Abstract
Various members of the enteric bacteria in recent times are evolving diverse survival mechanisms for antibiotic therapy resulting in failure of treatment in infection and disease cases. The Enterococcus species are potential strains implicated in gastrointestinal tract infection and are recently evolving in the resistance mechanism. The study evaluates the occurrence of New Delhi Metallo-beta-lactamase 1 (NDM-1) amongst Enterococcus species using the phenotypic and genomic characterization of environmental strains in the Oghara water nexus. Presumptive isolates of Enterococcus species were retrieved from various sampled water sources and confirmed using polymerase chain reaction (PCR). Antibiotic susceptibility testing was conducted on confirmed isolates using Kirby-Bauer disk diffusion methods. The result reveals 63 genus isolates confirmed Enterococcus species, of which 42 (67%) were Enterococcus faecium, 15 (23%) were Enterococcus faecalis, and 6 (10%) were other Enterococcus species. Fourteen among the E. faecalis isolates show resistance to Ertapenem-EDTA, while 17 (44.8%) of the E. faecium show resistance to Ertapenem-EDTA to presumptively reveal their NDM-1 phenotype. The PCR detection of the NDM-1 gene further confirmed 23 (36.5%) isolates as positive genotypes amongst the isolates that previously showed presumptive NDM-1 phenotype. It was also observed that 10 (15.9%) of Enterococcus faecium members harbored the NDM-1 genotype, whereas 8 (12.7%) members of the Enterococcus faecalis harbored the NDM-1 genotype. The observation of such resistance determinants necessitates a call for the adroit application of relevant therapeutics in the management of related infections and an environmental health caution to prevent the spread of such resistance potential enteric bacteria pathogens.
Introduction
The water nexus has been described as a central distribution hub of diverse microbial pathogens in developing regions of the world with ardent incrimination of poor sanitation and hygiene.1-3 Various disease cases have been source-tracked with these concerns, yet insufficient attention and intervention strategies have been employed in these regards.4,5 Globally, an estimated 1.1to 2.6 billion people lack access to essential/safe water supply.5,6 Although the populace is aware of the water-related concerns, they have continued with diverse unhygienic practices—for example, washing meat with contaminated water, contact with effluents, animals exposed to contaminated water, indiscriminate disposal of waste etc. Such activities have created a significant challenge for water-related researchers as they encounter new daily problems as antibiotic-resistant organisms continue to emerge and re-emerge.7,8 In addition, there had been an imbalance in microbiota and the environment arising from unrestricted use of antimicrobials and wastewater release, giving rise to the emergence and spread of antimicrobial resistance.
One notable resistance reported in recent times is the New Delhi Metallo-β-lactamase 1 (NDM-1). It was first described by a Swedish national, Yong et al, 9 in the capital city of India. Klebsiella pneumoniae was the first organism reported to harbor the first documented NDM-1 gene. It is expressed in organisms that harbor them with the following features; a broad spectrum of aminoglycoside resistance enzymes such as acetyltransferases and methylases, and topoisomerase mutations leading to a high level of resistance5,10,11 resistance to carbapenem and β-lactam antibiotics were the produce carbapenemase and β-lactamase.5,11 NDM-1, like most enzymes, requires metal-based cofactors to carry out catalytic reactions (in the presence of the zinc ions), hence the term “Metallo.” 12 NDM-1 is involved in the inactivation of the beta-lactams ring while excluding aztreonam. Its monomeric molecular mass is 28 kda. The gene (blaNDM-1) is readily transferred from one bacterium to another via horizontal gene transfer. 13 Such metallo-based resistant phenotype/genotype has been reported amongst Gram-negative enteric bacteria (Klebsiella pneumonia, Pseudomonas aeruginosa, Escherichia coli, Morganella morganii, Acinetobacter baumannii, and Proteus mirabilis etc.) globally with high endemicity in the USA and Greece.5,14,15 Southern Europe, Germany and Asia have not been left out on the prevalence of NDM-1 and the metallo-based resistant determinants amongst Gram-negative strain. 16 The prevalence of NDM-1 has also been reported in the United Kingdom, Pakistan, India and other continents of the world as interest is growing to prevent its spread.14,16 In Africa, several investigators have reported the occurrence of NDM-1 amongst members of enteric bacteria, including Klebsiella species, Enterobacter cloacae, Citrobacter freundii, Serratia marcescens, Acinetobacter baumannii etc.17-21 in South Africa, Kenya, Ethiopia and other eastern/southern African countries. In Nigeria, NDM-1 has been reported amongst Gram-negative enteric bacteria in various geopolitical zone, including the Southwestern region (Lagos), Northeastern region (Maiduguri), South-southern region (Bayelsa), North-western region (Kano), and South-eastern region (Enugu) of the country22-29 however, non has reported NDM-1 amongst Gram-positive strains.
A study conducted in Spain by Nuñez et al 30 reveals high antibiotic resistance (to sulbactam, clavulanate, and tazobactam) associated with New Delhi Metallo-β -lactamase -1 (NDM-1) phenotype and genotype. Such resistant determinants have necessitated an increase in difficult-to-treat infections and disease cases which, according to some investigators, are on the rise amongst hospitalized patients 30 with a 10% increase in NDM-1 since 2003 in the US.
