Abstract
A serious concern of public-health proportion is rising from the carriage of antibiotic resistance determinant in Non-Lactose-Fermenting Bacteria and acquisition of virulence particularly in strains that are not routinely isolated or screened from common food animals. This study investigated the resistance profile and pathogenicity potential of selected Non-Lactose-Fermenting Bacteria isolated from 18 poultry farms. In total, we investigated the antibiotic susceptibility patterns of 25 Pseudomonas lactis and 71 Pseudomonas paralactis isolated from chicken faeces by testing them against 12 antibiotics. Resistance genes borne by the selected isolates were screened by sequencing the genetic location of resistance determinants was determined by plasmid curing. The virulence potential of the studied strains was determined phenotypically. Pseudomonas lactis isolates were mostly resistant to azetronam (93%), followed by trimethoprim (90%), cefotaxime (86%) and then amoxicillin/clavulanic acid (57%), while Pseudomonas paralactis. isolates were most resistant to azetronam (94%), trimethoprim (90%), cefepime (80%), piperacillin (75%) and amoxicillin/clavulanic acid (70%). The Multiple Antibiotic Resistance Index of Pseudomonas lactis and Pseudomonas paralactis isolates respectively ranged from 0.0 to 0.8 and 0.0 to 0.9. Polymerase Chain Reaction revealed the presence of antibiotic resistance factors such as blaCTX-M, qnrS, aac (6′)-lb-cr and blaSHV while plasmid curing revealed carriages of resistance determinants on Resistance Plasmid. Moreover, virulence enzymes such as alkaline protease and phospholipase C were found in 3% and 12% of Pseudomonas paralactis and Pseudomonas lactis, respectively. This study reports the first occurrence of Pseudomonas lactis and Pseudomonas paralactis strains from chicken faeces, and their antimicrobial resistance and relative virulence suggest the encroachment of food animals by the under-studied non-lactose-fermenting bacteria that should alert public health concerns.
Keywords
Introduction
Non-lactose-fermenting bacteria (NLFB) are mostly reported as aetiological agents of zoonotic infections and they are posing ever-increasing threats to socio-economies as newer strains emerge amongst them and wreak havoc on the health sector. 1 NLFB are unique in their ubiquity and distribution, being reported in niches from plants, water, soil, humans and food animals. Importantly, they are also implicated in food-borne infections which have grown to public health proportions. 2 Transference of infectious NLFB to humans is considerably common as a result of the dependence of the large human population on food animals, as reported from the consumption of undercooked meat and raw eggs from poultry animals. 2 Direct transference to humans aside from food contamination has also been reported. 3
The Pseudomonas species are a classic NLFB species, extremely diverse phylogenetically, and have been reported as agents of chicken putrefaction. 4 Depending on the substrate, different Pseudomonas species colonise different animal foods such as milk, fish and meat products. 5 Extensively diverse Pseudomonas spp. have been isolated from chickens, 4 and chickens are also high-risk food animals which are a reservoir of Pseudomonas-associated opportunistic diseases that spread widely through the flock and are therefore of significant zoonotic importance to humans. 6 Aside from their being a reservoir of diverse Pseudomonas spp., colonisation of animal food sources by previously-reported novel species (sp. nov.) of Pseudomonas strains is increasingly being reported. Pseudomonas carnis sp. nov. has been isolated from meat, 7 Pseudomonas paracarnis sp. nov. from refrigerated beef, 8 both Pseudomonas weihenstephanensis sp. nov. and Pseudomonas helleri sp. nov. from raw cow meat, 9 Pseudomonas sivasensis sp. nov. from fishery farm 10 and both Pseudomonas lactis sp. nov. and Pseudomonas paralactis sp. nov. from bovine raw milk. 11
Both Pseudomonas lactis sp. nov. and Pseudomonas paralactis sp. nov. were previously isolated as part of psychrotrophic bacterial from bovine raw milk and characterised as round, smooth, rod-shaped, Gram-negative, non-sulphide-producing, indole-negative, oxidase-positive, glucose-positive, mannitol-positive and sucrose-negative. Their 16S rRNA gene sequence GenBank accession numbers were also reported as NR_156986.1 and NR_156987.1, respectively. 11 For Pseudomonas lactis sp. nov., genomic characterisation on the bases of genomic size (Mb), DNA GC content (%), number of genes and number of coding sequences were 6.73 Mb, 60.0%, 6038 and 5811, respectively. 11 For Pseudomonas lactis sp. nov., the values for the respective genomic parameters were 6.02 Mb, 59.7%, 5341 and 5167. 11 Respectively, the draft genome sequence of Pseudomonas lactis sp. nov. and Pseudomonas paralactis sp. were registered in the GenBank with the accession numbers JYLO00000000 and JYLN00000000. 11 Unguided application of antimicrobial in bovine husbandry and medication had produced selective pressure in bovine bacteria to several antibiotics 12 and, generally, a commensal-to-pathogen transition of bacterial strains from nucleotide rearrangement is not uncommon. 13 However, currently, neither has the antibiotic resistance profile nor potential virulence of both Pseudomonas lactis and Pseudomonas paralactis been investigated and therefore are unknown.
In this study, with 16S rRNA gene sequencing similarity score of ⩾97% we report, for the first time, the isolation of both Pseudomonas lactis and Pseudomonas paralactis from chicken faeces from poultry farms in South West Nigeria.
The indiscriminate administration of antimicrobials in food animals has drastically raised the prevalence of antibiotic-resistant pathogens and commensal as well, increasing the dangers of the transmission of multidrug-resistant pathogens to humans by significant proportions. 14 Very importantly, the acquisition of resistance through abuse of antimicrobials and consequent horizontal spread of antibiotic resistance genes across the community by resistance plasmids pose serious problems because there is an established association between multidrug resistance and virulence, an observed phenomenon in which erstwhile commensal and innocuous bacteria acquire virulence factors as they develop mechanisms for multidrug resistance. 15
There are accounts of under-studied and less-common NLFB that have evolved as opportunistic pathogens of clinical significance. 1 Unfortunately, their mechanisms of resistance remain largely unknown as the strains are not usually covered in routine antibiotic resistance screening studies. The aim of this study was therefore to determine both the pathogenicity potentials and plasmid-mediation of antibiotic resistance pattern of the studied Pseudomonas lactis and Pseudomonas paralactis strains isolated from chicken faeces in the studied population.
Materials and Methods
Bacterial isolation and identification
A total of 337 fresh faecal samples were collected from 337 healthy chickens (broilers and layers) from 18 poultry farms in 6 states in South West Nigeria, namely Ogun, Osun, Oyo, Ekiti, Lagos and Ondo. Microbiological analysis was carried out at the laboratory of the Department of Pharmaceutical Microbiology, Faculty of Pharmacy, University of Ibadan. All possible non-lactose-fermenting colonies on MacConkey plates were selected based on differing morphology and subjected to biochemical characterisation while genomic DNA was isolated from colonies by boiling method as previously described. 16 Briefly, freshly grown colonies were suspended in 500 μL of sterile distilled water in an Eppendorf tube, vortexed and centrifuged for 3 minutes at 10 RCF, decanted and resuspended in 100 μL of sterile water and heated in a heating block at 100°C for 15 minutes. The bacterial solution was centrifuged at 10 RCF for 5 minutes, after which 50 μL of the supernatant was carefully collected with a micropipette into a sterile Eppendorf tube without picking the cell sediment or disturbing the suspension. The collected supernatant constitutes the genomic DNA and is stored at −20°C for further analysis. Identification of strains was carried out by PCR amplification and sequencing of the 16S rRNA gene of the extracted DNA template by PCR thermocycler condition previously described. 17 Primers for amplification of the 16S rRNA gene are indicated in Table 1. Amplicons of the expected size for the 16S rRNA gene were purified and sequenced using the 16S rRNA sequencing method (ABI PRISM 377 DNA Sequencer). Sequence results were evaluated with the BLAST suite against the National Centre of Biotechnology Information database ( http://blast.ncbi.nlm.nih.gov/Blast.cgi ). All identified Pseudomonas lactis and Pseudomonas paralactis were selected and subjected to antibiotic susceptibility testing.
The primer used for the amplification of 16S rRNA genes from the isolates.

Biochemical and growth characterisation of studied strains
Biochemical characteristics of the studied strains were described according to carbohydrate fermentation to selected sugars, indole, oxidase, Gram reaction tests and hydrogen sulphide production by the protocol described elsewhere. 18 The growth of studied strains at 4°C and 42°C were tested by inoculation on nutrient agar and incubation at the respective temperatures for 18 to 24 hours.
