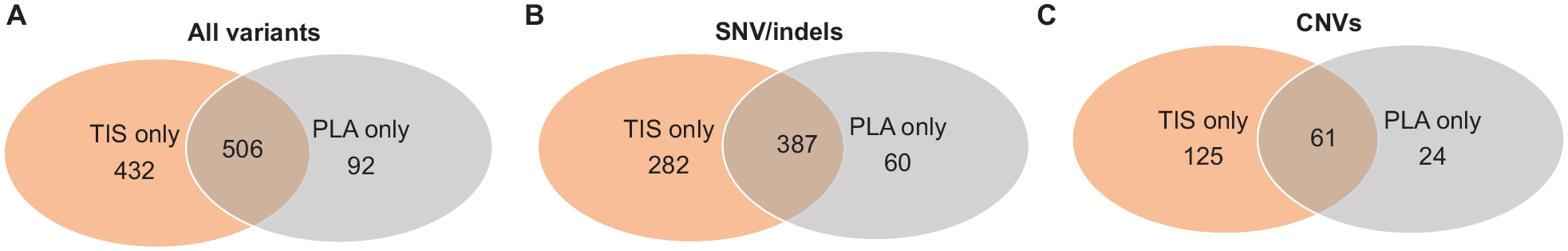
Concordance of molecular variants between tissue and plasma samples. (A) Concordance of all variants. (B) Concordance of SNVs/Indels. (C) Concordance of CNVs. PLA indicates plasma; TIS indicates tissue.
Then, the sensitivity of driver gene variants was explored, including EGFR, KRAS, MET, ALK, RET, and ROS1 variants (Figure 3A and B). Given that none of the variants were observed in 19 plasma samples, 126 paired positive tissue and plasma samples were performed for subsequent analysis. A total of 104 actionable variants were identified both in tissue and plasma samples, 19 variants in tissue only, and 6 variants in plasma only (Figure 3C). The sensitivity and PPV of actionable variants achieved 84.6% and 99.9%. EGFR is the most frequently altered gene in lung cancer. Sixty-three EGFR variants were detected both in tissue and plasma, 9 in tissue only, and 3 in plasma only (Figure 3D). The sensitivity of EGFR variants was 87.5% and its PPV was 95.5%. For KRAS and MET variants, the sensitivity was 84% (with a PPV of 89%) and 67% (with a PPV of 100%), respectively. ALK, RET, and BRAF achieved a 100% sensitivity and a 100% PPV.

Concordance in driver genes. (A) Genomic profiling of patients detected from tissue samples. (B) Genomic profiling of patients detected from plasma samples. (C) Concordance of all driver mutations. (D) Concordance of EGFR variants. PLA indicates plasma; TIS indicates tissue.
By-patient concordance between tissue and plasma samples
The by-patient concordance of variants between tissue and plasma was also analyzed (Figure 4A and B). In all variants, concordance were observed in 83.6% patients (Figure 4A). Tissue-specific variants were identified in 13% of patients (Figure 4A). In CNVs/LGRs, only 66.4% concordance were observed (Figure 4B), while 30.8% of patients harbored tissue-specific CNVs/LGRs (Figure 4A). The sensitivity of CNVs and LGRs detected in plasma was only 39.2% (with a PPV of 87.9%) and 33.3% (with a PPV of 100%), respectively (Table S2). The concordance of all variants between tissue and plasma increased to 83.6%, when CNVs/LGRs were excluded. In other variants, over 80% concordance were also observed (Figure 4B), such as SNVs/Indels and driver variants. EGFR variants between tissue and plasma were concordant in 90.4% patients. There were 9.6% of patients who showed different EGFR alteration status between tissue and plasma, including 8.2% of patients with tissue-specific EGFR variants and 1.4% of patients with plasma-specific EGFR variants.