Various members of the Gram-negative enteric bacteria have been reported to harbor NDM-1 resistance but seldom reported among Gram-positive. However, one Gram-positive strain that has shown multiple emerging resistance tendencies is the Enterococci members. Enterococcus species are anaerobic facultative Gram-positive cocci. 30 They are significant potential pathogens that inhabit the specific environment, and it has been a group of fecal contamination indicator organisms and gastrointestinal tract infection. 30 Due to their high virulence determinants and tendency to evade antibacterial agents, including imipenem, monobactam and vancomycin, it has been a notorious species implicated in hospital and community-acquired infections opportunistic. 30 In diverse environments, the Enterococci members are part of common flora, making it challenging to differentiate colonization from actual infection and non- pathogenic in typical environment.31,32
Reports on the prevalence of Enterococcus in water nexus and some hospital environment in Africa has been few 33 with most NDM-1 reports on Gram-negative strains.22,23,34
There had been few reports of various resistance determinants, for example vancomycin resistant enterococcus (VRE), cephalosporin and other β-lactamase resistance in Africa. However, no report of NDM-1 has been reported in the water nexus of the Western region of Delta State, Nigeria to the best of our knowledge. In addition, there is also a dearth of information as regards Nigeria reports, with few cases reported in other African countries.22,23,34 Each year in the US, delays in treatment of 9300 water-related infections are caused by NDM-1 resistance gene,35,36 and approximately 600 deaths result in common types of NDM-1. 33 The increasing reports of Enterococcus species associated infection with diverse life-threatening disease cases are also contributing factors. To this end, this study characterized antibiotic resistance profile and NDM-1 producing Enterococcus species obtained from water sources in Oghara nexus using molecular biology techniques with a view to source-tracking such emergence of resistance determinants.
Materials and Methods
Study area and sample collection
The study was conducted within the suburban area, the Western Delta region of the State, which consisted of 4 communities, including Ajagbodudu, Oghara-efe, Oghara-eki, and Otefe. These communities were selected based on a presumed interest and susceptibility to possible antibiotic resistance outbreaks of related strains and multiple antibiotic resistance strains currently on the rise. The region is located at latitude (E 057°C and W 047°C). Using Nalgene 1 l sterile glass bottles, water samples were aseptically collected (once-off) from surface water (River ethiope, abattoir lake, other lakes, and swamp). Collected samples were transported to Lahor research laboratories, Nigeria, within 6 hours in a cooler box with ice packs for processing (microbiological and molecular analysis) between June and December 2019. Serial dilutions (101-103) of collected water samples were carried out using sterile distilled water. Briefly, test tubes with 9 ml of sterile distilled water were used in dilution, followed by adding 1 ml raw sample onto 9 ml of the sterile distilled water, and the test tubes were labeled 10−1. Each sample serially diluted solution was mixed using a vortex mixer. After that, 3 prepared dilutions of all collected samples were made in tubes from 100 to 10−3, while a volume of 0.1 ml was dispensed onto each labeled agar plate corresponding to the dilution factor. A sterile glass spreader was used to spread the pre-prepared water inoculum onto selective agar plates (Bile Esculin Azide Agar) (Merck Germany, www.merck-chemicals.com). Agar plates were incubated at 37°C for 24 hours. One colony per plate with black cultural morphology and colonial characteristics on a culture medium (culturonomics) was selected as a presumptively positive strain, while a single purified colony was used for DNA extraction using the crude isolate boiling technique.31,37
Genetic confirmation of Enterococcus genus
The enterococci members were identified/confirmed genetically based on the detection of the genus-specific tuf-gene (product size 112 bp). E. feacalis ATCC 19 433 was used as a positive control. The reaction mixture consisted of a final volume of 25 µL, which contains 5 µL of DNA template, 12.5 µL of a GoTaq (P)G2 master mix (Promega Corporation USA www.promega.com), 0.5 µL each of primer ent1 and ent2, and 6.5 µL of nuclease-free water. The sequences of primers used are as follows Ent1 5′-TACTGACAAACCATTCATGATG-3′ and Ent2 5′AACTTCGTCACCAACGCGAAC-3′. These sequences were synthesized by Inqaba Biotechnical Industries (Pty) Ltd. (www.inqababiotech.co.za). The PCR cycling conditions consisted of an initial denaturation of 94°C/3 min, amplification in 30 cycles (94°C/30 s, 53°C/45 s, 72°C/60 s) and a final extension of 72°C/8 min. The amplicons were detected using gel electrophoresis in 2% agarose stained with ethidium bromide and visualized using a UV transilluminator and photographed.31,37
Genetic confirmation of pathogenic species
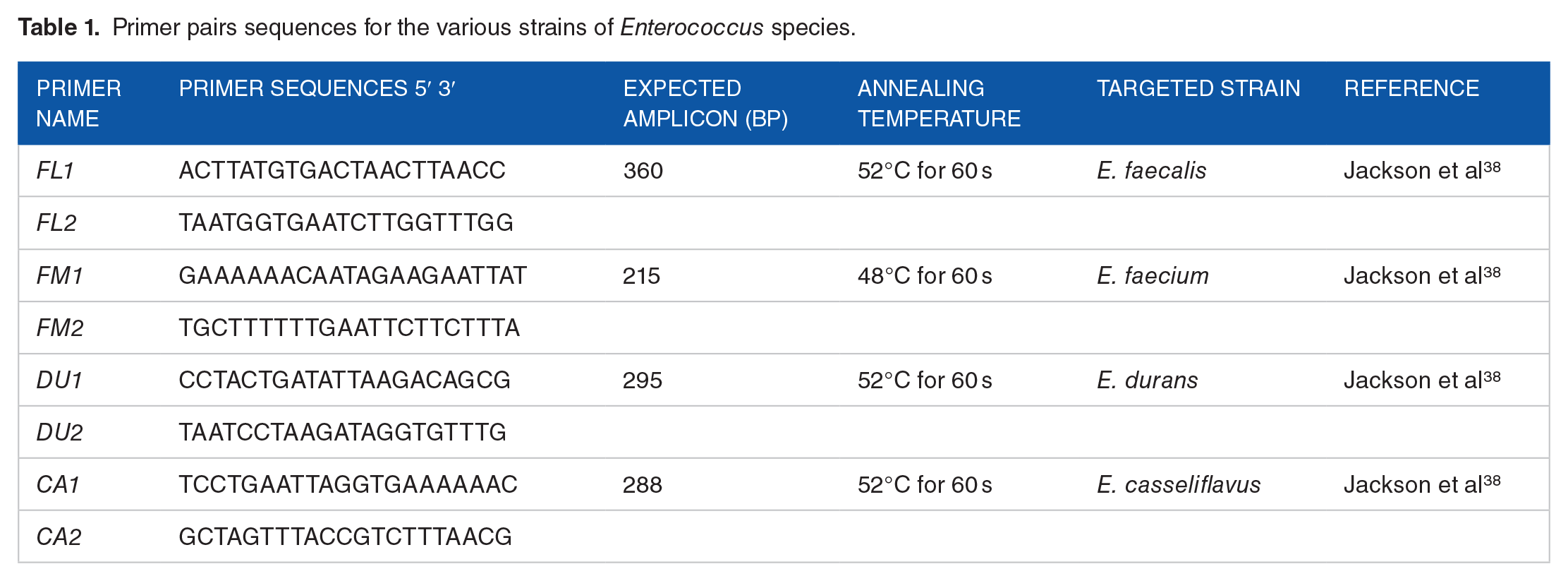
PCR technique was employed for the confirmation of Enterococcus (E) species. The species-specific primer pairs of E. faecalis, E. faecium, E. durans, and E. casseliflavus 35 were employed (Table 1). The reaction mixture consisted of a final volume of 25 µL, which contains 3 µL of DNA extract (as template nucleic acid), 12.5 µL of GoTaq master mix, 0.5 µL of 0.5 µM concentration of each primer pair, and 8.5 µL of sterile nuclease-free water. The working solutions of all employed molecular biology consumables and premix preparations were aliquot, secured and stored in a refrigerator before analysis, and thawed in ice boxes in a biosafety cabinet to prevent contamination during analysis.
