Abstract
Antimicrobial peptides and proteins (AMPs) are molecules that can interact with microbial cells and lead to membrane disruption or intracellular molecule interactions and death. Several molecules with antimicrobial effects also present other biological activities. One such protein group representing the duplicity of activities is the tachykinin family. Tachykinins (TKs) form a family of neuropeptides in vertebrates with a consensus C-terminal region (F-X-G-Y-R-NH2). Invertebrate TKs and TK-related peptides (TKRPs) are subfamilies found in invertebrates that present high homology with TKs and have similar biological effects. Several of these molecules have already been described but reports of TKRP in Hemiptera species are limited. By analyzing the Triatoma infestans hemolymph by reversed-phase high-performance liquid chromatography, biological assays, and mass spectrometry, two antimicrobial molecules were isolated and identified as TKRPs, which we named as TRP1-TINF and TRP2-TINF (tachykinin-related peptides I and II from T. infestans). TRP1-TINF is a random secondary structure peptide with 9 amino acid residues. It is susceptible to aminopeptidases degradation and is active mainly against Micrococcus luteus (32 μM). TRP2-TINF is a 10-amino acid peptide with a 310 helix secondary structure and is susceptible to carboxypeptidases degradation. It has major antimicrobial activity against both Pseudomonas aeruginosa and Escherichia coli (45 μM). Neither molecule is toxic to human erythrocytes and both present minor toxicity toward Vero cells at a concentration of 1000 μM. As the first description of TKRPs with antimicrobial activity in T. infestans, this work contributes to the wider comprehension of the insects’ physiology and describes pharmacological relevant molecules.
Introduction
Antimicrobial peptides and proteins (AMPs) are cationic and amphipathic molecules that can interact with molecules present on cell membranes and with intracellular pathways in microbes, leading to cell disruption and death. 1
These bioactive molecules were initially described in insects by Stephens 2 in 1962, followed by Hink and Briggs 3 in 1968, Powning and Davidson 4 in 1973, Boman et al 5 in 1974, and Faye et al 6 in 1975. Subsequent studies led to the characterization of the first 2 antimicrobial molecules in 1981 by Steiner et al. 7 These molecules were isolated from the moth Hyalophora cecropia, and thus were entitled cecropins, originating of one of the largest antimicrobial peptide groups. Antimicrobial peptides and proteins show a wide distribution and have been isolated from nearly all living organisms.8-21
Beyond their antimicrobial effects, several AMPs have been shown to have other biological activities.11,22-25 One such protein group that represents the duplicity of activities for the same molecules is the tachykinin family. Tachykinins (TKs) form a family of neuropeptides in vertebrates, represented by compounds such as Neurokinin A, 26 Neurokinin B, 27 hemokinin/endokinin K,28,29 and substance P (SP), which was the first TK discovered in alcoholic extracts of equine brain 30 and then isolated from the bovine hypothalamus. Its sequence was determined 40 years later. 31
After the discovery of SP, several other neuropeptides were identified,32-35 all of which contain a consensus C-terminal region (F-X-G-Y-R-NH2).28,36-39 Erspamer, 40 using similar methods as Von Euler and Gaddum, 30 detected a substance in the salivary glands of Eledone moschata that was capable of lowering blood pressure, increasing salivation, and stimulating smooth muscle in rabbits and dogs. This substance was named as eledoisin and was the first non-mammalian TK described.36,37,41,42 Other TKs from frogs and non-mammalian were subsequently described and isolated.32-34,43,44
Champagne and Ribeiro 45 identified 2 TKs from the salivary glands of the mosquito Aedes aegypti, confirming the presence of peptides related to vertebrate TKs in invertebrates. Following this study, 4 molecules from brain, corpora cardiaca, corpora allata, and subesophageal ganglion extracts of the nervous system of Locusta migratoria, named as locust atachykinins, were found to show 30% homology with vertebrate TKs and up to 45% similarity with fish and amphibian TKs.46-48
After these discoveries in invertebrates, 2 new molecule subfamilies were established: the invertebrate tachykinins (Inv-TKs) and tachykinin-related peptides (TKRPs). The main difference between these 2 subfamilies is their C-terminal amino acid composition. Tachykinin-related peptides are clearly related to vertebrate TKs, sharing the same C-terminal region (F-X-G-Y-R-NH2), whereas Inv-TKs differ in their last 2 residues, containing leucine and methionine (F-X-G-L-M-NH2). However, they still maintain high homology and exhibit similar biological effects.
Several Inv-TKs and TKRP have been described in species such as Agrotis ipsilon, 49 Apis mellifera, 50 Camponotus floridanus, 51 Drosophila melanogaster, 52 Drosophila pseudoobscura pseudoobscura, 53 Leucophaea maderae, 54 and Delia radicum. 55
Tachykinin-related peptides have been described in other insects in the order Hemiptera, including in Triatoma infestans, which was analyzed in the current study. Neupert et al 56 described 6 TKRPs produced by Nezara viridula, Banasa dimiata, Pyrrhocoris apterus, Oncopeltus fasciatus, Pentatoma rufipes, and Euschistus servus, and Ons et al 57 identified 8 TKRP sequences through neuropeptidome analysis of Rhodnius prolixus.
There are no further reports of TKRP in other Hemiptera species. Thus, the aim of this study was to isolate and characterize the first 2 TKRPs in the T. infestans hemolymph (Hemiptera: Reduviidae).
Materials and Methods
Microbial strains
The microorganisms Alcaligenes faecalis (ATCC 8750), Aspergillus niger (bread isolated), Candida albicans (IOC 4558), Candida parapsilosis (IOC 4564), Candida tropicalis (IOC 4560), Cladosporium sp. (bread isolated), Cladosporium herbarum (ATCC 26362), Cryptococcus neoformans, Enterobacter cloacae b-12, Escherichia coli (SBS363), Bacillus megaterium (ATCC 10778), Bacillus subtilis (ATCC 6633), Micrococcus luteus (strain A270), M. luteus (Nalidixic resistant), Paecilomyces farinosus (IBCB-215), Penicillium expansum (bread isolated), Pseudomonas aeruginosa (ATCC 27853), Saccharomyces cerevisiae, Serratia marcescens (ATCC 4112), and Staphylococcus aureus (ATCC 29213) were provided by the Special Laboratory of Applied Toxinology from their microorganism library (Butantan Institute, São Paulo, Brazil).
Animals
Both male and female T. infestans (Hemipterans: Reduviidae) were provided by the Ecolyzer Group—Entomology Laboratory (São Paulo, Brazil). The insects were maintained under controlled humidity and temperature in the biotherium of the Special Laboratory of Applied Toxinology (Butantan Institute, São Paulo, Brazil) and fed every 2 weeks with human blood (collected from a healthy volunteer donor and stored in citrate buffer [150 mM, pH 7.4]). 58 The study was conducted under exemption from the Institutional Ethics Committee (CEUAIB No. I-1354/15).