Concordance in all lung cancer patients. (A) Distribution of patients with concordant variants between tissue and plasma samples (Concordance), discordant variants both in tissue and plasma (Discordance_Both), tumor-specific variants (Discordance_onlyPLA), plasma-specific variants (Discordance_onlyTIS), and no variants detected both in tissue and plasma (Negative_Both). (B) Distribution of patients with concordant and discordant variants. Concordance presents patients with Concordance and Negative_Both in Figure 4A. Discordance presents patients with Discordance_Both, Discordance_onlyPLA, and Discordance_onlyTIS in Figure 4A. LGR indicates large genomic rearrangement; PLA, plasma; rmCNV: variants with excluding CNVs; TIS, tissue.
The concordance of variants was further explored in patients with different histological subtypes. All variants achieved a concordance of 86.3% in 126 non-small cell lung cancer (NSCLC) patients (111 LUAD patients and 15 LUSC patients) who had detectable variants both in tissue and plasma samples (Figure 5A). Concordance of either EGFR, ALK, or KRAS variants reached over 90% in NSCLC (Figure 5B). Similar results were also observed in LUAD and LUSC (Figure 5B). Discordance of CNVs/LGRs reached 31.75%, 32.43%, and 26.67% in NSCLC, LUAD, and LUSC, respectively (Figure 5B).
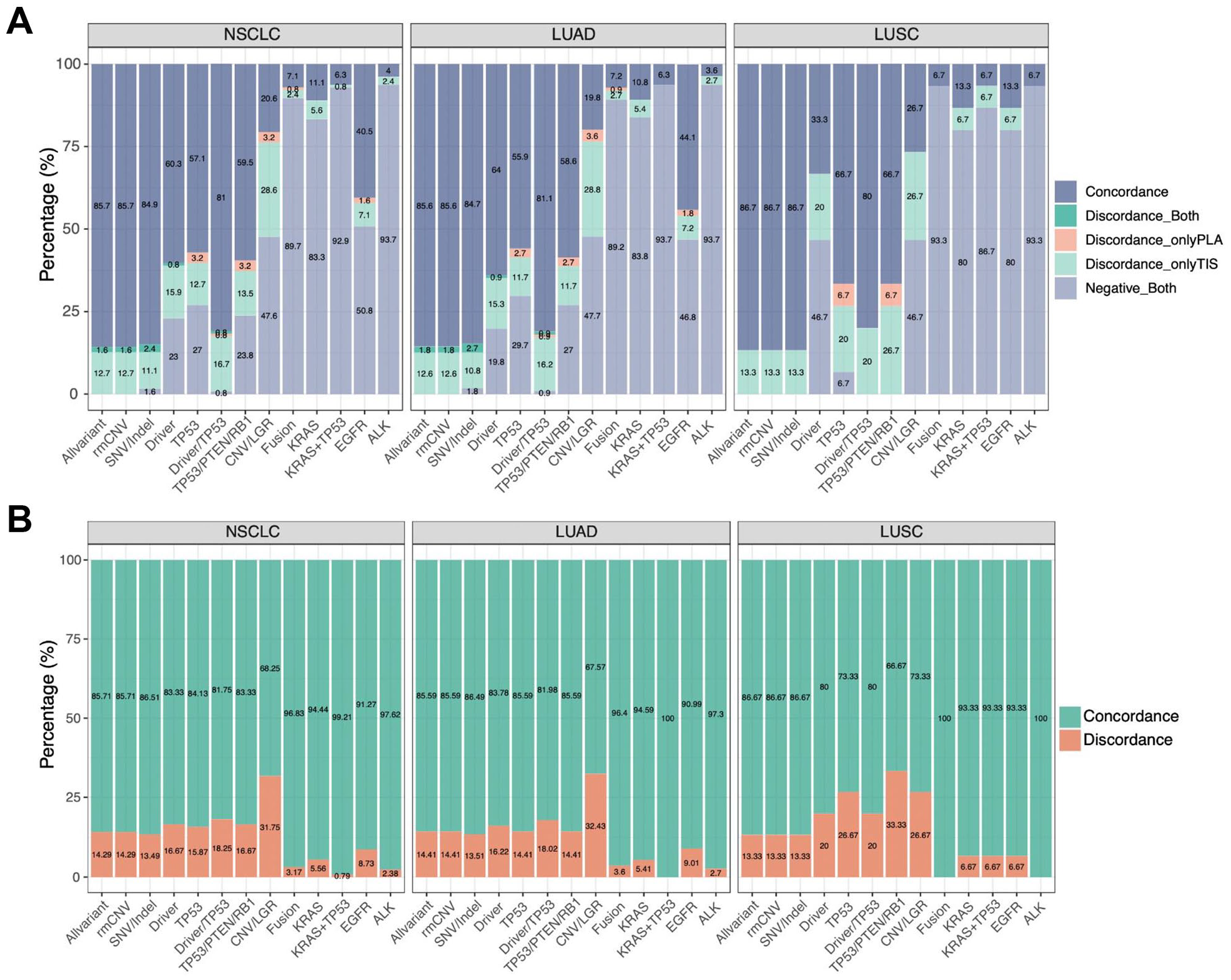
Concordance in NSCLC patients. (A) Distribution of patients with concordant variants between tissue and plasma samples (Concordance), discordant variants both in tissue and plasma (Discordance_Both), tumor-specific variants (Discordance_onlyPLA), plasma-specific variants (Discordance_onlyTIS), and no variants detected both in tissue and plasma (Negative_Both) in NSCLC, LUAD, and LUSC. (B) Distribution of patients with concordant and discordant variants. Concordance represents patients with Concordance and Negative_Both in Figure 5A. Discordance represents patients with Discordance_Both, Discordance_onlyPLA, and Discordance_onlyTIS in Figure 5A. LUAD indicates lung adenocarcinoma; LUSC, lung squamous cell carcinoma; NSCLC, non-small cell lung cancer; PLA, plasma; TIS, tissue; rmCNV: variants with excluding CNVs.
