Abstract
We built a novel Bayesian hierarchical survival model based on the somatic mutation profile of patients across 50 genes and 27 cancer types. The pan-cancer quality allows for the model to “borrow” information across cancer types, motivated by the assumption that similar mutation profiles may have similar (but not necessarily identical) effects on survival across different tissues of origin or tumor types. The effect of a mutation at each gene was allowed to vary by cancer type, whereas the mean effect of each gene was shared across cancers. Within this framework, we considered 4 parametric survival models (normal, log-normal, exponential, and Weibull), and we compared their performance via a cross-validation approach in which we fit each model on training data and estimate the log-posterior predictive likelihood on test data. The log-normal model gave the best fit, and we investigated the partial effect of each gene on survival via a forward selection procedure. Through this we determined that mutations at
Introduction
The recently completed The Cancer Genome Atlas (TCGA) program has provided a comprehensive molecular characterization of 33 cancer types from more than 10 000 patients, and the database remains a valuable public resource. 1 The different cancer types are generally defined by their tissue-of-origin. The project has revealed striking heterogeneity in the genomic and molecular profiles across patients within each cancer type. This heterogeneity is presented in several flagship publications (eg, see The Cancer Genome Atlas Network2,3 and Verhaak et al 4 ) in the form of molecularly distinct subtypes, and these subtypes often correlate with clinical end points. However, leveraging molecular heterogeneity for personalized risk prediction on a more granular scale is limited by the number of patients with reliable clinical data for each cancer type. 5 Moreover, the effects of individual molecular biomarkers on survival or other clinical outcomes are often small, and thus predictive analyses within a single type of cancer are underpowered. 6
In 2013, TCGA began the Pan-Cancer Analysis Project, motivated by the observation that “cancers of disparate organs reveal many shared features, and, conversely, cancers from the same organ are often quite distinct.”7,8 The Pan-Cancer Analysis Project can also reveal differences in the effect of genomic changes across different cancer types, as demonstrated by the NOTCH gene family. 7 This initiative has resulted in several studies across multiple cancer types that have revealed important shared molecular alterations for somatic mutations, 9 copy number, 10 messenger RNA, 11 and protein abundance. 12 If these potential biomarkers have similar clinical effects across multiple types of cancer, then predictive models that are estimated using pan-cancer data will have more power than models that are fit separately for each type of cancer. The TCGA Pan-Cancer Clinical Data Resource (TCGA-CDR) 5 has facilitated pan-cancer models by curating and standardizing available data for 4 clinical outcomes (overall survival [OS], disease-specific survival, disease-free interval, or progression-free interval) across all 33 TCGA cohorts.
We propose and implement a novel Bayesian hierarchical framework to predict survival from multiple molecular predictors that are shared across multiple cancer types. An important feature of this approach is that it allows for borrowing information across models for each cancer type under the assumption that shared molecular predictors are likely to have a similar effect on survival prognosis, but it also allows sufficient flexibility for the same biomarker to have different effects depending on the type of cancer. Moreover, the Bayesian framework provides a principled way to incorporate prior information from other studies or cohorts, which is well-motivated given the vast body of literature on molecular biomarkers in cancer.
Using our proposed framework, we developed a pan-cancer predictive model for OS, using the somatic mutation profile of each tumor as the primary predictors of interest. We used OS because it is unambiguously defined and is available for almost all types of cancer, despite short follow-up times. 5 We focus on somatic mutations because they play a critical role in the development of many cancer types, 13 are available for all cohorts in the TCGA database, and are straightforward to compare across different tissues of origin. A pan-cancer analysis of somatic mutations across 12 different cancer types from TCGA revealed several genes that have frequent mutations across multiple types of cancer. 9 Their analysis also considered the marginal effect of each gene on OS, via cox proportional hazards models for (1) each cancer type separately and (2) for a fully joint model with all cancer types together; our approach aims to compromise between these 2 strategies. For our application we consider the somatic mutation status of 50 genes, and survival data for 5698 patients comprising 27 different types of cancer. We evaluate and compare several different potential models via a robust cross-validation approach to assess predictive accuracy for this data set.
