Abstract
Signal detection and Bayesian inferential tools were applied to salivary biomarkers to improve screening accuracy and efficiency in detecting oral squamous cell carcinoma (OSCC). Potential cancer biomarkers are identified by significant differences in assay concentrations, receiver operating characteristic areas under the curve (AUCs), sensitivity, and specificity. However, the end goal is to report to individual patients their risk of having disease given positive or negative test results. Likelihood ratios (LRs) and Bayes factors (BFs) estimate evidential support and compile biomarker information to optimize screening accuracy. In total, 26 of 77 biomarkers were mentioned as having been tested at least twice in 137 studies and published in 16 summary papers through 2014. Studies represented 10 212 OSCC and 25 645 healthy patients. The measure of biomarker and panel information value was number of biomarkers needed to approximate 100% positive predictive value (PPV). As few as 5 biomarkers could achieve nearly 100% PPV for a disease prevalence of 0.2% when biomarkers were ordered from highest to lowest LR. When sequentially interpreting biomarker tests, high specificity was more important than test sensitivity in achieving rapid convergence toward a high PPV. Biomarkers ranked from highest to lowest LR were more informative and easier to interpret than AUC or Youden index. The proposed method should be applied to more recently published biomarker data to test its screening value.
Introduction
Purpose
The purpose of this study was to develop methods for improving salivary biomarker efficiencies in oral cancer screening with inferential tools adapted from clinical epidemiology and Bayesian analysis.
Problem
Even as screening and diagnostic technologies drop in price, methodological efficiencies can improve access and hasten implementation of salivary disease screening. Reducing test complexity, cost, and interpretation of results while increasing test information and robustness is a useful goal of any medical screening.
However, the current methods of biomarker discovery might limit their value in detecting disease. Potential salivary biomarkers are first observed as different concentrations in the saliva of healthy patients versus that of patients with cancer. From these samples, some or all of 4 attributes are commonly reported: sensitivities, specificities, areas under the curve (AUCs), and statistical significance in biomarker concentration between healthy and cancer groups.
However, from a clinical or policy decision’s perspective, more information is needed than just these 4 statistics. First, sensitivity and specificity are test characteristics and not related to patient health status. Statistical significance is subject to misinterpretation and often fails to meet underlying assumptions. 1 Finally, AUC is not easily computed and interpreted for use in medical decisions. Area under the curve may represent biomarker accuracy in a population but does not offer an estimate of personal risk given positive test results. However, the positive predictive value (PPV) includes test accuracy characteristics, accounts for disease prevalence, and states personal patient risk as a probability of having a disease.
Oral cancer
Oral cancer is a significant disease worldwide with up to 400 000 new cases and almost 130 000 deaths annually. 2 About 90% of oral cancers are oral squamous cell carcinoma (OSCC), and 80% occur in Southeast Asian countries. In the United States, the 5-year OSCC survival rate is 60%, but higher incidence and lower survival rates have been reported in some South Asian countries.3–5 Most OSCC lesions are small, asymptomatic, and easily overlooked or misjudged making early detection a challenge. Risk factors include alcohol and tobacco consumption, use of betel quid chew, and alcohol-based mouthwashes. 6 More than 95% of those with OSCC smoke tobacco, drink alcohol, or both. In recent decades, more cases of OSCC are emerging without known attributable risk factors. 7
Early detection of OSCC would improve survival, mortality, and morbidity. The gold standard for diagnosis is a biopsy of the suspicious lesion, but this test is not convenient for screening purposes because of its invasive nature, high cost, and need for specially trained medical personnel and equipment. However, saliva sampling to assess disease biomarkers is simple and less expensive, and multiple samples can be obtained for disease monitoring. Research suggests that salivary biomarkers have high discriminatory power for multiple diseases and cancers,8–12 and they have been well characterized and validated by comparisons of biomarker levels in healthy and cancerous oral epithelia. Development from normal to OSCC cells leads to altered expression of proteins and messenger RNA markers in saliva. As such, transcriptomic and proteomic approaches to biomarker development have had the most success in clinical testing. 13
The most informative screening tests are both highly sensitive and specific. 14 High Sensitivity with a Negative finding rules out disease, the SNout mnemonic. A highly Specific test with a Positive will rule-in disease, the SPin mnemonic. It may seem intuitive that more tests improve screening accuracy, but they do not, unless read as conditional probabilities where each biomarker test is read in context with others.
Independence of biomarker probabilities can only be incorrect because they are derived from the same biological system (person). For instance, 2 truly independent and positive cancer biomarkers (designated T+) in just 3% and 4% in healthy people would co-occur at just 0.12% (3% × 4%) in healthy individuals. The simultaneous occurrence of such rare biomarkers is prima facie support of a nonhealthy state, but such evidence neglects disease prevalence and does not indicate the personal risk of disease for the screened individual with both biomarkers.
Likelihood ratios reveal biomarker information values
The likelihood ratio (LR) describes a medical test’s information value. A biomarker’s ratio of true hit rate to false positive rate, is a positive LR. LR is used in signal detection research,14,15 with analogous application to cancer detection. The LR is an index of how well a biomarker can separate the signal from noise.
Each biomarker offers information about the presence or absence of a disease state with varying degrees of accuracy. This task is similar to that of psychological testing, where each test item reflects the extent of an underlying construct, generally called θ. With cancer as θ, biomarker LRs can improve screening accuracy with fewer biomarker items, i.e., improve screening efficiency.
Bayes factors as evidential support
Bayes factors (BF) estimate a biomarker’s strength of evidence or evidential support (ES).15–17 Mathematically, the BF is the ratio of probability of data occurring with and without a specific biomarker. Evidential support, the natural logarithm of the BF, expresses the contribution to screening information provided by the biomarker. 15
Correcting for inflated personal risk, PPV
Figure 1 illustrates the effects of prevalence on PPV under 6 hypothetical biomarkers, each of a different fixed sensitivity and specificity. The biomarker on the red line has 90% sensitivity and 90% specificity. Other lines in Figure 1 represent different combinations of sensitivity and specificity. In all cases, PPV increases with prevalence. For prevalence under 20% (an exaggerated maximum for disease prevalence), the 6 combinations of sensitivities and specificities show wide discrepancies in PPV. Therefore, all retrospective case-control studies in which 50% of cases have cancer and the other 50% are without cancer (and presumably healthy) inflate PPV estimates. In terms of sensitivity and specificity, some degree of each are required to improve up on prevalence as the a priori best estimate of personal cancer status.
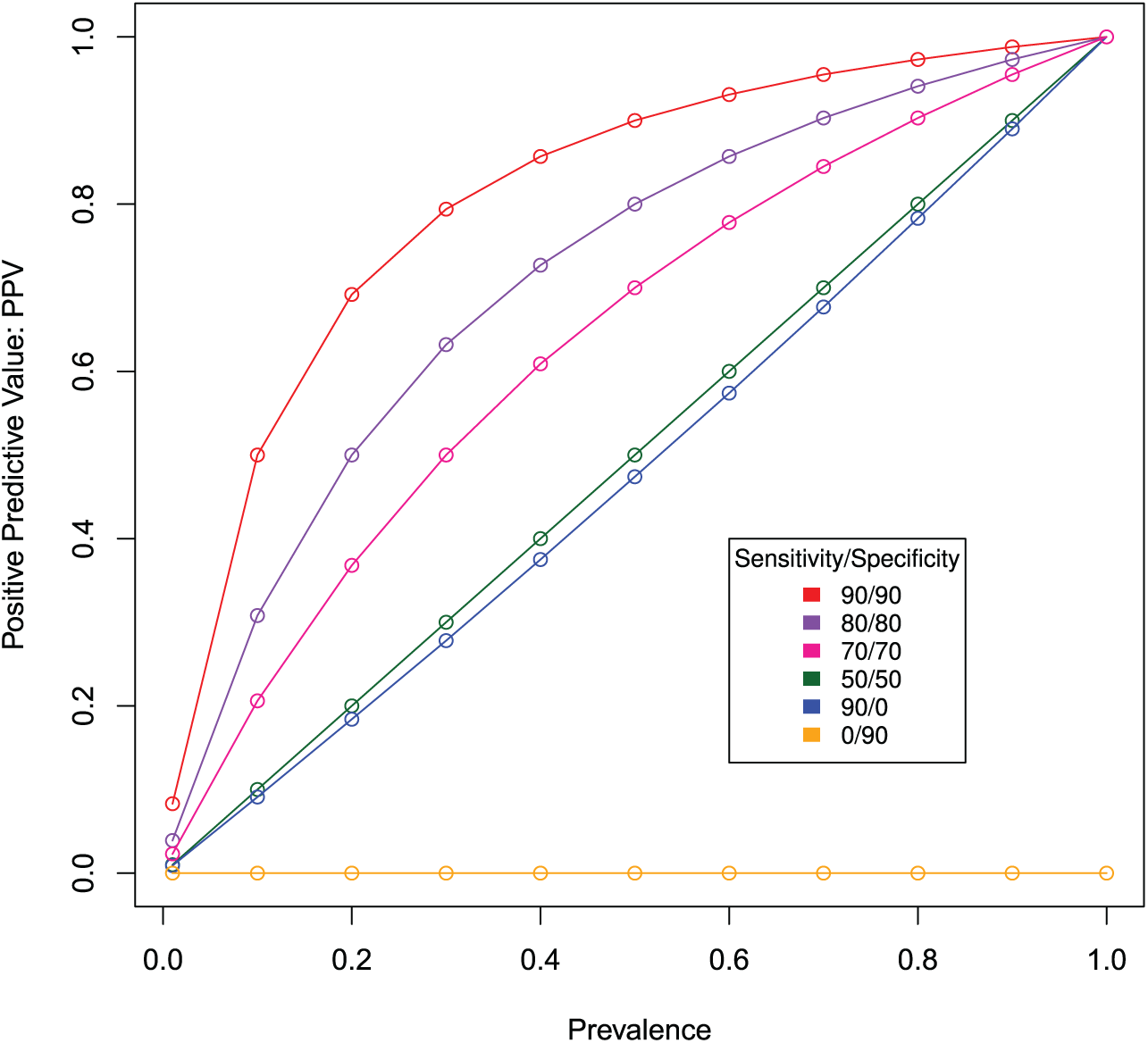
The effects of prevalence on positive predictive value (PPV): simulations comparing 6 sensitivity and specificity combinations.
