Abstract
The bacteriological safety of food/food products and the menace of antimicrobial resistance amongst enteropathogenic bacteria raise therapeutic management concerns within the public health system. Recently consumers of food/food products purchased from the public market of Bushenyi District presents with Enterobacteriaceae infection-associated symptoms and clinical conditions. We determine the molecular characterization and antibiotic signatures of some enteric bacterial recovered from foods/food products in markets of Bushenyi District, Uganda. Standard molecular biology techniques (Polymerase chain reaction PCR) and microbiological procedures were applied. Meat (MT) and milk (MK) samples were collected from 4 communities/town markets (Kizinda, Ishaka, Bushenyi, kashenyi) between April and September 2020 and analyzed. Our result reveals high differential counts of Salmonella species (175.33 ± 59.71 Log 10 CFU/100 ml) and Escherichia coli (53.33 ± 26.03 Log 10 CFU/100 ml) within the 4 markets with the count of Salmonella species higher than that of E. coli in each sampled market. The PCR further confirmed the detected strains (22.72% of E. coli and 54.29% of Salmonella species) and diverse multiple antibiotic-resistant determinants {TEM: (12 (23.1%) blaTEM-2 gene, 3 (5.8%) blaTEM gene}, 5 (9.6%) blaSHV gene, 3 (5.8%) bla-CTX-M-2, 1 (1.9%) bla-CTX-M-9 }. Other resistance genes detected were {10 (21.7%) strA gene} and 8 (17.4%) aadA gene} indicating a potential antibiotic failure. The need for alternative medicine and therapeutic measure is suggestive. Astute and routine surveillance/monitoring of potential pathogens and food products in the public market remains a core for maintaining future consumer safety.
Introduction
In humans, food-associated gastroenteritis is majorly caused by enteric bacterial pathogens. 1 They cause high morbidity and mortality in children and adults and pose significant public health risks, especially in low resource communities. These potential bacterial pathogens have been isolated from fresh and ready-to-eat food specimens (milk, meat, etc.), water and the environment. Enteric bacterial pathogens span several genera, including Salmonella, Shigella, Yersinia, Escherichia, Vibrio, Bacillus, Enterobacter, Plesiomonas, Klebsiella, Proteus, Serratia, Aeromonas etc. Primary infections associated with the aforementioned bacterial pathogens have remained a public health concern due to the multiple antibiotic resistance (MAR) revealed in the studies of various investigators.2,3
Generally, regions with a high prevalence of antibacterial-resistant pathogens and antibacterial-resistant determinants (genotypes and phenotypes) have been reported to arise from the indiscriminate use of antimicrobial agents as therapeutic control strategies, agricultural and veterinary purposes. These resistance determinants are significantly transferred to related and unrelated species in other niches, such as the soil, water and food, via horizontal gene transfer.4-6
In the middle and low-income communities, the rapid emergence and spread of multidrug-resistant Enterobacteriales pose a severe threat to the public health and management system, making infectious diseases difficult to treat.7-10 In the Bushenyi District of Uganda, there is a paucity of studies on the molecular characterization and assessment of resistance genotypes of Enterobacteriales recovered from dairy food products. To this end, this study aimed to evaluate the occurrence, antibiotic susceptibility, and genes encoding antibacterial resistance of Salmonella spp. and Escherichia coli from milk and meat sold in the Bushenyi District markets Western region of Uganda. This study appears to be the first report on this subject within the study district.
Methods
Study area
This study was conducted in the Bushenyi District of the Western Region, Uganda. Bushenyi District is one of Uganda’s highly populated and low-income communities where major public markets and dairy products are commercially available to the public. Samples of milk products and meat were collected (once-off) from open markets within the Bushenyi District of Uganda, as described below in Table 1.
The study area in Bushenyi district, Uganda.
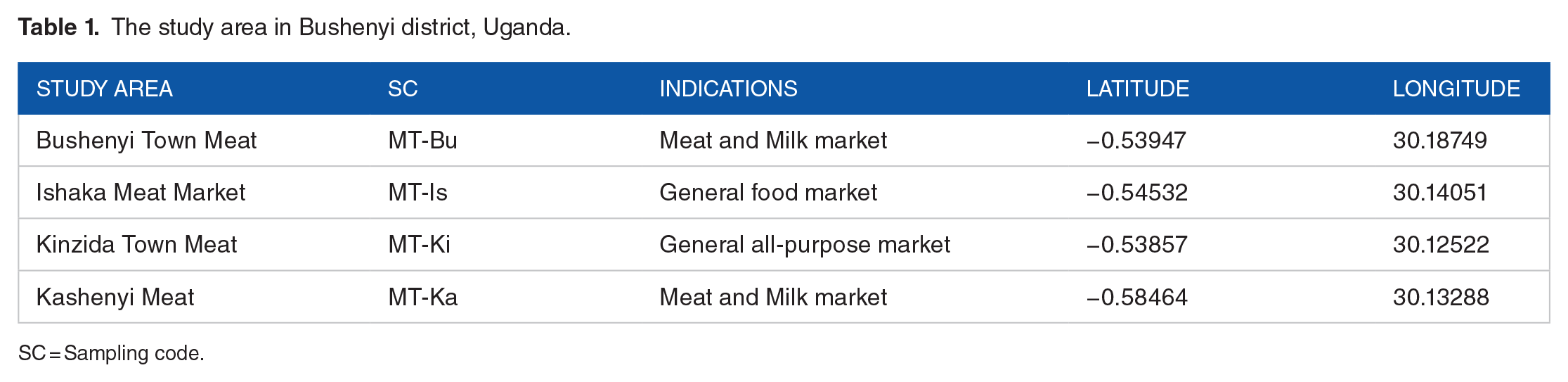

SC = Sampling code.
Bacterial isolation and identification
Briefly, meat (MT) and milk (MK) samples were collected from 4 community/town markets (Kizinda, Ishaka, Bushenyi, Kashenyi) and labelled as Ki, Is, Bu and Ka, respectively. The food samples were collected between April and September in sterile glass bottles, packed in an ice-box and transported to the Department of Microbiology laboratory, KIU Teaching Hospital, for culture within 3 hours of collection. The meat samples (1 g) were homogenized and pre-enriched in 5 ml of peptone water for 2 hours, subcultured onto pre-prepared McConkey agar (MCA) and Salmonella Shigella agar (SSA) agar plates were incubated at 37°C (following manufacturer’s instructions Merck Germany, www.merck-chemicals.com). Culture plates were incubated at 37°C for 24 to 48 h for the isolation of Salmonella spp. and E. coli, respectively. On the other hand, 1 ml of milk samples were pre-enrichment on 4 ml peptone water, subcultured onto MacConkey Agar (MCA), Deoxycholate agar (DCA) and Salmonella Shigella Agar (SSA) as cultured agar plates were incubated at 370°C for 24 to 48 hours. 11 Discrete colonies from the enumeration with black colouration on SSA and pink on MCA media were considered presumptive Salmonella spp. and E. coli. These isolates were then stored in 25% glycerol stock and kept at −200 C before biochemical and molecular characterization.