Bacteria inoculation and hemolymph collection
Adult T. infestans were injured with thin needles soaked in a pool of culture of Enterobacter cloacae and M. luteus in the logarithmic phase. At 72 hours after induction, hemolymph samples were obtained by excising the metathoracic legs of the insects and pressing gently on the abdomen. Drops were collected with micropipettes and pooled in sterile Eppendorf tubes on ice containing phenylthiourea to avoid activation of the phenol oxidase cascade. 5 The solution was stored at −80°C until use.
Sample fractionation
Intracellular content extraction
To release the intracellular components from the hemolymph, the sample was subjected to acid extraction in the presence of acetic acid (2 M). The supernatant was obtained by centrifugation at 16.000 ×g for 30 minutes at 4°C and injected directly into 2 coupled Sep-Pack C18 cartridges (Waters, Milford, MA, USA) equilibrated in 0.1% trifluoroacetic acid (TFA). The sample was eluted with 3 different concentrations of acetonitrile (ACN) in water (5%, 40%, and 80%) and then concentrated in a vacuum centrifuge and reconstituted with ultrapure water.
Reversed-phase high-performance liquid chromatography
Separation steps were performed using high-performance liquid chromatography (HPLC) using a reverse-phase semi-preparative C18 column (Jupiter, 10 × 250 mm; Phenomenex, Torrance, CA, USA) equilibrated in 0.05% TFA. The elution gradient for the fraction eluted with 5% ACN was from 2% solution A (0.05% [v/v] TFA in water) to 20% solution B (0.10% [v/v] TFA in ACN). For the 40% ACN fraction, the elution gradient was 2% to 60% solution B (0.10% [v/v] TFA in ACN). For the 80% ACN fraction, the elution gradient was 20% to 80% solution B (0.10% [v/v] TFA in ACN) in solution A. Each run was performed over 60 minutes at a flow rate of 1.5 mL/min.
Effluent absorbance was monitored at 225 nm, and fractions corresponding to the absorbance peaks were hand-collected, concentrated under vacuum, and reconstituted in ultrapure water.
When necessary, a second chromatography step was performed. The gradient for the second chromatography was determined by evaluation of the molecule’s retention time and performed on a VP-ODS analytic C18 column (Shim-pack; Shimadzu, Kyoto, Japan) at a flow rate of 1.0 mL/min for 60 minutes.
Liquid growth inhibition assay
All fractions obtained by HPLC and synthetic peptides were evaluated by antimicrobial screening against the microorganism strains (see section “Microbial Strains”)8,21 using poor broth nutrient medium (PB: 1.0 g peptone in 100 mL of water containing 86 mM NaCl at pH 7.4; 217 mOsM) and Müller-Hinton medium (peptone 5.0 g/L; casein peptone 17.5 g/L; agar 15.0 g/L; Ca2+ 20.0-25.0 mg/L; Mg2+ 10.0-14.5 mg/L; pH 7.4) for bacteria and poor dextrose broth (1/2 PDB: 1.2 g potato dextrose in 100 mL of H2O at pH 5.0; 79 mOsM) and RPMI 1640 (Roswell Park Memorial Institute medium) medium with MOPS 0.165 mol/L (RPMI without bicarbonate 10.4 g/L; MOPS [3-(n-morpholino) propanesulfonic acid] 34.53 g/L; pH 7.0) at half half-strength.59,60
Antimicrobial activity was determined in a 5-fold microliter broth dilution assay in 96-well sterile plates at a final volume of 100 µL. The mid-log phase microbial culture was diluted to a final concentration of 1 × 105 colony-forming units per milliliter.59,61,62
Lyophilized fractions were dissolved in 500 µL ultrapure water, and 20 μL peptide solution was aliquoted into each well containing 80 μL of the bacterial dilution, giving a final volume of 100 μL. To determine the minimal inhibition concentration, 20 μL peptide stock solution was added to each well of the microtiter plate at a 2-fold serial dilution and added to 80 μL of the bacterial dilution. Sterile water and PB were used as growth controls, and streptomycin was used as a growth inhibitor control.
The microtiter plates were incubated for 18 hours at 30°C, and then growth inhibition was determined by measuring the absorbance at 595 nm.63,64 To determine the minimal inhibition concentration, the bacterial growth rates were measured after 18 hours of incubation. To determine the minimal bactericidal concentration, the bacterial growth rates were measured at 595 nm after 96 hours.63,64 All antimicrobial experiments were performed in triplicate.
Mass spectrometry
Active antibacterial fractions were analyzed by liquid chromatography-tandem mass spectrometry on an LTQ-Orbitrap Velos (Thermo Fisher Scientific, Waltham, MA, USA) coupled to an Easy-nLCII liquid nano-chromatography system (Thermo Fisher Scientific). For chromatography, 5 µL of each sample was automatically separated on a C18 pre-column (100 mm I.D. × 50 mm; Jupiter 10 mm; Phenomenex) coupled to a C18 analytical column (75 mm I.D. × 100 mm; ACQUA 5 mm; Phenomenex). The eluate was electro-sprayed at 2 kV and 200°C in positive ion mode. Mass spectra were acquired with a Fourier transform-based mass spectrometer; full scan (MS1) involved using 200-2000 m/z (60 000 resolution at 400 m/z) with a mass scan interval in data-dependent acquisition mode. The 5 most intense ions in each scan were selected for fragmentation by collision-induced dissociation. The minimum threshold for selecting an ion for a fragmentation event (MS2) was set to 5000. The dynamic exclusion time was 15 seconds, repeating at 30-second intervals.
Computational analysis
Mass spectrometry data were analyzed using Mascot Daemon software (version 2.2.2) through database searching using the Swiss-Prot, NCBInr, Hemipteras, and Triatomineos banks for comparison. Homology searching was performed using the PepBank peptide database, 65 Signal Peptide Database, 66 Vector Base, 67 APD3: Antimicrobial Peptide Calculator and Predictor, 68 and BLAST: Basic Local Alignment Search Tool. 69 The results were considered valid only when they were reproducible by different analyses.
Primary structure alignment
Alignment of primary sequences was performed using ClustalW2 version 2 online software 70 with default parameters.
Solid-phase peptide synthesis
Peptides were synthesized using the solid-phase method 71 using methylbenzhydrylamine resin (MBHAR) and employing the t-Boc strategy. For synthetic peptide entitled TRP2-TINF, the methionine (second amino acid in the C-terminal region) was changed to norleucine, reducing the molecular weight to 1006 Da. After cleaving the peptides from the resin, the peptides were purified from the lyophilized crude solutions by HPLC on a C18 column. To guarantee high purity and to characterize the peptides, LC-ESI-MS equipment was used.