Antibiotic susceptibility testing
Antibiotic susceptibility testing of studied strains was carried out by the Kirby-Bauer disc diffusion method and data was interpreted based on CLSI guidelines. 19 The 12 antibiotics used belonged to different antimicrobial classes, namely: fluoroquinolone (ciprofloxacin, 5 µg), monobactam (Aztreonam, 30 µg), aminoglycoside (gentamycin, 10 µg), carbapenem (ertapenem, 10 µg), generations of cephalosporin (cefoxitin, 30 µg; ceftazidime, 30 µg; cefotaxime, 30 µg and cefepime 30 µg), penicillin (piperacillin, 30 µg), β-lactam inhibitors (amoxicillin/clavulanic acid, 30 µg and piperacillin/tazobactam, 110 µg). Pseudomonas aeruginosa ATCC 27853 was used as the control. Data for antibiotic susceptibility testing are included in Supplemental Tables S1a and S1b.
Multidrug-resistant (MDR) isolates were defined as those resistant to more than 3 different classes of antibiotics. For each isolate, the multiple antibiotic resistance index (MAR) was calculated as previously described 20 by the formula:

Phenotypic evaluation of ESBLs production by the studied strains
ESBL producers were screened amongst the studied strains by the Double Disc Synergy Test as described elsewhere. 21 In summary, 0.5 McFarland standard of the studied isolate was inoculated evenly on sterile Mueller Hinton agar, allowed to aseptically air-dry in a laminar flow cabinet, and antibiotics discs of cefotaxime (30 µg) and ceftazidime (30 µg) placed 15 mm apart each other and synergistic effect of either of the cephalosporin observed with amoxycillin/Clavulanic acid combination disc (20:10 µg). Enhancement of the zone of inhibition of either of the cephalosporins towards the beta-lactam/beta-lactamase indicator was indicative of a positive result.
Detection of resistance determinants
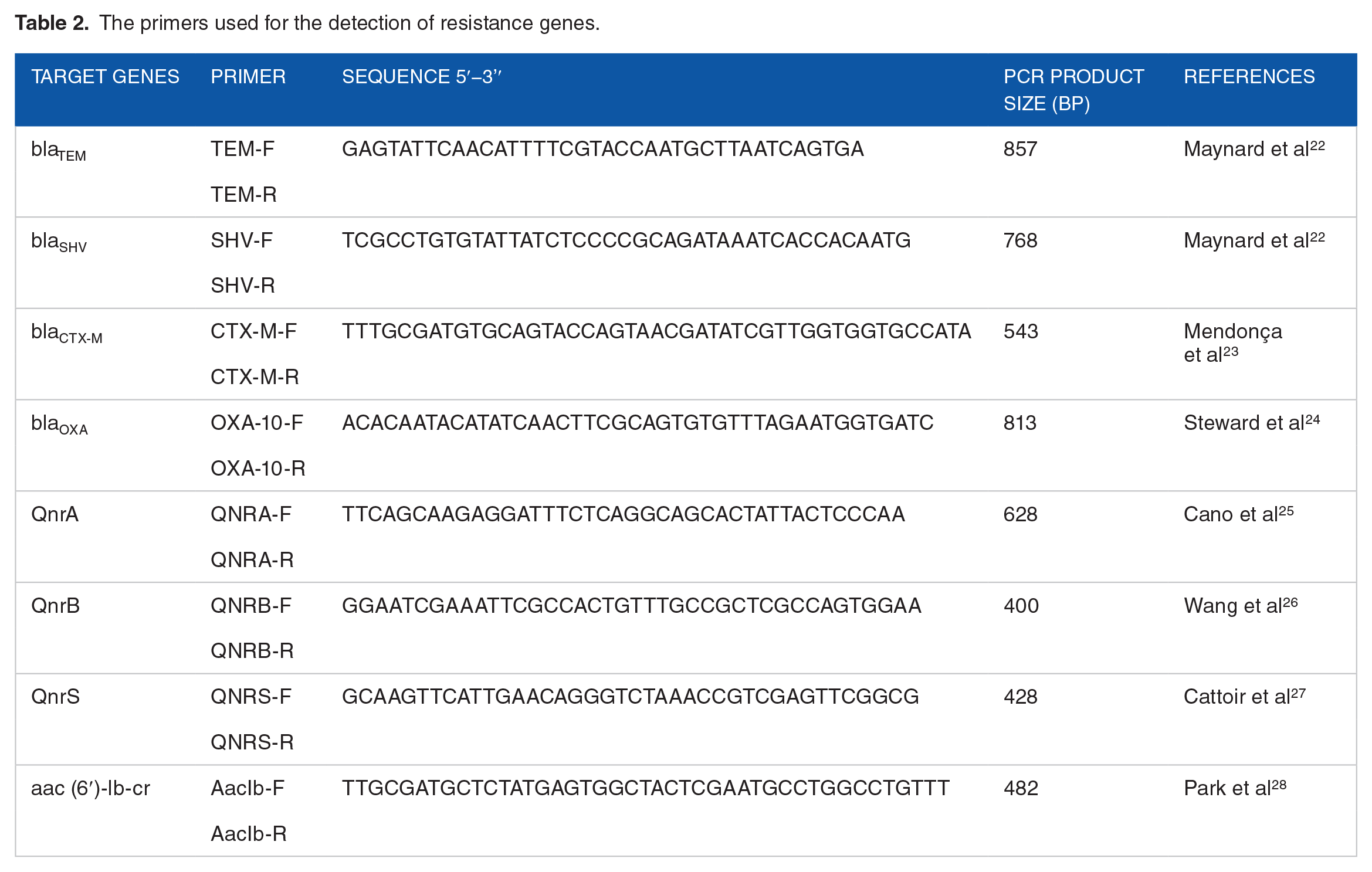
Screening for various categories of resistance factors, namely ESBLs (blaCTX-M and blaSHV, blaOXA-10 and blaTEM), quinolone (qnrS, qnrB and qnrA) and aminoglycoside [aac (6′)-lb-cr] by PCR and the quantities (µL) of primers, polymerase, dNTPs, sterile molecular grade water and DNA template making up the PCR reaction mix, as well as PCR thermocycler conditions for denaturation, annealing and elongation stages of amplification of each gene were as described.22-28 Primers used for the amplification of the respective genes are indicated in Table 2.
The primers used for the detection of resistance genes.
Phenotypic evaluation of virulence in the studied strains
Productions of alkaline protease and phospholipase were, respectively, investigated by skim milk agar and blood agar as previously described by Mohammad. 29 In summary, freshly prepared bacterial culture was inoculated on skim milk agar and blood agar and incubated at 37°C for 24 hours to investigate the production of virulence factors alkaline protease and phospholipase, respectively. Pseudomonas aeruginosa virulent strain ATCC 27853 was used as the reference strain. Clearance of the zone around isolated indicated production of the respective virulence factors.
Plasmid profiling and curing of selected isolates from the studied strains
Selected strains were subjected to an antibiotic susceptibility test with gentamicin, cefotaxime, ceftazidime and cefotaxime by investigating the minimum inhibitory concentrations (MICs) of the respective antibiotics method as previously described. Extraction of plasmid DNA from selected strains, determination of the molecular weight of plasmids and plasmid curing with ethidium bromide at 37°C were also carried out as described somewhere.30,31 In brief, 5 sub-inhibitory ethidium bromide concentrations were used for the curing assay. Each of the selected strains was sub-cultured into 5 test tubes of 10 mL nutrient broth containing sub-inhibitory concentrations 10, 5, 2.5, 1.25 and 0.625 μg/mL of ethidium bromide respectively and cultured for 37°C for 24 hours. Viable colonies of each strain were selected and antibiotic susceptibility to gentamicin, cefotaxime, ceftazidime and ciprofloxacin by MICs method was repeated. The MICs before and after curing were compared.
Results
Bacterial identification
A total of 122 morphologically-distinct colonies of non-lactose-fermenting bacteria were identified. The prevalence of the studied strains were 58.2% Pseudomonas paralactis (71 strains isolated from 16 farms) with associated GenBank accession number NR_156987.1 and 20.5% Pseudomonas lactis (25 strains isolated from 9 farms) with associated GenBank accession number NR_156986.1. The remaining 26 strains were strains of Serratia fonticola, Pseudomonas gessardii, Pseudomonas mucidolens, Lysinibacillus fusiformis, Pseudomonas stutzeri, Bacillus aryabhattai, Pseudomonas viridiflava and Bacillus megaterium.