Enumeration of total bacterial load
Salmonella spp from food samples (meat and milk)
Homogenized samples were serially diluted (10−1 to 10−3) and cultured onto Selenite broth, SSA and Desoxycholate agar. 11 The bacterial load was determined by counting the number of discrete colonies using the viable plate count method. 11 The results were expressed as colony-forming unit per millilitre (CFU/ml) for milk and colony-forming unit per gram (CFU/g) for meat while their expression in a log of CFU/ml or CFU/g was applied using the formulae:
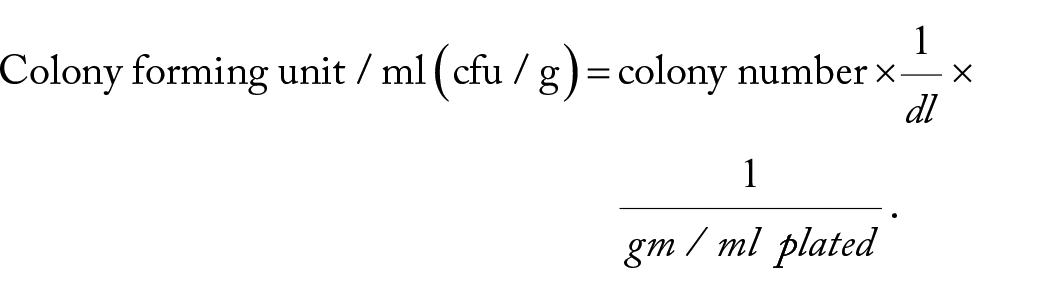
E. coli from food samples (meat and milk)
Homogenized samples were serially diluted (10−1 to 10−3) and cultured on MacConkey agar.12,13 The bacterial load was determined by counting the number of discrete colonies using the viable plate count method. 13 The colony-forming unit (CFU) was expressed in a log of CFU/ml or g by the formulae:
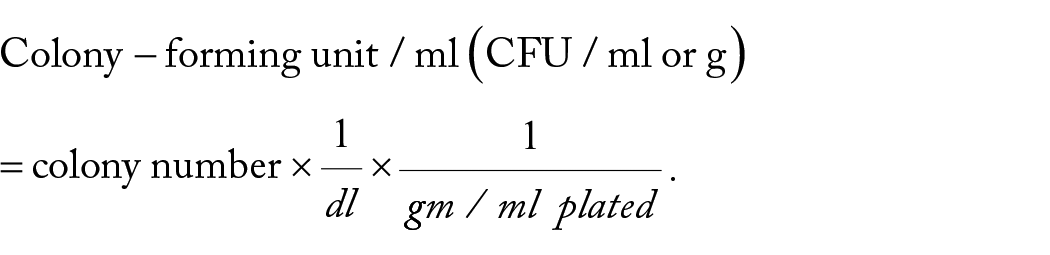

Phenotypic characteristic of the isolates (biochemical test)
Single colonies were further sub-cultured onto nutrient agar to obtain pure isolates. The pure isolates were used for phenotypic characteristics, including (Gram staining, Indole, Citrate, Urease, Hydrogen sulphide) to determine the bacteria morphology, biochemical and enzyme-based culture-dependent test described by Cheesbrough 14 .
Molecular confirmation of the presumptive isolates
The genomic DNA was extracted from the purified isolates using the boiling method described elsewhere15-17 as a DNA template for PCR reactions. The primers used for the PCR confirmation of the isolates were retrieved from previous studies and synthesized by Inqaba biotechnological Inc, Rausbauer, South Africa. The sequences are Ec-1 GACCTCGGTTTAGTTCACAGA (5′−3′), Ec-2 CACACGCTGACGCTGACCG (5′−3′), amplicon Size (bp) of 585, which was used to confirm the E. coli as described by El-Sayed et al 17 Salmonella spp. were confirmed using the primer SdiA1 AAT ATC GCT TCG TAC CAC (5′−3′), SdiA2 GTA GGT AAA CGA GGA GCA G (5′−3′) with amplicon size of 274 bp as described by. 18 For E. coli the PCR assay was done in 24 µL reaction volume as explained; (5 µL of nuclease-free water, 12 µL master mix, 2 µL of primer 1microMolar and 5 µL of template DNA). PCR cycling conditions was set as initial denaturation (93°C for 5 minutes); 35 cycles of : denaturation at 92°C for 40 seconds, annealing at 57°C for 1 minute, extension at 72°C for 2 minutes), and a final extension at 72°C for 2 minutes. The same reaction process was done for Samonella spp. with an annealing temperature of 52 0C. The amplified PCR products were electrophoresed using agarose gel electrophoresis (1.5% agarose) stained with ethidium bromide and visualized UV trans-illuminator. All positive isolates were stored in 25% glycerol stock at −200°C for subsequent analysis.
Antimicrobial susceptibility profile
The antimicrobial susceptibility test of the confirmed bacterial isolates was determined using the recommended Clinical and Laboratory Standard Institute (Clinical and Laboratory Standards Institute (CLSI) 19 guidelines for disc diffusion technique on Muller-Hinton agar (MHA) plates. Briefly, fresh culture from the glycerol stock was streaked onto nutrient agar plates, incubated at 37°C for 24 hours. Colonies were suspended in a 5 mL sterile standard saline test tube and adjusted to obtain turbidity corresponding with 0.5 Mc-Farland standards. The isolates were inoculated onto MHA plates, and disks impregnated with antibacterial agents/disc were dispensed onto the inoculated plates. The plates were incubated at 37°C for 18 to 24 hours, and zones of inhibition were measured after incubation. The susceptibility reports of the isolates were interpreted as resistant, intermediate or sensitive the commercial antibiotic discs used were: Cefotaxime (CTX) 30 μg, Amoxicillin (AM) 25 μg, Polymixin B (PB) 300 μg, Nitrofurantoin (NI) 200 μg, Ciprofloxacin (CIP) 5 μg, Nalidixic Acid (NA) 30 μg, Gentamicin (CN) 10 µg, Cefazolin. (CZ) 30 µg, Kanamycin (K) 30 µg.
Molecular characterization of antibiotic resistance genes
The PCR technique was used to characterize the resistance genes from the phenotypic antibiotic-resistant isolates. 3 The resistance genes explored in this study include β-lactamases-encoding genes (blaCTX-M2, blaCTX-M9, blaTEM, blaTEM-2) and aminoglycosides resistant genes (aadA, aac(3)-IIa (aacC2-a), strA). Table 2 includes the primers, the targetted genes, sequence, and PCR conditions. The PCR products were electrophoresed, as explained earlier.
Antibiotic resistance genes, primer sequences and PCR conditions.