Synthetic peptide concentration
Peptide concentrations were determined using the Lambert-Beer law using the molar extinction coefficient at 205 nm absorption, 72 obtained using the tool available at http://nickanthis.com/tools/a205.html.
Peptide stability in serum
Aliquots of 20 µL peptide solution (10 mg/mL) were added to 1 mL of 25% non-heat inactivated horse serum (Sigma-Aldrich, St. Louis, MO, USA) in phosphate-buffered saline (PBS) and incubated at 37°C in triplicate for different time intervals (0, 10, 30, 60, and 120 minutes). During incubation, 100 µL of sample was withdrawn incubated with 10 µL of pure TFA at 5°C for 15 minutes. The resulting mixtures were centrifuged at 300 ×g for 5 minutes. A volume of 30 µL of the supernatants was injected in an online HPLC coupled to a mass spectrometer (LC-ESI-MS). Peptide consumption was evaluated using a linear gradient of ACN in acidified water from 3% to 57% in 30 minutes at a flow rate of 0.4 mL/min, followed by the measurement of the area decrease of the peak that corresponds to the peptide in the chromatogram. 73
Circular dichroism
Circular dichroism (CD) measurements were performed on a Jasco J-815 circular dichroism spectropolarimeter (Jasco Corp., Tokyo, Japan). For UV measurements (190-250 nm), CD spectra were recorded after 4 accumulations at 20°C using a 0.5-mm pathlength quartz cell between 250 and 195 nm at 50 nm/min with a bandwidth of 0.5 nm. Both peptides were analyzed in 0%, 10%, 30%, and 50% v/v solutions of 2,2,2 trifluoroethanol in water. Fast Fourier transform was applied to minimize background effects. 74
Cytotoxic assays
The toxicity of the peptides against Vero cells (African green monkey kidney fibroblasts) was evaluated. Cells were obtained from the American Type Culture Collection (ATCC CCL81; Manassas, VA, USA) and maintained at 37°C in T culture flasks (25 cm2) containing 5 mL Leibovitz medium (L-15) supplemented with 10% fetal bovine serum, both from Cultilab (Campinas, Brazil) or in a 96-well microplate. Toxicity was determined using the MTT colorimetric assay. Briefly, the cells were seeded into 96-well plates (2 × 105 cells/well) and cultured for 24 hours at 37°C in a humidified atmosphere containing 5% CO2. Twelve 2-fold serial dilutions of both peptides were performed with L-15 to give solutions with final concentrations ranging from 1.95 to 1000 μM. Varying concentrations were added and allowed to react with the cells for 24 and 48 hours, followed by addition of 20 μL MTT (5 mg/mL in PBS) and incubation for another 4 hours at 37°C. Formazan crystals were dissolved by adding 100 μL dimethyl sulfoxide and incubation at room temperature until all crystals were dissolved. Absorbance at 550 nm was measured using a microplate ELISA reader (1420 Multilabel Counter/Victor3; Perkin Elmer, Waltham, MA, USA). Cell survival was calculated using the following formula: survival (%) = (A550 of peptide-treated cells/A550 of peptide-untreated cells) × 100. 20
Hemolytic assay
Human erythrocytes from a healthy donor were collected in 0.15 M citrate buffer (pH 7.4) washed 3 times by centrifugation (700 ×g, for 10 minutes at 4°C) with 0.15 M PBS containing 137 mM NaCl, 2.7 mM KCI, 10 mM Na2HPO4, 1.76 mM KH2PO4 (pH 7.4) and resuspended in PBS to a final concentration of 3% (v/v). The peptides (serial 2-fold dilutions in PBS) were added to 80 µL erythrocyte suspension to a final volume of 100 μL and incubated for 1 hour at 37°C. Hemoglobin release was monitored by measuring the supernatant absorbance at 405 nm with a Microplate ELISA Reader (1420 Multilabel Counter/Victor3). The hemolysis percentage was expressed in relation to a 100% lysis control (erythrocytes incubated with 0.1% Triton X-100); PBS was used as a negative control.12,75,76
Results and Discussion
Sample fractionation and peptide purification
The hemolymph was processed and separated on a Sep-Pack C18 column using 5%, 40%, and 80% ACN concentrations to elute the main sample into three different fractions (see section “Sample fractionation”). During purification of the 5% ACN sample by RP-HPLC, the fraction eluted at 67.3 minutes (sample 1; Figure 1A) showed antimicrobial against E. coli, whereas the fraction eluted at 56.2 minutes during 40% ACN sample purification by RP-HPLC (sample 2; Figure 2A) showed antimicrobial activity against M. luteus. Both were completely isolated in the second chromatography step (Figures 1B and 2B).
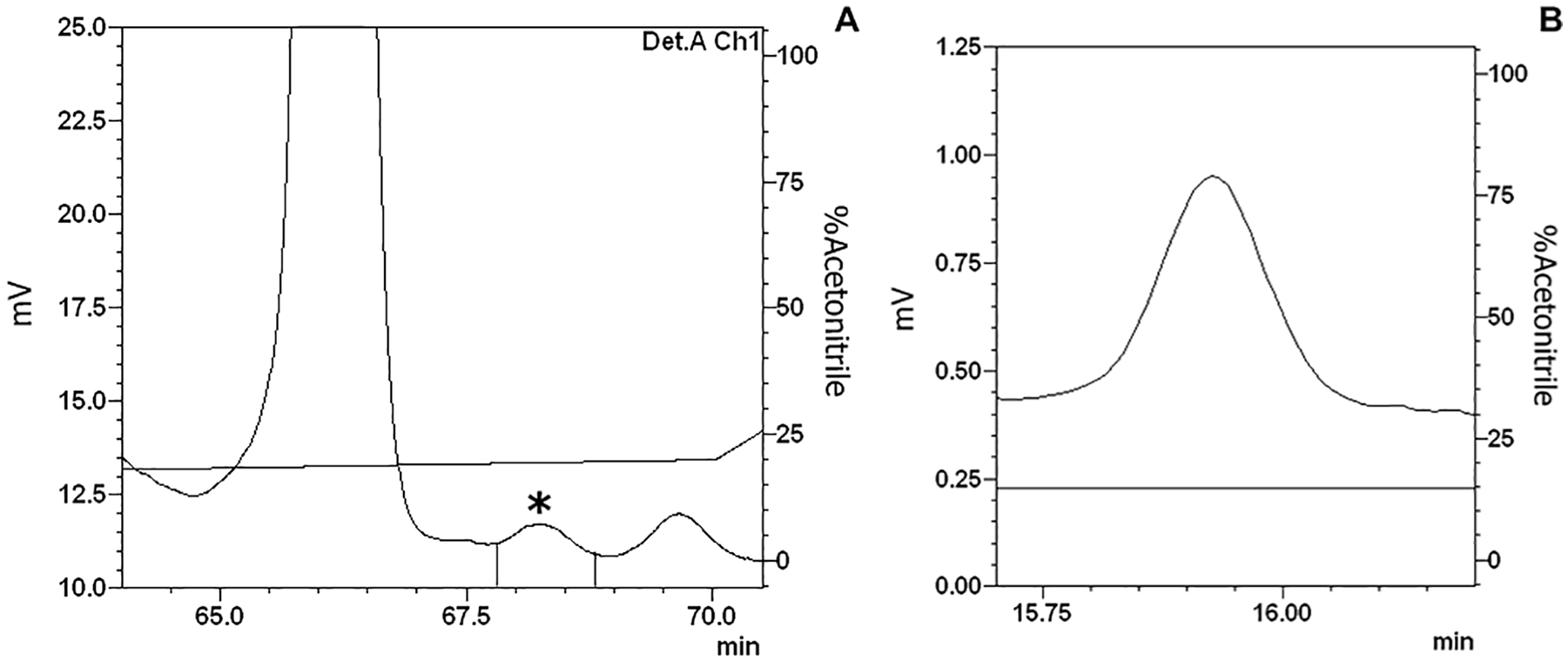
5% ACN active fraction purification—sample 1: (A) The 5% ACN fraction isolated from Triatoma infestans hemolymph was separated by RP-HPLC using a C18 column, eluted over a linear gradient from solution A from 0% to 20% of solution B in 60 minutes. The labeled fraction (*), eluted at 67.3 minutes, exhibited antimicrobial activity and was subjected to a second chromatography step. (B) The second RP-chromatographic step on an analytic VP-ODS C18 column, with an ACN gradient from 13% to 23% solution B in 60 minutes, to guarantee fraction homogeneity.
