Abstract
Background
Non-small cell lung cancer (NSCLC) patients who have mutations in their epidermal growth factor receptor (EGFR) gene respond more favorably to tyrosine kinase inhibitors (TKIs) than to standard chemotherapy. However, tissue biopsy-based EGFR testing is invasive, costly, and technically challenging. Plasma-derived circulating tumor DNA (ctDNA) offers a minimally invasive and cost-efficient alternative for mutation profiling. This study assessed the agreement between EGFR mutation status in plasma-derived ctDNA and tissue biopsy in NSCLC patients from tertiary care hospitals in Bangladesh.
Methods
In this cross-sectional analytical study, we recruited 32 patients with NSCLC before EGFR-TKI treatment. EGFR mutations in ctDNA samples were identified using the Amplification Refractory Mutation System (ARMS) polymerase chain reaction method. Tissue biopsy results were obtained from routine diagnostic procedures. Agreement between ctDNA and tissue biopsy results was assessed using kappa statistics, and diagnostic performance metrics were calculated.
Results
Most of our study participants were male (75%) and had stage IV lung adenocarcinoma (72%). We observed substantial agreement between plasma-derived ctDNA samples and tissue biopsies (kappa, κ = 0.683). This agreement was almost perfect (κ = 0.826) when calculated for patients with stage IV disease. The overall concordance was 84.4%. Compared with tissue biopsy, ctDNA testing yielded a sensitivity of 73.3% and a specificity of 94.1%.
Conclusion
Plasma-derived ctDNA demonstrates substantial agreement with tissue biopsy for EGFR mutation detection in patients with NSCLC, particularly those with advanced-stage disease. These findings support ctDNA as a viable alternative for molecular profiling in settings where tissue biopsy is limited or impractical.
Introduction
Cancer remains a leading cause of global mortality, accounting for 16.8% of all deaths and 22.8% of deaths related to non-communicable diseases in 2022. Among these, lung cancer is the most lethal malignancy, with approximately 2.5 million incident cases and 1.8 million deaths annually. 1 In Bangladesh, lung cancer is the third most frequently diagnosed cancer and the second leading cause of oncological mortality. 2
Non-small cell lung cancer (NSCLC) accounts for approximately 85% of all lung cancer cases. 3 To ensure accurate categorization and the selection of appropriate targeted therapies, the 2021 World Health Organization classification of thoracic tumors mandates molecular profiling for biomarkers including EGFR, KRAS, ALK, ROS1, RET, NTRK1-3, BRAF, MET, and ERBB2. 4 The epidermal growth factor receptor (EGFR), located on chromosome 7p11.2, is a single-pass transmembrane protein that triggers downstream signaling cascades—such as Ras/Raf/MAP kinase and PI3K/Akt/mTOR—to promote cell proliferation. Activating mutations or disruptions in these cascades result in constitutional activation of the receptor without ligand-induced stimulation, driving neoplastic transformation.5–8
Among the various mutations identified in NSCLC, the EGFR mutations are the most prevalent among Asian patients. They are detected in approximately 55% of cases in East Asia and between 23% and 44% of cases in India, whereas their frequency among Caucasian patients is considerably lower, ranging from 10% to 15%.9–11 Importantly, about 93% of EGFR mutations are detected within exons 18–21, which encode the tyrosine kinase domain. 5 NSCLC patients harboring mutations in this domain are eligible for targeted therapy with tyrosine kinase inhibitors (TKIs). The advent of TKIs has transformed treatment paradigms, replacing conventional chemotherapy and nearly doubling progression-free survival. 10
In real-world clinical practice, particularly within resource-limited settings like Bangladesh, EGFR mutation detection faces significant logistical and biological challenges. Tissue biopsy remains the gold standard 12 ; it is invasive, and often associated with complications such as pneumothorax, hemoptysis, and infection. 13 Moreover, obtaining adequate tumor material is frequently difficult, particularly in patients with advanced disease who are unfit for repeat procedures. Even when tissue is available, sampling from a single tumor site may not capture the full molecular heterogeneity, 14 leading to incomplete mutation profiling. Furthermore, pathological evaluation reveals malignant cells in approximately 73% of specimens, 15 often necessitating repeat biopsies to obtain enough tissue for genetic testing. These limitations can delay treatment initiation and increase patient burden, especially in resource-constrained healthcare systems. These limitations can be overcome by liquid biopsy, a non-invasive, repeatable procedure that utilizes plasma-derived circulating tumor DNA (ctDNA). ctDNA is a fraction of the total cell-free DNA released into the bloodstream from both primary and metastatic sites via apoptosis, necrosis, or active secretion.16,17 Therefore, it captures mutations from multiple metastatic sites. Also, ctDNA has a half-life of approximately 114 min. 18 So, it serves as a dynamic biological marker for real-time monitoring of disease status and drug responsiveness.