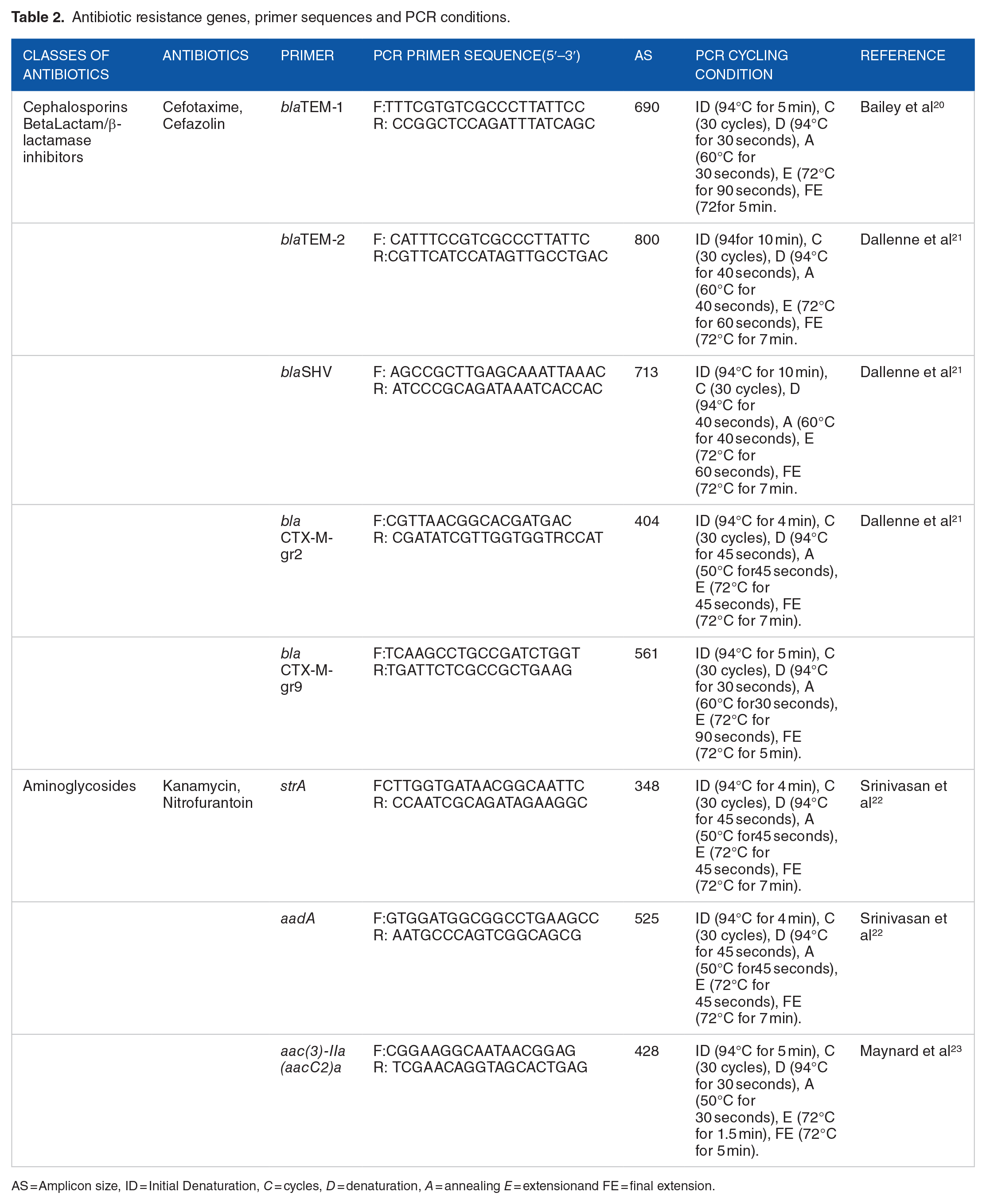
AS = Amplicon size, ID = Initial Denaturation, C = cycles, D = denaturation, A = annealing E = extensionand FE = final extension.
Patterns of multiple antibiotic resistance genotypes (MARGs)
The Multiple Antibiotic Resistance Genotypes (MARGs) patterns of the confirmed potential pathogens with ⩾2 multiple resistance genes was evaluated as described by. 24
Statistical method and analysis
All experiments were performed in triplicates, expressed in mean and standard error difference adopting the ANOVA (using graph prism version 8.0.2). This was followed by Fisher’s least level of significant difference (LSD) to test significant differences in bacterial counts in each sampling point and the months at p < .05 with regression analysis. RStudio version 3.5 1 was used to develop the multivariate heatmap analysis.
Results
Enumeration of total bacterial load
Lower density of E. coli. than Salmonella spp. count in meat from various locations of Uganda in April and September
In Kizinda, there was neither a difference in the Salmonella species count during April (114.33 ± 48.89 Log 10 CFU/100 ml) and September (175.33 ± 59.71 Log 10 CFU/100 ml) sampling periods, nor was there a difference in the E. coli during April (53.33 ± 26.03 Log 10 CFU/100 ml) and September (36.00 ± 11.59 Log 10 CFU/100 ml) sampling periods. However, the E. coli count was significantly lower than the Salmonella species count during both periods (Figure 1a).
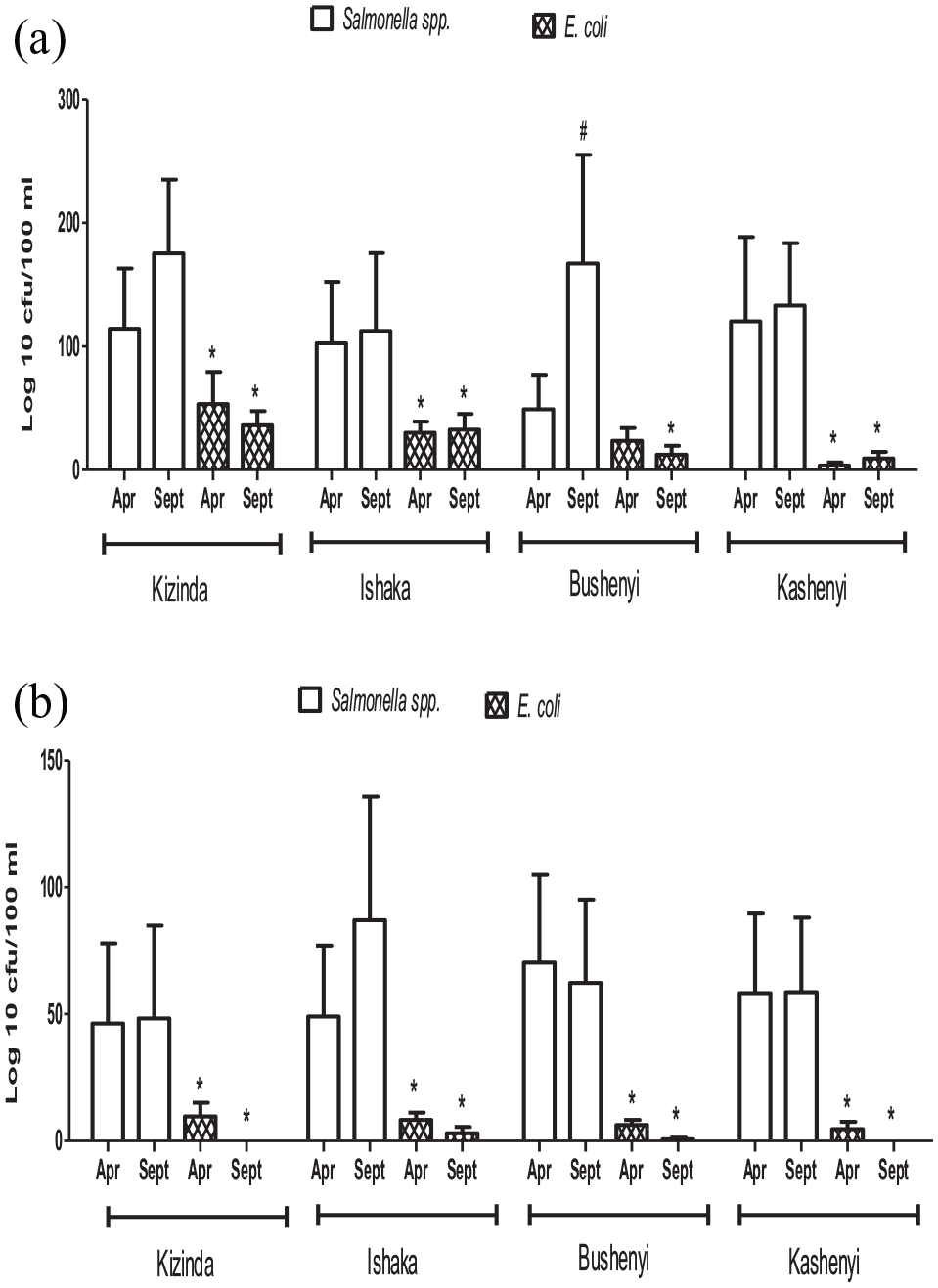
Salmonella species and E. coli counts in meat (A) and milk (B) from various locations of Uganda in May and September.