Materials and Methods
Studies chosen
The authors selected 137 analyses with keywords—biomarkers, oral cancer, oral squamous cell carcinoma, sensitivity, and specificity—in 16 papers from a total of 80 reviewed.3,4,7,11–13,16–86 All studies were chosen from searches in PubMed and Web of Knowledge databases in 2014. Accepted papers reported at least 2 separate estimates of sensitivity, specificity, AUC, or statistical significances between diseased and healthy groups for each biomarker. Two biomarker parameter estimates allow some degree of reliability for analysis. Twenty-six biomarkers met the criteria.
The final 26 selected biomarkers and associated panels are summarized in Appendix 1. Outcomes were transferred to Meta-DiSc and R language for computation of LRs. Biomarkers selected were: absent in melanoma 1 (AIM1), calcitonin-related polypeptide alpha (CALCA), cyclin-A1 (CCNA1), cyclin-D2 (CCND2), cadherin-1 (CDH1), death-associated protein (DAPK), deleted in colorectal carcinoma (DCC), erythrocyte sedimentation rate (ESR), hypermethylated in cancer 1 (HIC1), homeobox protein Hox-A9 (HOXA9), interleukin 1B (IL1B), interleukin 8 (IL8), O-6-methylguanine-DNA methyltransferase (MGMT), munc-18-interacting 1 (MINT1), munc-18-interacting 31 (MINT3), miR31, OAX1, p16, protein gene product 9.5 (PGP9.5), retinoic acid receptor beta (RARB), Ras association domain-containing protein 1 (RASSF1A), RNA-binding motif protein 6 (RBM6), retinoblastoma-interacting zinc-finger protein 1 (RIZ1), S100A2, sulfate adenylyltransferase (SAT), transforming growth factor beta receptor 2 (TBFBR2), and metalloproteinase inhibitor 3 (TIMP3).
Evidential support and likelihood ratio calculations
Probabilities of data under the biomarker absent (or “null”) condition over the biomarker present probability of data were obtained via an hierarchical data analytic approach described elsewhere.89,90 The ratios are Bayes Factors; the logarithms of these ratios express the ES’s. In a sense, the BF is a “between” condition likelihood ratio, addressing how much more information is provided by a biomarker’s presence versus its absence.
The complement is a “within” index, the LR describes biomarker accuracy or efficiency. LR is the ratio of true positives to false positives. LR times prevalence in odds form yields a PPV in odds form. LR was then adjusted for sample prevalence in case-control studies by dividing by prevalence to obtain an unbiased LR. Biomarker diagnostic accuracy was visualized using the “mada” package in R as shown in Figure 2.91-93
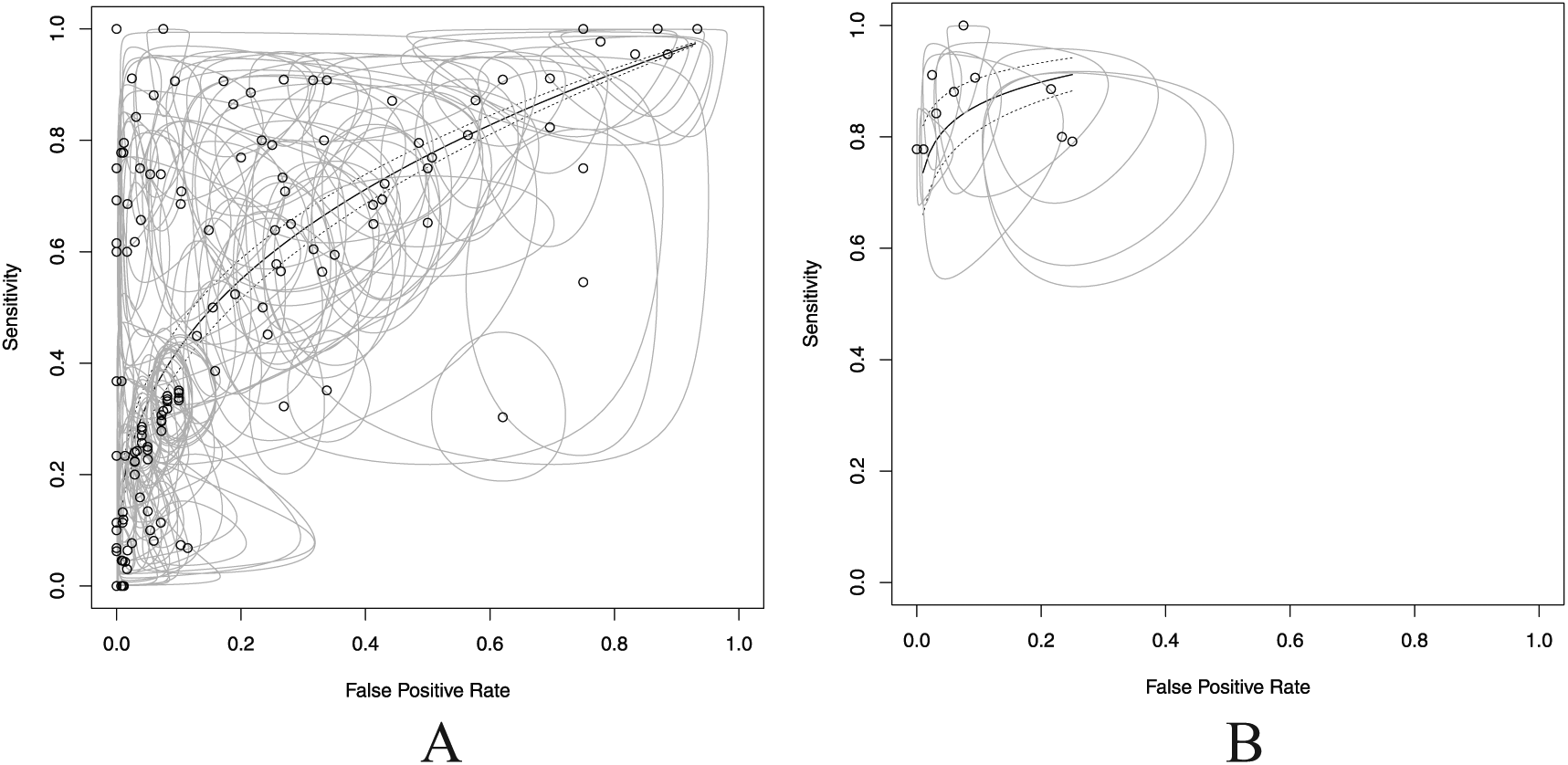
Biomarkers equal better test efficiencies with fewer versus more biomarkers—illustration that fewer biomarkers can have a higher positive predictive value and decreased error variance. Fewer number of biomarkers on the right (B) have a higher discriminating ability measured in area under the curve, AUC = 0.818, than the more complex “scattershot” approach on the left (A), where the AUC = 0.760.
Results
Table 1 presents the 26 selected biomarkers from the 16 accepted studies listed individually and in panels with associated sensitivities (Se), specificities (Sp), AUCs, or statistical significance in biomarker amount between healthy individuals and patients with OSCC. The 5 most sensitive biomarkers were CALCA (Se = 1.00), RBM6 (0.97), S100A2 (0.96), HOXA9 (0.81), and IL8 (0.74). The 5 most specific biomarkers were RASSF1A (Sp = 0.993), AIM1 (0.991), ESR (0.986), p16 (0.950), and DCC (also 0.950). In Figure 2 (scatterplot), all OSCC biomarkers and panels reviewed in the current evidence synthesis are plotted as AUCs for detecting OSCC. The scatterplot in Figure 2A, based on 137 data points of 26 biomarkers, has an AUC of 0.760. The 9 most informative LR biomarkers in Figure 2B have an AUC of 0.818.
