Abstract
Introduction:
Most of the subjects undergoing diagnostic colonoscopy do not have neoplastic bowel lesions. Potentially, some of the symptoms may therefore be caused by extracolonic malignancy, and subjects with persisting symptoms may need subsequent examinations. Blood-based, cancer-associated biomarkers may aid in directing the examinations for other specific malignant diseases.
Methods:
EDTA plasma samples available from a previous prospective study of subjects undergoing diagnostic colonoscopy were used for analysis of 18 protein biomarkers. The study population of 3732 subjects included 400 patients with colorectal cancer (CRC) and 177 patients with extracolonic malignancies. Univariable analysis of the association of specific biomarkers and extracolonic cancers included those with 10 or more cases. Subsequently, reduced models of 4 or 6 biomarkers, respectively, were established by choosing those with the highest likelihood; age and sex were included as well.
Results:
Univariable analyses showed that CyFra21-1 had an area under curve (AUC) of 0.87 for lung cancers (n = 33), CA19-9 had an AUC of 0.85 for pancreatic cancer (n = 22), CA125 had an AUC of 0.95 for ovary cancer (n = 16), B2M had an AUC of 0.81 for non-Hodgkin lymphoma (n = 12), and total prostate-specific antigen had an AUC of 0.99 for prostate cancer (n = 10). The multivariable analysis of 4 or 6 biomarkers plus age and sex as explanatory variables showed AUCs of 0.82 to 0.85 both for extracolonic cancers and CRC. The 4 biomarkers included in the model for detection of extracolonic cancers were CA125, hsCRP, CA19-9, and CyFra21-1; the 2 additional for the 6 biomarkers model were CEA and Galectin-3. Similarly, the 4 biomarkers included in the model for detection of CRC were CEA, CyFra21-1, Ferritin, and HE4; the two additional for the 6 biomarkers model were hsCRP and Pepsinogen 2.
Conclusions:
Results of this study indicate that it may be possible to detect subjects that have an increased risk of extracolonic cancer following a colonoscopy without findings of neoplastic lesions. Combinations of various protein biomarkers may direct subsequent examination after colonoscopy with clean colorectum. The results, although preliminary, may form the basis for additional research directed both for primary examinations of subjects with symptoms of malignancy and subsequent examinations after colonoscopy.
Introduction
Worldwide, colorectal cancer (CRC) is a global challenge for the health care systems with more than 1.4 million primary cancers diagnosed every year; annually, approximately 695 000 patients die of the disease. 1 Screening of the average risk populations may be one of the options to reduce the incidence and mortality of CRC,2,3 presumably due to detection of adenomas and CRC at early stages.2–4 At present, fecal immunochemical test (FIT) screening appears as the leading screening option with a substantial sensitivity and specificity 5 and at a low cost benefit as well. 6 Compliance issues may, however, reduce the clinical sensitivity to less than 50% resulting in more than 50% of those subjects with nonsymptomatic neoplastic bowel lesions will not be identified by FIT screening. 7 Therefore, present research has focus on developments of blood-based biomarkers that in various combinations may improve the current screening for CRC. 8 Recent results have shown that a combination of 8 protein biomarkers associated with CRC may detect large bowel neoplasia. 9 These results are retrieved from a cohort of subjects offered diagnostic colonoscopy and are currently validated in a major prospective screening-associated study. 8
Previous results have indicated that increased protein biomarker levels in association with colonoscopy for large bowel neoplasia are, however, not only uniformly related to bowel lesions but may also be shown among subjects with clean colorectum.10–12 Such facts may be a major challenge in validation and clinical implementation of blood-based biomarkers, as it may lead to both specificity and sensitivity issues that may reduce the acceptability in future screening for bowel neoplasia. Consequently, that may prompt research on identification, evaluation, and validation of biomarkers that may direct subsequent examination of subjects offered colonoscopy due to increased biomarker levels, but who are identified with clean colorectum.
A variety of blood-based biomarkers, including proteins is not uniformly associated with a specific malignant disease, but may be more or less associated with malignancy in general. 12 It was shown recently that subjects with increased biomarker levels and clean colorectum at diagnostic bowel endoscopy for CRC had an increased risk of having or developing extracolonic cancers within few years.13,14 The biomarkers included in those reports were TIMP-1, CEA, CA19-9, and YLK-40. In the recent major study of protein biomarkers in subjects scheduled for diagnostic colonoscopy, 177 patients with various extracolonic cancers were also diagnosed after clean colorectum, but with subsequent examination due to persisting symptoms. 9 Indeed, some protein biomarkers are more associated with one malignancy than with others, such as prostate-specific antigen (PSA) to prostate cancer, CA19-9 to pancreatic cancer, and HE4 to ovarian cancer.15–17 Additional analysis of some of these biomarkers in samples collected from that particular study 9 could potentially have been a help in directing subsequent examination for some of those extracolonic cancers.