Biochemical and growth characterisation of studied strains
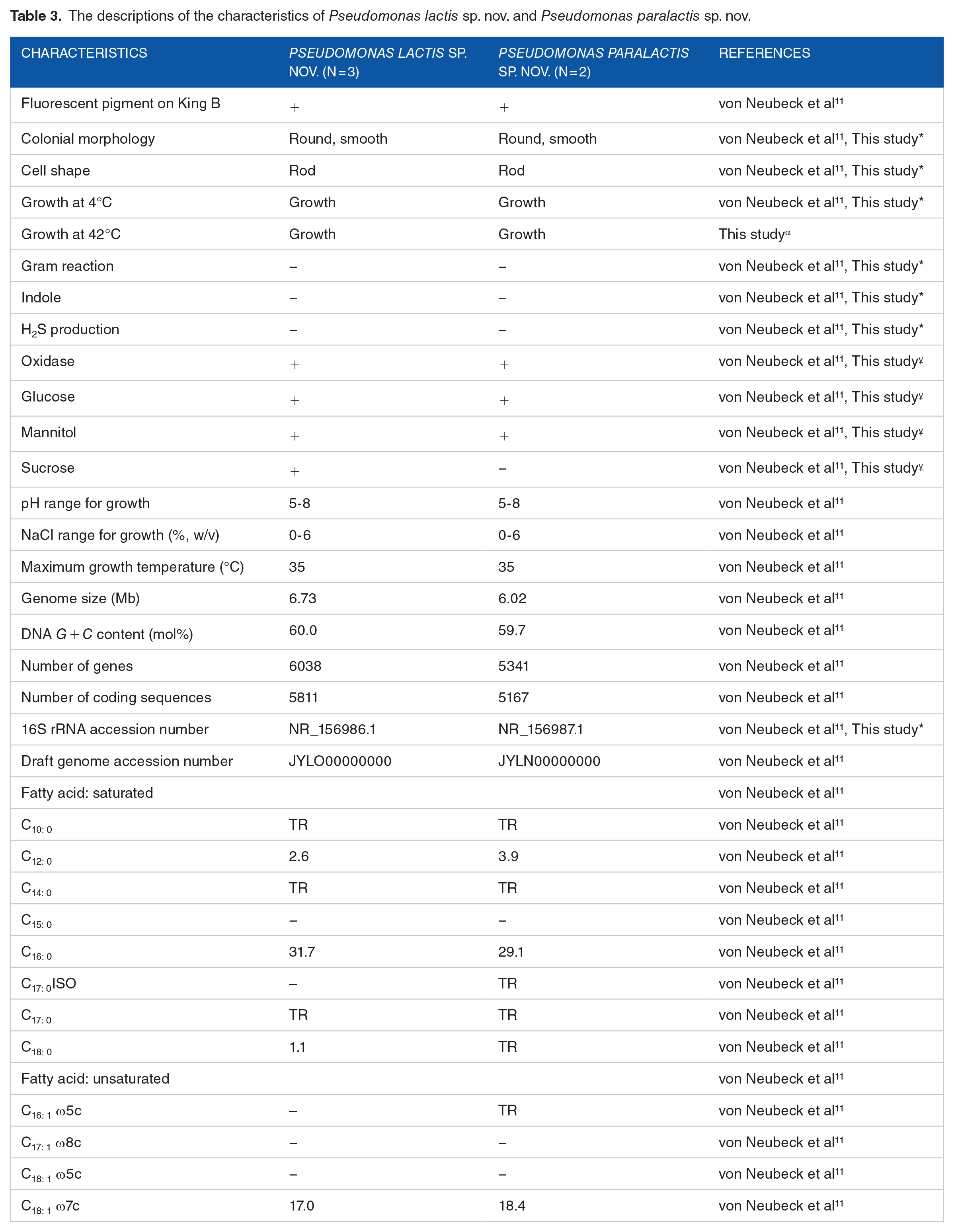
Results of reactions of studied organisms to carbohydrate fermentation with mannitol, glucose, sucrose and reactions to oxidase, indole, sulphide and Gram test as well as growth at 4°C and 42°C are as shown in Table 3
The descriptions of the characteristics of Pseudomonas lactis sp. nov. and Pseudomonas paralactis sp. nov.
Antibiotic susceptibility profiles of studied strains
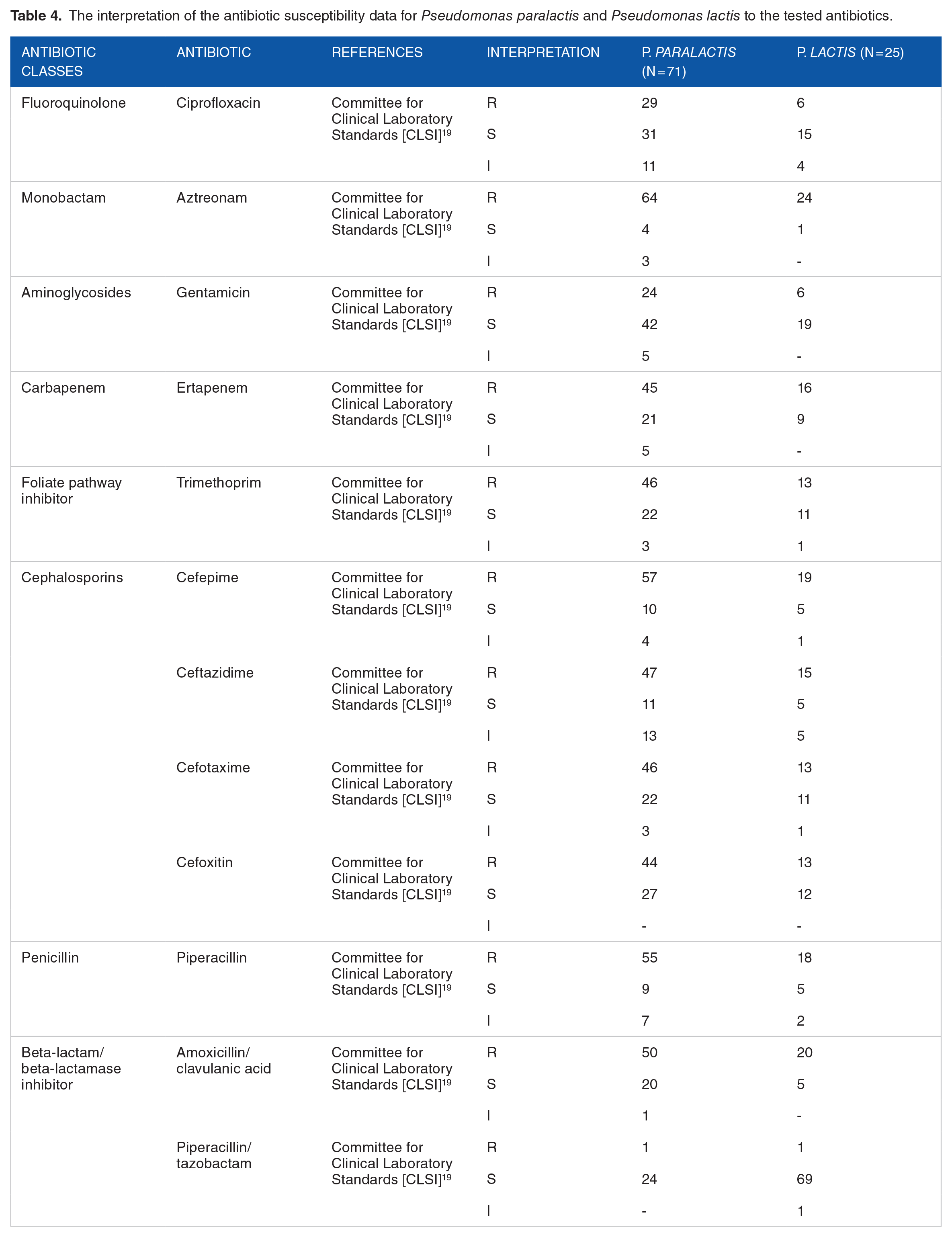
Results of antibiotic susceptibility testing of isolates were classified into 3 different interpretive categories, namely susceptible (S), intermediate (I) or resistant (R) according to the Clinical and Laboratory Standards Institute. 19 Table 4 shows strains of P. lactis and P. paralactis grouped into the antibiotic susceptibility interpretive categories. Except for gentamycin and piperacillin/tazobactam, the susceptibility of P. paralactis strains to the tested antibiotics was considerably low, ranging from 5.6%-45.1%, with the lowest susceptibilities observed in aztreonam (5.6%), ceftazidime (14.1%), piperacillin (14.1%) and cefepime (12.7%). Considerably high antibiotic imperceptibilities were observed in gentamicin (61%) and piperacillin/tazobactam (98.9%). However, the highest resistance was observed in Aztreonam (91.5%) while the percentage of resistant strains decreased through piperacillin (76.1%), cefepime (81.7%) ertapenem (63.4%) and the lowest observed in piperacillin/tazobactam (1.4%). On the other hand, a relatively high percentage of strains of P. paralactis were susceptible to gentamicin (76%), piperacillin/tazobactam (96%) and ciprofloxacin (60%), while relatively lower susceptibility was observed in cefepime (20%), piperacillin (20%), ceftazidime (20%), amoxicillin/clavulanic acid (20%) and ertapenem (30%). However, resistance ranged from the highest observed in aztreonam (96%) through cefoxitin (52%) and least in piperacillin/tazobactam (4%).
The interpretation of the antibiotic susceptibility data for Pseudomonas paralactis and Pseudomonas lactis to the tested antibiotics.