Summary accuracy statistics for 26 oral salivary biomarkers.
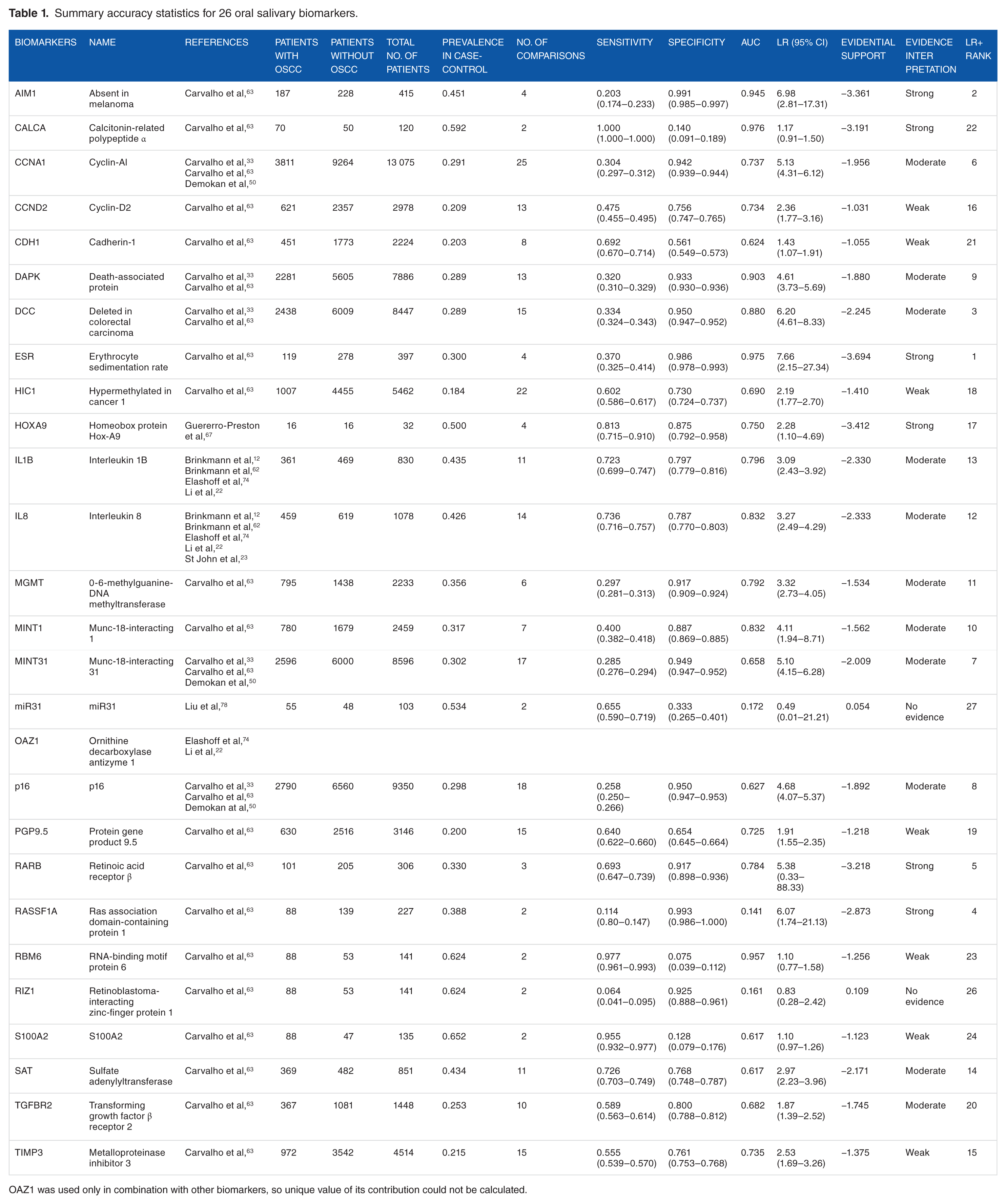
OAZ1 was used only in combination with other biomarkers, so unique value of its contribution could not be calculated.
Information value (LR) and PPV
The 5 highest LRs from reviewed studies were ESR (LR = 7.66), AIM1 (6.98), DCC (6.20), RASSFIA (6.07), and RARB (5.38). When reading biomarkers conditionally from lowest to highest LR that have prevalence levels of 2%, 0.2%, 0.02%, 0.002%, 0.0002%, the number of biomarkers needed to achieve a PPV 100% were 11, 14, 15, 16, 18, and 20, respectively (see Figure 3). In Figure 3, the dotted line shows the number of biomarkers plotted against the PPV on the y axis using conditional probabilities, with green area representing 95% confidence bands needed to screen for OSCC. The blue area and long dashed line represent the Youden index plot and the pink area and solid line represent the AUC analysis. Both the Youden and AUC plots are the number of biomarkers required to approach a PPV of 100%.
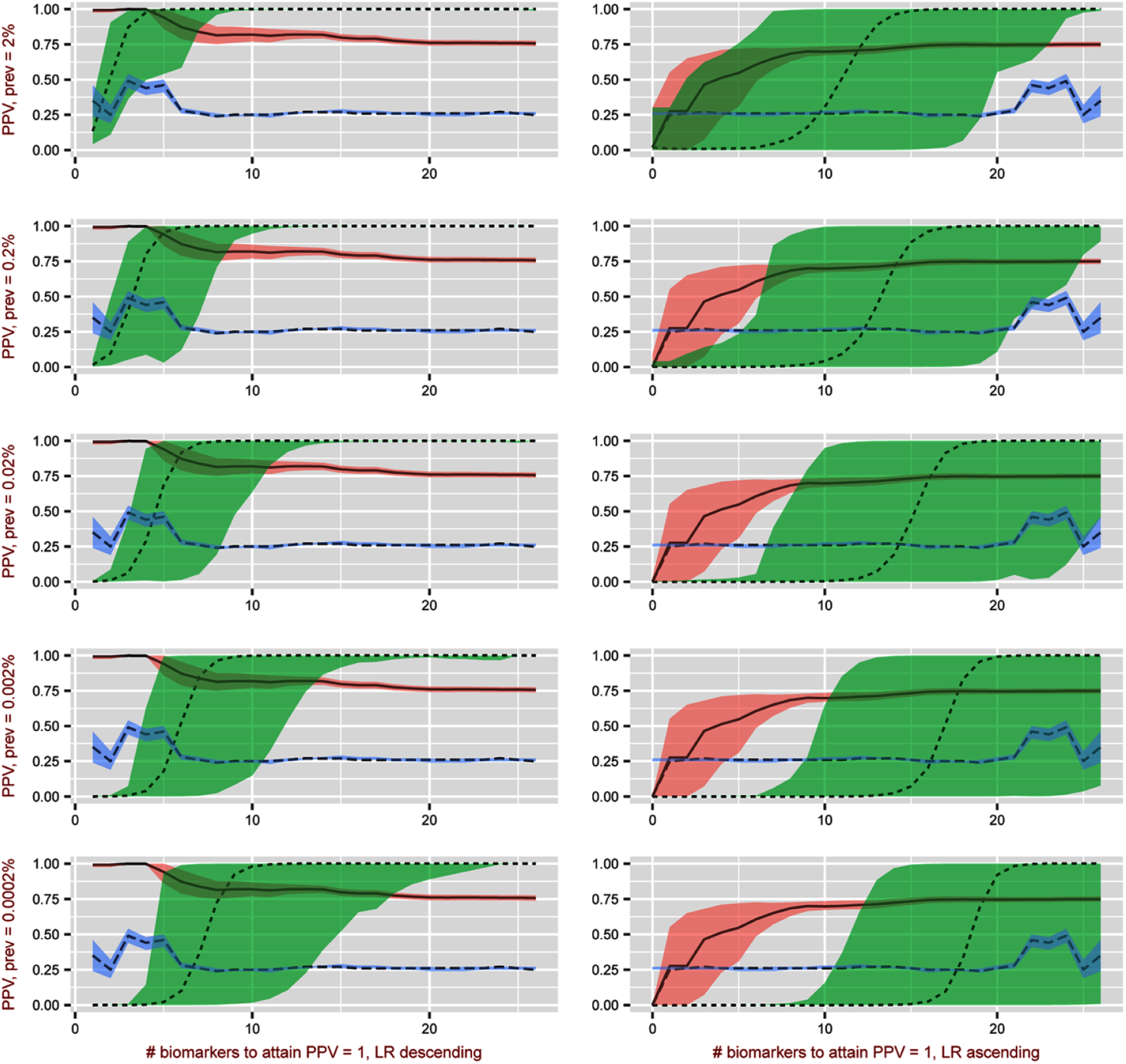
Number of biomarkers needed to approximate a positive predictive value of 1.00. Rows represent example prevalences with 2% at the top, decreased by a factor of 0.1 to 0.0002% prevalence at the bottom row.