Concordance of actionable variants between tissue and plasma samples
Next, the concordance of actionable variants between tissue and plasma samples was explored. As shown in Table 2, the sensitivity of EGFR 19del, ALK fusion, KRAS p.G12C, EGFR p.L858R, and MET exon 14 skipping mutation achieved 90%, 85.7%, 85.7%, 77.3%, and 75%, respectively. Furthermore, the sensitivity of EGFR p.T790M, EGFR exon 20 insertions, EGFR p.S768I, EGFR p.L861Q, RET fusion, BRAF p.V600E achieved 100%, respectively. Only 1 patient harbored ROS1 fusion detected from the tissue sample rather than the plasma sample. NTRK1/2/3 fusions were not defected in this work. Of note, 1 patient harbored EGFR p.L858R detected from the plasma rather than the tissue sample.
Concordance of actionable alterations between tissue and plasma samples in lung cancer patients.

Abbreviations: FN, false negative; FP, false positive; NA, not applicable; PPV, positive predictive value; TP, true positive.
In this study, 111 LUAD patients and 15 LUSC patients were enrolled. Based on tumor-based NGS, 74 (74/111, 66.7%), 53 (53/111, 47.7%), 7 (7/111, 6.3%), 4 (4/111, 3.6%), 3 (3/111, 2.7%), 2 (2/111, 1.8%), 2 (2/111, 1.8%), and 1 (1/111, 0.9%) LUAD patient(s) harbored TP53 variants, EGFR variants, KRAS p.G12C, ALK fusions, MET exon 14 skipping mutations, BRAF p.V600E, RET fusions, and ROS1 fusion, respectively. Among 53 patients with EGFR variants, 25, 20, and 3 patients harbored EGFR 19del, EGFR p.L858R, EGFR p.L861Q; 2 patients harbored EGFR p.T790M, EGFR p.S768I, EGFR exon 20 insertion, and EGFR p.G719X, respectively. For LUSC, 13, 1, and 1 patient(s) harbored TP53 variants, EGFR variant (EGFR 19del), and ALK fusion, respectively. None of LUSC patients harbored EGFR p.L858R, EGFR p.T790M, EGFR p.S768I, EGFR exon 20 insertion, EGFR p.L861Q, EGFR p.719X, ROS1 fusion, RET fusion, MET exon 14 skipping mutation, or BRAF p.V600E. The difference of concordance of molecular variants between tissue and plasma in LUAD vs LUSC was explored. We found a similar sensitivity of plasma in detecting tumor protein p53 (TP53) variants, EGFR variants and ALK fusions from LUAD vs LUSC, respectively (Table 3).