Evaluating the concordance between ctDNA and tissue biopsy is critical to determine whether ctDNA can serve as a reliable substitute in clinical practice. Research comparing ctDNA and tissue biopsies in the South Asian region remains sparse, with no published data currently available from Bangladesh. While the 2021 WHO classification and international guidelines mandate broader molecular profiling of druggable genetic drivers, 4 this study focused specifically on EGFR due to its exceptionally high prevalence in Asian NSCLC patients to validate a cost-effective liquid biopsy approach. In this study, we assessed the agreement between EGFR mutation profiles in ctDNA and tissue biopsies in NSCLC patients at tertiary care hospitals in Bangladesh. Our aim was to establish whether ctDNA can serve as a viable alternative for molecular profiling in settings where tissue biopsy is limited by clinical complications, financial constraints, or inadequate sample yields.
Materials and methods
Study design and participants
This cross-sectional analytical study was conducted at Bangladesh Medical University (formerly Bangabandhu Sheikh Mujib Medical University) over a 1-year period from March 2024 to February 2025. Ethical clearance for this study was granted by the university's Institutional Review Board. The sample size was calculated specifically for an agreement test of a two-rater study on a dichotomous variable. To ensure statistical significance, we targeted a kappa value (κ) of 0.5 with a power of 80% and a two-tailed alpha (α) of 0.05, which yielded a minimum required sample size of 32 participants. 19 Patients were recruited using a consecutive sampling approach from the oncology inpatient and outpatient departments of several tertiary care hospitals across the country. Participants were aged ≥ 40 years and had histologically confirmed NSCLC. The eligibility criteria included no prior treatment with systemic anti-cancer drugs, including TKIs, and the availability of an EGFR mutation test result from a tissue biopsy within 1 week of blood sample collection. After obtaining informed written consent from the study participants, an initial evaluation was performed by history taking. The clinical history of the participants and EGFR mutation test results from tissue biopsy were subsequently recorded in a pre-formed data collection sheet.
cfDNA extraction and EGFR mutation analysis
From each study participant, eight mL of blood was collected in K2EDTA tubes via antecubital venipuncture for ctDNA extraction. Following the collection, the samples were processed within two hours. Initially, they were centrifuged at 3000 × g for 10 min to separate plasma. Subsequently, a microcentrifugation at 11,000 × g for 10 min was performed to produce clear samples for mutation analysis. Plasma was stored at −80°C until DNA extraction using the CatchGene® Catch-cfDNA Serum/Plasma 1000 Kit (CatchGene Co. Ltd., New Taipei City, Taiwan). DNA quality and concentration were verified using a NanoDrop™ ND 2000 Spectrophotometer (Thermo Fisher Scientific Inc., Waltham, MA, USA), accepting an A260/A280 ratio of 1.7–2. 20
EGFR mutation status was assessed using the AmoyDx EGFR 29 Mutations Detection Kit (Amoy Diagnostics, Xiamen, China) on a Rotor-Gene Q (72 wells) real-time polymerase chain reaction (PCR) machine (Qiagen, Hilden, Germany). This assay utilizes Amplification Refractory Mutation System (ARMS) technology, where mutant templates match the 3ʹ end of the primer for selective amplification, which is then detected via FAM-labeled fluorescent probes. It qualitatively identifies 29 somatic mutations in the EGFR gene, categorized into seven distinct reaction groups targeting exons 18 through 21. These include G719X substitutions (exon 18), in-frame deletions (exon 19), the T790 M resistance mutation, S768I, and various insertions (exon 20), as well as the L858R and L861Q point mutations (exon 21).