By ordering from highest to lowest LR, the required number of biomarkers required to approach a PPV 100% for prevalence levels of 2%, 0.2%, 0.02%, 0.002%, and 0.0002% were 5, 8, 9, 11 and 12, respectively. Area under the curve appears to be more efficient than the LR conditional approach because it nears a PPV of 100% by starting with a high LR biomarker. For all sample prevalence conditions, 8 fewer biomarkers were needed to achieve the same OSCC screening accuracy. When the 4 least informative LR biomarkers are omitted from analysis, the number of biomarkers needed to achieve a PPV near 1.00 drops further to 3, 5, 7, 8, and 10 for each prevalence level.
The AUC and Youden index do not improve PPV by adding biomarkers. The Youden index is asymptotic at PPV = 25%, representing uncertainty for 75% of patients screened. Area under the curve was similar by increasing uncertainty: maximum screening accuracy achieved as PPV was 75%, leaving 25% error or uncertainty.
ES results
Six biomarkers showed strong ES: ESR (−3.694), AIM1 (−3.361), CALCA (−3.191), HOXA9 (−3.412), RARB (−3.218), and RASSF1A (−2.873). Another 11 showed moderate evidence: IL8 (−2.333), IL1B (−2.330), DCC (−2.245), SAT (−2.171), MINT31 (−2.009), CCNA1 (−1.956), p16 (−1.892), DAPK (−1.880), TGFBR2 (−1.745), MINT1 (−1.562), and MGMT (−1.534). Seven biomarkers only were supported by only weak evidence: HIC1 (−1.410), TIMP3 (−1.375), RBM6 (−1.256), PGP9.5 (−1.218), S100A2 (−1.123), CDH1 (−1.055), and CCND2 (−1.031). The 2 biomarkers lacking evidential support were miR31 and RIZ1.
Plots in Figure 4 show the individual biomarkers and their associated confidence intervals by sensitivity and specificity with color coding for ES. Plot points were color coded according to ES: green = strong, gold = moderate, red = weak, and black = lacks evidence. The blue dots near the bottom right represent biomarkers with sensitivity over 0.75 and specificity over 0.75. The pink dots at lower right plots biomarkers with sensitivities over 0.75 or specificities over 0.75.
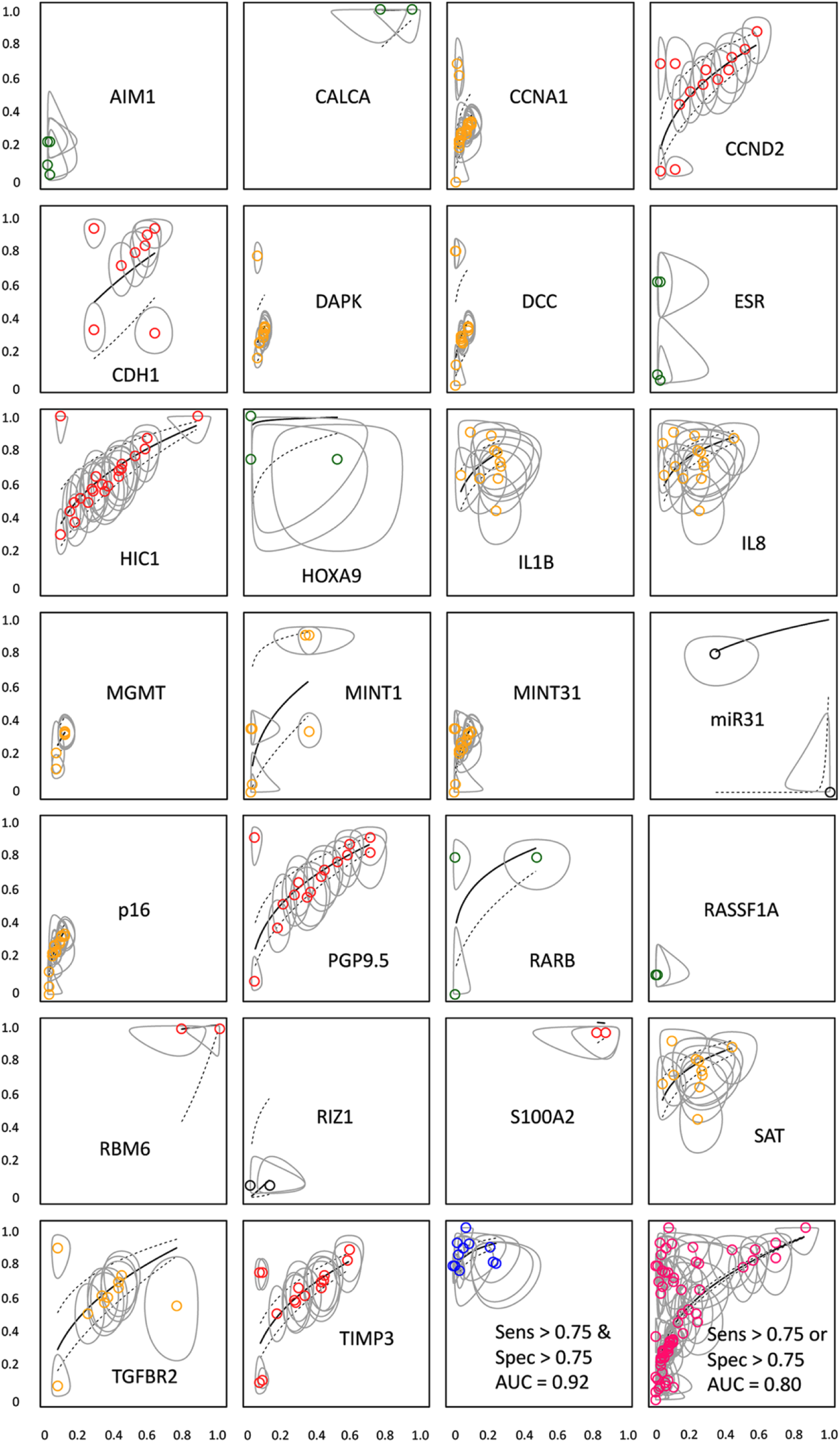
Individual biomarker AUC plots coded by color for evidential support: green = strong evidence, gold = moderate, red = weak, black = lack of evidence Panels in bottom right illustrate 0.75 sensitivity and 0.75 specificity biomarkers (blue) and 0.75 sensitivity or 0.75 specificity biomarkers (pink) with their respective AUC’s.
Based on the summary variable estimates of n = 26, ES correlated moderately with LR, (Pearson r = .66) and PPV (r = .67). Area under the curve correlated with ES at (r = .52), LR (r = .29), and PPV (r = .42). Prevalence is not associated with ES but mildly with LR (−0.35).
Discussion
Detecting cancer early has life - and cost-saving advantages over treating advanced stages. Candidate cancer biomarkers are identified by differences in biomarker concentrations between healthy individuals and patients with OSCC. Four commonly derived indices of biomarkers—sensitivity, specificity, statistical significance, and AUC—do not yield a personal risk of disease—a key feature in the future of personalized medicine. To refine screening results to derive personal probability of disease, PPVs are the metric of choice.
Retrospective case-control methods artificially inflate prevalence and thus make it easier to overestimate efficiency in biomarker disease detection by inflating PPV estimates. LRs should be adjusted for exagerated high prevalences before estimating biomarker accuracy at lower disease prevalences.
In the final step after adjusting for prevalence, PPVs of biomarkers can be combined with other biomarkers in panels to be more informative with fewer biomarkers. Passing along positive test results from one biomarker to the next refines and augments information as to true disease status of a patient.
In all, 24 of 26 studied biomarkers demonstrated at least weak evidence for OSCC detection. The other 2 biomarkers indicated that their inclusion reduced PPV, ie, increased screening uncertainty. When weak and low information value biomarkers are eliminated the number required to screen will decrease, improving efficiency and affordability.
Area under the curve was an inferior metric as compared to LR in for achieving an acceptable PPV with fewer biomarkers. In Figure 3, when using the highest LR biomarkers first, AUC accuracy tops Bayesian accumulation and Youden index. Few biomarker’s high performance measured by AUC is expected, since high AUC means high sensitivity and specificity. Such ideal biomarkers may act as initial screens for disease, but may not identify the disease for which the screening is intended. This finding illustrates Kraemer’s assertion that more tests about a condition does not improve diagnostic and screening accuracy. 14
Value of biomarker information
Biomarker LR is a convenient expression of information value that can be manipulated mathematically to optimize biomarker usage. The number of biomarkers required to screen for a disease is a function of LR along with an estimated disease prevalence. When testing for life-threatening disease, the risk preference is to accept more false positives at the expense of false negatives.