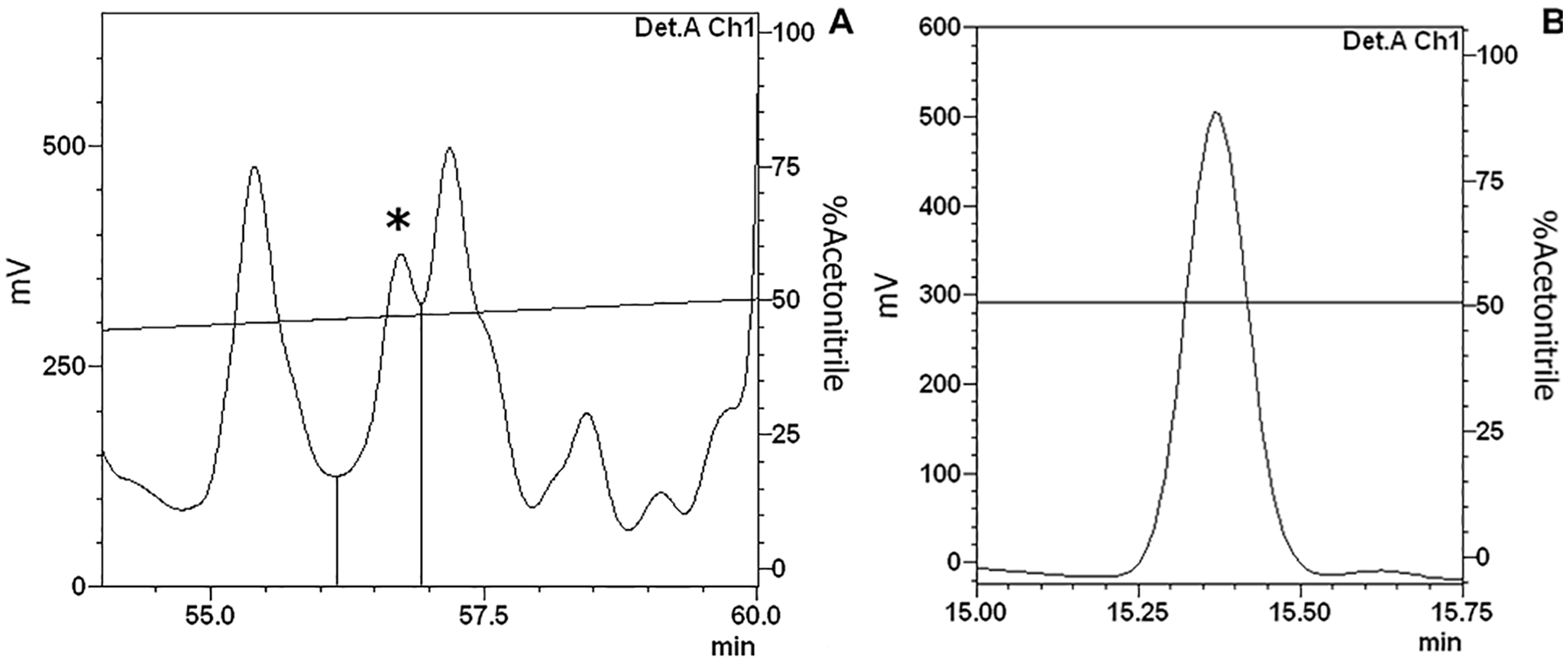
40% ACN active fraction purification—sample 2: (A) The 40% ACN fraction isolated was separated by RP-HPLC using a C18 column, eluted over a linear gradient from solution A from 2% to 60% of solution B in 60 minutes. The labeled fraction (*), eluted at 56.2 minutes, exhibited antimicrobial activity and was subjected to a second chromatography step. (B) The second RP-chromatographic step on an analytic VP-ODS C18 column with an ACN gradient from 49% to 59% solution B in 60 minutes to guarantee its homogeneity.
Peptide identification
After purification, both samples were examined by mass spectrometry and the data were analyzed in MagTran 1.02. We verified that samples 1 and 2 had approximated molecular weights of 951 and 1035 Da, respectively (Figure 3).