The difference of concordance of molecular alteration between tissue and plasma in LUAD and LUSC.

Abbreviations: LUAD, lung adenocarcinoma; LUSC, lung squamous cell carcinoma; NA, not applicable; TP, true positive.
indicates combined with EGFR 19del, EGFR p.L858R, EGFR p.T790M, EGFR p.S768I, EGFR p.L861Q, EGFR p.G719X, and EGFR exon20 insertion.
In this study, a total of 23 patients carried EGFR p.L858R. There were 4 patients with stage III disease and 19 patients with stage IV disease. Of these 23 patients, 17 patients harbored EGFR p.L858R detected both in tissue and plasma (G1), 5 patients harbored EGFR p.L858R detected in tissue only (G2), and 1 patient harbored EGFR p.L858R in plasma only. The difference of clinicopathological characteristics between G1 and G2 was preliminarily explored. A similar median age (70 vs 71 years old, P = .169) and distribution pattern of gender (female/male, 4/1 vs 6/11, P = .135) was observed between G1 and G2. Further study investigating the clinicopathological characteristics of patients with tissue-specific variants, or plasma-specific variants are needed in a large cohort of lung cancer patients. These studies might enable clinical physician to identify patients who are suitable for tumor- or ctDNA-based NGS.
The outcomes of patients who underwent plasma-based NGS
In this work, 4 patients who treated with targeted agents had available survival data. These 4 patients were diagnosed with treatment-naïve EGFR-mutant LUAD, furthermore, EGFR mutations were detected both in plasma and paired tissue samples. Both Case 1 and Case 2 who harbored EGFR p.L858R benefited from first-line gefitinib with Case 1 having a PFS of 365 days and Case 2 having a PFS of 315 days. Case 3 who harbored EGFR 19del achieved a PFS of 394 days with first-line gefitinib. In addition, Case 4 carrying EGFR 19del benefited from first-line icotinib with a PFS of 455 days (Table 4).
The outcomes of patients who underwent ctDNA-based NGS.

Abbreviations: ctDNA, circulating tumor DNA; EGFR, epidermal growth factor receptor; NGS, next-generation sequencing; PFS, progression-free survival; PR, partial response; SD, stable disease; TKI, tyrosine kinase inhibitor.
Discussion
Molecular testing regarding driver variants in lung cancer patients could guide decision-making in treatment, such as EGFR sensitizing mutations. Although liquid biopsy has been increasingly used in lung cancer patients, the concordance of variants between tissue and plasma remains to be a major concern in clinical practice. 11 In this study, we performed capture-based targeted sequencing using a panel consisting of 168 cancer genes on matched tissue and plasma samples derived from 146 Chinese lung cancer patients. The genomic profiling of enrolled lung cancer patients were described and the concordance of variants between tissue and plasma samples was explored.
Consistent with previous studies on the genomic profiling of lung cancer, TP53 and EGFR mutations were the most frequently altered genes. Other actionable variants in driver genes such as MET, ALK, RET, and ROS1 were also observed in this work. A study has revealed 98% concordance of variants in 39 genes between tissue and matched plasma samples derived from 82 NSCLC patients, with 71% sensitivity and 100% specificity. 19 Recent studies in treatment-naïve NSCLC patients reported the concordance of NGS results between tissue and plasma ranged from 62.2% to 77.6%.20,21 Although the concordance of variants between tissue and plasma has been reported in several studies,19-21 it varies across different studies.
In our study, more than 80% patients harbored at least a concordant variant both in tissue and plasma including SNVs/Indels, CNVs/LGRs and fusions. Among 13% patients with discordant variants, most patients had tissue-specific variants. Moreover, 3.4% patients harbored a concordant altered gene but with distinct variants in tissue and plasma. When focusing on the variants, a total of concordant 506 variants were identified both in tissue and plasma samples. Compared with tissue, all variants in plasma showed a sensitivity of 53.9% and a PPV of 84.6%.