The probability of having the disease given positive test results, PPV, is the ultimate objective of good medical tests. Biomarkers with both the highest sensitivity and specificity are the most accurate ones. 14 Biomarkers are more informative when included in panels including other biomarkers with high LR that incrementally increase true positives and eliminate true negatives from the screening pool. 16 High biomarker specificity appeared as more efficient than high sensitivity for screening.
Both AUC and Youden’s index are hard to interpret and do not improve accuracy with additional biomarkers (Figure 3).
When screening a population where disease prevalence is unknown, prevalence might be estimated from groups with similar genetic predispositions, behavioral and dietary risks, cultural risks such as chewing betel nut, tobacco use, alcohol consumption, and other health habits and beliefs.
Interestingly, the most informative biomarkers (ones with the highest LR) turned out to have the highest specificities. Mathematically, this can be explained as the denominator of the LR—the proportion of false positives, or 1–specificity. Large specificities reduce the denominator and increase the overall LR.
The method of ES, where diagnostic accuracy with a biomarker is compared to accuracy without it, appears to ignore some promising biomarkers shown in Figure 4. CCND2, HIC1, PGP9.5, TGFBR2, and TIMP3 have more consistent grouping of data points and therefore are more reliable estimates of their AUC. However, since none of their regression lines pull towards the upper left-hand corner of high accuracy, such biomarkers are reliable but not accurate. They may be used alone or in combinations with others to achieve the best accuracy per unit costs of deployment, preservation, and transportation.
Limitations of method
The estimated LR of biomarkers in panels was probably diluted by other biomarkers also in the panel. If so, then we may expect biomarker accuracy to be lower than that estimated for single biomarkers in isolation. Nonetheless, biomarkers may interact synergistically in their ability to detect disease in which cases the simultaneous presence of such biomarkers has a higher LR than that expected from each individual one.
Where physicians, patients, and public health officials have budgetary constraints for biomarkers in field settings, the number of biomarkers required for screening may be reduced by obtaining a more focused at-risk population by first screening for behaviors and family history. The risk of making errors as false negatives versus false positives must always be balanced.
Data used herein were limited to studies published through 2014. Data collection ended in 2014 so that the method could be developed without introducing new information. Information published since then may update findings and be useful to test the method.
Some biomarkers tested are conceptually too general and will not serve to identify a specific disease. The ESR is one of those biomarkers. Elevated ESR might serve as a first-level general biomarker screen to be passed along to other biomarkers for more information. Elevated ESR indicates many diseases, not just cancer. Thus, ESR’s strong ES and high sensitivity may make it ideal for ruling out cancer with a negative ESR result.
Finally, assays that came from the same authors or same labs must be considered as sources of bias. One paper 63 reported all the CCND2 biomarker results, and perhaps a single laboratory was responsible for all CCND2 analyses. If so, a bias must be ruled out in future studies.
Conclusions
The same diagnostic and screening information may be obtained from fewer but more informative biomarkers rather than many weak to moderately informative ones.
By converting summary salivary biomarker data to positive likelihood ratios and then reading them as conditional probabilities ordered from highest to lowest LR, higher efficiency of OSCC screening may be obtained with fewer biomarkers. Sensitivity, specificity, and AUC do not allow the aggregation and ordering needed to improve screening accuracy.
The future and promise of personalized medicine is exactly the problem tackled here—how to translate multiple indicators of population-level disease and personal characteristics into a particular patient’s probability of disease? Individual patient test results can then inform treatment and prevention strategies in the context of culture, behaviors, genetic predispositions, and economic and caretaker burdens.
Footnotes
Appendix
All biomarker panels culled from 16 evidence summary papers.
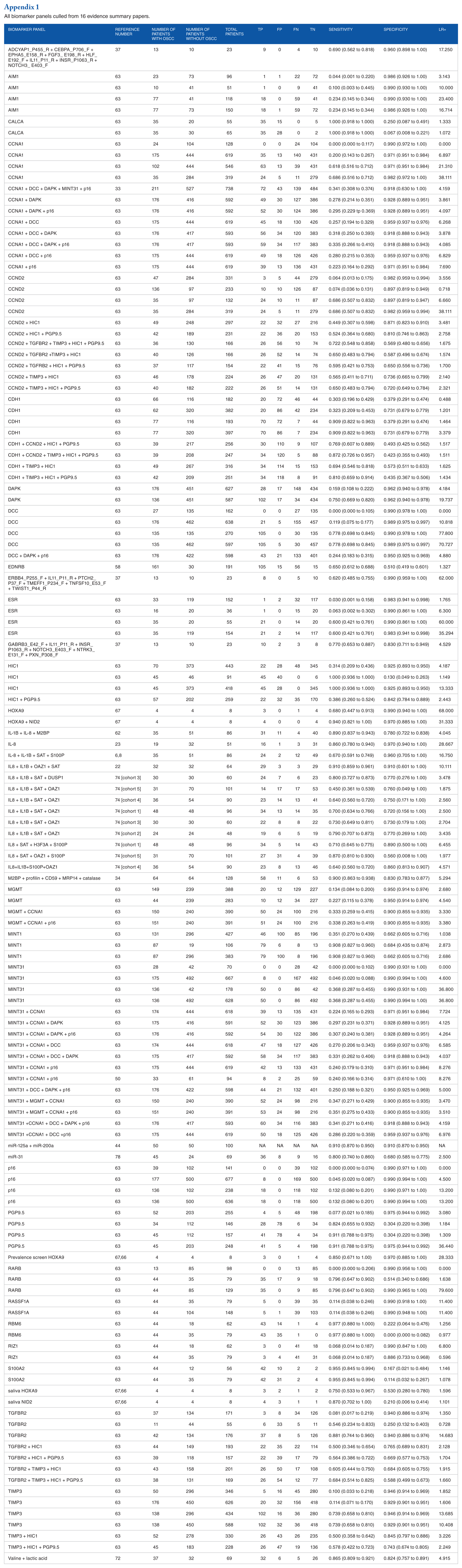
| Biomarker panel | Reference Number | Number of patients with OSCC | Number of patients without OSCC | Total patients | TP | FP | FN | TN | Sensitivity | Specificity | LR+ |
|---|---|---|---|---|---|---|---|---|---|---|---|
| ADCYAP1_P455_R + CEBPA_P706_F + EPHA5_E158_R + FGF3_ E198_R + HLF_E192_F + IL11_P11_R + INSR_P1063_R + NOTCH3_ E403_F | 37 | 13 | 10 | 23 | 9 | 0 | 4 | 10 | 0.690 (0.562 to 0.818) | 0.960 (0.898 to 1.00) | 17.250 |
| AIM1 | 63 | 23 | 73 | 96 | 1 | 1 | 22 | 72 | 0.044 (0.001 to 0.220) | 0.986 (0.926 to 1.00) | 3.143 |
| AIM1 | 63 | 10 | 41 | 51 | 1 | 0 | 9 | 41 | 0.100 (0.003 to 0.445) | 0.990 (0.930 to 1.00) | 10.000 |