Also, in Ishaka, there was neither a difference in the Salmonella species count during April (102.67 ± 49.60 Log 10 CFU/100 ml) and September (112.67 ± 63.04 Log 10 CFU/100 ml) period of sampling, nor was there a difference in the E. coli during April (30.00 ± 9.17 Log 10 CFU/100 ml) and September (36.67 ± 12.81 Log 10 CFU/100 ml) period of sampling. However, the E. coli count was significantly lower than the Salmonella species count during both periods (Figure 1a).
In Bushenyi, there was a higher Salmonella species count during September (167.00 ± 88.07 Log 10 CFU/100 ml) than April (49.00 ± 28.04 Log 10 CFU/100 ml) sampling period, but there was no difference in the E. coli during April (23.67 ± 10.11 Log 10 CFU/100 ml) and September (12.33 ± 7.22 Log 10 CFU/100 ml) sampling periods. However, the E. coli count was significantly lower than the Salmonella species count during September but not April (Figure 1a).
Similarly, in Kashenyi, there was neither a difference in the Salmonella species count during April (120.33 ± 68.22 Log 10 CFU/100 ml) and September (133.00 ± 50.72 Log 10 CFU/100 ml) sampling periods, nor was there a difference in the E. coli during April (3.67 ± 2.33 Log 10 CFU/100 ml) and September (9.33 ± 5.21 Log 10 CFU/100 ml) sampling periods. However, the E. coli count was significantly lower than the Salmonella species count during both sampling periods (Figure 1a).
Nevertheless; the bacterial enumeration of Salmonella spp. and E. coli were significantly abundance in meat when compare with milk samples.
Lower Density of E. coli than Salmonella spp., count in milk from various locations of Uganda in May and September
In Kizinda, there was no difference in the Salmonella species count during April (46.33 ± 31.59 Log 10 CFU/100 ml) and September (48.33 ± 36.55 Log 10 CFU/100 ml) sampling periods, but there was a difference in the E. coli during April (9.67 ± 5.36 Log 10 CFU/100 ml) and September (0.00 ± 0.00 Log 10 CFU/100 ml) sampling periods. However, the E. coli count was significantly lower than the Salmonella species count during both sampling periods (Figure 1b).
Also, in Ishaka, there was neither a difference in the Salmonella species count during April (49.00 ± 28.04 Log 10 CFU/100 ml) and September (87.00 ± 48.85 Log 10 CFU/100 ml) sampling periods, nor was there a difference in the E. coli during April (8.33 ± 2.91 Log 10 CFU/100 ml) and September (3.00 ± 2.52 Log 10 CFU/100 ml) sampling periods. However, the E. coli count was significantly lower than the Salmonella species count during both sampling periods (Figure 1b).
In Bushenyi, there was neither a difference in the Salmonella species count during April (70.33 ± 34.64 Log 10 CFU/100 ml) and September (62.33 ± 32.82 Log 10 CFU/100 ml) sampling periods, nor was there a difference in the E. coli during April (6.33 ± 2.03 Log 10 CFU/100 ml) and September (0.67 ± 0.67 Log 10 CFU/100 ml) sampling periods. However, the E. coli count was significantly lower than the Salmonella species count during both sampling periods (Figure 1b).
Similarly, in Kashenyi, there was no difference in the Salmonella species count during April (58.33 ± 31.39 Log 10 CFU/100 ml) and September (58.67 ± 29.38 Log 10 CFU/100 ml) periods, but there was a difference in the E. coli during April (4.67 ± 2.91 Log 10 CFU/100 ml) and September (0.00 ± 0.00 Log 10 CFU/100 ml). However, the E. coli count was significantly lower than the Salmonella species count during both sampling periods (Figure 1b).
Molecular characterizations/confirmations of presumptive isolates
The target-specific oligonucleotide primer sets produced the expected amplicon sizes at 585 base pairs, which indicated that E. coli was detected from the study (Figure 2a). Ten E. coli isolates were confirmed by molecular detection (PCR); 6 strains were detected/observed in meat and 4 strains detected/observed in milk (Table 3). Similarly, Salmonella spp., was also confirmed during the study as revealed in the PCR detection (Figure 2b), which produced the expected amplicon sizes at 274 base pairs. Nineteen Salmonella spp. strains were confirmed, amongst which 10 strains were confirmed in meat, and 9 strains were confirmed in milk (Table 3).