Compared with SNVs/Indels or fusions, the concordance of CNV between tissue and plasma were poor. These results were consistent with previous studies.22,23 To date, identifying CNVs in ctDNA has proven to be challenging.
Data suggest that almost 20% to 30% of NSCLCs are driven by an activating KRAS mutations. 24 Point mutations in KRAS most commonly occur at codon 12. The KRAS p.G12C mutation occurs in 13% of NSCLC.25,26 The National Comprehensive Cancer Network recommends sotorasib as a subsequent therapy option for patients with metastatic NSCLC and KRAS p.G12C mutations who have disease progression after treatment with platinum-based chemotherapy (±immunotherapy) based on clinical trial data and Food and Drug Administration approval. 5 In this study, the sensitivity of KRAS p.G12C between tissue and plasma achieved 85.7% with a PPV of 100%. Other driver variants, including EGFR 19del, ALK fusion, RET fusion, BRAF p.V600E, also had a high sensitivity.
There are some limitations associated with our study. The analyses on correlation of clinicopathological characteristics with concordance of genetic variants between tissue and plasma samples might be helpful in identifying patients who are or not suitable for plasma-based NGS in detecting status of molecular alterations. However, due to the fact that this is a retrospective study, most clinicopathological characteristics are missing besides sex, age, tumor stage, and histological subtype. A prospective, large cohort study is warranted to explore the clinicopathological characteristics of lung cancer patients who had high concordance of genetic alterations between tissue and plasma samples.
Conclusion
Our study performed NGS on paired tissue and plasma samples from Chinese patients with lung cancer. High concordances of lung cancer driver variants were observed both at patient and variant levels. ctDNA-based NGS might be a reliable alternative approach for tissue biopsy in patients who had unavailable tissue samples.
Supplemental Material
sj-docx-1-onc-10.1177_11795549221116834 – Supplemental material for Concordance of Genomic Profiles in Matched Tissue and Plasma Samples From Chinese Patients With Lung Cancer
Supplemental material, sj-docx-1-onc-10.1177_11795549221116834 for Concordance of Genomic Profiles in Matched Tissue and Plasma Samples From Chinese Patients With Lung Cancer by Yueming He, Weifeng Guo, Meng Xu, Junling Huang, Xiange Zhang, Huanzhang Su, Dongxia Hong and Qun Liu in Clinical Medicine Insights: Oncology
Supplemental Material
sj-xlsx-2-onc-10.1177_11795549221116834 – Supplemental material for Concordance of Genomic Profiles in Matched Tissue and Plasma Samples From Chinese Patients With Lung Cancer
Supplemental material, sj-xlsx-2-onc-10.1177_11795549221116834 for Concordance of Genomic Profiles in Matched Tissue and Plasma Samples From Chinese Patients With Lung Cancer by Yueming He, Weifeng Guo, Meng Xu, Junling Huang, Xiange Zhang, Huanzhang Su, Dongxia Hong and Qun Liu in Clinical Medicine Insights: Oncology
Footnotes
Acknowledgements
Genomic profiling of the patient was performed at Burning Rock Biotech (Guangzhou, China). The authors thank the patients and their families, medical and research staff who participated in this study.
Declaration of Conflicting Interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding:
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Author Contributions
Yueming He: Conceptualization, Methodology, Data Curation, Resources, Writing - Original Draft; Weifeng Guo: Methodology, Data Curation, Writing - Review & Editing; Meng Xu: Methodology, Data Curation, Writing - Review & Editing; Junling Huang: Methodology, Data Curation, Writing - Review & Editing; Xiange Zhang: Methodology, Writing - Review & Editing; Huanzhang Su: Methodology, Writing - Review & Editing; Dongxia Hong: Methodology, Writing - Review & Editing; Qun Liu: Conceptualization, Methodology, Resources, Supervision, Writing - Review & Editing.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