| AIM1 | 63 | 77 | 41 | 118 | 18 | 0 | 59 | 41 | 0.234 (0.145 to 0.344) | 0.990 (0.930 to 1.00) | 23.400 |
| AIM1 | 63 | 77 | 73 | 150 | 18 | 1 | 59 | 72 | 0.234 (0.145 to 0.344) | 0.986 (0.926 to 1.00) | 16.714 |
| CALCA | 63 | 35 | 20 | 55 | 35 | 15 | 0 | 5 | 1.000 (0.918 to 1.000) | 0.250 (0.087 to 0.491) | 1.333 |
| CALCA | 63 | 35 | 30 | 65 | 35 | 28 | 0 | 2 | 1.000 (0.918 to 1.000) | 0.067 (0.008 to 0.221) | 1.072 |
| CCNA1 | 63 | 24 | 104 | 128 | 0 | 0 | 24 | 104 | 0.000 (0.000 to 0.117) | 0.990 (0.972 to 1.00) | 0.000 |
| CCNA1 | 63 | 175 | 444 | 619 | 35 | 13 | 140 | 431 | 0.200 (0.143 to 0.267) | 0.971 (0.951 to 0.984) | 6.897 |
| CCNA1 | 63 | 102 | 444 | 546 | 63 | 13 | 39 | 431 | 0.618 (0.516 to 0.712) | 0.971 (0.951 to 0.984) | 21.310 |
| CCNA1 | 63 | 35 | 284 | 319 | 24 | 5 | 11 | 279 | 0.686 (0.516 to 0.712) | 0.982 (0.972 to 1.00) | 38.111 |
| CCNA1 + DCC + DAPK + MINT31 + p16 | 33 | 211 | 527 | 738 | 72 | 43 | 139 | 484 | 0.341 (0.308 to 0.374) | 0.918 (0.630 to 1.00) | 4.159 |
| CCNA1 + DAPK | 63 | 176 | 416 | 592 | 49 | 30 | 127 | 386 | 0.278 (0.214 to 0.351) | 0.928 (0.889 to 0.951) | 3.861 |
| CCNA1 + DAPK + p16 | 63 | 176 | 416 | 592 | 52 | 30 | 124 | 386 | 0.295 (0.229 tp 0.369) | 0.928 (0.889 to 0.951) | 4.097 |
| CCNA1 + DCC | 63 | 175 | 444 | 619 | 45 | 18 | 130 | 426 | 0.257 (0.194 to 0.329) | 0.959 (0.937 to 0.976) | 6.268 |
| CCNA1 + DCC + DAPK | 63 | 176 | 417 | 593 | 56 | 34 | 120 | 383 | 0.318 (0.250 to 0.393) | 0.918 (0.888 to 0.943) | 3.878 |
| CCNA1 + DCC + DAPK + p16 | 63 | 176 | 417 | 593 | 59 | 34 | 117 | 383 | 0.335 (0.266 to 0.410) | 0.918 (0.888 to 0.943) | 4.085 |
| CCNA1 + DCC + p16 | 63 | 175 | 444 | 619 | 49 | 18 | 126 | 426 | 0.280 (0.215 to 0.353) | 0.959 (0.937 to 0.976) | 6.829 |
| CCNA1 + p16 | 63 | 175 | 444 | 619 | 39 | 13 | 136 | 431 | 0.223 (0.164 to 0.292) | 0.971 (0.951 to 0.984) | 7.690 |
| CCND2 | 63 | 47 | 284 | 331 | 3 | 5 | 44 | 279 | 0.064 (0.013 to 0.175) | 0.982 (0.959 to 0.994) | 3.556 |
| CCND2 | 63 | 136 | 97 | 233 | 10 | 10 | 126 | 87 | 0.074 (0.036 to 0.131) | 0.897 (0.819 to 0.949) | 0.718 |
| CCND2 | 63 | 35 | 97 | 132 | 24 | 10 | 11 | 87 | 0.686 (0.507 to 0.832) | 0.897 (0.819 to 0.947) | 6.660 |
| CCND2 | 63 | 35 | 284 | 319 | 24 | 5 | 11 | 279 | 0.686 (0.507 to 0.832) | 0.982 (0.959 to 0.994) | 38.111 |
| CCND2 + HIC1 | 63 | 49 | 248 | 297 | 22 | 32 | 27 | 216 | 0.449 (0.307 to 0.598) | 0.871 (0.823 to 0.910) | 3.481 |
| CCND2 + HIC1 + PGP9.5 | 63 | 42 | 189 | 231 | 22 | 36 | 20 | 153 | 0.524 (0.364 to 0.680) | 0.810 (0.746 to 0.863) | 2.758 |
| CCND2 + TGFBR2 + TIMP3 + HIC1 + PGP9.5 | 63 | 36 | 130 | 166 | 26 | 56 | 10 | 74 | 0.722 (0.548 to 0.858) | 0.569 (0.480 to 0.656) | 1.675 |
| CCND2 + TGFBR2 +TIMP3 + HIC1 | 63 | 40 | 126 | 166 | 26 | 52 | 14 | 74 | 0.650 (0.483 to 0.794) | 0.587 (0.496 to 0.674) | 1.574 |
| CCND2 + TGFRB2 + HIC1 + PGP9.5 | 63 | 37 | 117 | 154 | 22 | 41 | 15 | 76 | 0.595 (0.421 to 0.753) | 0.650 (0.556 to 0.736) | 1.700 |
| CCND2 + TIMP3 + HIC1 | 63 | 46 | 178 | 224 | 26 | 47 | 20 | 131 | 0.565 (0.411 to 0.711) | 0.736 (0.665 to 0.799) | 2.140 |
| CCND2 + TIMP3 + HIC1 + PGP9.5 | 63 | 40 | 182 | 222 | 26 | 51 | 14 | 131 | 0.650 (0.483 to 0.794) | 0.720 (0.649 to 0.784) | 2.321 |
| CDH1 | 63 | 66 | 116 | 182 | 20 | 72 | 46 | 44 | 0.303 (0.196 to 0.429) | 0.379 (0.291 to 0.474) | 0.488 |
| CDH1 | 63 | 62 | 320 | 382 | 20 | 86 | 42 | 234 | 0.323 (0.209 to 0.453) | 0.731 (0.679 to 0.779) | 1.201 |
| CDH1 | 63 | 77 | 116 | 193 | 70 | 72 | 7 | 44 | 0.909 (0.822 to 0.963) | 0.379 (0.291 to 0.474) | 1.464 |
| CDH1 | 63 | 77 | 320 | 397 | 70 | 86 | 7 | 234 | 0.909 (0.822 to 0.963) | 0.731 (0.679 to 0.779) | 3.379 |
| CDH1 + CCND2 + HIC1 + PGP9.5 | 63 | 39 | 217 | 256 | 30 | 110 | 9 | 107 | 0.769 (0.607 to 0.889) | 0.493 (0.425 to 0.562) | 1.517 |
| CDH1 + CCND2 + TIMP3 + HIC1 + PGP9.5 | 63 | 39 | 208 | 247 | 34 | 120 | 5 | 88 | 0.872 (0.726 to 0.957) | 0.423 (0.355 to 0.493) | 1.511 |
| CDH1 + TIMP3 + HIC1 | 63 | 49 | 267 | 316 | 34 | 114 | 15 | 153 | 0.694 (0.546 to 0.818) | 0.573 (0.511 to 0.633) | 1.625 |
| CDH1 + TIMP3 + HIC1 + PGP9.5 | 63 | 42 | 209 | 251 | 34 | 118 | 8 | 91 | 0.810 (0.659 to 0.914) | 0.435 (0.367 to 0.506) | 1.434 |
| DAPK | 63 | 176 | 451 | 627 | 28 | 17 | 148 | 434 | 0.159 (0.108 to 0.222) | 0.962 (0.940 to 0.978) | 4.184 |
| DAPK | 63 | 136 | 451 | 587 | 102 | 17 | 34 | 434 | 0.750 (0.669 to 0.820) | 0.962 (0.940 to 0.978) | 19.737 |
| DCC | 63 | 27 | 135 | 162 | 0 | 0 | 27 | 135 | 0.000 (0.000 to 0.105) | 0.990 (0.978 to 1.00) | 0.000 |
| DCC | 63 | 176 | 462 | 638 | 21 | 5 | 155 | 457 | 0.119 (0.075 to 0.177) | 0.989 (0.975 to 0.997) | 10.818 |
| DCC | 63 | 135 | 135 | 270 | 105 | 0 | 30 | 135 | 0.778 (0.698 to 0.845) | 0.990 (0.978 to 1.00) | 77.800 |
| DCC | 63 | 135 | 462 | 597 | 105 | 5 | 30 | 457 | 0.778 (0.698 to 0.845) | 0.989 (0.975 to 0.997) | 70.727 |
| DCC + DAPK + p16 | 63 | 176 | 422 | 598 | 43 | 21 | 133 | 401 | 0.244 (0.183 to 0.315) | 0.950 (0.925 to 0.969) | 4.880 |
| EDNRB | 58 | 161 | 30 | 191 | 105 | 15 | 56 | 15 | 0.650 (0.612 to 0.688) | 0.510 (0.419 to 0.601) | 1.327 |
| ERBB4_P255_F + IL11_P11_R + PTCH2_P37_F + TMEFF1_P234_F + TNFSF10_E53_F + TWIST1_P44_R | 37 | 13 | 10 | 23 | 8 | 0 | 5 | 10 | 0.620 (0.485 to 0.755) | 0.990 (0.959 to 1.00) | 62.000 |