Each PCR run included nine samples, comprising seven patient samples, one positive control, and one non-template control and was subjected to 46 cycles of amplification. For each sample, eight reaction tubes were utilized: tubes one through seven featured mutation detection systems with FAM-labeled probes to detect EGFR mutations and internal control systems with HEX-labeled probes to identify the presence of PCR inhibitors to ensure experimental validity. The eighth tube contained an external control reaction mix with an FAM-labeled probe targeting a region of genomic DNA to assess the quality of the template DNA. Analysis of the mutation was performed by calculating the ΔCt value (ΔCt = Mutant FAM Ct – External Control FAM Ct). A sample was determined to be positive for a mutation if the mutant FAM Ct was < 29 and the ΔCt was < 13, while samples exhibiting no amplification or a ΔCt ≥ 13 were classified as negative or below the kit's limit of detection (LOD).
Statistical analysis
Data from this study were analyzed using the Statistical Package for Social Sciences (ver. 27) software. Chi-square and Fisher's exact tests were applied to compare the EGFR mutation status across various clinicopathological characteristics. Agreement between plasma-derived ctDNA samples and tissue biopsies was assessed using the kappa statistic
Results
Clinicopathological characteristics of study participants
This study included a total of 32 NSCLC patients. Key clinicopathological parameters of the participants are summarized in Table 1. The mean age of the participants was 59.2 (59.2 ± 10.9) years. Most of them were male (n = 24, 75%) and had adenocarcinoma histology (n = 30, 94%). The tumor node metastasis (TNM) staging of the tumor was determined using the International Association for the Study of Lung Cancer's 9th edition of the lung cancer TNM staging system. Among the study participants, 23 (72%) had stage IV disease.
Clinicopathological characteristics and EGFR mutation frequency of the study subjects (n = 32).
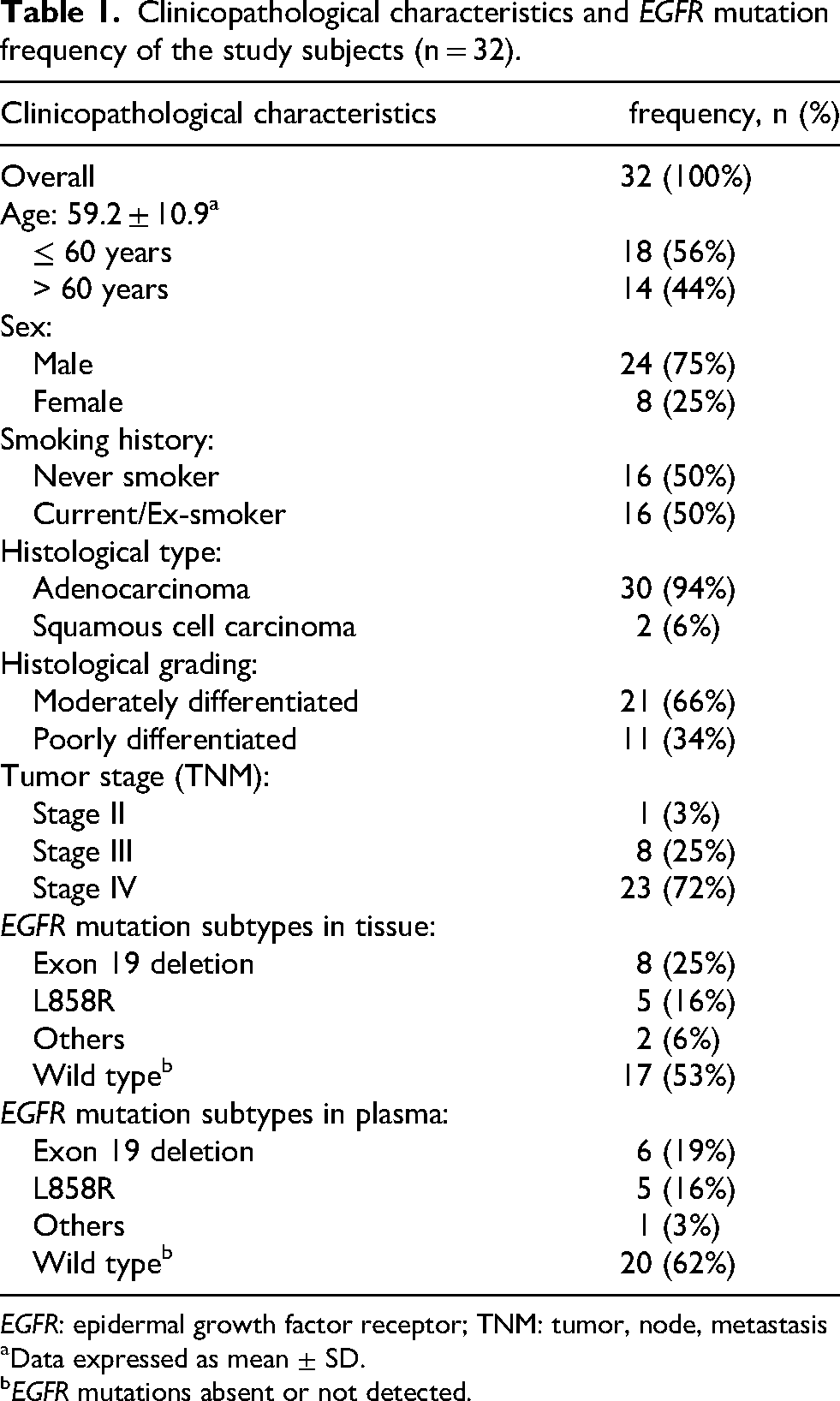
EGFR: epidermal growth factor receptor; TNM: tumor, node, metastasis
Data expressed as mean ± SD.
EGFR mutations absent or not detected.
EGFR mutation status