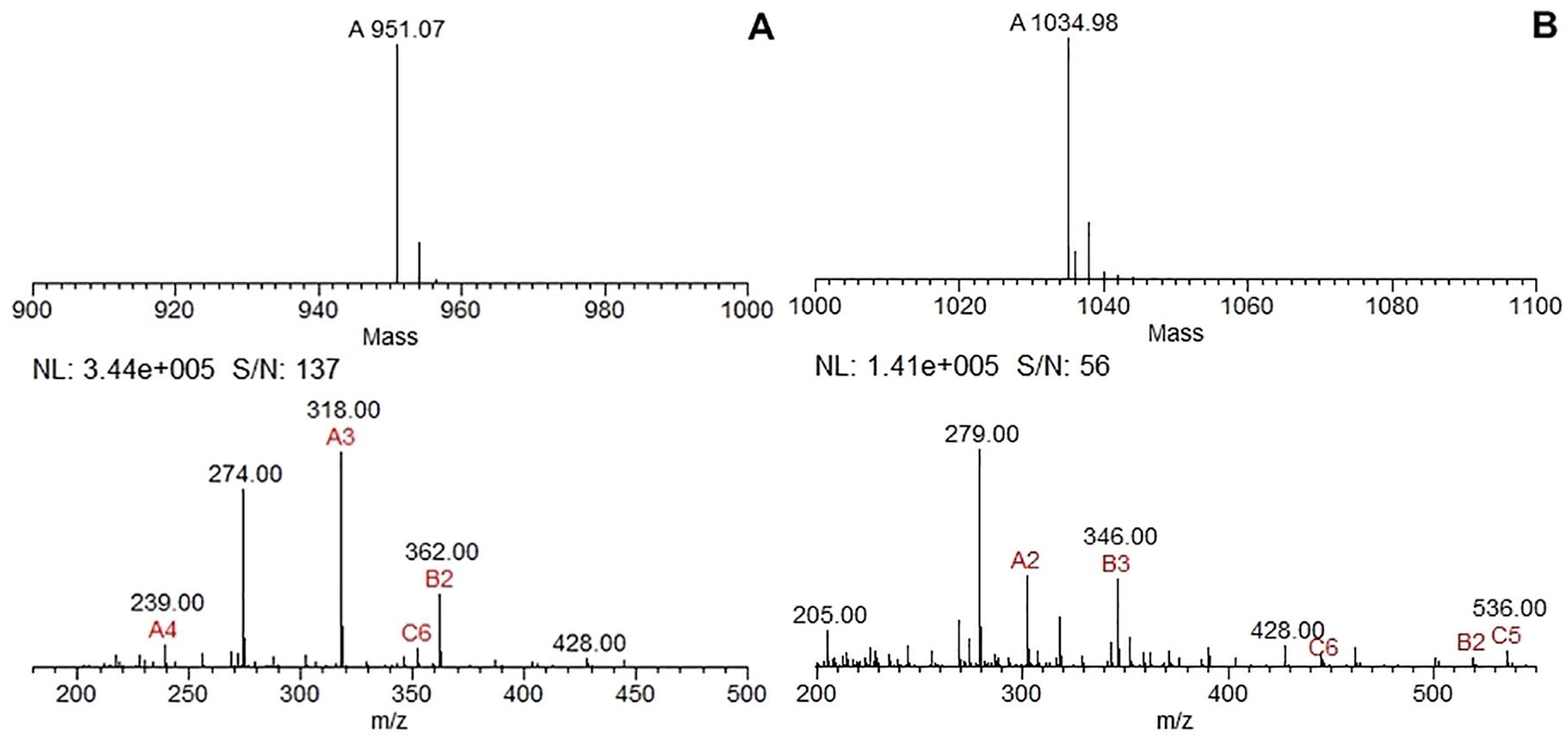
Mass determination using MagTran software: (A) Sample 1 had an approximate mass of 951.07 Da. (B) Sample 2 had an approximate mass of 1034.98 Da.
To determine the amino acid content and perform protein identification, mass spectrometry data were analyzed with Mascot software with the Fingerprint tool and using the Hemiptera database (118 063 sequences). Both samples showed similarities with tachykinin-related peptides I and II (TRP1 and TRP2), respectively, from different insect species (Figure 4).
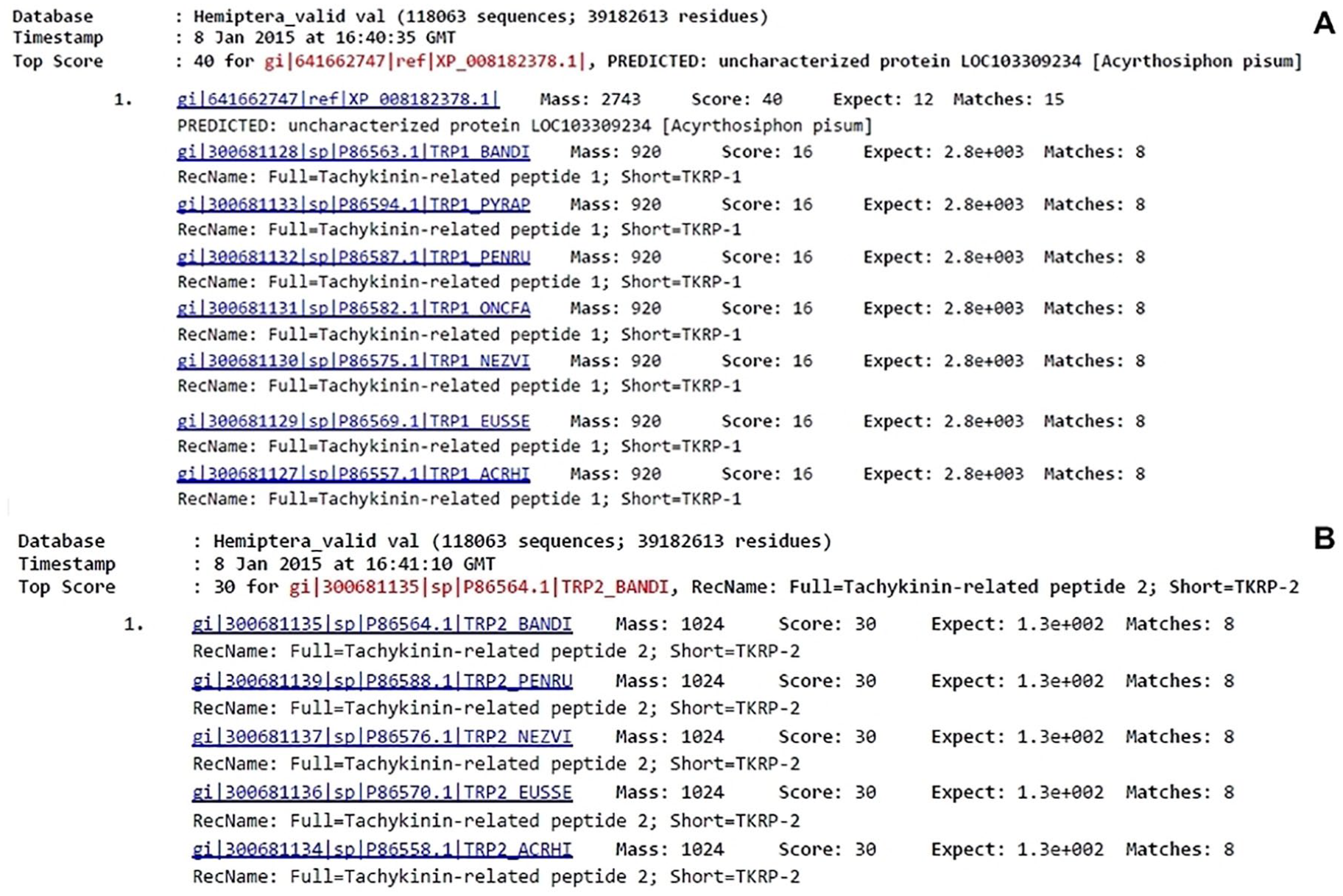
Mass spectrometry analysis using MASCOT software: (A) Sample 1 mass spectrometry data compared with a Hemiptera database (118 063 sequences). Sample showed similarity with tachykinin-related peptide I from Banasa dimiata, Pyrrhocoris apterus, Pentatoma rufipes, Oncopeltus fasciatus, Nezara viridula, Euschistus servus, and Acrosternum hilare. (B) Sample 2 mass spectrometry data compared with a Hemiptera database (118 063 sequences). Sample showed similarity with tachykinin-related peptide II from B. dimiata, P. rufipes, N. viridula, E. servus, and A. hilare.