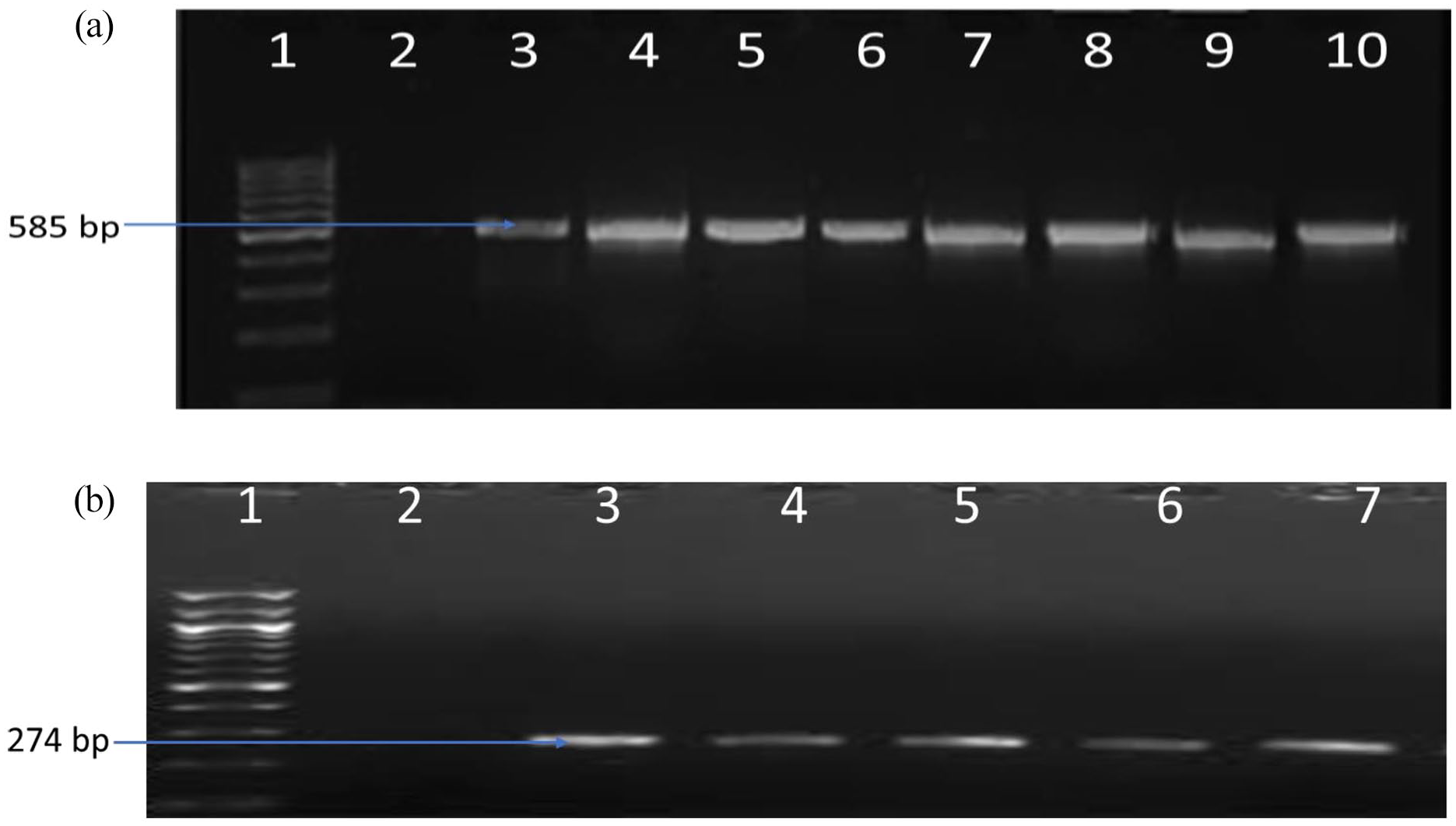
(a) Gel photo representing molecular confirmation of E. coli: Lane 1: Molecular Marker (1 kbp); Lane 2: Negative control; Lane 3: positive control E. coli, Lane 4 to 8 positive confirmed Isolates). (b) Gel picture representing molecular confirmation of Salmonella spp.: Lane 1: Molecular Marker (2 kbp); Lane 2: Negative control; Lane 3: positive control Salmonella, Lane 4 to 7 positive confirmed Isolates).
A genotypic and molecular characterization of presumptive isolates results.

Antibiogram of isolates
Antibiotic Susceptibility Profile of Isolates using various antibiotics
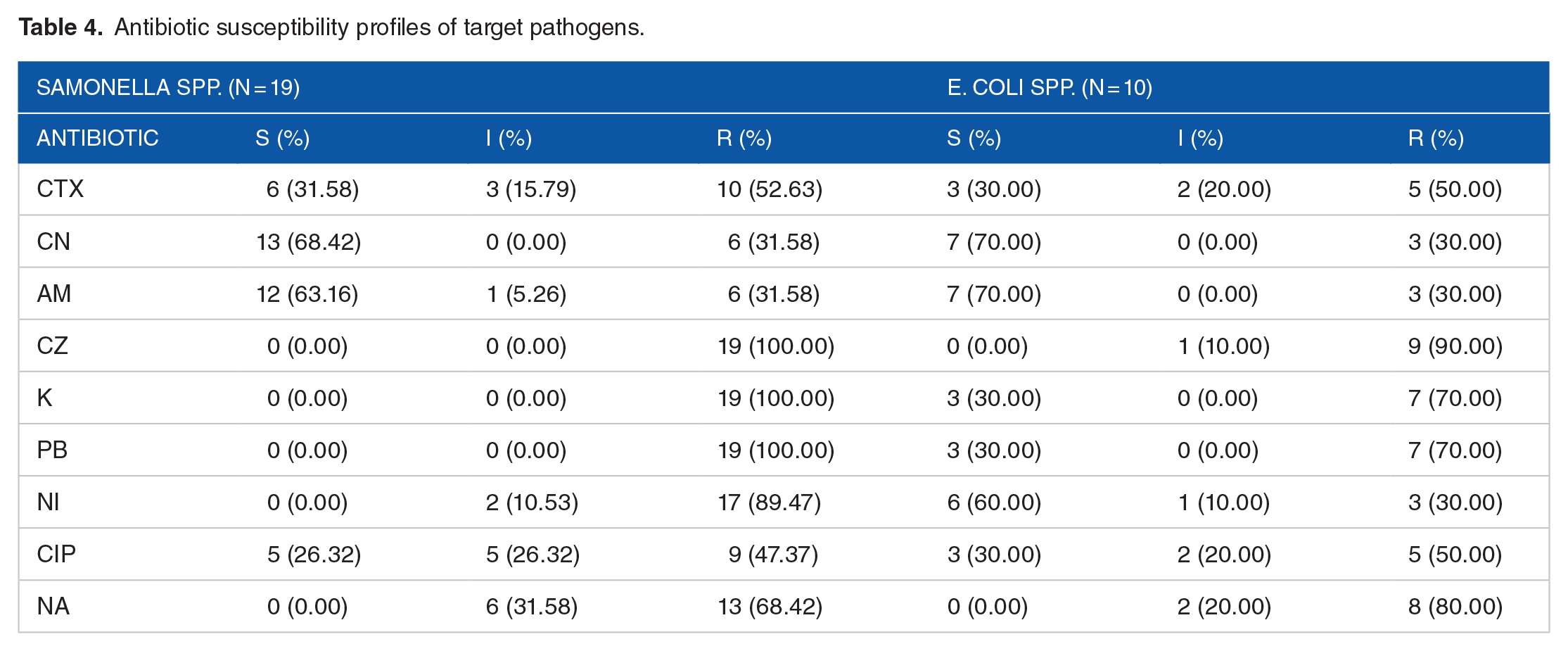
The results depict that of 19 isolates of the Salmonella spp. Nineteen (100.00) were resistant to Cefazolin (CZ) of Cephalosporins, Kanamycin (K) of Aminoglycosides, and Polymixin B (PB; aka Colistin) family of Lipopeptides. Also, a high resistance level of 17 (89.47) was observed against Nitrofurantoin (NI) of the Nitrofuran family, as shown in Table 4. In the case of the E. coli potential pathogens, of out of the 10 isolates profiled for antibiogram and antibiotic signature, 9 (90.0%) of them were resistant to Cefazolin (CZ) of Cephalosporins, followed by 8 (80.00) of them being resistant against Nalidixic Acid (NA) of Fluoroquinolones (Table 4).
Antibiotic susceptibility profiles of target pathogens.