Resistance patterns observed across the 12 antibiotics for both P. lactis and P. paralactis are indicated in Figures 1 and 2. For P. lactis, 92% were resistant to 3 or more antibiotic classes and therefore termed as MDR strains, while strains resistant to less than 3 antibiotic classes were non-MDR. Amongst the non-MDR strains, no isolates were resistant to 0 or 1 antibiotic class, however, 8% were resistant to 2 classes. Amongst the MDR strains, the number of antibiotic classes to which isolates were resistant ranged from 4 to 7.
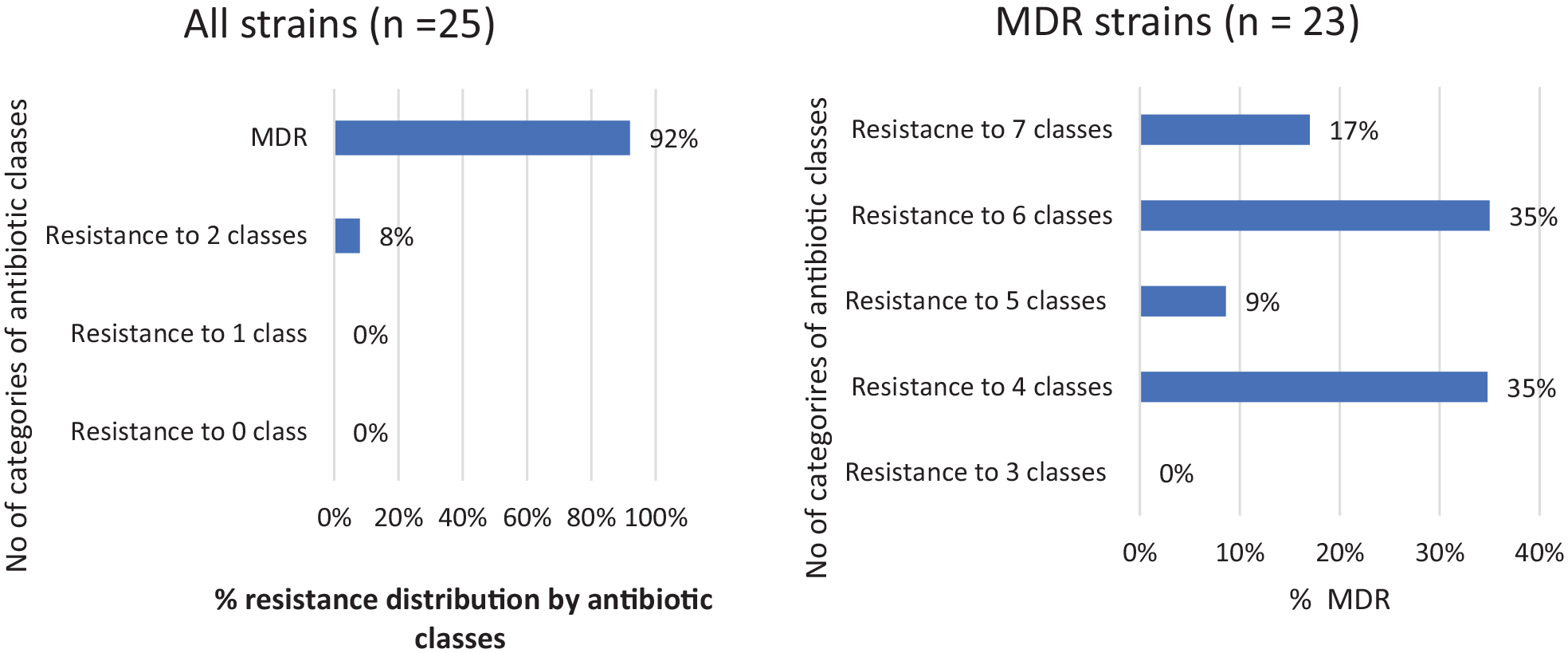
The distribution of resistance patterns of P. lactis strains.
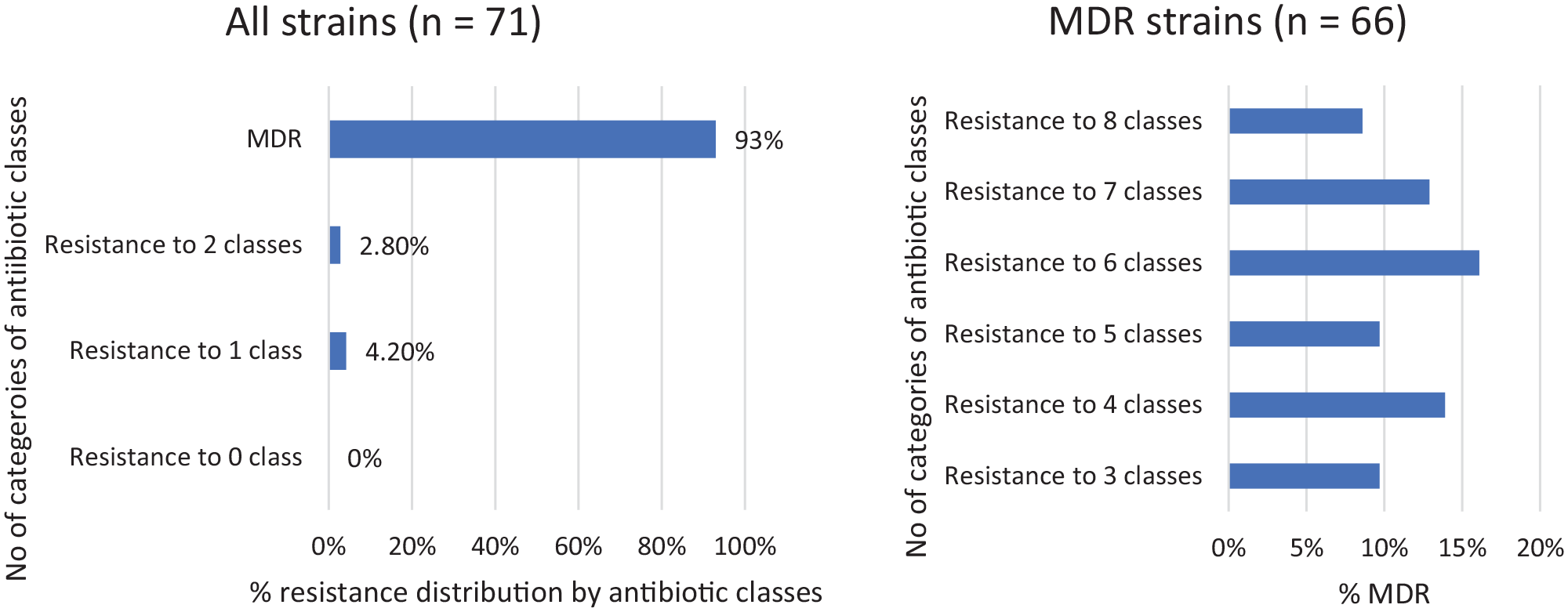
The distribution of resistance patterns of P. paralactis strains.
For P. paralactis, 93% of the strains were MDR, with the number of antibiotic classes ranging from 3 to 8. Amongst non-MDR strains, 4.2% and 2.8% of isolates were resistant to 1 antibiotic class and 2 antibiotic classes, respectively. The MAR index values for P. lactis strains ranged from 0.2 to 0.8, while 23/25 (92%) of the strains had values greater than 0.2 (Supplemental Table S2a). However, MAR index values for P. paralactis strains ranged from 0.1 to 0.7, while 68/71 (96%) of the strains had values greater than 0.2 (Supplemental Table S2b).
ESBL production and detection of resistance genes in the studied isolates
In the results of DDDT for ESBL production on all Pseudomonas lactis and Pseudomonas paralactis isolates indicated that 6 (24%) and 23 (32%) of the respective species were ESBL producers.
Resistance determinants in the studied strains
In total, 3 classes of resistance determinants (ESBLs, aminoglycoside and quinolone resistance genes) were detected in the studied isolates (Figure 3). Twenty-five and 6 strains of P. paralactis and P. lactis, respectively, carried at least 1 resistance gene. The aminoglycoside-resistance genes, aac (6′)-lb-cr, was detected in 1/25 (4%) of P. lactis strains. The quinolone resistance gene, qnrS, was also found in 3/25 (12%) of the P. lactis strains, while the 2 ESBLs genes, blaCTX-M and blaSHV were found respectively in 1/25 (4%) and 1/25 (4%) of the P. lactis strains. In total, cross-linkage of 2 resistance genes was found in 2 P. lactis strains: both blaCTX-M and qnrS were detected in 1 strain while both aac (6′)-lb-cr and qnrS were found in another strain. The aac (6′)-lb-cr gene was detected in 5/71 (7%) of P. paralactis strains, while the qnrS, blaCTX-M and blaSHV were found in 6/71 (9%), 5/71 (7%) and 9/71 (13%) respectively. In total, cross-linkage of 2 resistance genes, blaCTX-M and blaSHV, were found in 1 P. paralactis strain.