Of 32 study subjects, EGFR mutations were present in 15 (47%) tissue biopsy samples, and mutations were identified in 12 (38%) plasma-derived ctDNA samples, with exon 19 deletion being the most frequent alteration in both specimen types (Table 1). One plasma mutation was a TKI-resistant mutation (T790 M), whereas two of the tissue mutations were TKI-resistant (T790 M and exon 20 ins9). Table 2 outlines the relationship between EGFR mutation status and various clinicopathological parameters of the patients. In plasma, EGFR mutation rates were significantly higher in patients aged > 60 years and in never smokers (p-values of 0.043 and 0.028, respectively). However, sex distribution, histological grade, and tumor stage did not differ significantly across the groups. In tissue samples, mutations were significantly higher in never smokers (p-values of 0.013).
Relationship between EGFR mutation status and clinicopathological characteristics (n = 32).
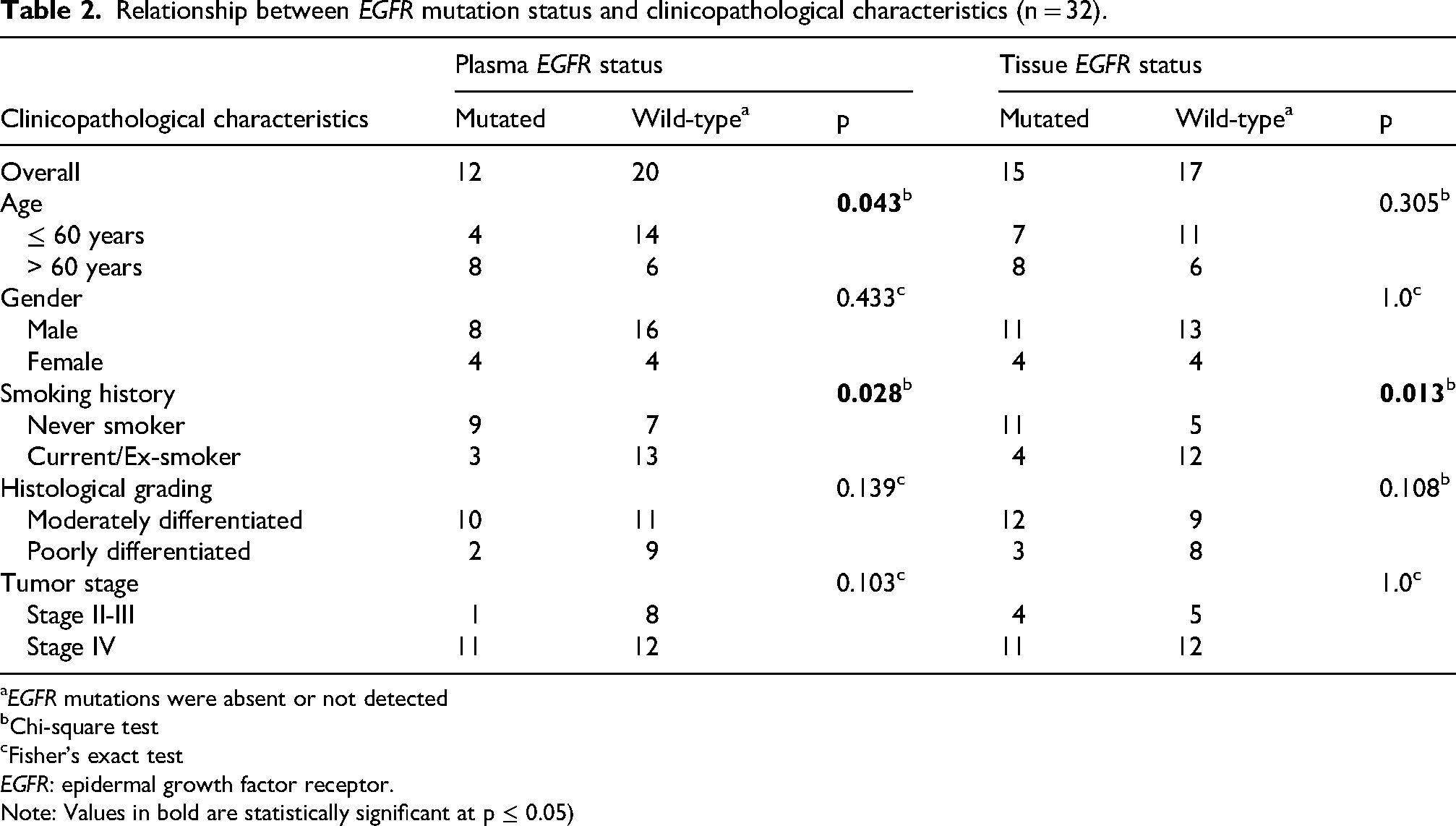
EGFR mutations were absent or not detected
Chi-square test
Fisher's exact test
EGFR: epidermal growth factor receptor.
Note: Values in bold are statistically significant at p ≤ 0.05)
Agreement on EGFR mutation status
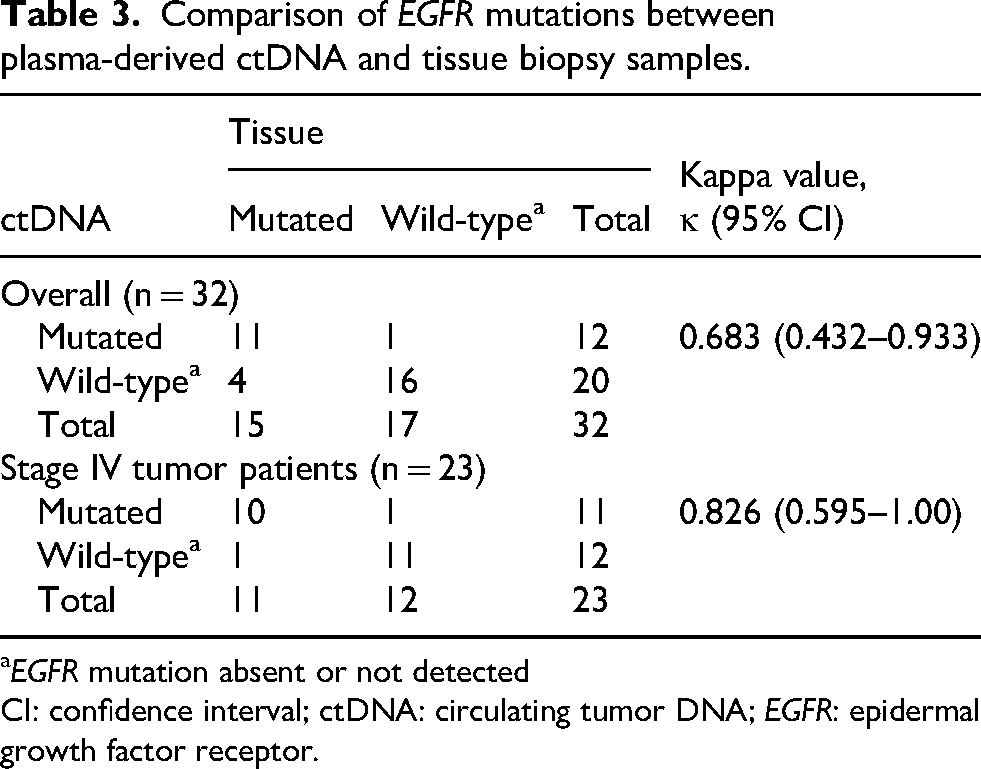