Primer pairs sequences for the various strains of Enterococcus species.
Antibiotic susceptibility testing of Isolates
Confirmed isolates were subjected to antibiotic susceptibility testing using the carbapenem penicillin and cephalosporin groups of antibiotics vis: Carbapenems: {Ertapenem (ETP-10 μg)}, {Imipenem (Imi-30 μg)}, {Doripenem (Dor-10 μg)}, {Meropenem (Mem-10 μg)}, Penicillins: {Ampicillin (Amp-10 μg)}, Betalactam/β-lactamase inhibitors: Piperacillin-Tazobactam (PTZ-110 μg)}, Cephalosporins or Cephem: {Cefotaxime (CTX-30 µg)}, {Ceftazidime (CAZ-30 µg)}, {Ceftriaxone (CRO-30 µg)}, {Cefuroxime (CXM-30 µg)}, {Cefazolin (CZ-30 µg)}, {Cephalexin (CFX-30 µg)}, {Cefepime (CPM-30 µg)} {Cephalothin (KF-30 µg)}.These antibiotics disks were purchased from the Davis diagnostics (Pty) Ltd, 141 Oak Avenue, Ferndale, Randburg, 2194, Gauteng, South Africa (www.daviesdiagnostics.co.za).The antibiotic susceptibility test was interpreted as sensitive (S), Resistance (R) and intermediate (I) using the CLSI (Clinical and Laboratory Standards Institute) guidelines. 39
Strains phenotypic confirmation of NDM-1 phenotype/carbapenem resistance
A further test to determine the NDM-1 phenotypic resistance was applied using the Modified Hodge Test (MHT) method and the EDTA-Ertapenem synergy test.32,39 First, isolates which were resistant to Ertapenem (ETP-10), and cephalosporin such as cefotaxime (CTX-30), ceftazidime (CAZ-30), and ceftriaxone (CRO-30) and those strains that show ⩾4 mm zone of clearance with EDTA enhanced disk were presumptively selected to harbor carbapenemase phenotype. The EDTA-Ertapenem synergy test was then conducted to detect carbapenemase in a secondary detection as those strains which show ⩾4 mm zone of clearance with EDTA enhanced disk reveals an indication for carbapenemase production. This was further confirmed by the MHT method as previously described by various investigators.40-42
Genetic confirmation of blaNDM-1 resistance amongst Enterococcus spp
All presumptive carbapenemase or Metallo- β -lactamase-producing Enterococcus strains were genetically analyzed for blaNDM-1 resistance determinants using PCR. A 25 µl final volume reaction mixtures which contain 12.5 µL GoTaq PCR master mix reagents (www.promega.com), 0.5 µL volume of 0.5 µM concentration of each primer pair as blaND-MβL-1-forward and blaND-MβL-1- reverse primers{(F: 5′ GGG CAG TCG CTT CCAACG GT 3′ while R: 5′ GTA GTG CTC AGT GTC GGC AT 3′}, 3 µL of DNA extract (as a template nucleic acid/DNA), and 8.5 µL of sterile nuclease-free water, was observed to produce an expected amplicon size of 475 bp. 43 The reaction mixture was prepared in an ice box in a biosafety cabinet, aliquot into a nuclease-free 200 μL microfuge tube and arranged in a thermal cycler (Bio-Rad T100™ thermal cycler, www.lasec.co.za, SA). The reaction cycling condition was as follows: initial denaturation at 94°C for 2 minutes, followed by 35 cycles at 94°C for 30 seconds, 55°C for 30 seconds, and 72°C for 30 seconds. A final extension step of 72°C for 10 minutes was conducted and stored at 4°C before detection. The PCR products or amplicons were then electrophoresed (machine CLS-AG100, Warwickshire, UK) in 1.5% (wt/vol) agarose gel incorporated with 4 μL/100 mL of 0.5 mg/L ethidium bromide, followed by mixing while avoiding the formation of air bubbles and pouring/casting in the mini gel casting (Anachem, UK) tray and visualized/detected 44 in gel documentation system (Bio-Rad, USA). The control strain Escherichia coli ATCC 25 922 and Klebsiella pneumoniae (CCBH13327) were used as negative and positive control strains during gene detection.