The rest of this article is organized as follows. In the “Methods” section, we discuss how the data were collected and our approach to filtering out cancer types and genes. We also describe our modeling framework, how we selected the survival distribution we chose to use in our analysis, and our Gibbs sampling algorithm to calculate the posteriors of the model parameters. In the “Results” section, we discuss the results of our model selection procedure and our approach to determining which genes were most predictive of survival. Using the results from these 2 processes, we show the resulting credible interval estimates of our parameters and display survival curves. In the “Discussion” section, we discuss our results and future work that builds on this study for pan-cancer survival modeling.
Methods
Data acquisition and processing
We acquired clinical data for each patient via the TCGA-CDR,
5
which includes data for 33 cancer types and more than 11 160 patients. We acquired genome-wide somatic mutation data through the TCGA2STAT package for R,
14
which gathers data from the Broad Institute GDAC Firehose. The curated mutation data were available for 27 305 genes and 5793 patients with a binary indicator for whether there was a somatic nonsilent mutation (
We first matched the observations in the CDR data set with the mutation data obtained through TCGA2STAT. If any patients were present in one data set but not the other, that observation was removed from the study. We also removed any observations that had a negative survival time, a survival time entered as
Cancer types included in study with their corresponding sample sizes.
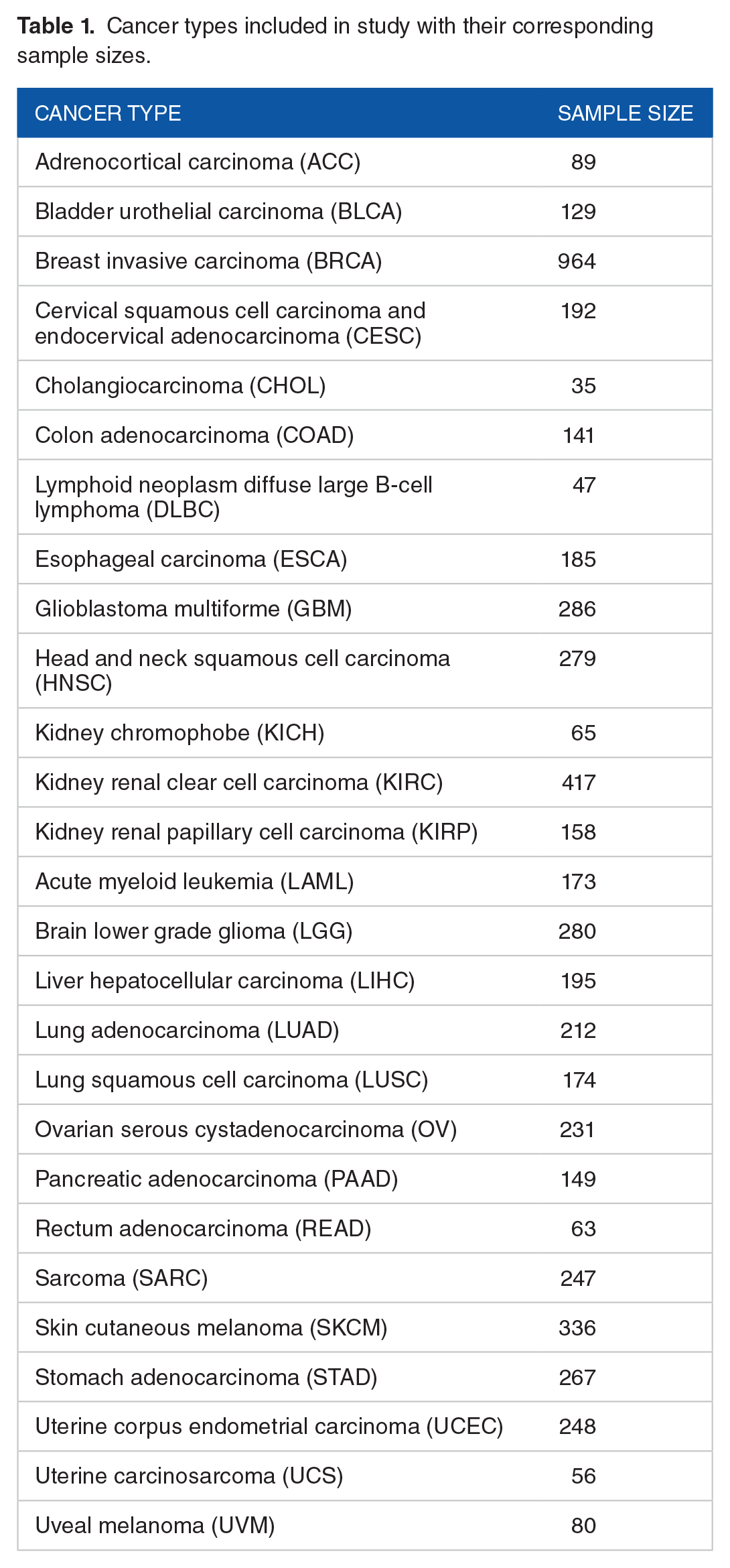
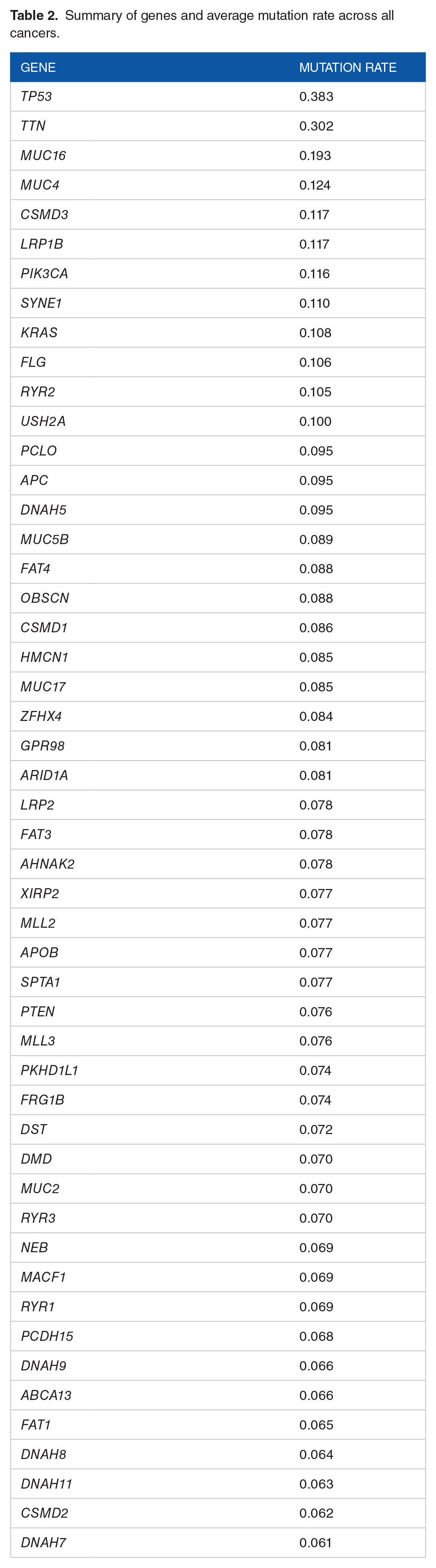
We filtered the genes based on average mutation rate across the 27 cancer types, selecting the top 50 mutated genes. To determine the average mutation rate, we calculated the mutation rate of each gene for each cancer type and took the average by gene across all cancers. In this way, each cancer type was weighted the same in calculating the mean mutation rate. This also ensured that each gene would be represented across most cancers, not just within a few. As a result, certain genes that are highly mutated in particular cancers but not in others were excluded. The genes we incorporated in our study can be found in Table 2 with their corresponding average mutation rates. In total, we used mutation data from 5698 patients.