Of the 32 samples, 27 were concordant (11 positive and 16 negative), yielding a concordance rate of 84.4% (Table 3 and Table 4). Among the 15 mutations present in the tissue sample, 11 were also positive in the plasma, resulting in a sensitivity of 73.3%. The overall specificity was 94.1%, with a positive predictive value (PPV) of 91.7% and a negative predictive value (NPV) of 80%. Concordance and sensitivity increased to 91.3% and 90.9%, respectively, when calculated from patients with stage IV disease. We also assessed the agreement between these two sample types (Table 3). Overall, we found a κ value of 0.683, indicating substantial agreement between the two sample types. κ increased to 0.826 and reached almost perfect agreement when calculated from participants with stage IV disease. 21
Comparison of EGFR mutations between plasma-derived ctDNA and tissue biopsy samples.

EGFR mutation absent or not detected
CI: confidence interval; ctDNA: circulating tumor DNA; EGFR: epidermal growth factor receptor.
Diagnostic performance of plasma-derived ctDNA samples compared to tissue biopsy samples for detecting EGFR mutation in NSCLC.
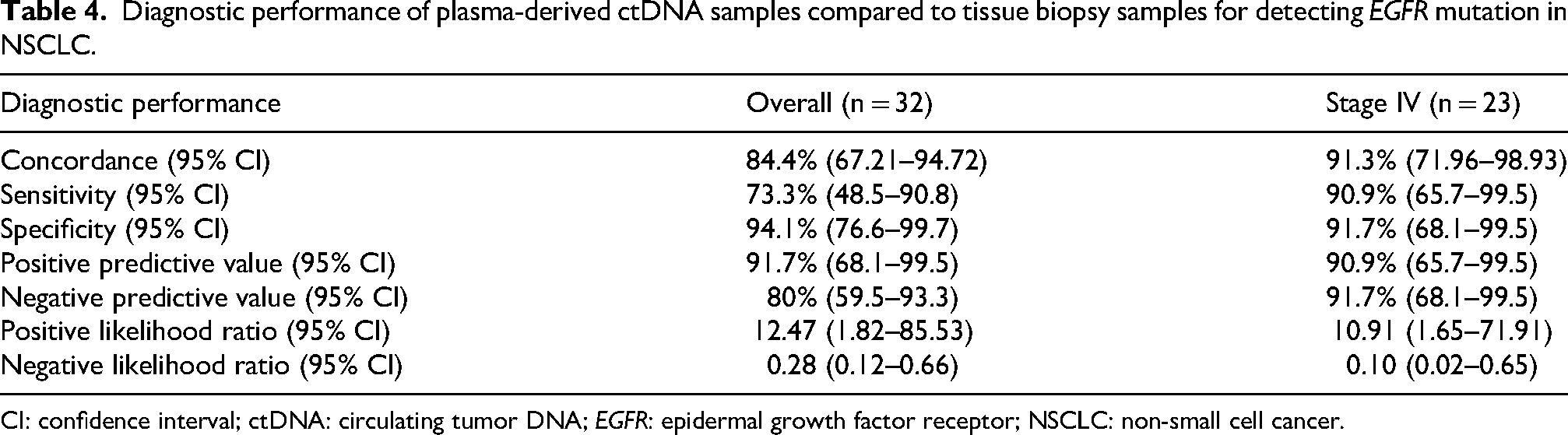
CI: confidence interval; ctDNA: circulating tumor DNA; EGFR: epidermal growth factor receptor; NSCLC: non-small cell cancer.