The limitations of the study
The study did not employ the micro broth dilution method for MIC of antibiotics. However, the Kirby-Bauer disk diffusion test method was applied to determine antibiotic susceptibility testing. The choice of applied method was informed by access to the available materials at our disposal during the study. The study did not confirm the horizontal transferability of such resistant genes amongst other environmental strains. However, our ongoing studies in our lab would access the presence of plasmid and the sequence of positively detected strains with diverse resistance markers and plasmid types. Whole genome sequencing and/or partial genome of the positive strains were not conducted since the private/individual funds available to us during the study would not be able to take care of such aspects of the study. External funds are currently being sourced to continue in this study area. We hope to show the reports in the near future publicly
Results
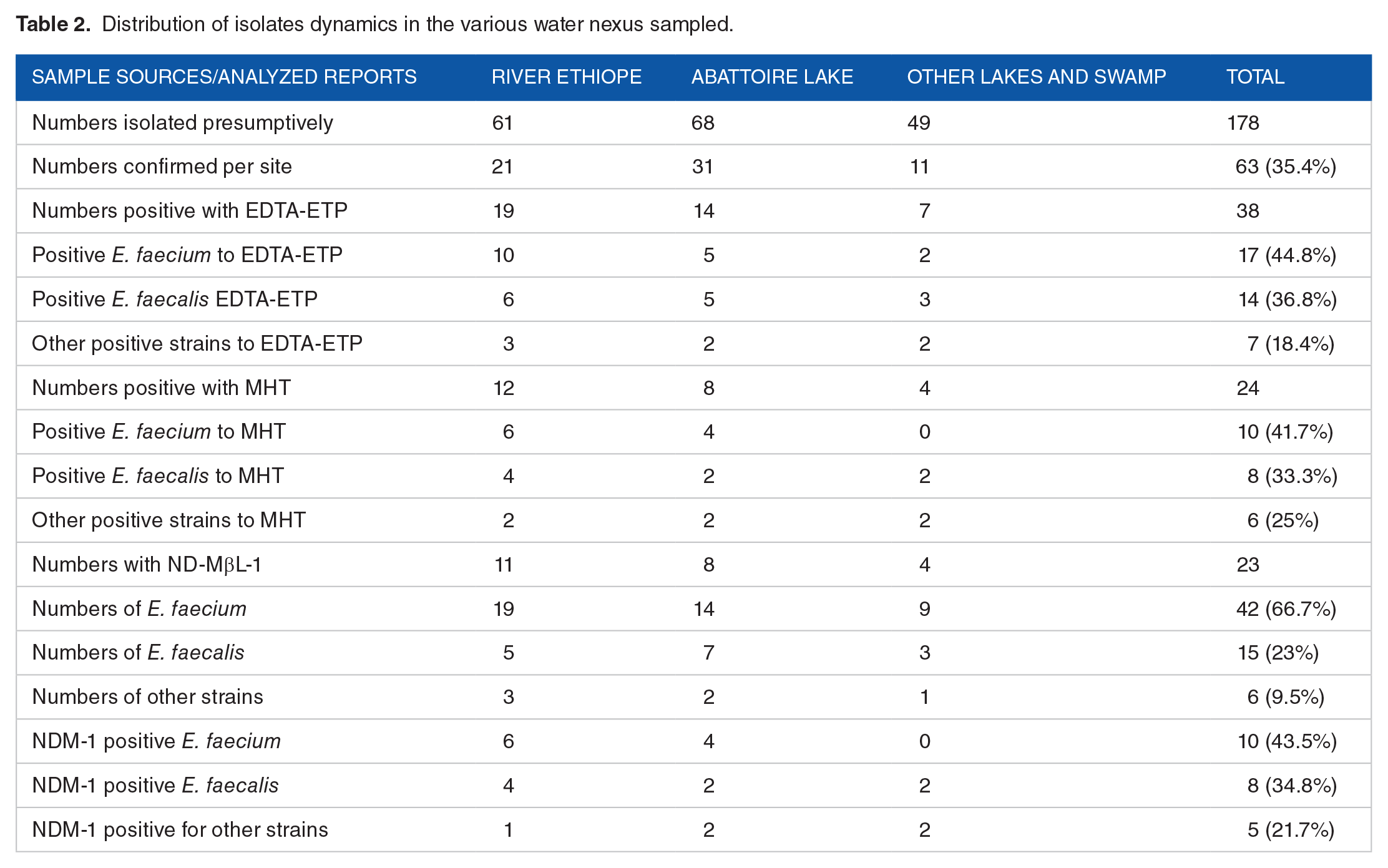
Our study recovered 178 presumptive isolates of Enterococcus specie from the various surface water samples collected as follows: River ethiope (61 isolates), abattoir lake (68 isolates), other lakes and swamp (49 isolates), with 63 (35.4%) confirmed E. species. The presumptive isolates were selected based on their culturonomics in the employed media detailed in Table 2. Amongst the confirmed isolates, 42 (66.67%) were E. faecium, 15 (23.81%) were E. faecalis, and 6 (9.52%) were other Enterococcus species. Figures 1 to 3 show various expected amplicon size photographs of the agarose detection. High antibiotic resistance to ampicillin was observed amongst E. faecium compared to E. faecalis strains. Other resistance signature include: piperacillin/tazobactam (PTZ-110 μg) (21.4%), ampicillin (Amp-10 μg) (100%), ertapenem (ETP-10 μg) (57.1%), cefotaxime (CTX-30 μg) (47.6%), ceftazidime (CAZ-30 μg) (33.3%), and ceftriaxone (CRO-30 μg) (42.9%) amongst E. feacium as depicted in Table 3 below, while piperacillin/tazobactam (PTZ-110 μg) (26.7%), ampicillin (Amp-10 μg) (66.7%), ertapenem (ETP-10 μg) (46.7%) cefotaxime (CTX-30 μg) (53.3%), ceftazidime (CAZ-30 μg) (46.7%), and ceftriaxone (CRO-30 μg) (53.3%) were observed amongst E. faecalis as shown in Figure 4 below. The PCR of ND-MβL-1 gene detection confirmed 23 (36.5%) isolates in Figure 5, which were previously positive for the ND-MβL-1 phenotype and carbapenem resistance.
Distribution of isolates dynamics in the various water nexus sampled.