| ESR | 63 | 33 | 119 | 152 | 1 | 2 | 32 | 117 | 0.030 (0.001 to 0.158) | 0.983 (0.941 to 0.998) | 1.765 |
| ESR | 63 | 16 | 20 | 36 | 1 | 0 | 15 | 20 | 0.063 (0.002 to 0.302) | 0.990 (0.861 to 1.00) | 6.300 |
| ESR | 63 | 35 | 20 | 55 | 21 | 0 | 14 | 20 | 0.600 (0.421 to 0.761) | 0.990 (0.861 to 1.00) | 60.000 |
| ESR | 63 | 35 | 119 | 154 | 21 | 2 | 14 | 117 | 0.600 (0.421 to 0.761) | 0.983 (0.941 to 0.998) | 35.294 |
| GABRB3_E42_F + IL11_P11_R + INSR_P1063_R + NOTCH3_E403_F + NTRK3_E131_F + PXN_P308_F | 37 | 13 | 10 | 23 | 10 | 2 | 3 | 8 | 0.770 (0.653 to 0.887) | 0.830 (0.711 to 0.949) | 4.529 |
| HIC1 | 63 | 70 | 373 | 443 | 22 | 28 | 48 | 345 | 0.314 (0.209 to 0.436) | 0.925 (0.893 to 0.950) | 4.187 |
| HIC1 | 63 | 45 | 46 | 91 | 45 | 40 | 0 | 6 | 1.000 (0.936 to 1.000) | 0.130 (0.049 to 0.263) | 1.149 |
| HIC1 | 63 | 45 | 373 | 418 | 45 | 28 | 0 | 345 | 1.000 (0.936 to 1.000) | 0.925 (0.893 to 0.950) | 13.333 |
| HIC1 + PGP9.5 | 63 | 57 | 202 | 259 | 22 | 32 | 35 | 170 | 0.386 (0.260 to 0.524) | 0.842 (0.784 to 0.889) | 2.443 |
| HOXA9 | 67 | 4 | 4 | 8 | 3 | 0 | 1 | 4 | 0.680 (0.447 to 0.913) | 0.990 (0.940 to 1.00) | 68.000 |
| HOXA9 + NID2 | 67 | 4 | 4 | 8 | 4 | 0 | 0 | 4 | 0.940 (0.821 to 1.00) | 0.970 (0.885 to 1.00) | 31.333 |
| IL-1B + IL-8 + M2BP | 62 | 35 | 51 | 86 | 31 | 11 | 4 | 40 | 0.890 (0.837 to 0.943) | 0.780 (0.722 to 0.838) | 4.045 |
| IL-8 | 23 | 19 | 32 | 51 | 16 | 1 | 3 | 31 | 0.860 (0.780 to 0.940) | 0.970 (0.940 to 1.00) | 28.667 |
| IL-8 + IL-1B + SAT + S100P | 6,8 | 35 | 51 | 86 | 24 | 2 | 12 | 49 | 0.670 (0.591 to 0.749) | 0.960 (0.705 to 1.00) | 16.750 |
| IL8 + IL1B + OAZ1 + SAT | 22 | 32 | 32 | 64 | 29 | 3 | 3 | 29 | 0.910 (0.859 to 0.961) | 0.910 (0.601 to 1.00) | 10.111 |
| IL8 + IL1B + SAT + DUSP1 | 74 [cohort 3] | 30 | 30 | 60 | 24 | 7 | 6 | 23 | 0.800 (0.727 to 0.873) | 0.770 (0.276 to 1.00) | 3.478 |
| IL8 + IL1B + SAT + OAZ1 | 74 [cohort 5] | 31 | 70 | 101 | 14 | 17 | 17 | 53 | 0.450 (0.361 to 0.539) | 0.760 (0.049 tp 1.00) | 1.875 |
| IL8 + IL1B + SAT + OAZ1 | 74 [cohort 4] | 36 | 54 | 90 | 23 | 14 | 13 | 41 | 0.640 (0.560 to 0.720) | 0.750 (0.171 to 1.00) | 2.560 |
| IL8 + IL1B + SAT + OAZ1 | 74 [cohort 1] | 48 | 48 | 96 | 34 | 13 | 14 | 35 | 0.700 (0.634 to 0.766) | 0.720 (0.156 to 1.00) | 2.500 |
| IL8 + IL1B + SAT + OAZ1 | 74 [cohort 3] | 30 | 30 | 60 | 22 | 8 | 8 | 22 | 0.730 (0.649 to 0.811) | 0.730 (0.179 to 1.00) | 2.704 |
| IL8 + IL1B + SAT + OAZ1 | 74 [cohort 2] | 24 | 24 | 48 | 19 | 6 | 5 | 19 | 0.790 (0.707 to 0.873) | 0.770 (0.269 to 1.00) | 3.435 |
| IL8 + SAT + H3F3A + S100P | 74 [cohort 1] | 48 | 48 | 96 | 34 | 5 | 14 | 43 | 0.710 (0.645 to 0.775) | 0.890 (0.500 to 1.00) | 6.455 |
| IL8 + SAT + OAZ1 + S100P | 74 [cohort 5] | 31 | 70 | 101 | 27 | 31 | 4 | 39 | 0.870 (0.810 to 0.930) | 0.560 (0.008 to 1.00) | 1.977 |
| IL8+IL1B+S100P+OAZ1 | 74 [cohort 4] | 36 | 54 | 90 | 23 | 8 | 13 | 46 | 0.640 (0.560 to 0.720) | 0.860 (0.813 to 0.907) | 4.571 |
| M2BP + profilin + CD59 + MRP14 + catalase | 34 | 64 | 64 | 128 | 58 | 11 | 6 | 53 | 0.900 (0.863 to 0.938) | 0.830 (0.783 to 0.877) | 5.294 |
| MGMT | 63 | 149 | 239 | 388 | 20 | 12 | 129 | 227 | 0.134 (0.084 to 0.200) | 0.950 (0.914 to 0.974) | 2.680 |
| MGMT | 63 | 44 | 239 | 283 | 10 | 12 | 34 | 227 | 0.227 (0.115 to 0.378) | 0.950 (0.914 to 0.974) | 4.540 |
| MGMT + CCNA1 | 63 | 150 | 240 | 390 | 50 | 24 | 100 | 216 | 0.333 (0.259 to 0.415) | 0.900 (0.855 to 0.935) | 3.330 |
| MGMT + CCNA1 + p16 | 63 | 151 | 240 | 391 | 51 | 24 | 100 | 216 | 0.338 (0.263 to 0.419) | 0.900 (0.855 to 0.935) | 3.380 |
| MINT1 | 63 | 131 | 296 | 427 | 46 | 100 | 85 | 196 | 0.351 (0.270 to 0.439) | 0.662 (0.605 to 0.716) | 1.038 |
| MINT1 | 63 | 87 | 19 | 106 | 79 | 6 | 8 | 13 | 0.908 (0.827 to 0.960) | 0.684 (0.435 to 0.874) | 2.873 |
| MINT1 | 63 | 87 | 296 | 383 | 79 | 100 | 8 | 196 | 0.908 (0.827 to 0.960) | 0.662 (0.605 to 0.716) | 2.686 |
| MINT31 | 63 | 28 | 42 | 70 | 0 | 0 | 28 | 42 | 0.000 (0.000 to 0.102) | 0.990 (0.931 to 1.00) | 0.000 |
| MINT31 | 63 | 175 | 492 | 667 | 8 | 0 | 167 | 492 | 0.046 (0.020 to 0.088) | 0.990 (0.994 to 1.00) | 4.600 |
| MINT31 | 63 | 136 | 42 | 178 | 50 | 0 | 86 | 42 | 0.368 (0.287 to 0.455) | 0.990 (0.931 to 1.00) | 36.800 |
| MINT31 | 63 | 136 | 492 | 628 | 50 | 0 | 86 | 492 | 0.368 (0.287 to 0.455) | 0.990 (0.994 to 1.00) | 36.800 |
| MINT31 + CCNA1 | 63 | 174 | 444 | 618 | 39 | 13 | 135 | 431 | 0.224 (0.165 to 0.293) | 0.971 (0.951 to 0.984) | 7.724 |
| MINT31 + CCNA1 + DAPK | 63 | 175 | 416 | 591 | 52 | 30 | 123 | 386 | 0.297 (0.231 to 0.371) | 0.928 (0.889 to 0.951) | 4.125 |
| MINT31 + CCNA1 + DAPK + p16 | 63 | 176 | 416 | 592 | 54 | 30 | 122 | 386 | 0.307 (0.240 to 0.381) | 0.928 (0.889 to 0.951) | 4.264 |
| MINT31 + CCNA1 + DCC | 63 | 174 | 444 | 618 | 47 | 18 | 127 | 426 | 0.270 (0.206 to 0.343) | 0.959 (0.937 to 0.976) | 6.585 |
| MINT31 + CCNA1 + DCC + DAPK | 63 | 175 | 417 | 592 | 58 | 34 | 117 | 383 | 0.331 (0.262 to 0.406) | 0.918 (0.888 to 0.943) | 4.037 |
| MINT31 + CCNA1 + p16 | 63 | 175 | 444 | 619 | 42 | 13 | 133 | 431 | 0.240 (0.179 to 0.310) | 0.971 (0.951 to 0.984) | 8.276 |
| MINT31 + CCNA1 + p16 | 50 | 33 | 61 | 94 | 8 | 2 | 25 | 59 | 0.240 (0.166 to 0.314) | 0.971 (0.610 to 1.00) | 8.276 |
| MINT31 + DCC + DAPK + p16 | 63 | 176 | 422 | 598 | 44 | 21 | 132 | 401 | 0.250 (0.188 to 0.321) | 0.950 (0.925 to 0.969) | 5.000 |