Discussion
Based on the currently available data, no prior study from Bangladesh has compared EGFR mutation detection between tissue biopsies and ctDNA samples. Our study showed good agreement and concordance between these two.
The majority of the participants were aged 55–70 years, with a mean age of 59.2 years, consistent with previous studies.11,22 The PIONEER study, the most extensive epidemiological study of EGFR mutations in Asia, also found a similar mean age. 23 Most of the subjects in our study were male (75%), and no statistically significant difference was found between sex distribution and mutation status in the plasma or tissue (p-values of 0.433 and 1.0, respectively). This result may reflect our relatively small sample size and imbalanced sex ratio (male:female = 3:1). Similar results were observed in some other studies.22,24–26 However, in large databases, female sex has been significantly associated with EGFR mutations, as in the IGNITE study. 27
Smoking is the most important factor for EGFR mutation detection, as EGFR mutations are more common in never smokers. 23 In our study, half of the participants were never smokers (50%). However, 75% of the mutations in plasma (9 out of 12) and 73% of the mutations in tissue (11 out of 15) were detected in never smokers. In the IGNITE study, 27 Han et al. observed similar results. In their study, 67% of mutations in tissue samples and 64% of mutations in plasma samples were found in never smokers. In our study, 72% of the study participants had stage IV NSCLC. In ctDNA samples, we found that the EGFR mutation was strongly positive in this group (11 out of 23 stage IV subjects) compared to stages II–III. These findings were similar to those reported by Wulandari et al. 22 In the metastatic stage, the ctDNA content in the plasma sample of the patient becomes very high compared to that in the earlier stage, resulting in a high mutation detection rate in the metastatic stage. 22
In our study, we found good agreement and concordance between the tissue biopsy and ctDNA samples. We obtained an overall κ-value of 0.683, which increased to 0.826 when calculated from stage IV disease subjects. Our results were supported by some small- and large-scale studies.28–30 Soria-Comes et al. 28 reported an overall κ-value of 0.6, which increased to 0.7 when calculated from subjects with stage IV disease. Veldore et al. 29 published their study on 132 Indian patients having stage IV NSCLC with distal metastasis, where they used next-generation sequencing (NGS) to test plasma samples. Therefore, they found almost perfect agreement (κ = 0.931) between the plasma and tissue samples. In the IFUM study, we calculated a κ-value of 0.755 from the given data. In this study, the authors included patients with locally advanced or metastatic NSCLC, which was reflected in their high κ-values. 30
In our study, we also found a high concordance between tissue and plasma samples. The overall concordance rate was 84.4%. Compared with tissue biopsy, ctDNA demonstrated a sensitivity of 73.3%, specificity of 94.1%, PPV of 91.7%, and NPV of 80%. Concordance and sensitivity substantially increased when calculated from subjects with stage IV disease (from 84.4% to 91.3% and from 73.3% to 90.9%, respectively). Our study was also supported by some studies conducted in India, the Asia-Pacific region, and Europe.11,24,28–30 Three studies reported similar results in the Indian population. Joshi et al. 24 and Prabhash et al. 11 reported concordance of 83% and 82.9%, respectively. Veldore et al. 29 did their study on stage IV disease subjects and found high concordance (97%), high sensitivity (91.1%), and perfect specificity (100%), which was very close to our results from stage IV disease. Soria-Comes et al. 28 published a study on the Spanish population, which provided results similar to those of our study (concordance, 87.4%; sensitivity, 70.6%; and specificity, 91.7%). The large IFUM study on the Caucasian population also reported similar results (concordance, 94.3%; sensitivity, 65.7%; specificity, 99.8%; and PPV, 98.6%). 30 Other studies from Asia have also reported similar concordance (ranging from 80% to 90%).26,27,31,32
In our study, we observed 5 discordant cases out of 32 (15.6%). Specifically, 4 patients were positive for mutations in tissue but negative in plasma (false negatives), while 1 was positive in plasma but negative in tissue. The biological basis for false negatives in plasma is often related to the shedding of tumor DNA into the bloodstream.16,17 We observed that concordance and sensitivity substantially increased in stage IV patients (91.3% and 90.9%, respectively) compared to the overall group. This is likely because, in metastatic disease, the ctDNA content in the plasma becomes significantly higher 17 than in earlier stages, where a lower tumor burden may result in ctDNA levels falling below the LOD of our ARMS-PCR kit. Furthermore, the short half-life of ctDNA 18 and its rapid clearance by the liver and kidneys 33 mean that low-shedding tumors may not consistently provide enough genetic material for detection by ARMS-PCR.