Summary of genes and average mutation rate across all cancers.
We considered the correlation of mutation status between genes across all cancers, for exploratory purposes and to investigate potential issues of multicollinearity for polygenic models. Figure 1 shows a pairwise correlation plot for all genes considered using mutation status across all patients included in this study. The Pearson correlation coefficients between genes were uniformly positive but relatively weak, ranging from

Correlation plot for correlations between somatic mutation statuses across tissue types. Blank squares indicate nonsignificant correlation between 2 genes. Squares that contain a circle indicate significant correlations and the magnitude of the correlation is indicated by the color opacity.
Model
We propose a Bayesian hierarchical model for patient survival that incorporates binary mutation status variables and age across 27 cancer types. We centered the age covariate by subtracting out the average age for each cancer type. This reduces collinearity of the age coefficient with the intercept terms, ensuring that the estimated effect of age is not influenced by one cancer having generally older patients and another cancer having generally younger patients. The multilayer nature of our model allows the effect of a mutation at each gene to vary by cancer type while simultaneously inferring the mean and variance of these effects. Thus, the model facilitates the borrowing of information across cancer types by shrinking the estimated effects toward a common mean. Our model can also accommodate censored observations, as discussed in the following subsection. We use the following notation in our framework:
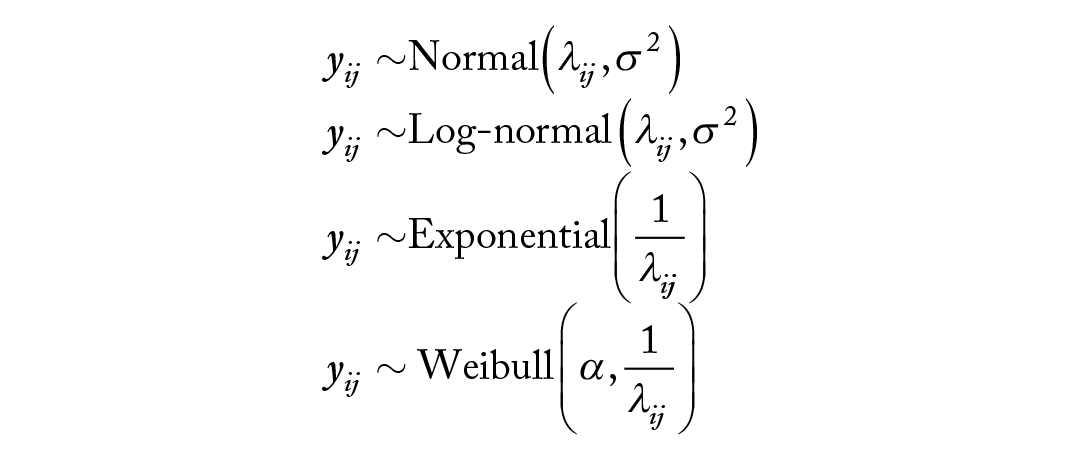
For each likelihood we consider a hierarchical linear model for

where
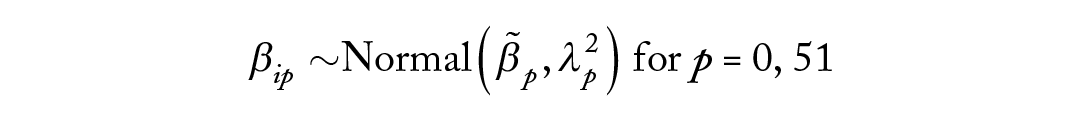
where
Parameter estimation
Depending on the survival model, we employed a different approach to infer the posterior for model parameters. The log-normal and normal models were fit in R using an in-house Gibbs sampler. The sampler is described below; however, more details can be found in Appendix 1.