Therefore, the primary aim of this study was to perform analyses of 10 protein biomarkers in relation to specific malignancies and add those to the 8 protein biomarkers determined previously 9 to identify single biomarkers or combinations that may be useful in directing examination subsequent to a colonoscopy with clean colorectum. The secondary aim was to improve an already established pilot biomarker profile for CRC. 9
Materials and Methods
Study population
A total of 4698 subjects scheduled for first time ever colonoscopy due to symptoms of colorectal neoplasia were prospectively included from May 1, 2010, to November 30, 2012, at 7 collaborating hospitals in Denmark. 9 Biobanked samples from 3732 subjects were available for this study; samples from 177 patients with extracolonic malignancies diagnosed after the primary colonoscopy were available (Figure 1).
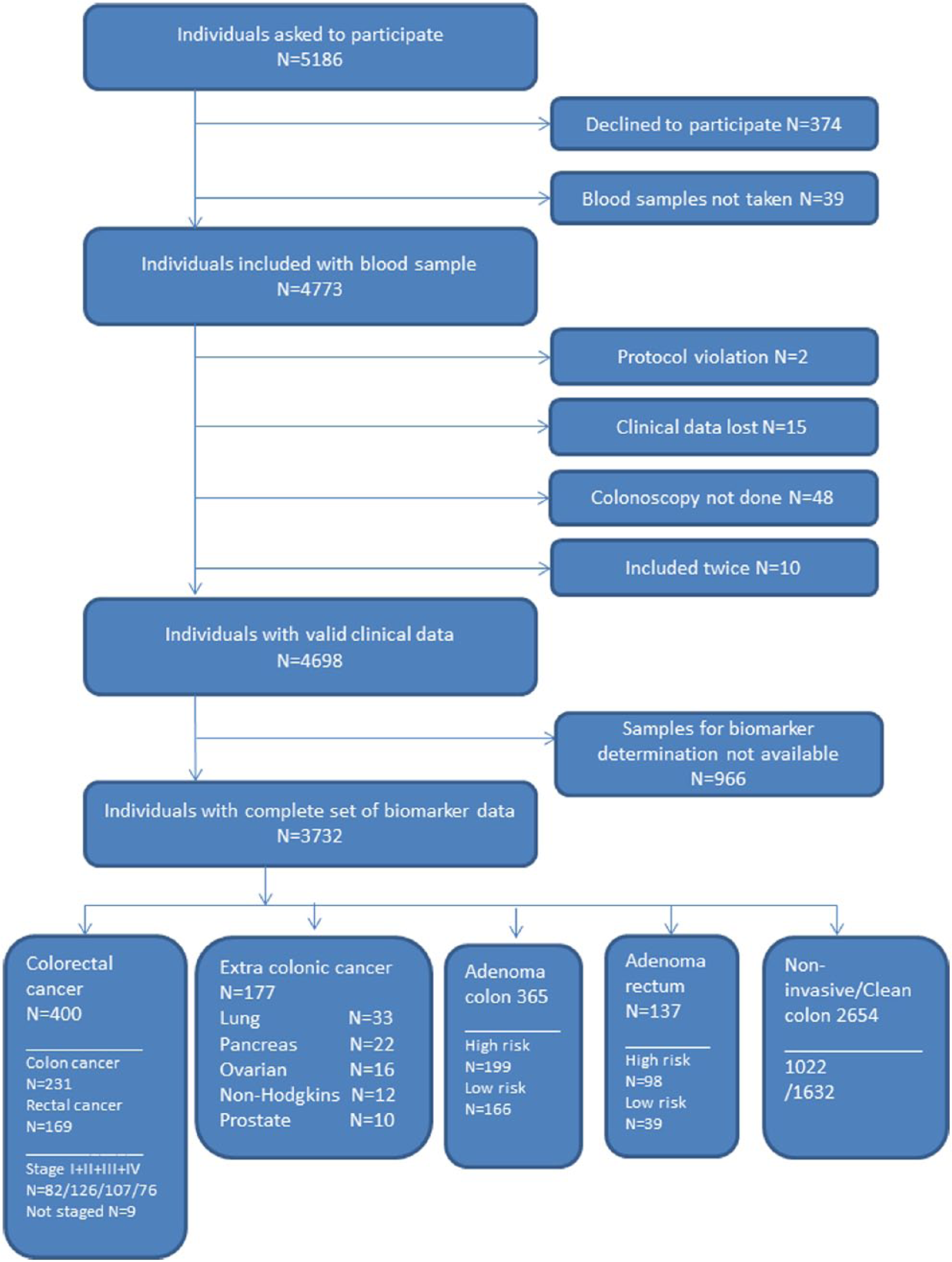
Flowchart of the entire study.
Demographic data including comorbidity were recorded before colonoscopy. Subjects unable to undergo complete colonoscopy and subjects with complete colonoscopy, but without bowel pathology and persisting symptoms, were offered additional examination, ie, gastroscopy, X-ray with barium enema, ultrasonography, computer-assisted tomography, and/or magnetic resonance imaging. All malignancies were histologically confirmed and stages classified according to relevant staging systems, such as TNM for solid tumors, Ann Arbor staging for Lymphoma, RAI staging for chronic lymphocytic leukemia, and International Staging System for Multiple Myeloma. Patients were followed-up using the computerized, unique central personal registration number (cpr.nr.) given to all Danish citizens. Due to Danish legislation, all diagnoses of extracolonic malignancies were established and verified within 1 month after colonoscopy.