According to the official TKRP nomenclature, these peptides were named as TRP1-TINF and TRP2-TINF (tachykinin-related peptides I and II from T. infestans). As observed for TRP1-TINF, the first match corresponded to an uncharacterized protein from Acyrthosiphon pisum. However, this protein showed an expected threshold P-value > .05, and thus this result was discarded. Showing 100% coverage with the corresponding sequences, TRP1-TINF is composed of 9 amino acid residues with the sequence GPSGFLGMR and RP2-TINF is composed of 10 amino acid residues with the sequence APAAGFFGMR.
Tachykinins are proteins that comprise a superfamily initially described by Von Euler and Gaddum 30 and have neurotransmitter/neuromodulator activities as their main physiological functions. A subfamily discovered in insects, named as tachykinin-related peptides (or tachykinin-like peptides), has been widely studied.36,37 The production of TKRP by species in the order Hemiptera has been described, and all sequences are summarized in Table 1.
Relationships with TKRPs from Hemipterans.
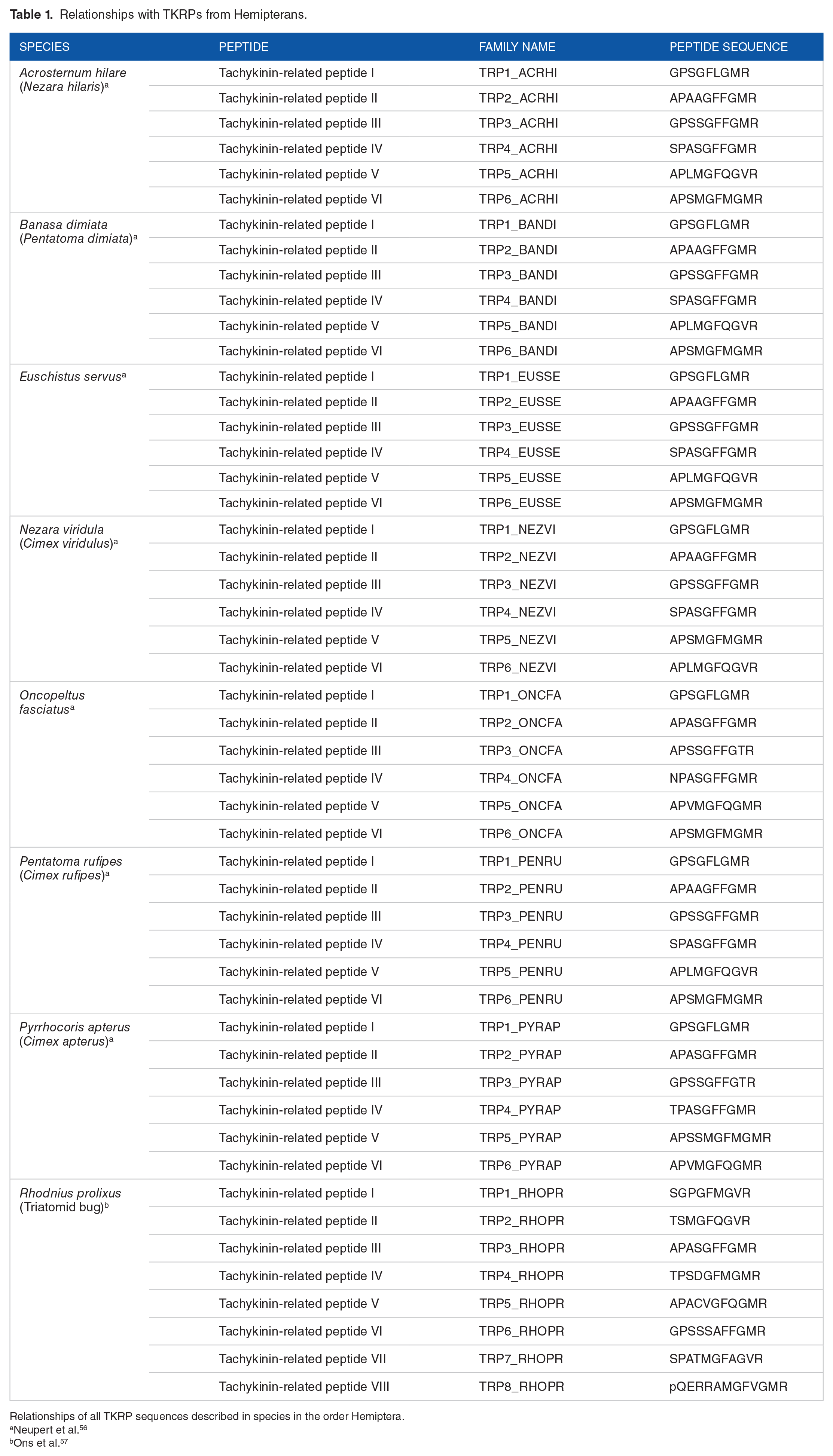
Relationships of all TKRP sequences described in species in the order Hemiptera.
Neupert et al. 56
Ons et al. 57
As Rhodnius prolixus is phylogenetically close to T. infestans, primary sequence alignment was performed to compare their most similar sequences (Figure 5), which revealed a close resemblance.

Primarily sequence alignment. Tachykinin-related peptides I and II from Triatoma infestans (TRP1-TINF and TRP2-TINF) compared with tachykinin-related peptides I and III from Rhodnius prolixus (TRP2_RHOPR and TRP3_RHOPR). Amino acid residues highly conserved between the sequences are highlighted in yellow. (*) Position with a single and fully conserved amino acid residue; (:) Position with amino acid residues conserved between groups of strong similar properties; (.) Position with amino acid residues conserved between groups of weakly similar properties.