Initialize
For samples
Draw
Draw
Draw
Draw
Generate survival times for censored observations using ● If assuming the data follows a normal distribution, generate survival times for censored observations from a normal distribution with mean β(t)i 0+ ● If assuming the data follows a log-normal distribution, generate survival times from a normal distribution with mean
We ran the sampler for 20 000 iterations and used a 10 000 iteration burn-in to ensure convergence of the parameters. In Appendix 2 of this article, we describe how we validated this model fitting procedure.
Posterior samples for the exponential and Weibull models were obtained using the Just Another Gibbs Sampler (JAGS) software. 16 For these models, we ran the sampler for the same number of iterations and employed the same burn-in as for the normal and log-normal models above. In all calculations based on the posteriors of our Gibbs sampler and our JAGS models, we thinned by every 10th iteration to speed up computing time and memory efficiency.
Model selection
To assess which of the normal, log-normal, exponential, and Weibull models was the best fit for the data, we calculated the log out-of-sample posterior predictive likelihood in a fivefold cross-validation procedure, as described below. Consider the

where
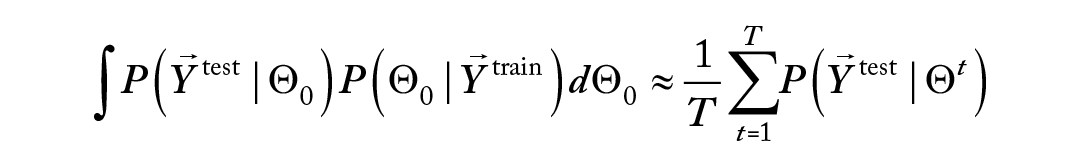
where
Forward selection
To assess the partial improvement of each gene in predicting survival, we constructed a forward selection approach that would allow us to see which genes were most important in predicting survival across all cancer types. In this way, we were able to determine the relative importance of each gene, and achieve a more parsimonious predictive model. For our forward selection approach, we used the same out-of-sample log-posterior likelihood metric described in the previous section on model comparison. Our forward selection method proceeded as follows:
Calculate the average log-posterior predictive likelihood for the null model (with age and cancer-type intercepts, but no genes) fit on each of the fivefolds.
For each gene, consider the model with only the intercept, age, and that gene included. Calculate the log-posterior predictive likelihood under this model using fivefold cross-validation. (a) Select the gene that produced the model with the highest log-posterior predictive likelihood. Call this model (b) Compare the likelihood for this model with the null model; if the likelihood has increased, proceed by adding this gene to the model. Otherwise, stop.
For each remaining gene, add that gene separately to model (a)● Select from the resulting 2 gene models the gene that maximized the log-posterior likelihood. Call this model (b)● Compare the log-posterior likelihood for
Continue until the log-posterior predictive likelihood ceases to increase. At this point, the final model has been found.
The order of the genes added to the model depended on which genes maximized the posterior likelihood at each step. Once a final model had been found, we investigated the effects of each mutation on survival through credible interval plots and survival curves, described further in the “Results” section.
Results
Model selection
The results of the comparison described in the “Methods” section are shown in Table 3. We found that the log-normal model had the highest out-of-sample log-posterior likelihood out of the normal, exponential, and Weibull models. We assumed a log-normal distribution of survival for the remainder of our investigation.
Out-of-sample posterior predictive likelihood.

In this model, the coefficients are inferred by borrowing information across cancer types. However, we considered an analogous log-normal model in which the coefficients were inferred independently for each cancer type to compare with the model we selected here. We used the same uninformative prior on each coefficient as the one assumed for
Forward selection
Table 4 displays the mean log-posterior predictive likelihoods for each step in the forward selection procedure. Every model is adjusted for patient age.
Results from forward selection procedure.

The log-posterior likelihood for the null model, meaning the model with only age as a predictor, was
We also used our forward selection approach without including
Results from forward selection procedure without

Without including
Credible intervals for model coefficients
To understand the magnitude and direction of the partial effect of age and a mutation at a gene on patient survival, we computed and visualized the
Credible intervals for model coefficients across each cancer type.