Sample collection, storage, handling, and processing
Blood samples were collected from an antecubital vein just before colonoscopy according to a validated standard operating procedure (SOP). The blood samples were collected by dedicated research nurses, handled into serum, EDTA plasma, and buffy-coat samples, respectively, and finally stored at −80°C as previously described; the entire procedure was finalized within 2 hours. 9
When all samples were collected, EDTA plasma levels of protein biomarkers associated with CRC were determined on the entire cohort of 4698 subjects. 9 Subsequently, the levels of β2-microglobulin (B2M), cancer antigen 125 (CA125), cancer antigen 15-3 (CA15-3), human epidermis antigen 4 (HE4), neuron-specific enolase (NSE), Pepsinogen 1 and 2, squamous cell cancer antigen (SCC), progastrin-related peptide (ProGRP), and total PSA (tPSA) were determined in EDTA plasma samples from the 3732 selected subjects using the Abbott ARCHITECT automated immunoassay and clinical chemistry platform. The rationale for inclusion of the specific biomarkers is shown in Table 1. Testing was performed at the Abbott Center of Excellence at Vrie University Medical Center (VUMC), Amsterdam, The Netherlands.
Overview of individual markers.

Ethics
The Ethics Committee of The Capital Region of Denmark (H-3-2009-110) and The Danish Data Protection Agency (2008-41-2252) approved the study, which was performed according to the Helsinki II declaration. The REMARK (REporting recommendations for tumor MARKer prognostic studies) guidelines for running and reporting the study were followed whenever applicable. 25
Statistics
All end points considered are binary and have been described in detail previously. 9