The multi-way heatmap cluster analysis reveals the antibiotic susceptibility profile of 10 E. coli and 19 Salmonella spp. PCR confirmed isolates with the sample locations concerns the interpretation of antibiogram signatures or fingerprints in Figure 3 A & B. The various colours read as follows: yellow = Susceptible range, pink = dose-dependent (intermediate), black = resistance. Based on the signature, the isolates were shown to arise from 2 major clades with multiple progenies amongst the E. coli and Salmonella spp. The profile also showed the multiple antibiotic-resistant nature of the various isolates, indicating that perhaps, if implicated in an infection case, it may be difficult to treat using the selected antibiotics that are routinely used to treat Salmonellosis other enteric pathogenic cases.
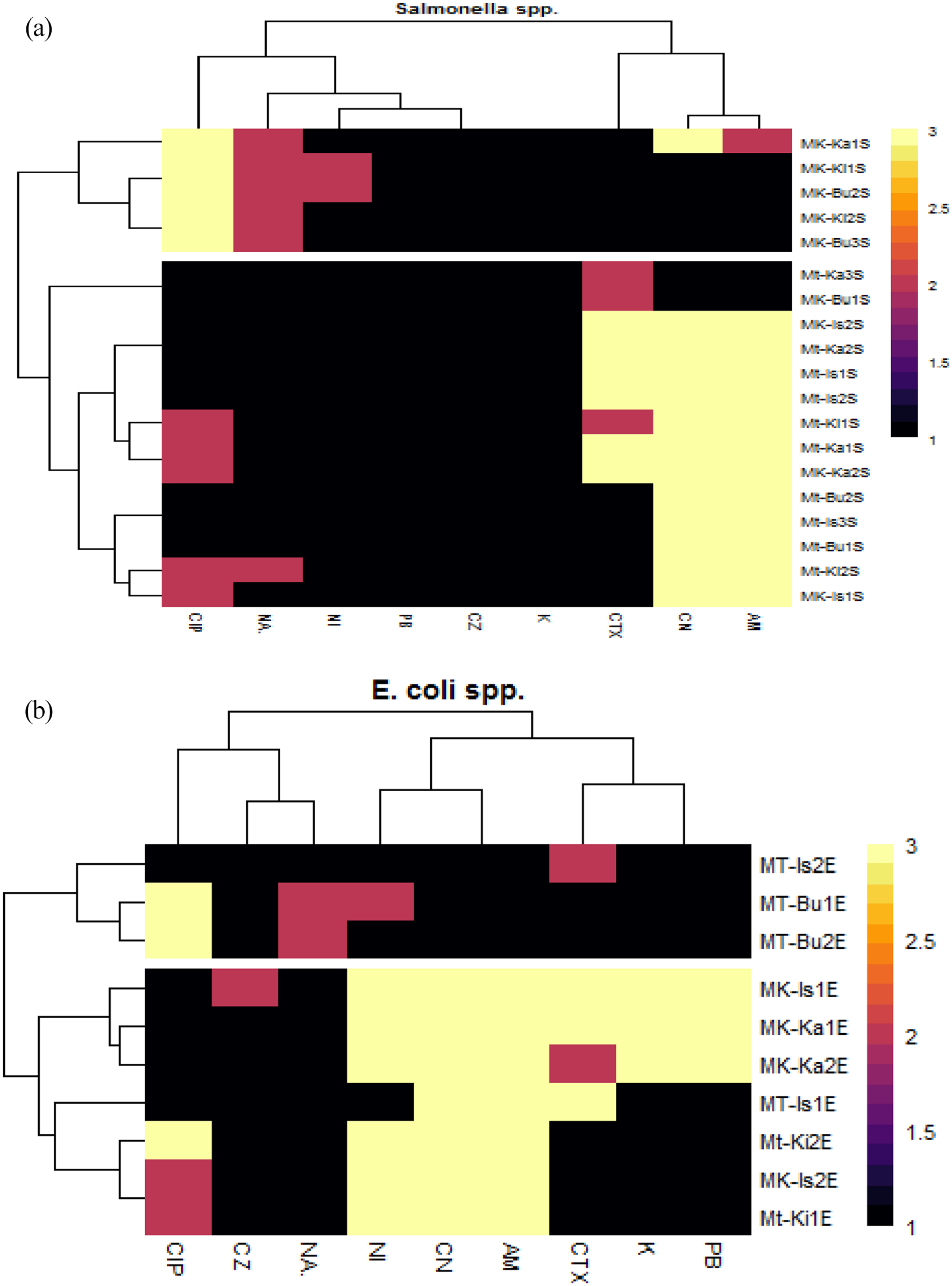
susceptibility of the isolates to antibiotics.
Patterns of multiple antibiotic resistance phenotypes (marps) and multiple antibiotic resistance indices (MARIs)
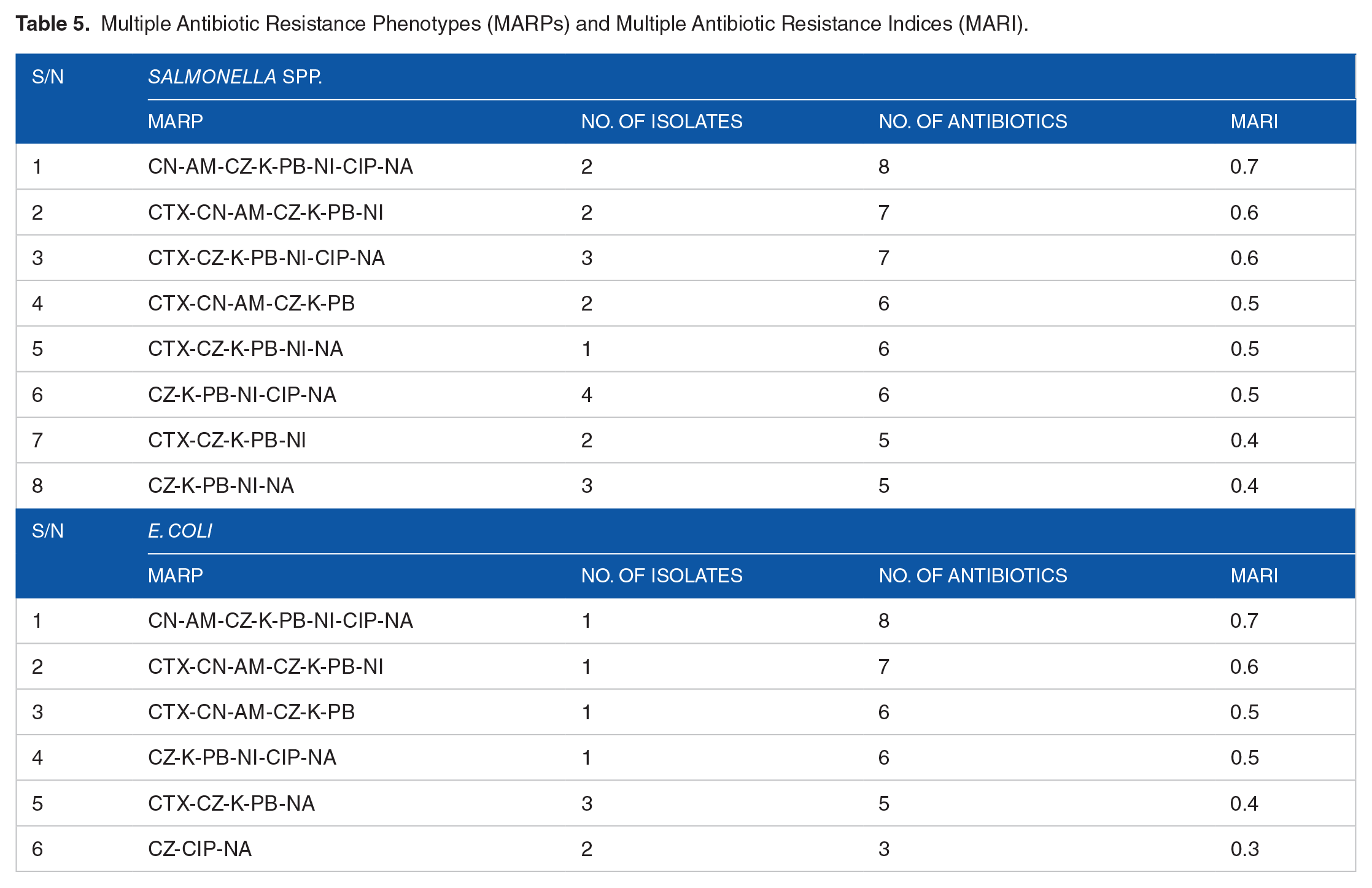
We observed a varied MARP and MARI distribution pattern in Table 5. which implies that the potential pathogens harboured resistance against more than 4 test antibiotics. For Salmonella spp., the CN-AM-CZ-K-PB-NI-CIP-NA MAR-phenotypes was detected in 2 isolates with MARI 0.7 and CZ-K-PB-NI-CIP-NA MAR-phenotypes from 4 isolates with MARI of 0.5. E. coli showed MAR-phenotypes CN-AM-CZ-K-PB-NI-CIP-NA in one isolate with MARI of 0.7 and CTX-CZ-K-PB-NA MAR-phenotypes in 3 isolates with MARI of 0.4, as revealed in Table 5.