Recent studies have shown that a few components of the vertebrate tachykinin family have antimicrobial activity against some bacterial strains such as S. marcescens and P. aeruginosa.77-79 Although none of the TKRP sequences mentioned above were evaluated in antimicrobial assays, urechistachykinin I and urechistachykinin II, which are tachykinin-related neuropeptides isolated from the Echiuroid worm 44 and share the C-terminal region, presented antimicrobial activity against different Gram-positive and Gram-negative bacteria and different fungi. 80
As a main characteristic of the peptide family, all sequences share the same C-terminal region (F-X-G-Y-R-NH2); thus, it is expected that they have similar biological effects.
Characterization of isolated TKRPs
Bioassays
TRP1-TINF and TRP2-TINF were artificially synthetized. For synthetic TRP2-TINF, the methionine (second amino acid in the C-terminal region) was changed to norleucine, reducing the molecular weight to 1006 Da. For comparison, the synthetic peptides were also tested against M. luteus and E. coli. Both peptides were active against the two microorganisms, indicating the antimicrobial potential of the synthetic molecules.
The synthetic peptides were tested against a broad range of bacterial, yeast, and fungal species and their antimicrobial activities are reported in Tables 2 and 3. The initial concentration of TRP1-TINF was 128 µM and that of TRP2-TINF was 164 µM.
Synthetic TRP1-TINF antimicrobial activities against bacterial and fungal strains.
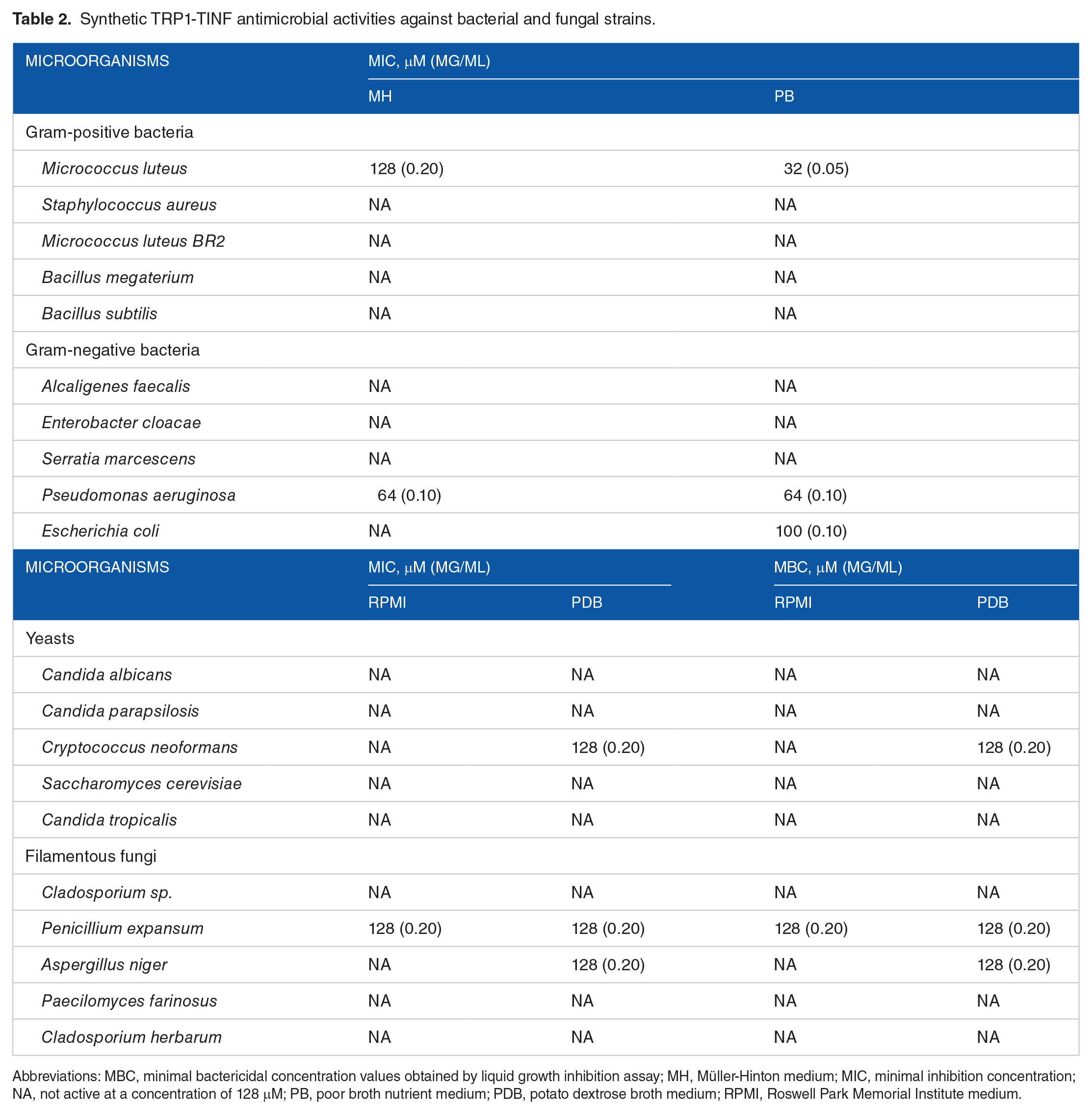
Abbreviations: MBC, minimal bactericidal concentration values obtained by liquid growth inhibition assay; MH, Müller-Hinton medium; MIC, minimal inhibition concentration; NA, not active at a concentration of 128 μM; PB, poor broth nutrient medium; PDB, potato dextrose broth medium; RPMI, Roswell Park Memorial Institute medium.
Synthetic TRP2-TINF antimicrobial activities against bacterial and fungal strains.