Figure 2 displays the credible intervals across cancer types for each parameter in the model. Panel 2A compares the baseline survival across the different cancer types. Panel 2B reveals the generally deleterious effect of age on patient survival, as indicated by the highlighted orange intervals. For most of the cancers, an increase in age led to a decrease in survival; however, the extent to which age has an effect is not homogeneous or precisely identified for every cancer. For breast cancer (BRCA), the impact of age is more certain, indicated by a narrower credible interval, compared with the effect of age on patients with, eg, uterine carcinosarcoma. Similarly, Figure 2C shows the estimated effect of a

Credible intervals by cancer type for the intercept (panel A), Age effect (panel B), TP53 effect (panel C), and FAT4 effect (panel D).
Credible intervals for mean mutation effect
We also studied the credible intervals for the mean effect of each covariate. The mean effect for each predictor,
Credible intervals for mean of model coefficients.

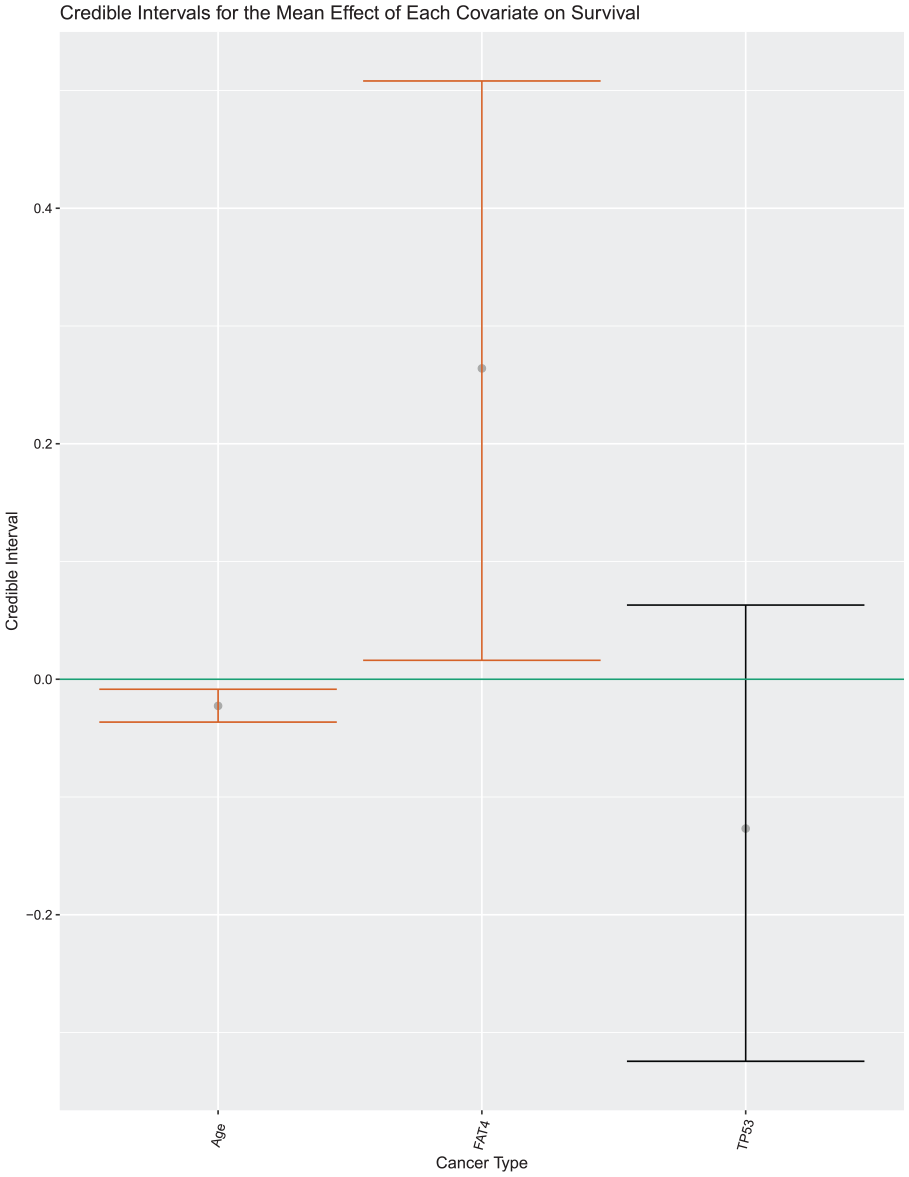
Visual display of the credible intervals for
These results indicate the effect of each covariate averaged across cancers; therefore, if a predictor was more potent in one cancer and less so in another, this may not necessarily be represented in estimates for the mean effect. This substantiates why we chose to allow the effect of each covariate to differentiate by tumor type. The credible intervals for age and
Survival plots
To visualize the impact of age and a mutation at each of

Survival curves under different covariate combinations for adrenocortical carcinoma with ages overlaid. Estimates for 30-year-old patients are shown in black, 50-year-olds in orange, and 80-year-olds in blue. The 95% error bounds are shown in dotted lines.
Discussion
In this article, we propose a novel Bayesian hierarchical model for survival of patients with cancer based on age and mutation status. This model is unique in its ability to allow the effect of each covariate to vary by cancer type. This framework is motivated by the assumption that similar genetic profiles may have similar, though not necessarily identical, effects on patient survival across tissues of-origin. This work may be extended to allow for other clinical covariates to be added to the model, such as stage and grade, and allows the user to adjust the effect of each predictor by cancer type as informed by prior knowledge.
To determine which genes were most important in survival prediction, we used a forward selection procedure that added