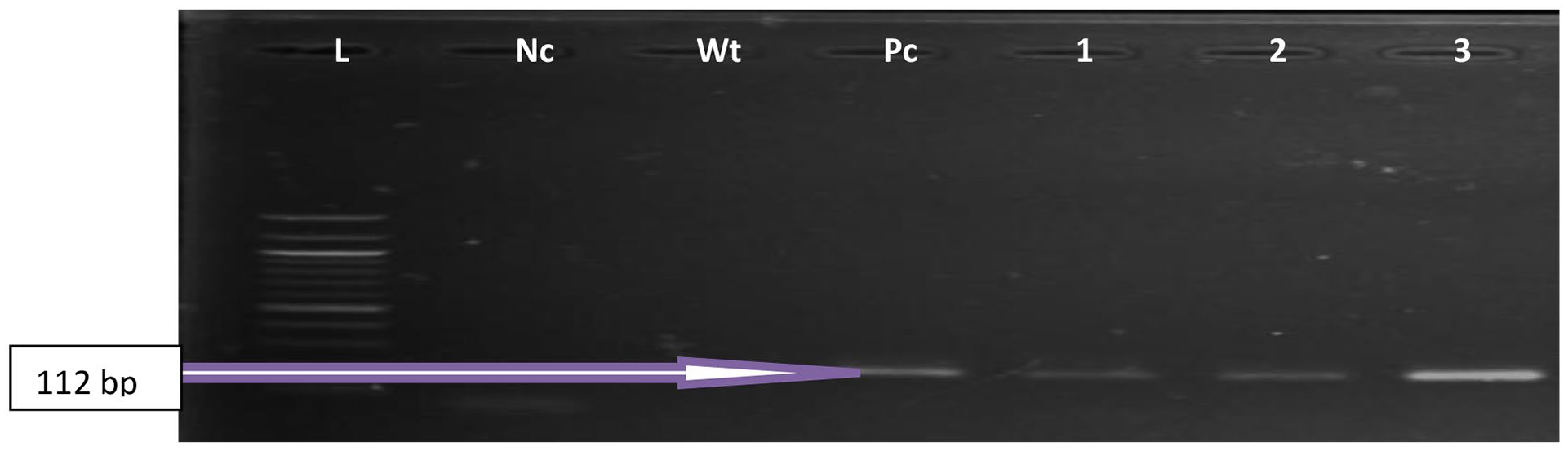

Shows an agarose gel analysis of the PCR products of confirmed Enterococcus species targeting the 16S rRNA gene or ENT1/ENT2 (112 bp). Lane L: DNA Ladder (100 bp), Lane Pc: The positive control, Lane Nc is the negative control, Lane Wt is sterile water, and Lanes 1 to 3 are the representative samples.
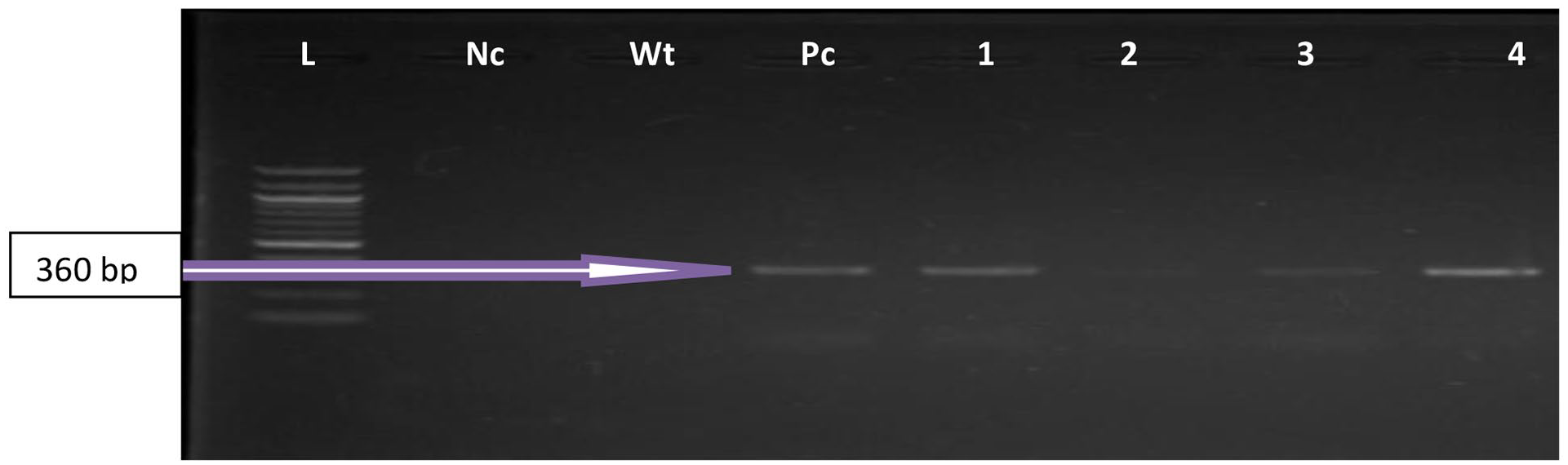
Shows an agarose gel electrophoresis photo of the PCR products of some confirmed Enterococcus feacalis strains (FL1/FL2) at 360 bp. Lane L: DNA Ladder (100 bp). Lane Pc: The positive control, Lane Nc is the negative control. Lane Wt is sterile water while, Lanes 1 to 4 are the representative positive strains.