Briefly, the 2 outcomes are CRC versus all others, but excluding extracolonic cancers, and extracolonic cancers versus all others, but excluding CRC. Univariable analyses were performed for each of the 18 explanatory variables, and then multivariable analysis was performed including all explanatory variables as well as age and sex. Both univariable and multivariable analyses were done using logistic regression models with the explanatory variables (not age and sex) log transformed. Analyses of specific cancers other than CRC were done for those malignancies with 10 or more cases except for unknown primary tumors as they were considered too heterogeneous. These analyses included the addition of biomarkers known to be associated with the specific cancer type. The results are presented by the receiver operating characteristic curves with area under curve (AUC) as a measure of discrimination. In addition, specificities at predefined sensitivities (70%, 80%, and 90%) are presented. In addition, a model reduction was done for each end point by selecting the best 4 or 6 biomarkers and age and sex in a backwards fashion identifying those with the highest likelihood score. The
Results
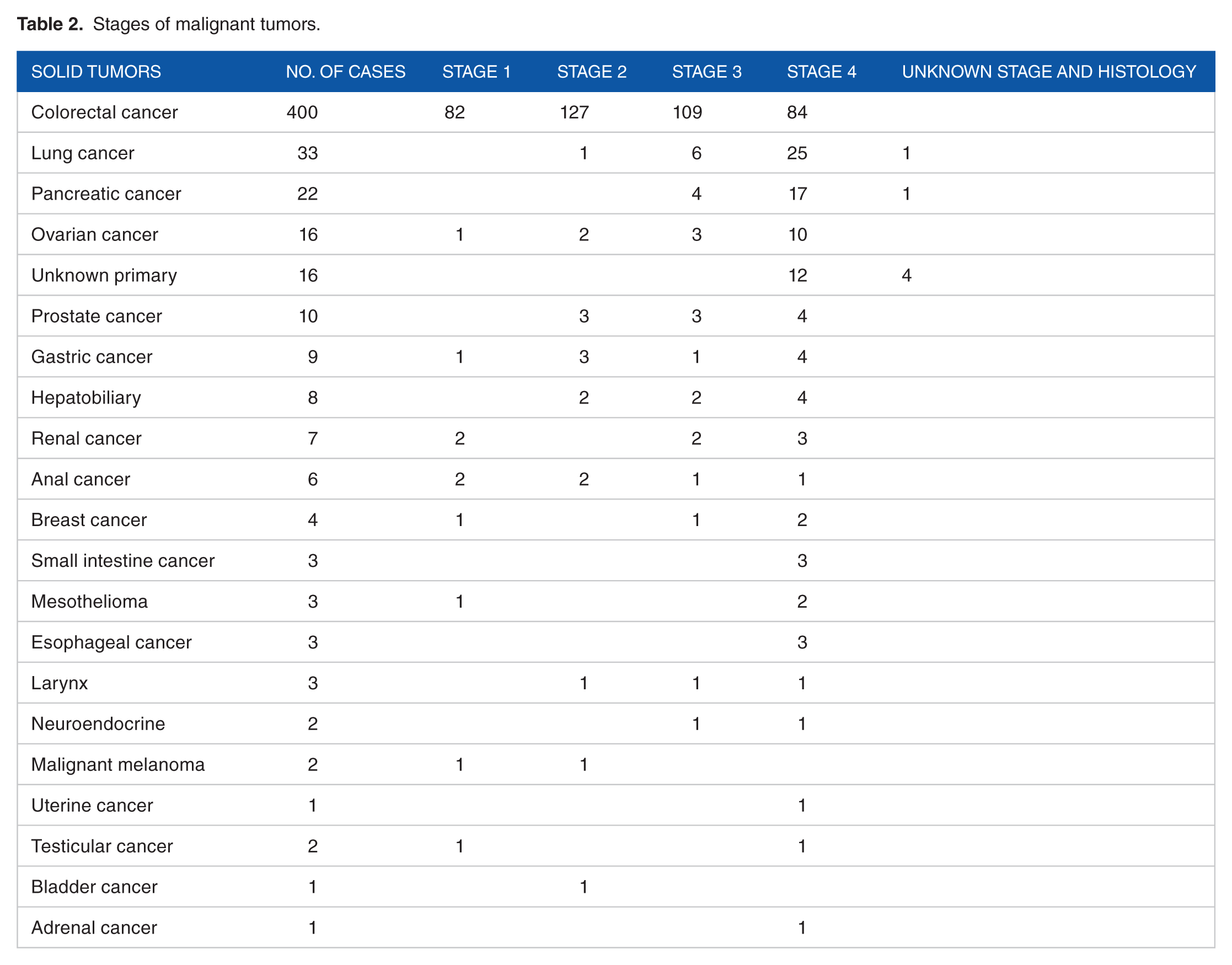
Diagnoses and findings at colonoscopy or subsequent examinations of the 3732 subjects were as follows: 400 with CRC, 177 with extracolonic cancers, 502 with adenomas, 1021 with other benign bowel lesions, and 1632 with clean bowel. Stages for CRCs and extracolonic cancers are shown in Table 2. Most of the extracolonic cancers were diagnosed at late stages as 58% (104/177) had stage IV disease (Tables 2 -5).
Stages of malignant tumors.
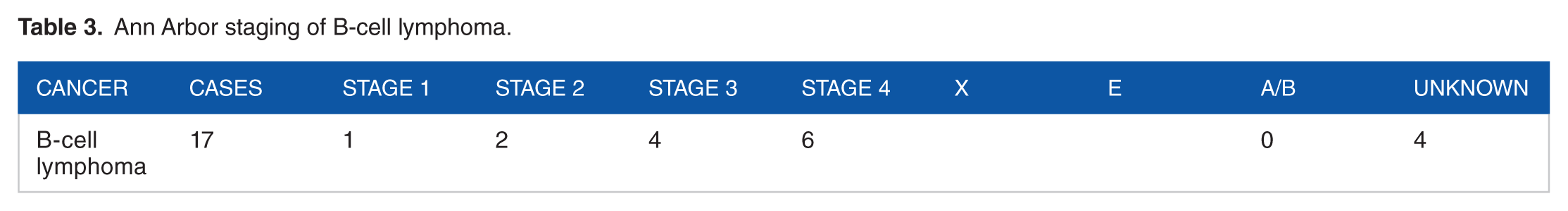
Ann Arbor staging of B-cell lymphoma.
International Staging System for Multiple Myeloma.

Chronic lymphocytic leukemia (CLL) (RAI stages).

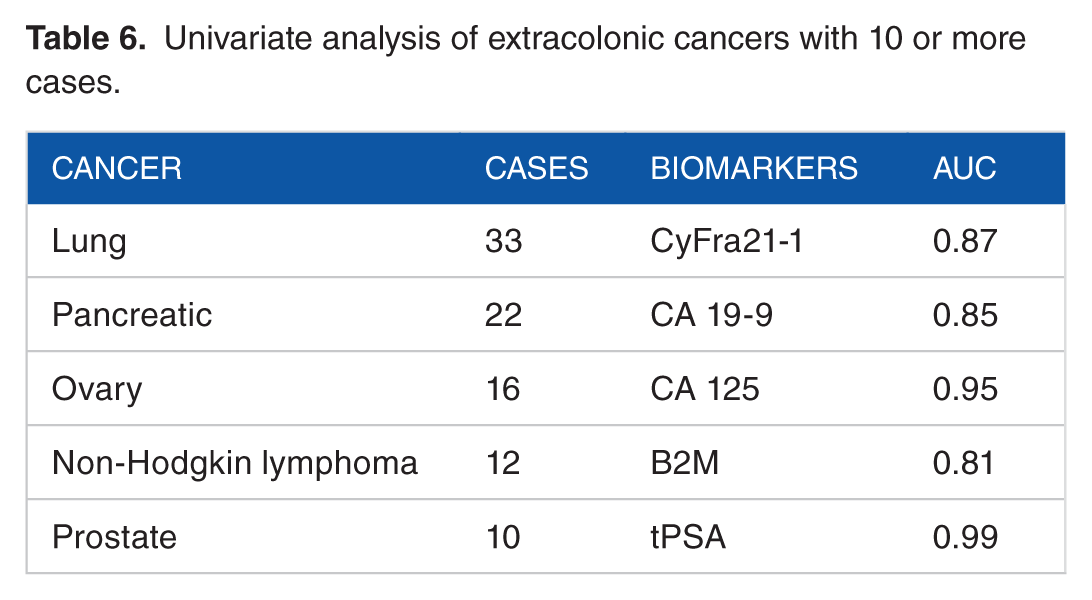
Univariable analyses of the single biomarkers to detect the specific extracolonic cancers of more than 10 cases showed that CyFra21-1 had an AUC of 0.87 for lung cancers (n = 33), CA19-9 had an AUC of 0.85 for pancreatic cancer (n = 22), CA125 had an AUC of 0.95 for ovary cancer (n = 16), B2M had an AUC of 0.81 for non-Hodgkin-lymphoma (n = 12), and tPSA had an AUC of 0.99 for prostate cancer (n = 10) (Table 6).