| MINT31 + MGMT + CCNA1 | 63 | 150 | 240 | 390 | 52 | 24 | 98 | 216 | 0.347 (0.271 to 0.429) | 0.900 (0.855 to 0.935) | 3.470 |
| MINT31 + MGMT + CCNA1 + p16 | 63 | 151 | 240 | 391 | 53 | 24 | 98 | 216 | 0.351 (0.275 to 0.433) | 0.900 (0.855 to 0.935) | 3.510 |
| MINT31 +CCNA1 + DCC + DAPK + p16 | 63 | 176 | 417 | 593 | 60 | 34 | 116 | 383 | 0.341 (0.271 to 0.416) | 0.918 (0.888 to 0.943) | 4.159 |
| MINT31 +CCNA1 + DCC +p16 | 63 | 175 | 444 | 619 | 50 | 18 | 125 | 426 | 0.286 (0.220 to 0.359) | 0.959 (0.937 to 0.976) | 6.976 |
| miR-125a + miR-200a | 44 | 50 | 50 | 100 | NA | NA | NA | NA | 0.910 (0.870 to 0.950) | 0.910 (0.870 to 0.950) | NA |
| miR-31 | 78 | 45 | 24 | 69 | 36 | 8 | 9 | 16 | 0.800 (0.740 to 0.860) | 0.680 (0.585 to 0.775) | 2.500 |
| p16 | 63 | 39 | 102 | 141 | 0 | 0 | 39 | 102 | 0.000 (0.000 to 0.074) | 0.990 (0.971 to 1.00) | 0.000 |
| p16 | 63 | 177 | 500 | 677 | 8 | 0 | 169 | 500 | 0.045 (0.020 to 0.087) | 0.990 (0.994 to 1.00) | 4.500 |
| p16 | 63 | 136 | 102 | 238 | 18 | 0 | 118 | 102 | 0.132 (0.080 to 0.201) | 0.990 (0.971 to 1.00) | 13.200 |
| p16 | 63 | 136 | 500 | 636 | 18 | 0 | 118 | 500 | 0.132 (0.080 to 0.201) | 0.990 (0.994 to 1.00) | 13.200 |
| PGP9.5 | 63 | 52 | 203 | 255 | 4 | 5 | 48 | 198 | 0.077 (0.021 to 0.185) | 0.975 (0.944 to 0.992) | 3.080 |
| PGP9.5 | 63 | 34 | 112 | 146 | 28 | 78 | 6 | 34 | 0.824 (0.655 to 0.932) | 0.304 (0.220 to 0.398) | 1.184 |
| PGP9.5 | 63 | 45 | 112 | 157 | 41 | 78 | 4 | 34 | 0.911 (0.788 to 0.975) | 0.304 (0.220 to 0.398) | 1.309 |
| PGP9.5 | 63 | 45 | 203 | 248 | 41 | 5 | 4 | 198 | 0.911 (0.788 to 0.975) | 0.975 (0.944 to 0.992) | 36.440 |
| Prevalence screen HOXA9 | 67,66 | 4 | 4 | 8 | 3 | 0 | 1 | 4 | 0.850 (0.671 to 1.00) | 0.970 (0.885 to 1.00) | 28.333 |
| RARB | 63 | 13 | 85 | 98 | 0 | 0 | 13 | 85 | 0.000 (0.000 to 0.206) | 0.990 (0.956 to 1.00) | 0.000 |
| RARB | 63 | 44 | 35 | 79 | 35 | 17 | 9 | 18 | 0.796 (0.647 to 0.902) | 0.514 (0.340 to 0.686) | 1.638 |
| RARB | 63 | 44 | 85 | 129 | 35 | 0 | 9 | 85 | 0.796 (0.647 to 0.902) | 0.990 (0.965 to 1.00) | 79.600 |
| RASSF1A | 63 | 44 | 35 | 79 | 5 | 0 | 39 | 35 | 0.114 (0.038 to 0.246) | 0.990 (0.918 to 1.00) | 11.400 |
| RASSF1A | 63 | 44 | 104 | 148 | 5 | 1 | 39 | 103 | 0.114 (0.038 to 0.246) | 0.990 (0.948 to 1.00) | 11.400 |
| RBM6 | 63 | 44 | 18 | 62 | 43 | 14 | 1 | 4 | 0.977 (0.880 to 1.000) | 0.222 (0.064 to 0.476) | 1.256 |
| RBM6 | 63 | 44 | 35 | 79 | 43 | 35 | 1 | 0 | 0.977 (0.880 to 1.000) | 0.000 (0.000 to 0.082) | 0.977 |
| RIZ1 | 63 | 44 | 18 | 62 | 3 | 0 | 41 | 18 | 0.068 (0.014 to 0.187) | 0.990 (0.847 to 1.00) | 6.800 |
| RIZ1 | 63 | 44 | 35 | 79 | 3 | 4 | 41 | 31 | 0.068 (0.014 to 0.187) | 0.886 (0.733 to 0.968) | 0.596 |
| S100A2 | 63 | 44 | 12 | 56 | 42 | 10 | 2 | 2 | 0.955 (0.845 to 0.994) | 0.167 (0.021 to 0.484) | 1.146 |
| S100A2 | 63 | 44 | 35 | 79 | 42 | 31 | 2 | 4 | 0.955 (0.845 to 0.994) | 0.114 (0.032 to 0.267) | 1.078 |
| saliva HOXA9 | 67,66 | 4 | 4 | 8 | 3 | 2 | 1 | 2 | 0.750 (0.533 to 0.967) | 0.530 (0.280 to 0.780) | 1.596 |
| saliva NID2 | 67,66 | 4 | 4 | 8 | 4 | 3 | 1 | 1 | 0.870 (0.702 to 1.00) | 0.210 (0.006 to 0.414) | 1.101 |
| TGFBR2 | 63 | 37 | 134 | 171 | 3 | 8 | 34 | 126 | 0.081 (0.017 to 0.219) | 0.940 (0.886 to 0.974) | 1.350 |
| TGFBR2 | 63 | 11 | 44 | 55 | 6 | 33 | 5 | 11 | 0.546 (0.234 to 0.833) | 0.250 (0.132 to 0.403) | 0.728 |
| TGFBR2 | 63 | 42 | 134 | 176 | 37 | 8 | 5 | 126 | 0.881 (0.744 to 0.960) | 0.940 (0.886 to 0.974) | 14.683 |
| TGFBR2 + HIC1 | 63 | 44 | 149 | 193 | 22 | 35 | 22 | 114 | 0.500 (0.346 to 0.654) | 0.765 (0.689 to 0.831) | 2.128 |
| TGFBR2 + HIC1 + PGP9.5 | 63 | 39 | 118 | 157 | 22 | 39 | 17 | 79 | 0.564 (0.386 to 0.722) | 0.669 (0.577 to 0.753) | 1.704 |
| TGFBR2 + TIMP3 + HIC1 | 63 | 43 | 158 | 201 | 26 | 50 | 17 | 108 | 0.605 (0.444 to 0.750) | 0.684 (0.605 to 0.755) | 1.915 |
| TGFBR2 + TIMP3 + HIC1 + PGP9.5 | 63 | 38 | 131 | 169 | 26 | 54 | 12 | 77 | 0.684 (0.514 to 0.825) | 0.588 (0.499 to 0.673) | 1.660 |
| TIMP3 | 63 | 50 | 296 | 346 | 5 | 16 | 45 | 280 | 0.100 (0.033 to 0.218) | 0.946 (0.914 to 0.969) | 1.852 |
| TIMP3 | 63 | 176 | 450 | 626 | 20 | 32 | 156 | 418 | 0.114 (0.071 to 0.170) | 0.929 (0.901 to 0.951) | 1.606 |
| TIMP3 | 63 | 138 | 296 | 434 | 102 | 16 | 36 | 280 | 0.739 (0.658 to 0.810) | 0.946 (0.914 to 0.969) | 13.685 |
| TIMP3 | 63 | 138 | 450 | 588 | 102 | 32 | 36 | 418 | 0.739 (0.658 to 0.810) | 0.929 (0.901 to 0.951) | 10.408 |
| TIMP3 + HIC1 | 63 | 52 | 278 | 330 | 26 | 43 | 26 | 235 | 0.500 (0.358 to 0.642) | 0.845 (0.797 to 0.886) | 3.226 |
| TIMP3 + HIC1 + PGP9.5 | 63 | 45 | 183 | 228 | 26 | 47 | 19 | 136 | 0.578 (0.422 to 0.723) | 0.743 (0.674 to 0.805) | 2.249 |
| Valine + lactic acid | 72 | 37 | 32 | 69 | 32 | 6 | 5 | 26 | 0.865 (0.809 to 0.921) | 0.824 (0.757 to 0.891) | 4.915 |
Peer review:
Two peer reviewers contributed to the peer review report. Reviewers’ reports totaled 417 words, excluding any confidential comments to the academic editor.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was initiated with generous support from the International Medical University research grant no. IMUJC 290813 (71st meeting), project ID no. IMU 286/2013. The authors are very grateful for International Medical University’s support of this project.
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