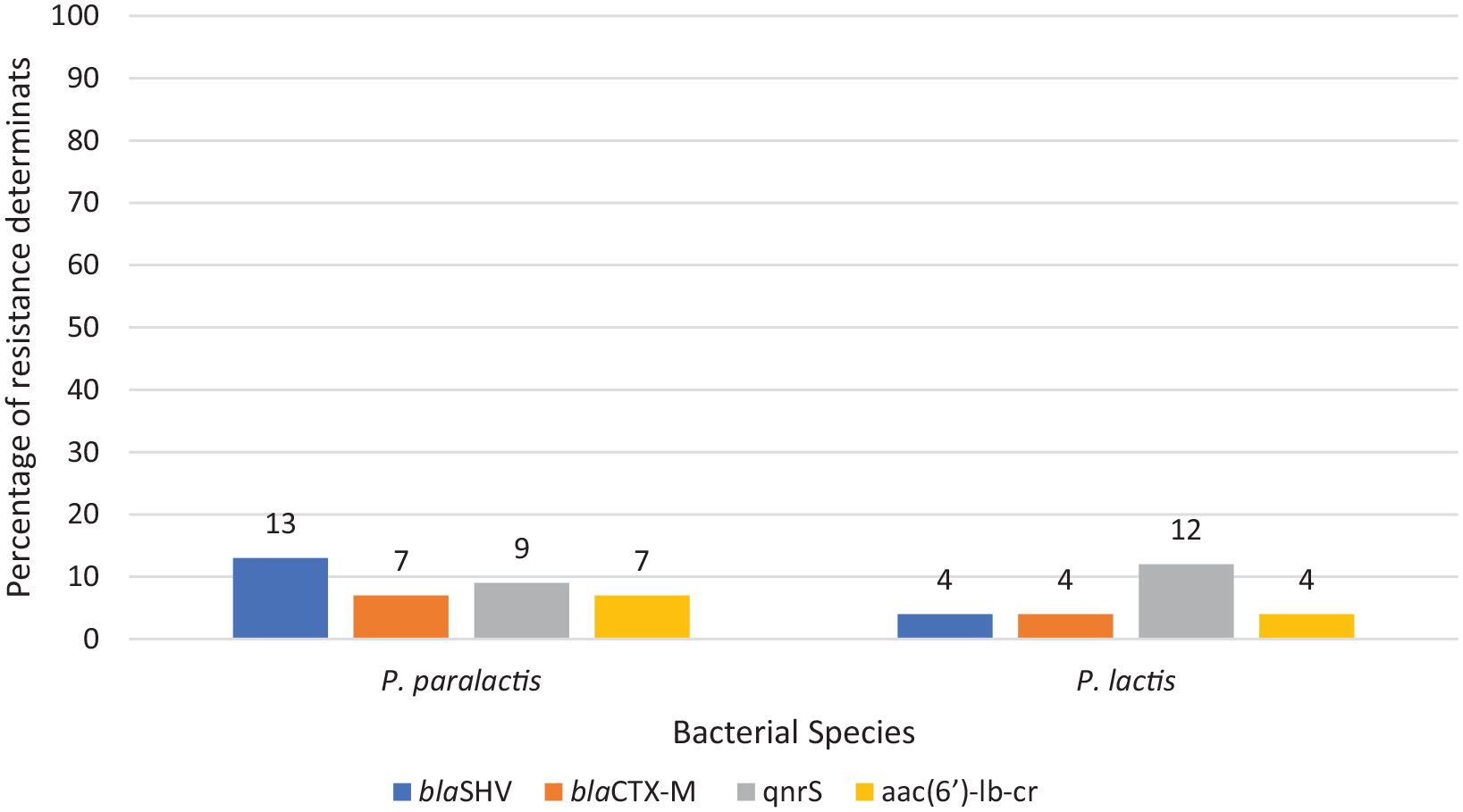
The distribution of resistance determinants amongst the isolates.
BlaTEM, qnrA, qnrB and blaOXA-10 genes were not found in any of the studied strains.
Phenotypic evaluation of virulence in studied strains
All Pseudomonas lactis and Pseudomonas paralactis strains were tested for the production of alkaline protease and phospholipase. The results showed that 3% and 3% of Pseudomonas lactis were alkaline protease and phospholipase producers respectively while 12% and 4% of Pseudomonas paralactis were alkaline protease and phospholipase producers respectively (Figure 4).
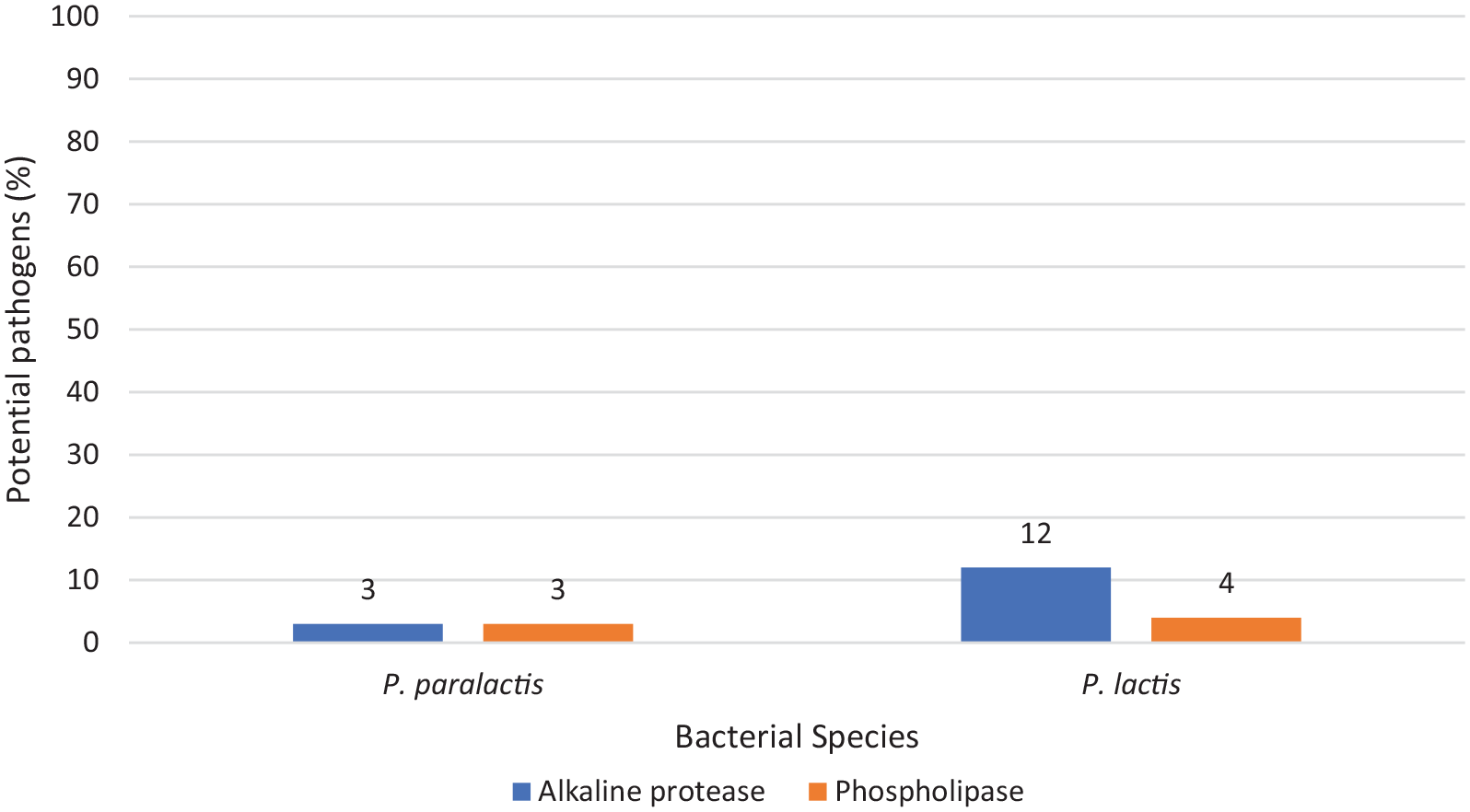
The pathogenicity profiles of the isolates.