In addition, we considered the possibility of collinearity between mutation status variables, prompting us to investigate the correlation levels between variables across all cancer types (Figure 1). Based on this plot, we concluded mutation statuses across all cancer types were not highly associated. However, the results may look different if one is not considering the cancers in aggregate. With that in mind, we propose in future work sorting genes differently to meet the interest of the researchers, ie, exploring genes that are known to be highly mutated in a set of related cancers. In such an instance, it may be necessary to consider collinearity between variables and adjust accordingly.
We also assessed the predictive quality of our model by calculating survival curves for each cancer type based on combinations of age and mutation status based on our model. These plots demonstrated the largely negative effect of increasing age and a
In our study, we used the TCGA2STAT R package to import TCGA somatic mutation data due to its convenience in data dissemination, providing somatic mutation data in a ready-to-use format for statistical analysis. Although the package does offer convenience, we were not able to acquire data for MESO, which is available through the TCGA Multi-Center Mutation Calling in Multiple Cancers (MC3) data set and through the National Cancer Institute’s (NCI) Genome Data Commons. In the interest of ease of future replication of this study, using the TCGA2STAT data set may make it simpler for scientists to acquire the same data we used. We also did not distinguish the genes we used based on driver mutation status, false-positive status, or otherwise.
In future studies, it would be interesting to apply the model to a subset of the 27 cancer types we selected to group cancers that may be more similar in genetic nature or otherwise. This may elucidate genes unique to predicting patient survival outcome to specific cancer groupings. Interactions between covariates could also be included in the model to assess their relationship to OS. It would also be interesting to investigate alternate approaches to selecting genes to incorporate in the model, possibly incorporating prior knowledge on driver mutation status or false-positive status, or achieving variable selection through a sparsity inducing prior, as is done in Maity et al 23 using horseshoe priors.
Footnotes
Appendix 1
Appendix 2
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Institutes of Health (NIH) National Cancer Institute (NCI) grant R21CA231214-01.
Declaration of Conflicting Interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
SS performed all analyses and drafted the manuscript. KAH provided helpful information and insight regarding the TCGA data and interpretation of the results. EFL conceived of the idea and developed the method with SS. All authors read and edited the manuscript.