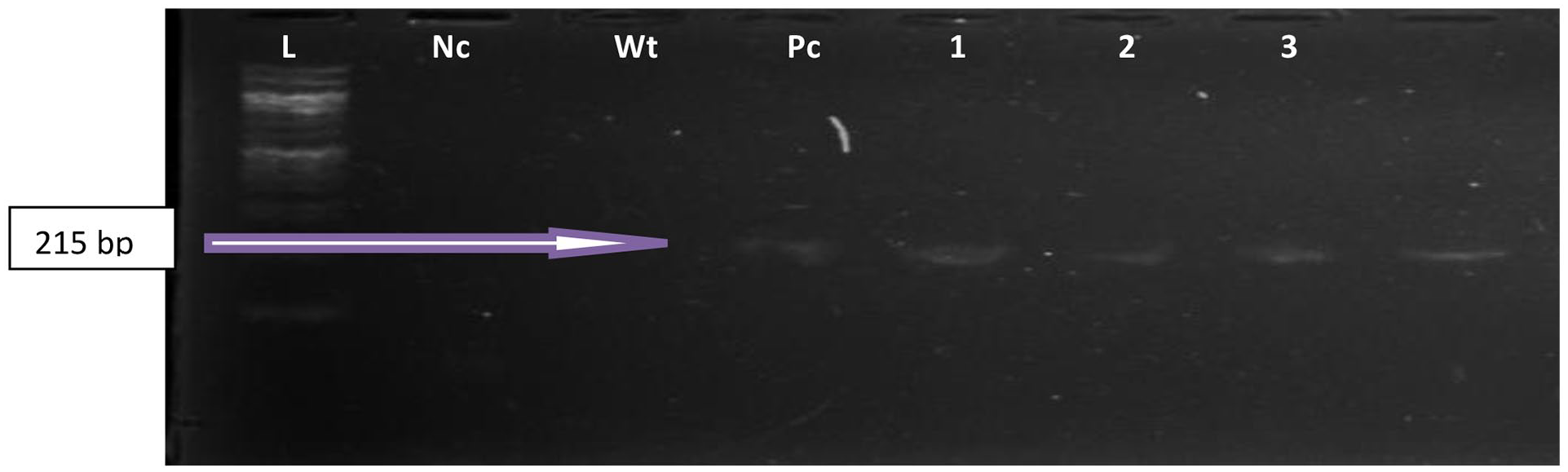
Shows an agarose gel electrophoresis photo of the PCR amplicons (products) of representative confirmed Enterococcus feacium strains (FM1/FM2) at 215 bp. Lane L: DNA Ladder (100 bp), Lane Pc: The positive control, Lane Nc is the negative control, Lane Wt is sterile water, and Lanes 1 to 4 are the samples.
The prevalence and phenotypic profile of E. faecium isolate.
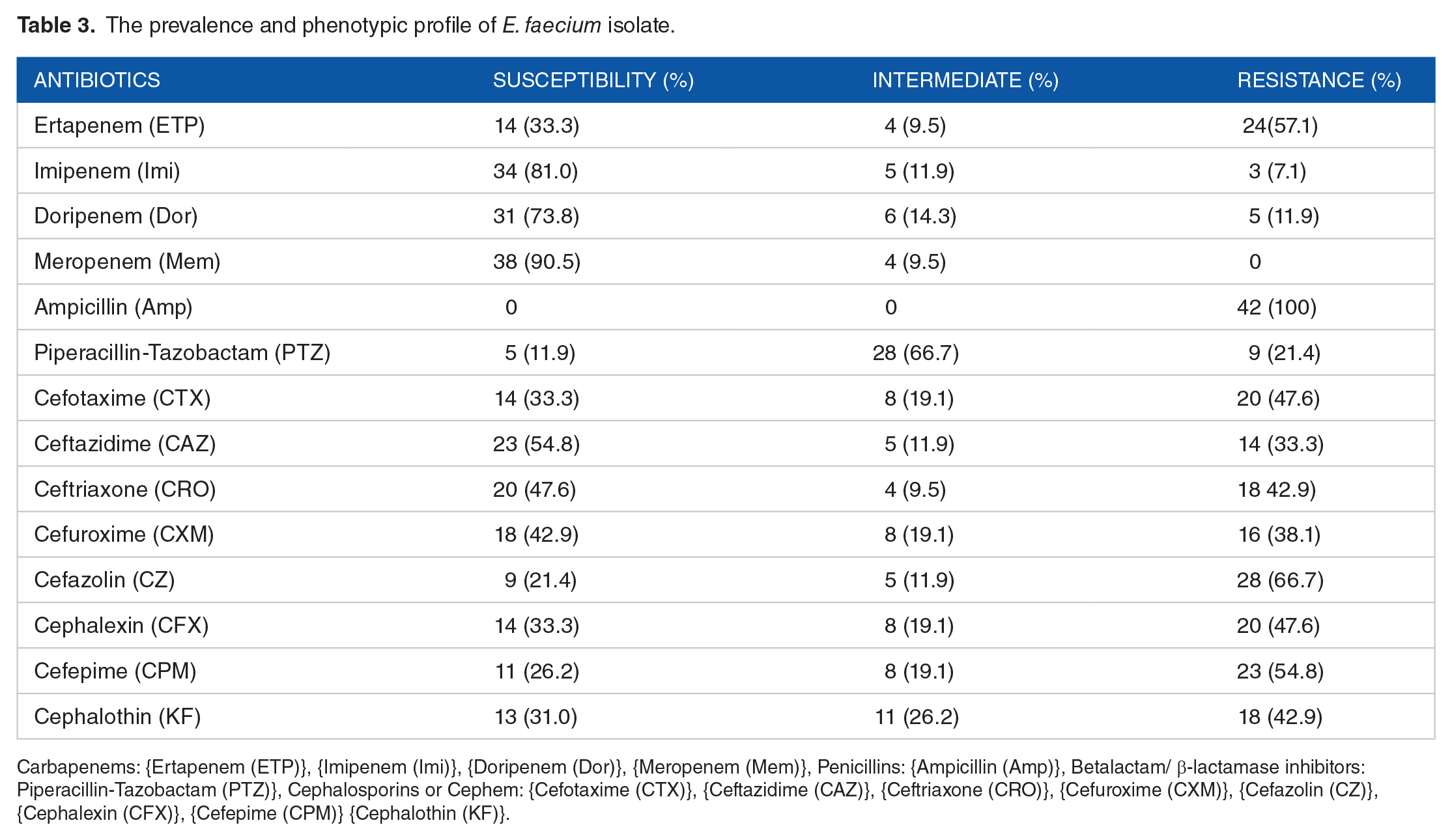
Carbapenems: {Ertapenem (ETP)}, {Imipenem (Imi)}, {Doripenem (Dor)}, {Meropenem (Mem)}, Penicillins: {Ampicillin (Amp)}, Betalactam/ β-lactamase inhibitors: Piperacillin-Tazobactam (PTZ)}, Cephalosporins or Cephem: {Cefotaxime (CTX)}, {Ceftazidime (CAZ)}, {Ceftriaxone (CRO)}, {Cefuroxime (CXM)}, {Cefazolin (CZ)}, {Cephalexin (CFX)}, {Cefepime (CPM)} {Cephalothin (KF)}.