Plasmid profiling and curing of selected isolates from studied strains
Candidate strains for plasmid profiling were those that had all the listed properties: phenotypically resistant to at least one of third-generation cephalosporin (namely, cefotaxime and ceftazidime), resistant to at least one of the aminoglycoside and quinolone classes, and carriage of at least one resistance gene detected by PCR. 13 and 3 strains respectively of P. paralactis and P. lactis met the criteria. Plasmid DNA was detected in only 2 of the 3 selected strains of P. lactis and detected in only 8 of the 13 strains of P. paralactis selected. One plasmid with molecular weight >10 Kb was isolated in each of the P. lactis strains, while the plasmid number ranged from 1 to 6 in P. paralactis and sizes ranged from 1.5 to >10 Kb (Supplemental Table S3a and 3b). The results indicated the effect of plasmid curing on the resistance profiles of selected strains (Figures 5 and 6; Supplemental Table S3a and 3b). In treatment with cefotaxime and as observed from the MICs, for the selected Pseudomonas lactis strains, resistance was eliminated and decreased from 100% in wild strains to 66% in the cured strains, while amongst P. paralactis resistance was eliminated and decreased from 100% to 75%. For gentamicin and ciprofloxacin, resistance was eliminated from 88%, 50%, 38% and 13% respectively amongst the P. paralactis strains; while the percentage reductions of resistant strains for the respective antibiotics amongst P. lactis strains were 100%, 33%, 67% and 33%. Generally, loss of resistance to the cephalosporins after curing amongst both P. lactis and P. paralactis ranged from 25% to 34% (25% loss of resistance to both cefotaxime and ceftazidime in P. paralactis strains and 34% loss of resistance to cefotaxime in P. lactis strains).
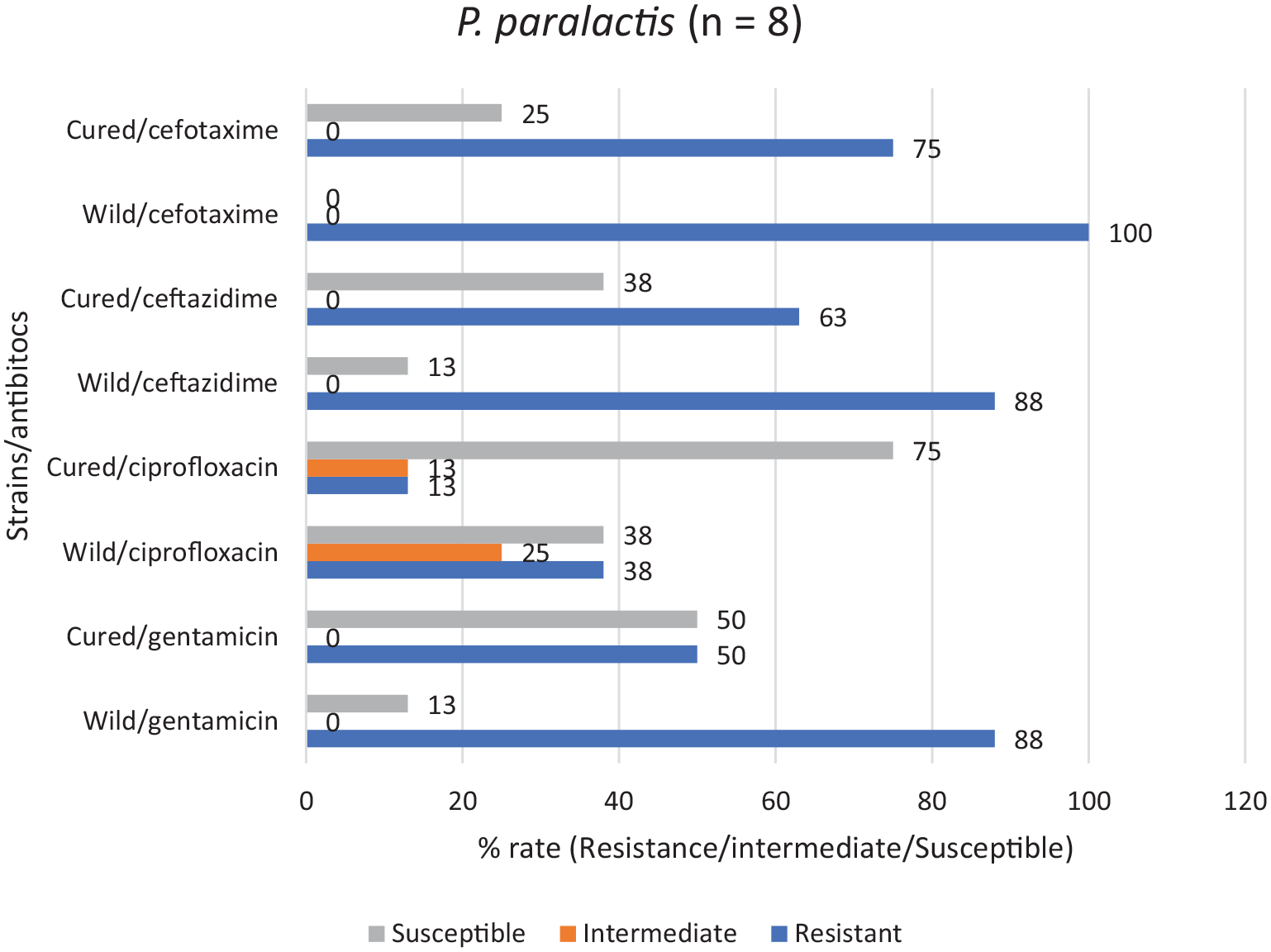
The effect of curing on the P. paralactis strains.
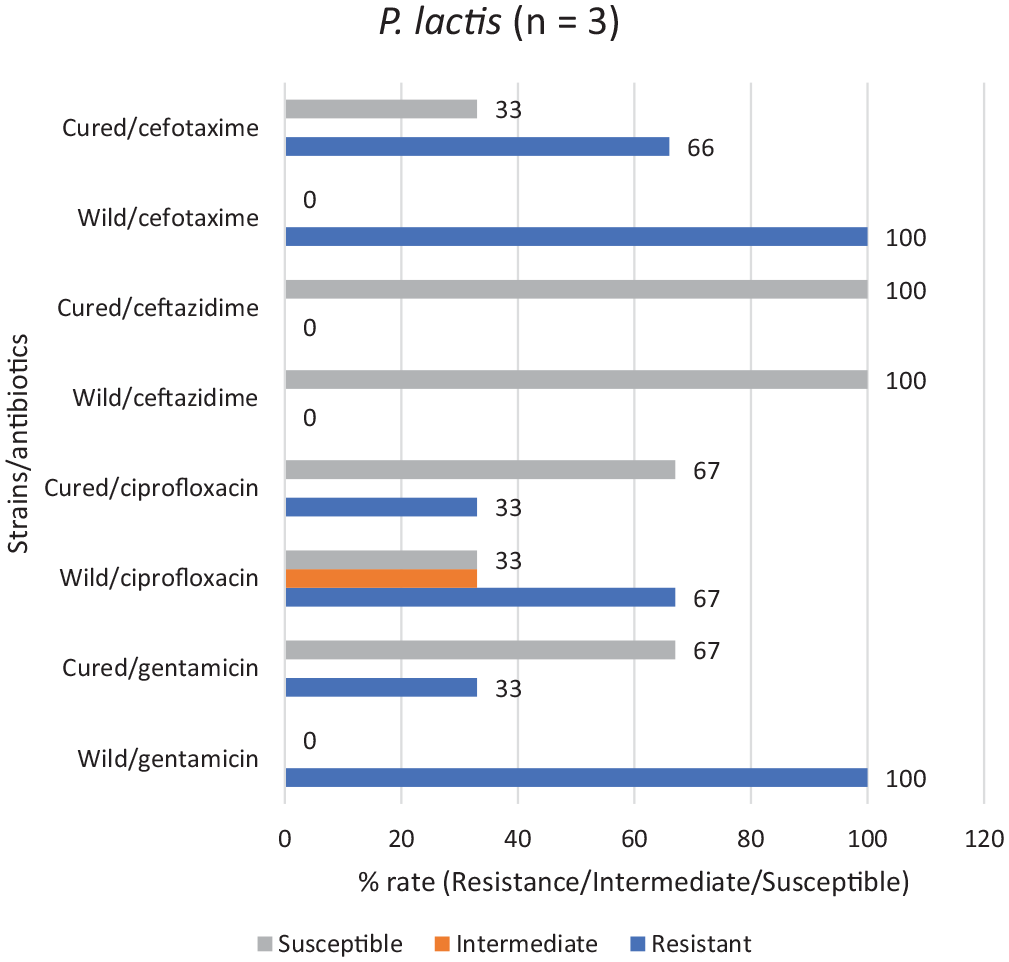
The effect of curing on the P. lactis strains.