Univariate analysis of extracolonic cancers with 10 or more cases.

Application of univariable analysis of the 18 single biomarkers on the entire group of patients with extracolonic cancers showed AUCs of 0.77 for CA125, CyFra21-1, and TIMP-1, respectively. Furthermore, HE4 had an AUC of 0.76, whereas AFP had the lowest ability to discriminate at an AUC of 0.52. Similar application of univariable analysis on the group of patients with CRC showed that CEA and CyFra21-1 had AUCs of 0.73, respectively. The only markers of the 10 additional single biomarkers that had discriminatory effects on CRC were B2M and HE4 with AUCs of 0.63, respectively.
Subsequently, specificities at fixed sensitivities for univariable analyses were assessed both for extracolonic cancers and CRC, respectively (Table 7). Among the included biomarkers, CyFra21-1 appeared as the best single protein for detection of both extracolonic cancers and CRC. To some extent, the chosen biomarkers appeared to be correlated; ie, Pepsinogen 1 and Pepsinogen 2 had a rank correlation of 0.69, and HE4, Pro-GRP, B2M, TIMP-1, Galectin-3, TIMP-1, hs-CRP, and CyFra 21-1 had rank correlations higher than 0.40 (data not shown), which suggests a need for specific considerations in interpretation of results of the full model. Therefore, the results of multivariable analyses are presented as a reduced model of 4 or 6 biomarkers, respectively. The single biomarker proteins with the highest detection rate was selected from that analysis.
Specificities at fixed sensitivities for univariate analysis of CRC and extracolonic cancers.
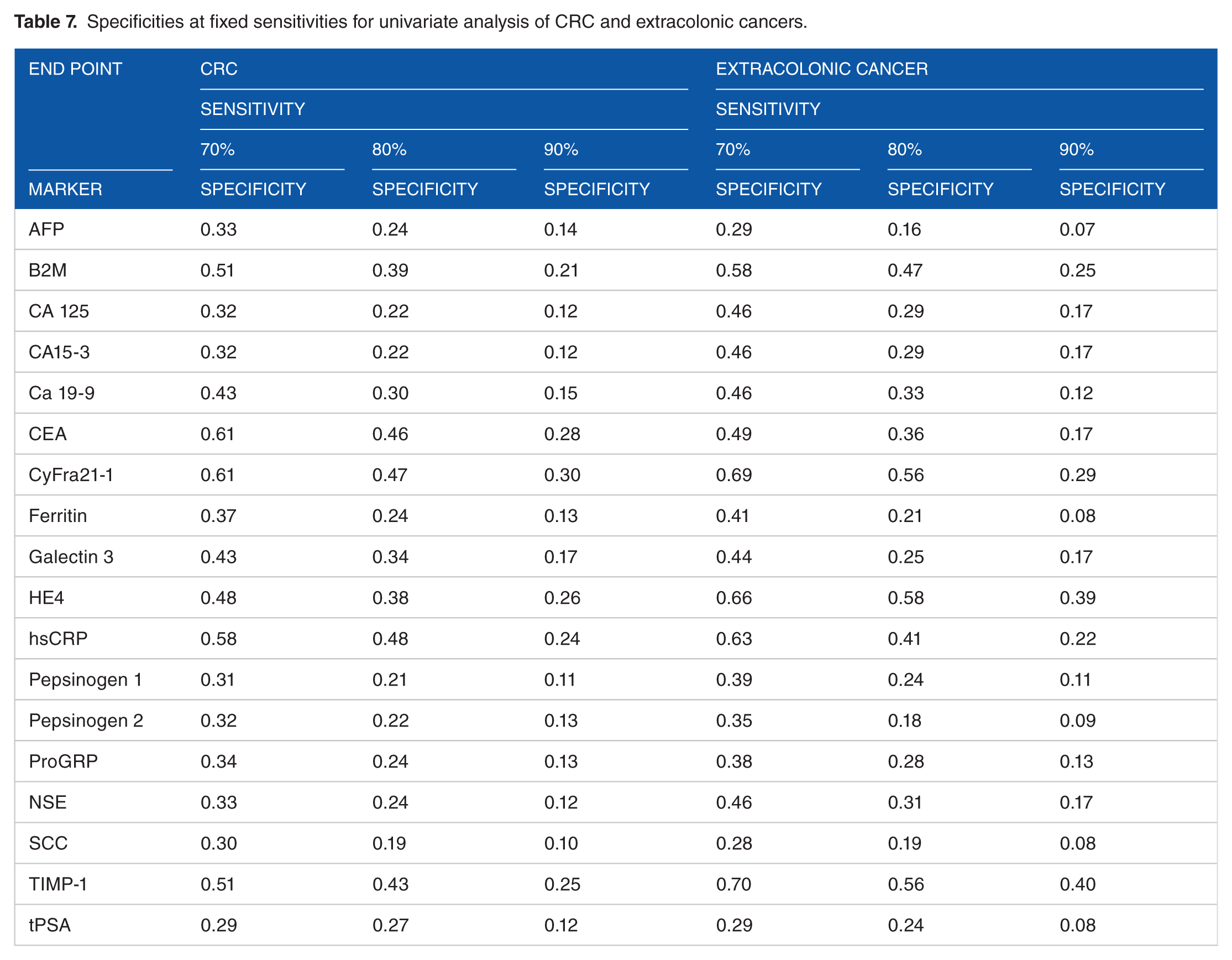
The multivariable analysis of 4 or 6 biomarkers plus inclusion of age and sex as explanatory variables showed AUCs of 0.82 to 0.85 both for extracolonic cancers and CRC (Table 8). The 4 biomarkers included in the model for detection of extracolonic cancers were CA125, hsCRP, CA19-9, and CyFra21-1; the 2 additional for the 6 biomarkers model were CEA and Galectin-3. Similarly, the 4 biomarkers included in the model for detection of CRC were CEA, CyFra21-1, Ferritin, and HE4; the 2 additional for the 6 biomarkers model were hsCRP and Pepsinogen 2.
Multivariable analysis of various end points including 4 or 6 best protein biomarkers plus age and sex as covariates.
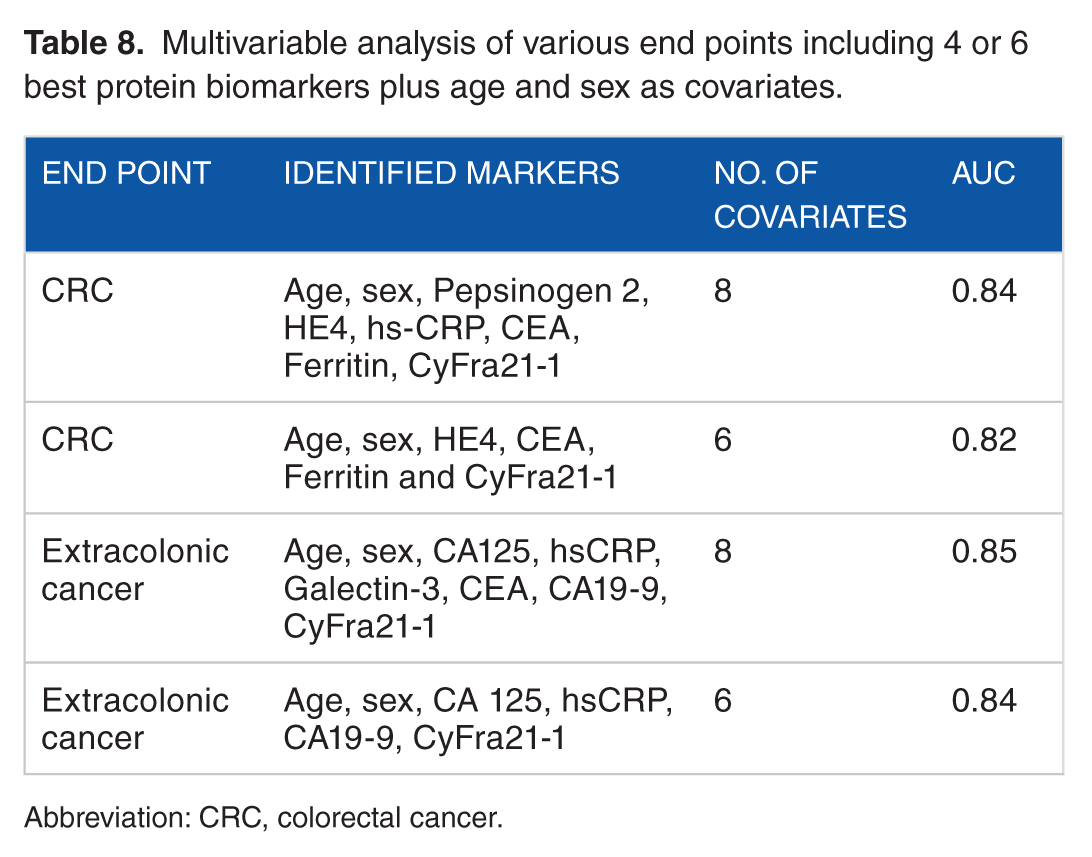
Abbreviation: CRC, colorectal cancer.