Multiple Antibiotic Resistance Phenotypes (MARPs) and Multiple Antibiotic Resistance Indices (MARI).
The potential pathogens which show phenotypic resistance to amoxicillin was 9 (31.03); cefotaxime were 15 (51.72); cefazolin were 28 (96.55) of the classes of penicillins and cephalosporins. Resistance to nitrofurantoin antibiotics was 20 (68.97), while those resistant to kanamycin were 26 (89.66) of the class aminoglycosides. The various isolates were pooled together for the PCR based determination of resistance genes.
Amongst the pooled isolates tested for extended-spectrum beta-lactamase genes (ESBL), the TEM (blaTem-2) has a distribution of 12 (23.1%), 5 (9.6%) for the blaSHV gene. The most distributed genes after the molecular detection were members of the aminoglycoside resistance genes {10 (21.7%)} while other genes were 8 (17.4%) for aadA, (5.8%) bla-CTX-M-gr2, (1.9%) bla-CTX-M-gr9 out of the pooled isolates as represented in Figure 4 and the gel pictures in Supplementary 1 A-D.
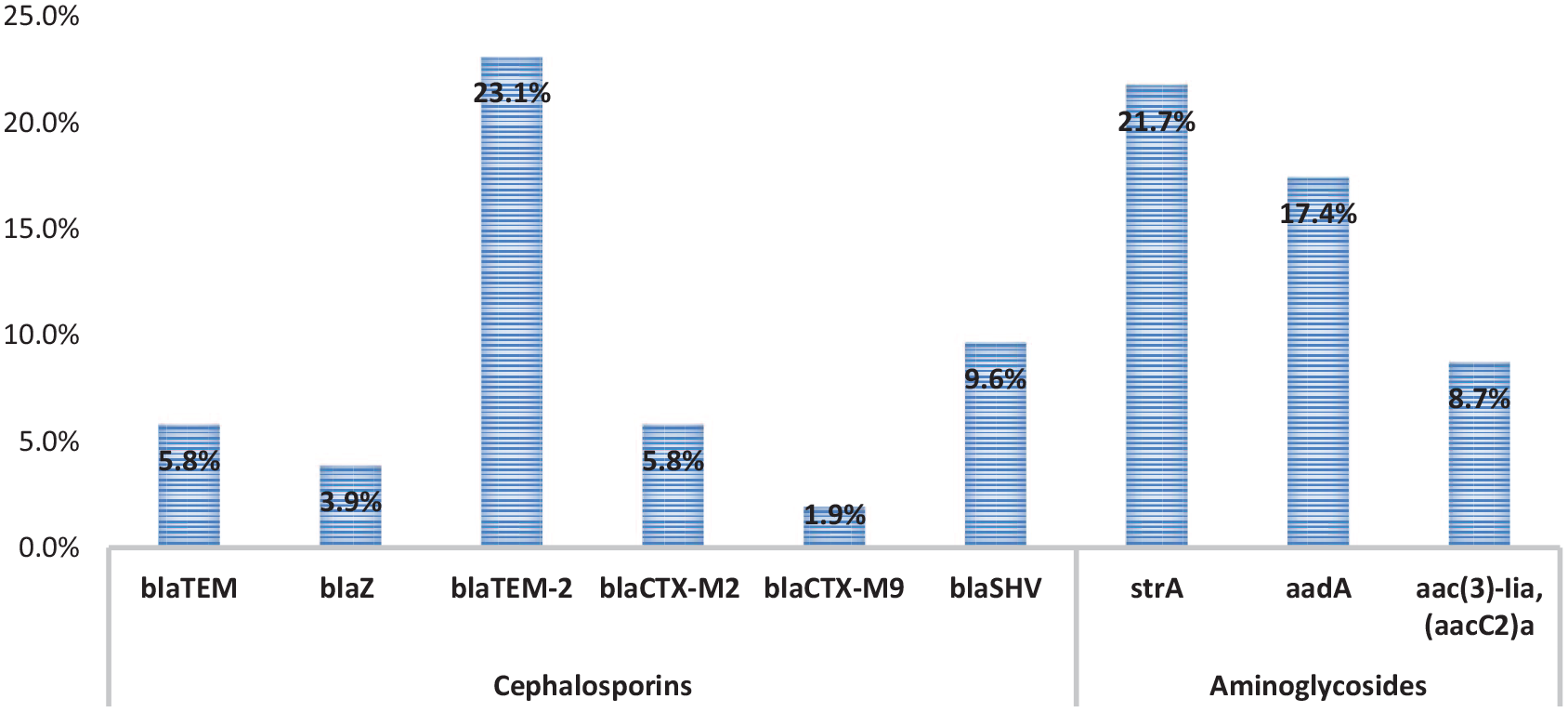
Antibiotic resistance genes of the phenotypic resistant pathogens.