Discussion
Antibiotic resistance amongst bacteria is undoubtedly a global concern, spanning human cultural practices such as farming and agriculture, clinical and environmental. The correlation between human/veterinary health and elimination of pathogens has caused researchers to prioritise resistance studies in pathogenic bacteria. 32 The focus on well-established pathogenic bacteria has recorded a considerable improvement in the fight against pathogenic inversion of animals and humans, as more methods for strengthening pathogens detection in the laboratory are increasingly suggested. 33 However, this approach would continue to yield considerable results only to the extent of researchers’ knowledge of the contribution of commensals and under-studied bacteria in the population to the global distribution of resistance genes. The distribution of antibiotic resistance genes is continually being reported amongst food animals, particularly livestock. 34 Selection of strains on species-specific selective and differential media is standard practice for bacterial isolation from cultures. However, isolation of bacteria on such selective and differential agar is only relevant for bacteria with already known, characterised and well-studied cultural characteristics. Therefore, only species-specific selective and differential agar is insufficient in isolating bacteria that have uncommon or un-studied characteristics. In contrast, MacConkey agar is more inclusive, permitting the evaluation of a variety of non-lactose-fermenting bacterial species, distinguished by their opaque and off-white colouration on incubation on MacConkey agar at 37°C for 18 to 24 hours, some of which are usually the better-known and commonly isolated species, but the possibility of entirely novel strains growing on the MacConkey agar cannot be ruled out, 35 before strain identification through molecular analysis. Concerning food animals, previously unknown or unidentified bacteria are increasingly being reported from chickens in particular. 36 Therefore, MacConkey agar was used in the present study to isolate as many possible non-lactose-fermenting bacterial isolates that can be isolated from chickens for the assessment of their antimicrobial resistance determinants.
There are recurrent reports of the use of beta-lactam antibiotics in poultry feeds, 37 therefore the present research accordingly incorporated beta-lactam antibiotics (such as cefotaxime, cefepime, ertapenem, azetronam, cefoxitin, piperacillin, ceftazidime) and other categories like fluoroquinolone (ciprofloxacin) and aminoglycoside (gentamicin) to the antibiotic susceptibility testing.
Poultry constitutes a microbial community that is a reservoir of extremely diverse and phenotypically rare bacterial strains, most of which have been reported as potentially novel with unusual metabolic activities, such as antimicrobial activity as observed in the biosynthesis of an anticlostridial factor by a novel and poultry-associated Bacillus subtilis strain that inhibited the pathogen Clostridium perfringens, 31 and other reported instances of novel genes detected from rare chicken gut bacteria isolates. 38
Isolation of rare non-lactose-fermenting strains from poultry samples is not uncommon, and this current study isolated, and interestingly in relatively more abundant proportion concerning other non-lactose-fermenting bacteria species in the population, 2 Pseudomonas spp. strains – rod-shaped, Gram-negative, phenotypically smooth, round colony and identified molecularly by 16S rRNA gene sequencing as Pseudomonas lactis and Pseudomonas paralactis with respective 16S rRNA GenBank accession numbers of NR_156986.1 and NR_156987.1. This study interestingly added to the repertoire of knowledge that chicken faecal samples are potential sources of rare species, but in addition, the study presents the first report of the occurrence of the 2 Pseudomonas spp. in the poultry bacterial population. Pseudomonas lactis sp. nov. and Pseudomonas paralactis sp. nov. were first proposed in 2017 11 for 2 Pseudomonas genera isolated from bovine raw milk and their phenotypic and genetic characteristics elucidated based on bacterial polar lipid and DNA GC content respectively. The descriptions from the first isolation of the 2 species novae, in comparison to isolation in our study, are shown in Table 3. The description of isolates from the current study, as shown in the table are; (*) all species in this study showed the same characteristics reported in the first isolation for the given test; (α) some strains of the species gave positive reactions to the specific biochemical test while some gave positive reaction – positive and negative test reaction not consistent amongst the strains of a given species; (ұ) some strains of the given species gave the same positive reaction like the first isolation while other strains otherwise gave a negative reaction, and vice versa. Since these species are native to the milk bacterial population (from their name, lactis, indicative of the raw milk from where they were originally isolated), investigating their antimicrobial resistance profiles – which are not available until our study – is essential in keeping up with other research work focused on the evaluation of antibiotic resistance from milk-associated bacteria, 39 and to expand our understanding on antibiotic resistance in psychrophilic milk strains such as Pseudomonas spp., which are potential milk spoilers.
The initial isolation of both species from the psychrophilic milk environment was scanty (only 3 strains of P. lactis sp. nov. and 2 strains of P. paralactis sp. nov.). 11 Both species have neither been reported in other food animals since their first report in the bovine nor has their antibiotic susceptibility been characterised. Interestingly, our isolation of the 2 species supports the growing reports of isolation of previously-uncharacterised bacterial isolates from poultry – particularly chickens. 40 We equally found it interesting that, while the 2 species novae were scantily isolated from their initial detection in cow milk, they were isolated in relatively high abundance in our study from the total poultry bacterial population isolated: 58% and 20% of the total non-lactose-fermenting bacterial population were P. paralactis and P. lactis, respectively.
In this study, the isolates of the 2 species uniformly gave negative biochemical results for Gram reaction, indole and hydrogen sulphide – which were the same results reported from the bovine Pseudomonas lactis sp. nov. and Pseudomonas paralactis sp. nov. strains by von Neubeck et al.
11
However, results varied for carbohydrate fermentation. Conventionally, bacterial strains are characterised biochemically and microbiologically as positive (or negative) for a particular test or physiological feature according to the descriptive symbols
We propose that, where discrepancies in biochemical tests exist between our biochemical test reports and that of the initial report on the bovine samples, this may likely have occurred because the number of strains from which biochemical data were obtained was significantly scanty in the bovine sample. As noted, the general assignment of a positive or negative report for tests on previously unclassified species is usually based on the relative percentage quantity of the strains under observation concerning the studied traits. 42 Moreover, strains of a bacterial species that present differing reactions to biochemical tests – symbolised as (d) – are not uncommon. A previous study by Cai et al 41 on veterinary bacterial isolates reported most species as difficult-to-characterise phenotypically because strains of the isolates showed varying reactions to biochemical reagents and most could not be identified either to genus or species level because of the observed biochemical variations, therefore identification was resolved by 16S rRNA gene sequencing. The workers explained that the unusual and discrepant reactions were observed because some of the strains were either biochemically inert to the available test reagents or they presented unusual biochemical reactivity that could not be tested by available biochemical assays, therefore they were difficult to identify by conventional biochemical identification method. Some of the strains could not be identified as well by commercial identification kits, suggesting that their reactions were not yet in the database of commercial identification systems.
As explained by Cai et al, 41 the differing results of biochemical tests in the current poultry study might suggest relative inertness of more strains of Pseudomonas lactis and Pseudomonas paralactis to the test reagents.
The Pseudomonas spp are extremely phylogenetically-complex bacterial species. The 16S rRNA gene sequencing method is considered the gold standard in species identification, but certain Pseudomonas genus has not been successfully identified to species level by 16S rRNA gene sequencing alone because of the complex diversity of Pseudomonas species, and in such cases, other housekeeping genes were used to complement 16S rRNA gene sequencing methods. 43 However, in reporting the identity of the 2 Pseudomonas species isolated from the poultry samples in the current study as Pseudomonas lactis and Pseudomonas paralactis, 16S rRNA gene sequencing was used for identification. The bacterial 16S rRNA gene sequence database confirms species-level identification of a sequence at ⩾97% similarity to sequence in the database, while 90%-96% is identification to genus level. 44 For strains having less than 97% 16S rRNA gene sequence identity to reference sequence in the database, strains are said to be unidentifiable to species level by 16S rRNA gene analysis alone, and other housekeeping genes – each having its database and percentage threshold values for species-level identification – are normally used to determine species-level identity. Interestingly, all the strains from poultry in the current study have 16S rRNA gene sequence database scores of ⩾97% (Supplemental Table S4), which is confirmatory for species-level identification therefore application of any other housekeeping gene sequence analysis for further confirmation was not necessary. Similarly, all strains of a given species in this study had a uniform 16S rRNA gene sequence accession number that exactly corresponds to the accession number of that species as reported in the 16S rRNA GenBank since their first isolation by von Neubeck et al, 11 which is NR_156987.1 for Pseudomonas paralactis sp. nov. and NR_156986.1 for Pseudomonas lactis sp. nov.
The existence of antibiotic-resistant bacterial strains from food animals is a serious concern and accounts for one of the most common channels for distribution of resistant strains globally. 45 In 2017, the World Health Organisation labelled cefotaxime as a critical and reference antibiotic for the treatment of human infections. 46 Studies have reported considerable genetic and clonal relatedness between resistant pathogenic bacteria from food animals – particularly poultry animals – and the human community, 47 in addition to reporting genetic relatedness between chicken bacterial isolates and those from outpatients and slaughterhouse personnel. Thorsteinsdottir et al 48 also reported similar antibiotic susceptibility patterns amongst the bacteria from these various sources. The genetic relatedness of pathogens from human and animal sources suggested the potential transfer of resistant pathogens between the poultry environment and humans.