These limitations highlight the importance of orthogonal technologies to resolve discordant cases in clinical practice. Our study used the AmoyDx ARMS-PCR kit, which targets 29 hotspot mutations but cannot detect rare variants outside its panel. Broader methods such as NGS and droplet digital PCR (ddPCR) can identify rare mutations and lower allele frequencies missed by PCR assays. Therefore, in cases where plasma is negative but clinical suspicion remains high, comprehensive NGS panels or ddPCR should be considered to resolve discrepancies and ensure accurate mutation profiling. However, these advanced methods remain costly and largely unavailable in Bangladesh. In low-resource settings, practical solutions include optimizing existing PCR assays, ensuring strict plasma handling, and considering repeat testing when clinical suspicion is high.
Conclusion
We observed substantial agreement, which increased to almost perfect agreement in patients with stage IV disease. This high level of agreement, along with high concordance, sensitivity, and specificity, strongly indicates that EGFR mutations can be reliably identified using ctDNA. These promising findings highlight the effectiveness of plasma-derived ctDNA as an alternative to tissue biopsy for detecting EGFR mutations in NSCLC patients in Bangladesh, especially when tissue biopsy is not feasible due to financial constraints or advanced disease stages, or when the tissue sample is inadequate for analysis.
Limitation
This study is subject to several limitations. First, the sample size was relatively small, and the predominance of male participants resulted in a non-homogeneous study population. Second, although broader biomarkers are clinically relevant, our analysis was restricted to EGFR given its high prevalence in Asian populations. Third, the use of singleplex ARMS-PCR confined mutation detection to 29 predefined hotspots, thereby excluding rare variants that could have been identified through sequencing-based approaches. Finally, tissue and plasma samples were processed at different centers, which may have introduced operator-related variability.
Recommendation
It is recommended that EGFR mutations be detected in tissue biopsies and ctDNA samples in the same center, possibly by the same operator, to minimize operator-related variability in the results. Future research should include a larger and more balanced number of male and female patients to avoid any gender bias. Additionally, we recommend integrating broader mutation panels and utilizing orthogonal technologies like NGS or ddPCR to resolve discordant cases.
Footnotes
ORCID iDs
Ethical considerations
Ethical approval for this study was granted by the Institutional Review Board of Bangabandhu Sheikh Mujib Medical University (Registration No. 5112, Date 13.07.2024). All procedures were conducted in accordance with the ethical standards outlined in the Declaration of Helsinki and relevant national guidelines. Patient data were anonymized prior to analysis to ensure confidentiality.
Consent to participate
Written informed consent was obtained from all study participants using a preformatted consent form, following a detailed explanation of the study's purpose and procedures.
Author contributions
Study conception and design: Md Kabirul Islam Soroar and Mohammad Masum Alam
Data acquisition and laboratory work: Md Kabirul Islam Soroar, Sharmistha Roy, and Hosne Ara
Statistical analysis and data interpretation: Md Kabirul Islam Soroar, Sharmistha Roy, Sharif Al Nur, Riyadh Arifin Akanda, and Mohammad Masum Alam
Study supervision: Mohammad Masum Alam
Manuscript drafting, critical revision for important intellectual content, and final approval of the version to be published: Md Kabirul Islam Soroar, Sharmistha Roy, Hosne Ara, Sharif Al Nur, Riyadh Arifin Akanda, and Mohammad Masum Alam
All authors have read and approved the final manuscript and agree to be accountable for all aspects of the work, ensuring its accuracy and integrity.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability
The datasets analyzed during the current study are available from the corresponding author upon reasonable request.