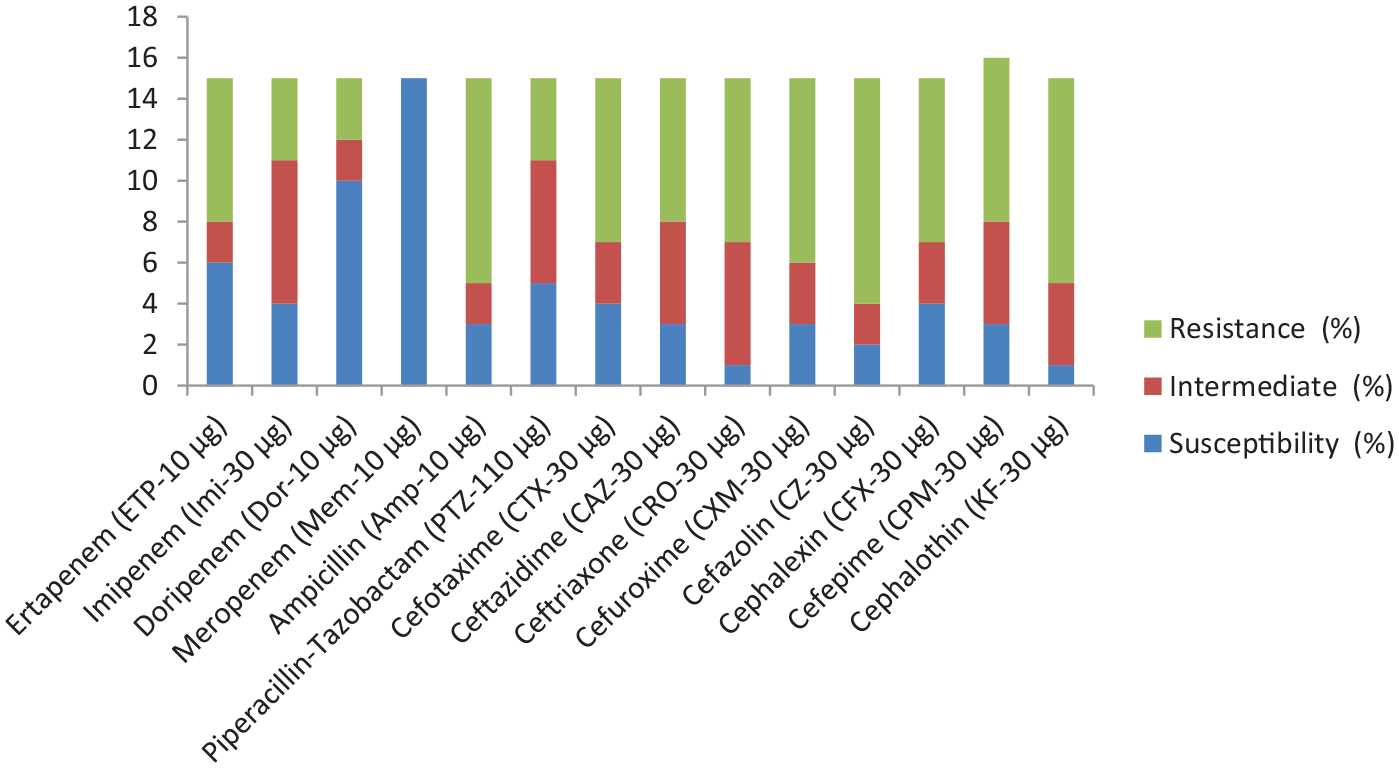
Showing the prevalence and phenotypic antibiotic profile of 21E. faecalis Isolates, S indicates susceptibility, I indicate intermediate, while R indicates resistance phenotype as show by the various strains.
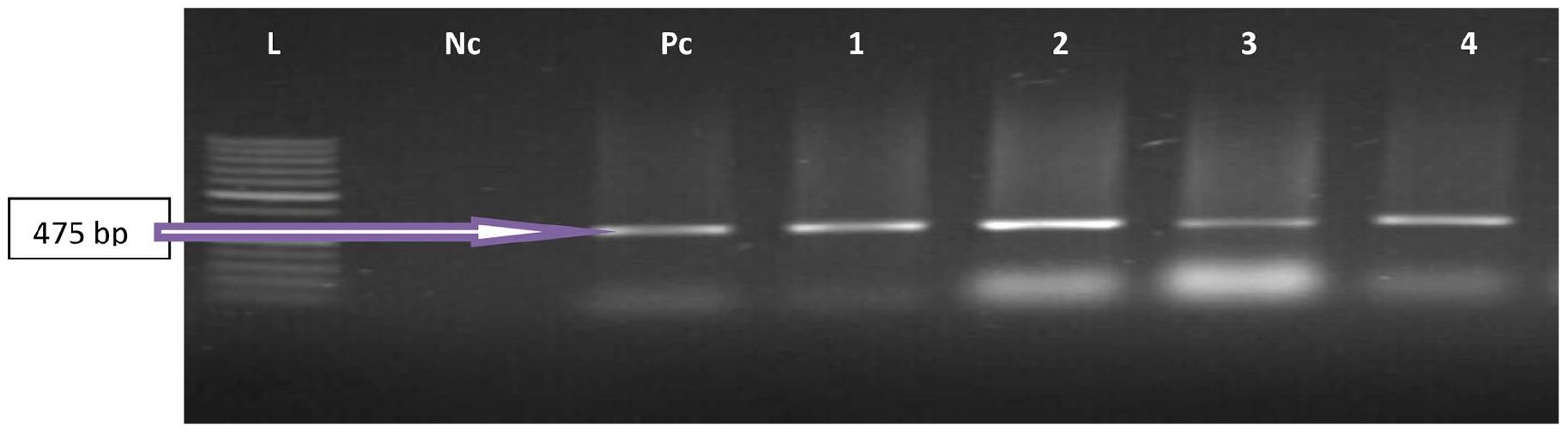

Shows an agarose gel photo of PCR products of representative confirmed New Delhi Metallo-beta-lactamase 1 (ND-MβL-1) amongst Enterococcus feacalis and Enterococcus feacium at 475 bp. Lane L: DNA Ladder (100 bp). Lane Pc: The positive control, Lane Nc is the negative control, and Lanes 1 to 4 are positive samples.