Over the years, Pseudomonas species borne by the food chain have been extensively researched not only because Pseudomonas species borne by food animals are increasingly emerging as important pathogens, but also because of the reputation of most bacterial isolates as being aetiological agents in the global spread of antibiotic resistance as measured by their resistance against several antimicrobial agents. Although not previously investigated until the current research, the high degree of multi-antibiotic resistance of Pseudomonas lactis and Pseudomonas paralactis strains in the current study to the tested antibiotic classes corroborated previous work on Pseudomonas species from chicken samples. 49 The workers also reported the highest resistance rate amongst the studied isolates to Azetronam, which corroborated the highest resistance of both Pseudomonas lactis and Pseudomonas paralactis to Azetronam.
The selection of resistance to different antibiotics by bacteria from food animals has been critically reviewed by different workers and proposed to continue to be on the rising because of poor antibiotic surveillance in livestock and other sources of food animals. 50 This is reflected in the current study on poultry bacterial isolates in which the majority of both Pseudomonas lactis and Pseudomonas paralactis were resistant to most of the tested antibiotics. Therefore, to address the continued spread of antibiotic resistance related to the food chain, there is a need for frequent assessment of antimicrobial susceptibility of commonly used antibiotics since even the sensitive strains of bacteria could acquire resistance to such antibiotics due to the transfer of resistance factors from resistant strains.
Phenotypic evaluation of virulence by detection of virulence enzymes such as phospholipase C and alkaline protease on diagnostic agar media such as skim milk agar and blood agar, respectively, has been used to screen for bacterial pathogenicity, and this method was particularly used in the work of Mohammad 29 to screen for virulent strains of Pseudomonas aeruginosa from clinical samples. Alkaline protease is produced by pathogenic strains of Pseudomonas spp as an antiphagocytic agent, while phospholipase C degrades the epithelial layer of the host’s cell.51,52
The patterns of resistance observed in both Pseudomonas lactis and Pseudomonas paralactis in this study showed the phenotypic pattern of antibiotic cross-resistance frequently noticed in bacteria species that are significantly resistant to beta-lactam antibiotics. Significant resistance of beta-lactam-resistant bacteria to other classes of antibiotics has been reported by other workers. 53 In this study, isolates that were phenotypically resistant to at least one β-lactam antibiotic were equally resistant to several other antibiotic classes, agreeing with another work. 54 A highly significant mechanism of bacterial resistance amongst the majority of the Pseudomonas spp is the development of resistance to an array of broad-spectrum cephalosporins (except cephamycins and carbapenems) and monobactam by the production of extended-spectrum β-lactamases (ESBLs).
Alarmingly, in our study, 92% of P. paralactis and 96% of P. lactis had MAR values >0.2, indicating a high degree of antibiotic contamination of the 2 species. The enzyme CTX-M has been reported in several geographical locations worldwide, African countries inclusive, particularly amongst isolates significantly resistant to cephalosporin such as cefotaxime. 54 In the current study, however, the prevalence of the CTX-M gene in both the Pseudomonas lactis and Pseudomonas paralactis was relatively low. For instance, amongst the P. paralactis and P. lactis strains, the CTX-M gene was detected in 7% and 4% strains respectively. Although the high occurrence of the CTX-M gene in ESBL-producing isolates is reportedly common,55,56 other workers have recorded relatively low-level occurrences of the gene in ESBL-producing bacterial strains. For instance, a relatively low prevalence of the blaCTX-M gene has been reported in ESBL-producing isolates in independent studies in Iran by Lee et al, 57 which corroborate s the finding in our study. Equally, Li et al 58 reported that, while the CTX-M gene is one of the most common genes encoding the ESBLs enzyme, there are several other mechanisms – such as the activity of families of genes including the CMY gene – that could generate ESBLs production aside the activity of CTX-M in bacteria. Interestingly, Carvalho et al 59 have also demonstrated a rare but very definite situation where none of the 3 common ESBL genes blaCTX-M, blaSHV and blaTEM was detected in isolates with phenotypes that suggested ESBL production.
Most of the plasmids isolated from the 2 species in the current study were of large sizes. This is consistent with previous reports that ESBL production is coded by genes that are prevalently located on large plasmids most of which are often larger than 10 Kb. 60 In this study, large molecular weight plasmids (>10 Kb) were isolated from 8 (62%) of the 13 selected ESBL-producing Pseudomonas paralactis isolates and in all the selected ESBL-producing Pseudomonas lactis isolates. ESBLs genes of the families blaSHV, blaCTX and blaTEM can reside on large plasmids and this has been particularly demonstrated for the CTX-M enzyme, although these genes can also be chromosomally-borne, in which case they reside on the chromosomes rather than on plasmids. 61 In the current study, the blaCTX gene was detected in 1 of the 2 isolates of Pseudomonas lactis isolates from which large molecular weight plasmids were isolated and 2 of the 8 Pseudomonas paralactis isolates had large molecular weight plasmids (Supplemental Table S3a and 3b). Multidrug resistance has often been reported in CTX-M producing isolates, owing to the possibility of other genes such as blaTEM, bla OXA, aac (6′)-lb and even Qnr determinants to be present on the same plasmid carrying the blaCTX gene. 60 Such multiple-carriage of resistance genes was observed in our study from CTX-M positive isolates that were equally carrying plasmids. Of the 8 Pseudomonas paralactis isolates from which plasmids were isolated, 2 isolates carried the CTX-M genes: 1 isolate co-haboured the blaSHVgene while the other isolate co-haboured the QnrS gene.
Plasmid curing essentially determines the genetic location of antibiotic resistance determinant – whether borne on plasmid or chromosome. Loss of resistance to a particular antibiotic by an isolate after curing is indicative that the gene coding for the resistance is borne on the plasmid, otherwise, the gene is chromosomally-borne. In the current study, loss of resistance to the test antibiotics after curing was observed in some strains, indicating the mediation of such resistance by an R-plasmid.
For Pseudomonas paralactis isolates, 25% of the plasmid-carrying isolates that were selected from the pool of strains resistant to third-generation cephalosporins cefotaxime and ceftazidime lost resistance to each of the 2 antibiotics after curing. This essentially implied that 75% of the observed resistance to the cephalosporins was chromosomally-mediated. For Pseudomonas lactis isolates, 34% of strains lost resistance to cefotaxime after curing, indicating 66% of strains habouring cefotaxime resistance chromosomally. Therefore, the selection of resistance for cefotaxime in the majority of the selected strains of the 2 Pseudomonas species isolated by cefotaxime-supplemented media in this study was considerably chromosomal, suggesting innate antibiotic resistance in the studied strains. Considerable chromosomal mediation of resistance in bacterial strains to beta-lactams has similarly been reported. 61
Conclusion
This study presented the isolation of 2 Pseudomonas species, previously termed novel strains, with considerably high resistance to third-generation cephalosporin, particularly cefotaxime. Third-generation cephalosporins are important antibiotics that are widely used in the treatment of serious human Gram-negative bacterial infections but are rarely reported in animal settings or food animals generally. On the World Health Organisation’s list of critical antibiotics used exclusively for the treatment of human infections is cefotaxime. The result of bacterial isolation with cefotaxime-supplemented media in the current study showed an overall 80% resistance to this antibiotic. Such a high level of resistance to cefotaxime among Pseudomonas paralactis and Pseudomonas lactis strains, both not commonly reported in food animals except for previous isolation from bovine milk, is alarming indeed as this might herald the emergence of cefotaxime-resistant bacterial strains with unstudied abilities to transfer resistance to other bacterial strains.
Knowledge of antibiotic susceptibility profiles from bacteria from food animals is highly urgent. Our study successfully determined the antibiotic resistance profiles of P. lactis and P. paralactis and also characterised their antibiotic resistance genes. Our results revealed a high prevalence of MRD as well as a high prevalence of clinically significant MAR values amongst the strains of the 2 species. Our study projects both species, currently understudied, as highly significant carriers of resistance determinants and having potential public health concerns. We, therefore, desire investigations on the genome-wide study of the resistance determinant of the species.
Supplemental Material
sj-xlsx-1-mbi-10.1177_11786361221130313 – Supplemental material for Occurrence of Pseudomonas lactis and Pseudomonas paralactis Amongst Non-Lactose-Fermenting Bacterial Isolates in Chickens and Their Antimicrobial Resistance Patterns
Supplemental material, sj-xlsx-1-mbi-10.1177_11786361221130313 for Occurrence of Pseudomonas lactis and Pseudomonas paralactis Amongst Non-Lactose-Fermenting Bacterial Isolates in Chickens and Their Antimicrobial Resistance Patterns by Samson A Olofinsae, Olufemi E Adeleke and Bartholomew O Ibeh in Microbiology Insights
Footnotes
Funding:
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Declaration of Conflicting Interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
SAO and OEA conceived of and designed the experiment. SAO wrote the first draft of the manuscript. SAO, OEA and BOI analysed the data. SAO, OEA and BOI contributed to the writing of the manuscript. All authors reviewed and approved of the final draft of the manuscript.
Ethical approval
This research was conducted with the knowledge and approval of the ethical committee of the University of Ibadan.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