Discussion
Major enterocyte infecting bacterial are currently thriving beyond the expectation even as various control measures are being put in place to combat enteric infections. This study genetically characterized potential pathogens antibiograms and genes encoding antimicrobial resistance of enteric bacterial recovered from foods specimen within the open markets of Bushenyi District, Uganda. The focus was to determine the bacteriological safety of such food specimens currently sold in the public market within Bushenyi District. The results indicate an improvement in hygienic practice as revealed by the total bacterial count for both food specimens (meat and milk). The side-by-side comparison of periodic total presumptive counts for Salmonella spp., and E. coli in both meat and milk reveals a high Salmonella spp. count both in April and September in all the sampling locations. There was a relatively low E. coli total presumptive count in meat and milk compared to that of Salmonella spp. in all sampling locations in September and April. Although there was a low E. coli count in milk collected in September at Ishaka and Bushenyi, there was no observed E. coli count in milk collected in September at Kashenyi and Kizinda during the study. This observation may be attributed to inappropriate hygienic practices in the marketplace and the possible sterility of collected milk.
The bacterial phenotypes also revealed an adverse gram staining, citrate utilization positive, hydrogen sulphide production positive (Salmonella spp.), and indole positive (E. coli). This is also similar to the previous reports of14,25 on the cultural morphology and biochemical identification of bacterial.
The molecular identification/characterization using PCR confirm the present of E. coli and Salmonella spp. in the food samples while other viral-relevance determinants were reported elsewhere. The result of PCR detection showed Salmonella spp. {19 (meat: 10; milk: 9); 54.29%} and E. coli {10 (meat: 6; milk: 4); 22.72%} with more Salmonella spp. detected than E. coli. Salmonella spp., and E. coli were previously reported in food specimen by various investigators from both West, East, North and Southern African countries.26-30 Such observation implies that these members of Enterobacteriaceae infecting potential pathogens thrive within the environment in the region. It also indicates that the food specimen both harbours and serve as carrier/distributor of these potential pathogens. A continuous surveillance/monitoring of food (meat and milk) and other food products, made commercially available in open public markets, remains a cardinal to ascertain the safety of such food products. This routine assessment of food products in markets should also ensure adequate inter-personal hygiene of such food product handlers and the environment.
It is important to note that these isolated strains of potential enteric pathogens of humans and animals are implicated in public health causing diarrhoea infection, enterotoxigenicity, enterohaemorrhagic and uropathogenicity etc. Observing them in food specimen is also a pointer that any human or animal that consumes such products have a predilection for being infected by the pathogens which arouse the need specific and potent for antibacterial therapy.
The antimicrobial susceptibility/antibiogram profile of the various isolates shows that 72.4% (21/29) of isolates were resistant to nalidixic acid, 34.5% (10/29) of isolates were resistant to cefotaxime, 82.8% (24/29) of isolates were resistant to cefazolin, 86.2% (25/29) of isolates were resistant to kanamycin etc. Such resistant phenotypes indicate antibiotic failure, should these members of antibiotics be used in the treatment of any disease case implicated by any of the characterized members of organisms. Similar resistant phenotypes have been previously reported by2,3,9 amongst some enteric associated bacterial pathogens, which show a failure of the various antibiotic regimen used, thereby arousing the need for effective alternative antibiotics. The antibacterial signatures also yielded multiple antibiotic-resistant phenotypes (MARP) with a profile of antibiotic resistance numbers ranging from 4 to 8 and multiple antibiotic-resistant indexes (MARI) range 0.4 to 0.7. Various investigators previously reported such multiple resistance phenotypes and high resistance index on enteric isolates retrieved from the water and food products in significant communities in Africa5,31,32 indicating that such high resistant signature is predominant amongst the enteric pathogens retrieved in African environment. It also indicates that these environments need apt attention in surveillance studies to avoid the various therapeutic concerns arising in the post-antibiotic era.
In a bid to ascertain the observed phenotypes, an antibiotic genotyping and profile was conducted using the PCR technique, which reveals resistance genes belonging to the extended-spectrum beta-lactamase, aminoglycoside resistance genes and other beta-lactamase resistance genes. Amongst the tested isolates, resistance genotypes observed were TEM 12 (23.1%) blaTem-2 gene and 3 (5.8%) blaTEM gene, 5 (9.6%) blaSHV gene, 3 (5.8%) bla-CTX-M-gr2, 1 (1.9%) bla-CTX-M-gr9. Other resistant genes detected were aminoglycoside/streptomycin resistance genes 10 (21.7%) strA and 8 (17.4%) aadA gene. Similar reports of resistant genes have been previously reported amongst multiple antibiotic-resistant Salmonella strains recovered from swine and package vegetables.4,6 According to the study of Iwu and his colleagues, it was reported that the Salmonella species recovered during the study showed high multiple antibiotic-resistant determinants, which may fail antibiotic therapy as well as thriving of the Isolates. Other investigators of related potential pathogens studied resistance characterization amongst Gram-negative isolates. However, their study employed the use of novel and existing clinically relevant resistance indicators. It was reported from their study that resistant genes ranging from aminoglycoside through fluoroquinolone, cephalosporins and other beta-lactam antibiotics were observed.5,9,33 Such observation reveals high resistant markers, which is similar to the report from our study. It is therefore possible that the potential pathogens may be sharing resistant genes via horizontal mechanism amongst other strain which may have informed the repeated resistance gene amongst the food and other food products.4-6,9,33 The need for astute and routine surveillance/monitoring of potential pathogens and food products in public market remains a core for maintaining safety.
Conclusion
This study emphasizes the microbiological safety of some food and food products, commercially available in open public open markets, occurrence of infectious Enterobacteriaceae pathogens and the relevance of molecular biology (PCR) technique in the detection/profiling of such bacterium resistance genes. The study revealed the statistical significance of the bacterial count and detected various strains of E. coli and Salmonella species which were shown to possess various Enterobacteriaceae resistance indices including multiple antibiotic resistance phenotypes and genotypes. Such resistant determinants are currently impacting therapeutic control strategies and public health system, which necessitate an astute surveillance of other related food and food products within the study environment.
Supplemental Material
sj-docx-1-mbi-10.1177_11786361221088992 – Supplemental material for Occurrence, Antibiotic Susceptibility and Genes Encoding Antibacterial Resistance of Salmonella spp. and Escherichia coli From Milk and Meat Sold in Markets of Bushenyi District, Uganda
Supplemental material, sj-docx-1-mbi-10.1177_11786361221088992 for Occurrence, Antibiotic Susceptibility and Genes Encoding Antibacterial Resistance of Salmonella spp. and Escherichia coli From Milk and Meat Sold in Markets of Bushenyi District, Uganda by Hope Onohuean and Bright E Igere in Microbiology Insights
Footnotes
Acknowledgements
The authors wish to appreciate Mr Theophilus Pius of the Department of Microbiology laboratory, KIU Teaching Hospital, for assistance during the laboratory analysis of the samples.
Funding:
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Authors’ Contributions
OH, conceived and designed the study; carried out and analyzed samples data; OH, and IBE interpreted the data. OH, drafted the manuscript. IBE and OH revised the manuscript; read and corrected the final copy of the manuscript.
Availability of Data and Materials
All data, machines, experiments and analysis sources were appropriately acknowledged as necessary while writing the manuscript. Other data generated during the study are attached as supporting documents while any other needed may be available on request.
Declarations
We declare that this study is our original work representing one of the first reported on food and food products within this study area.
Ethical Approval
The protocol for this study was approved by the Ethical Research Committee of the Kampala International University Western Campus, Uganda.
Consent for Publication
The various authors consent for publication was sorted while the journal option was discussed and agreed upon. However, there are no individual authors data/report included in the study
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