Multivariable analysis of the specificities at fixed sensitivities showed that the sensitivity for detection of the entire group of extracolonic cancers was 70% at 88% specificity, 80% at 72% specificity, and 90% at 44% specificity, respectively. Similarly, the sensitivity for detection of CRC was 70% at 82% specificity, 80% at 71% specificity, and 90% at 48% specificity, respectively.
Discussion
Frequently, subjects undergoing symptom-directed diagnostic colonoscopy appear to have clean colorectum. Among 3732 subjects included in this study 1021 had clean colorectum, whereas 400 had CRC, 502 had adenomas, and 1632 had other benign bowel lesions. Due to persistent symptoms, some of the subjects with nonmalignant bowel lesions or clean colorectum needed subsequent additional examinations, and thereby extracolonic malignancies were identified in 177 patients. This study focused on a selected part of the entire study cohort to address an important clinical question: Can blood-based biomarkers aid in directing subsequent examination for organ-specific malignancies after colonoscopy with finding of a clean colorectum? The present preliminary results indicate that it may be possible, but this needs to be validated in future and sufficiently powered studies.
The results achieved by univariable analysis of the 18 single protein biomarkers, on the entire cohort of patients with 24 different extracolonic malignancies (Tables 2 -5), indicate that CA125, CyFra21-1, and TIMP-1 may have fair detection rates as the AUCs were 0.77, respectively; lower AUCs were shown for some of the other 18 biomarkers. Application of additional univariable analysis indicates that specific, single biomarkers may be useful in directing subsequent examination. Although the cohorts of various extracolonic malignant diseases had limited numbers, the AUCs ranged from 0.99 (tPSA for prostate cancer) via 0.95 (CA125 for ovary cancer) and 0.87 (CyFra21-1 for lung cancer) and 0.85 (CA19-9 for pancreatic cancer) to 0.81 (B2M for non-Hodgkin lymphoma). It must be taken into consideration, however, that most of the patients diagnosed with extracolonic malignancies had disseminated disease. This fact may have led to the very high AUCs for single biomarkers of the specific diseases. Subjects with clean colorectum after diagnostic colonoscopy and with persisting symptoms may, however, benefit from a blood-based test that may aid in directing subsequent examination to identify the cause of the symptoms. Thereby, such subjects may avoid series of complex and invasive examinations. In its present form, the tests may not be applicable in directing examination for primary, low-stage malignancies. Such tests may be based on the addition of other biomarker entities, including cell-free DNA analyses,26,27 which recently have shown promising results. It is well known that cell-free DNA does not detect all malignancies with high sensitivity at high specificity, and therefore, combinations of circulating protein and DNA biomarkers may be a future option to improve the clinical accuracy of the specific cancer tests. 28
In multivariable analysis, panels of 4 or 6 biomarkers plus age and sex were identified. Regarding the primary end point, extracolonic cancers, the following markers were identified: CA125, hsCRP, Galectin-3, CEA, CA19-9, and CyFra21-1. With this panel, an AUC of 0.85 was achieved and thereby may aid as the diagnosis of patients with certain diseases. The specificities at fixed sensitivities ranged between 0.88 and 0.44 for this end point. In the secondary end point, an AUC of 0.84 was achieved and the following markers were identified: Pepsinogen 2, HE4, Hs-CRP, CEA, Ferritin, and CyFra21-1. The result was a slight improvement of the result shown in the previous study of primary CRC detection. 9 For this end point, the specificities at fixed sensitivities ranged between 0.82 and 0.48.
Current research of early detection of large bowel neoplasia in screening procedures focuses on biomarkers that may have the highest possible specificity to reduce the numbers of false positives and thereby reduce subsequent colonoscopy. Such achievements may have influenced the sensitivity, however, which often is somewhat lower and may lead to subjects that are classified as false negative. This study has focused on the sensitivity to direct examinations for extracolonic cancers subsequent to a colonoscopy, which had not led to identification of bowel neoplasia. As most of the examinations of subjects with symptoms of CRC are performed via the outpatient clinics, it is essential that examination procedures focus as much as possible on the risk of a malignant disease that may cause the symptoms. In particular, focused examinations may be urgent for subjects that have stage IV diseases as in this study. Due to the severity of disease, such patients need immediate initiation of the offered therapy. It is well known, however, that tests with high sensitivities often have lower specificities that may lead to more subjects, who are classified as false positives. Pros and cons on sensitivities and specificities are a subject of intense discussions, but the severity of the diseases must also be a part of the debate.