Discussion
Following the previous reports of researchers on the spreading nature of antibiotic resistance determinants and distribution in the water environment, there have been numerous emerging resistance determinants amongst potential pathogens of public health concern. Among such organisms are the enterocyte infecting members to which the Enterococcus species belong. This study has observed Enterococcus species in diverse surface water sources revealing its thriving tendency within the study area. It is important to note that Enterococcus species are bacteria reported as potential indicator strains of fecal contamination. Its presence in our study reflects a potential risk to human health and users of the sampled waters. Our results shown that amongst the 178 presumptive isolates of Enterococcus species recovered from the various surface water samples collected, 63 (35.4%) of them were confirmed E. species. The photograph of isolates agarose gel detection of amplicons confirmed isolates 66.7% (42) were Enterococcus faecium, 23.8% (15) were Enterococcus faecalis, and 6 (10%) were other Enterococcus species. Therefore, our study shows that E. faecium is the most prevalent, followed by E. faecalis. Such an evident report indicates that the environmental water nexus possesses enterococcal contaminants, which are traceable to fecal contamination.8,41
In an earlier study by Iweriebor et al, similar strains were reported, including other strains of Enterococcus species such as E. casseliflavus, E. avium, E. sgallinarum, Enterococcus durans, and E. raffinosu.31,33 Other investigators have also reported E. faecalis and E. faecium as the most prevalent species in both water environments and hospital or public health systems which may be source-tracked and/or traced with major nosocomial infections.16,37,45 During the study, multiple resistant phenotypes were observed amongst the confirmed isolates, which borders on the carbapenem group of antibiotics and the β-lactam antibiotics group. It was found that 92% of the confirmed isolates exhibited resistance toward ampicillin in addition to other antibiotics (carbapenem) resistance phenotype, including piperacillin/tazobactam, ertapenem, imipenem, doripenem, ceftazidime, ceftriaxone etc. Due to its broad-spectrum nature, such resistance phenotype was not reported for meropenem. However, the intermediate phenotypic range on meropenem and resistance in other carbapenem antibiotics indicates resistance determinants. Such a high level of antibiotic resistance phenotype to the carbapenem antibiotics has become a significant cause of health concern as it resulted in aggravated infection cases and a challenge to treat situation11,37 in a disease implication. Following the reports of Shah et al and his group on related studies, it was affirmed that such increased resistant phenotypes might result in increased volume and incidence of hospitalized patients. 46
The PCR and molecular gene detection of resistance to New Delhi Metallo-β-lactamase-1 (NDM-1) further confirmed 23 (36.5%) positive isolates to the targeted resistance gene, affirming the presence of the gene amongst environmental strains. This indicates the emergence of Enterococcus harboring strains in the water nexus of Oghara. It also implies the emergence of NDM-1 amongst Gram-positive strains in the study environment. Although reports of NDM-1 amongst Gram-positive strains are seldom documented, this study has shown the need for appropriate documentation and epidemiology of such resistance determinants, especially in the water environment. This is similar to Ranjan and Thatikonda 43 and Khan and Mustafa 44 in which the ND-MβL-1 resistance gene is spreading in the water environment, especially amongst Gram-negative strains. Although our study did not access the horizontal transferability of the resistance gene, studies by diverse investigators have also indicated that the water nexus is a potential hub for sharing such genotype even amongst other unrelated potential pathogens of environmental health relevance.42,44,47 Observing such resistance phenotype and genotype amongst environmental strains present the isolates as potential pathogens of environmental risk, as previously depicted in the reports of the center for disease control. 33 Such risk may be likened to the previously reported disease cases, which affected more than 9300 patients in the US, with 600 deaths due to ND-MβL-1 resistance gene.5,25,26,33,42,44,46 Therefore, it is imperative for respective personnel within the study region that uses these water sources for diverse activities to stop their usage without adequate treatment. It is also important to point out that good hygienic practices be implemented should there be any contact with the specified water sources.
Conclusion
The study evaluates the occurrence of New Delhi Metallo-beta-lactamase 1 (ND-MβL-1) amongst Enterococcus species in the water nexus of Oghara Delta State as the emergence of environmental ND-MβL-1-harboring enterococci bacterial strains. Diverse phenotypes and genotypes of the resistance determinant were observed amongst the isolates of Enterococcus, which were recovered from water bodies. Observing/reporting such resistance determinants amongst Enterococcus members necessitates a call for prompt control action, especially on carbapenem and β-lactam, with the need for further ardent studies in this area. It is important to note that the potential pathogen and its emerging resistant genotypes/phenotypes are spreading in the water environment. There is a need for routine surveillance and monitoring of the water nexus within the study area. It is also of paramount importance that appropriate implementation of hygienic practice is employed amongst the general public to nip the spread of such resistance phenotype and genotype currently spreading in our environment and/or water nexus.
Footnotes
Acknowledgements
Authors wish to appreciate the efforts of microbial researchers and investigators in Lahor Research Laboratory Benin City, Edo State and Western Delta University Oghara Delta State in ensuring adequate sample processing and molecular analysis of the isolates are conducted as required.
Funding:
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Declaration of Conflicting Interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
IBE conceived and designed the study, IBE, OH, and GO carried out the study, analyzed and interpreted the data, IBE drafted the manuscript, while IBE, OH, and GO revised the manuscript. All authors read and approved the final manuscript.
Ethics Approval Statement
The Ethics Research Committee of Western Delta University approved the protocol for this study.
Data Availability Statement
The datasets used for this study are available from the corresponding author on reasonable request.