Although the results were achieved based on a limited number of patients, the study findings may lead to consideration of whether this testing approach could potentially be applicable in the primary evaluation of subjects with symptoms of various malignancies to direct primary examination. Thereby, some subjects may not need bowel preparation and subsequent colonoscopy, which may be associated with serious discomfort and side effects, particularly for fragile subjects. Testing with blood-based biomarkers may have several advantages. First of all, blood collection for analysis of biomarkers appears to have high compliance rates.7,29 Second, testing with various blood-based protein biomarkers may lead to options of combinations, which appear to improve sensitivity and specificity as shown previously 9 as well as in this study. Third, new biomarker entities such as genomics,30–34 epigenomics,35,36 cell-free DNA,26,27 and metabolomics 37 may lead to combinations of biomarker panels, which may yield improved sensitivity and specificity. Fourth, due to noninvasive nature for analysis of blood-based biomarkers, testing can readily be repeated as needed; this may be an advantage in situations such as in monitoring treatment of subjects under surveillance due to risk of developing neoplastic diseases.13,14 Blood-based biomarker levels or presence may, however, be influenced by comorbidity and use of tobacco and/or alcohol. Application of combinations of various biomarker entities may reduce such influence and thereby improve interpretation of achieved results.
The strengths of this study include the total number of subjects recruited, the fact that the extracolonic cancers were identified within the same examination process due to the symptoms that led to colonoscopy, that none were lost to follow-up, that the study recruited subjects in 7 institutions of 2 regions in Denmark (1 rural and 1 nonrural), that the samples were collected and handled according to a validated SOP, and finally that the study adhered to guidelines including REMARK whenever possible. 25
The study has limitations, however, that must be taken into consideration in the interpretation. First of all, the numbers of specific extracolonic cancers were limited and therefore the statistical analysis was restricted to those with 10 or more cases. Second, the number of subjects selected for the study was reduced to 3732 from the original 4698 due to restrictions on samples availability; however, samples from all the 177 patients with extracolonic cancers were available. Third, the study was not initiated to identify biomarkers for detection of extracolonic cancers, and therefore, the extracolonic findings were achieved by subsequent follow-ups for those who had no colonic findings but had persistent symptoms.
Conclusions
The results of this study indicate that it may be possible to detect subjects that have an increased risk of extracolonic cancer following a clean colonoscopy as previously hypothesized. 13 It appears that combinations of various protein biomarkers may be useful to direct subsequent examination of patients with persistent symptoms, but who had clean colorectum at colonoscopy. These results, although preliminary, may form the basis for additional research directed both for primary examinations of subjects with symptoms of malignancy as indicated very recently by a multi-analyte blood test 28 and subsequent examinations to a clean bowel at colonoscopy. From a clinical perspective, such future results will be of major importance in subjects with symptoms of malignancy, as such testing may reduce the number of potential futile examinations.
Footnotes
Acknowledgements
The research nurses and biotechnicians at the participating hospitals are thanked for their skillful work with recruiting the subjects, collecting and processing the blood samples, and recording all the various data.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The study received financial support from The Augustinus Foundation, The Becket Fund, The Inger Bonnén Fund, The Hans & Nora Buchard Fund, CEO Jens Bærentsen (private donation), The Walter Christensen Family Fund, The P.M. Christiansen Family Fund, The Aase & Ejnar Danielsen Fund, The Erichsen Family Fund, The Knud & Edith Eriksen Fund, The Svend Espersen Fund, The Elna and Jørgen Fagerholt Fund, The Sofus Carl Emil Friis Fund, The Torben & Alice Frimodt Fund, The Eva & Henry Frænkel Fund, The Gangsted Fund, The Thora & Viggo Grove Fund, The Humanitarian Foundation, The Erna Hamilton Fund, The Sven & Ina Hansen Fund, The Søren & Helene Hempel Fund, The Henrik Henriksen Fund, The Jørgen Holm Family Fund, Foundation Jochum, The KID Fund, The Kornerup Fund, The Linex Fund, The Dagmar Marshall’s Fund, The “Midtjyske Bladfund,” The Axel Muusfeldt Fund, The Michael Hermann Nielsen Fund, The Arvid Nilsson Fund, The Obel Family Fund, The Krista & Viggo Petersen Fund, The Willy & Ingeborg Reinhard Fund, The Kathrine & Vigo Skovgaard Fund, The Toyota Fund, The Vissing Fund, The Wedell-Wedellsborg Fund and Hvidovre University Hospital (The Capital Region of Denmark). Abbott Laboratories Inc., Abbott Park, IL, USA, sponsored the protein analyses at their Center of Excellence, VUMC, Amsterdam, The Netherlands. The study was partly sponsored by an unrestricted grant from Abbott Laboratories Inc. to Hvidovre Hospital (H.J.N.).
Declaration of conflicting interests:
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: SG, XY, and GD are employees of Abbott Laboratories Inc. All others have no disclosures.
Author Contributions
Substantial contributions to the conception or design of the work; or the acquisition, analysis, or interpretation of data for the work: MW, lJC, LNJ, MRM, JV, MK, KTN, SL, GD, AMH, FM, HJN. Drafting the work or revising it critically for important intellectual content: MW, lJC, LNJ, MRM, SG, GD, HJN. Final approval of the version to be published: All the authors. Agreement to be accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved: All the authors.