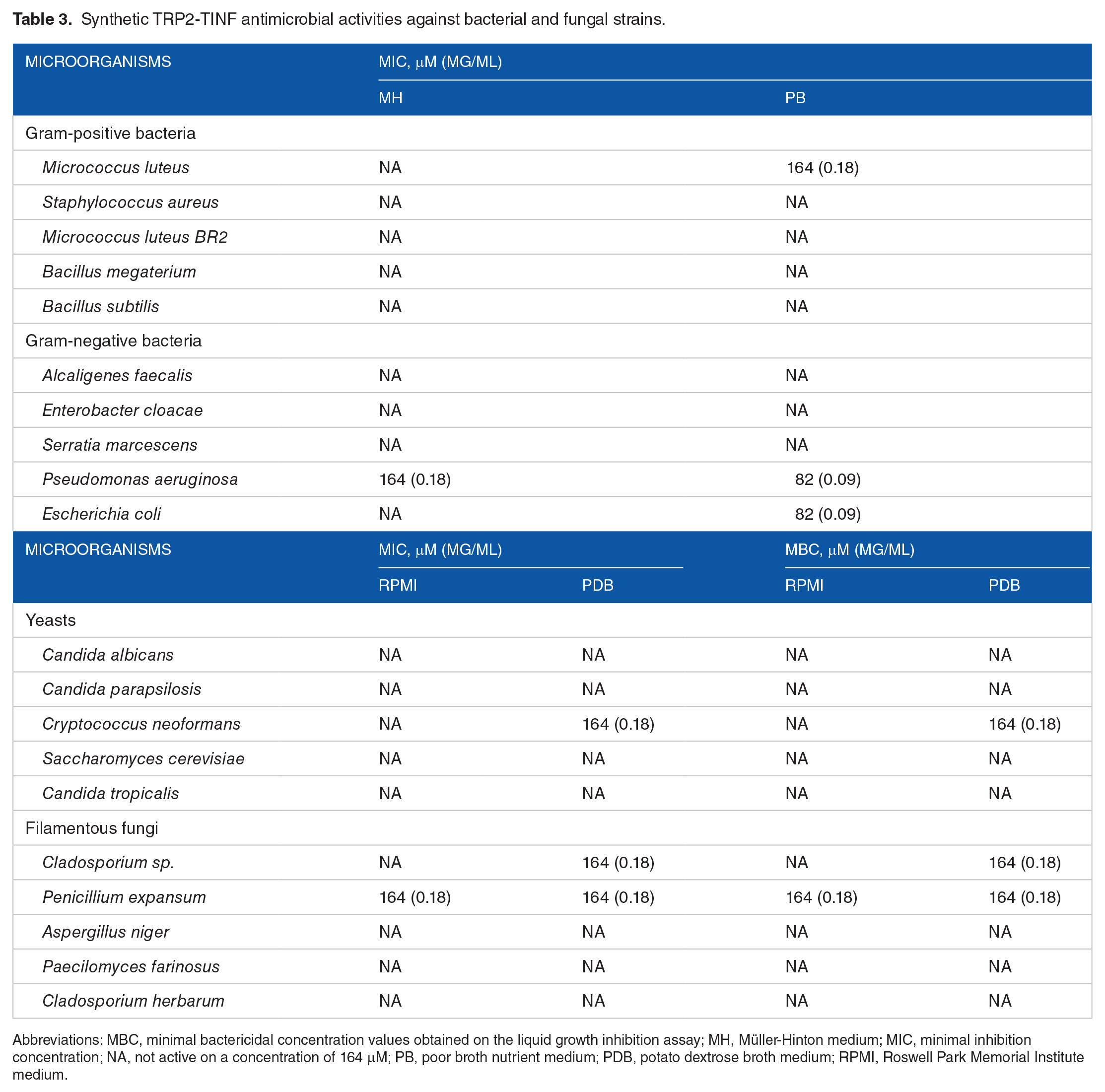
Abbreviations: MBC, minimal bactericidal concentration values obtained on the liquid growth inhibition assay; MH, Müller-Hinton medium; MIC, minimal inhibition concentration; NA, not active on a concentration of 164 μM; PB, poor broth nutrient medium; PDB, potato dextrose broth medium; RPMI, Roswell Park Memorial Institute medium.
Both TRP1-TINF and TRP2-TINF could partially or completely inhibit the growth of several bacteria and fungi. Neither peptide showed specific activities, although both could disrupt bacterial cells at a lower concentration than that required for disrupting fungi cells. For instance, TRP1-TINF was active against Gram-positive and Gram-negative bacteria (M. luteus with 32 µM and P. aeruginosa with 64 µM), and TRP2-TINF has predilection toward Gram-negative bacteria, presenting activity with lower concentrations toward P. aeruginosa and E. coli.
Hemolysis percentages of 0.9% and 0.6% were observed when red blood cells were incubated with 300 μM of TRP1-TINF and 360 μM of TRP2-TINF, respectively. Thus, the peptides showed no significant hemolytic activity (P < .05).
In contrast, when incubated Vero cells with the highest tested concentration of both peptides (1000 μM), cell viability was approximately 84% and 90% after exposure to TRP1-TINF and TRP2-TINF, respectively (Figure 6). The selectivity index was not calculated, as no median lethal dose values were found at the maximum evaluated concentrations.
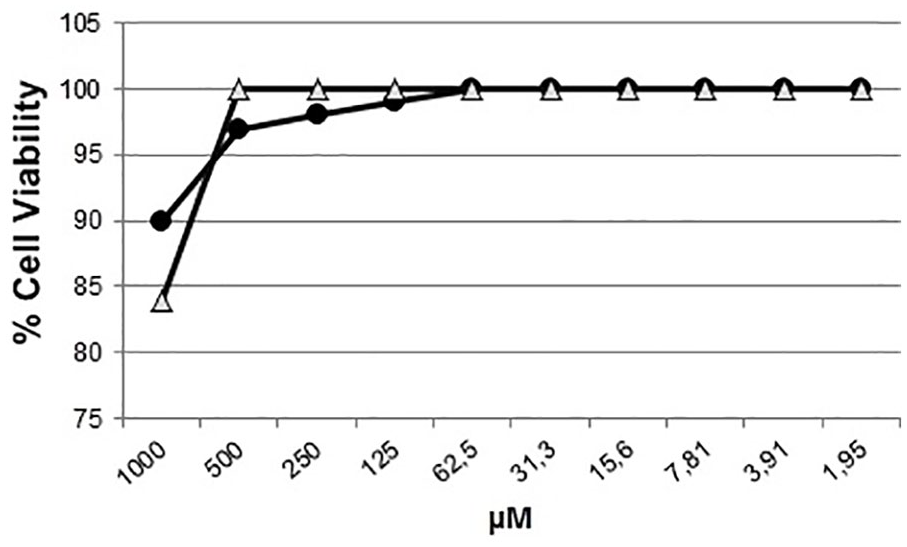
TRP1-TINF and TRP2-TINF toxicity assays. Cytotoxicity of TRP1-TINF (Δ) and TRP2-TINF (•) over Vero cells in a 2-fold series dilution at a maximum concentration of 1000 μM.
To investigate the physicochemical dynamics of the peptides, we performed tridimensional structure analysis and degradation analysis in plasma assays. In CD analysis, the structural conformation of asymmetric molecules was obtained from the patterns of absorption and refraction of the absorbance of right- and left-handed circularly polarized light81,82 as a hygroscopic liquid was added to the original solution.
As shown on the CD spectra, TRP1-TINF (Figure 7A) tended to assume a random coil conformation, whereas TRP2-TINF (Figure 7B) tended to assume a Poly (Pro) II (PPII) or helix-31 conformation.
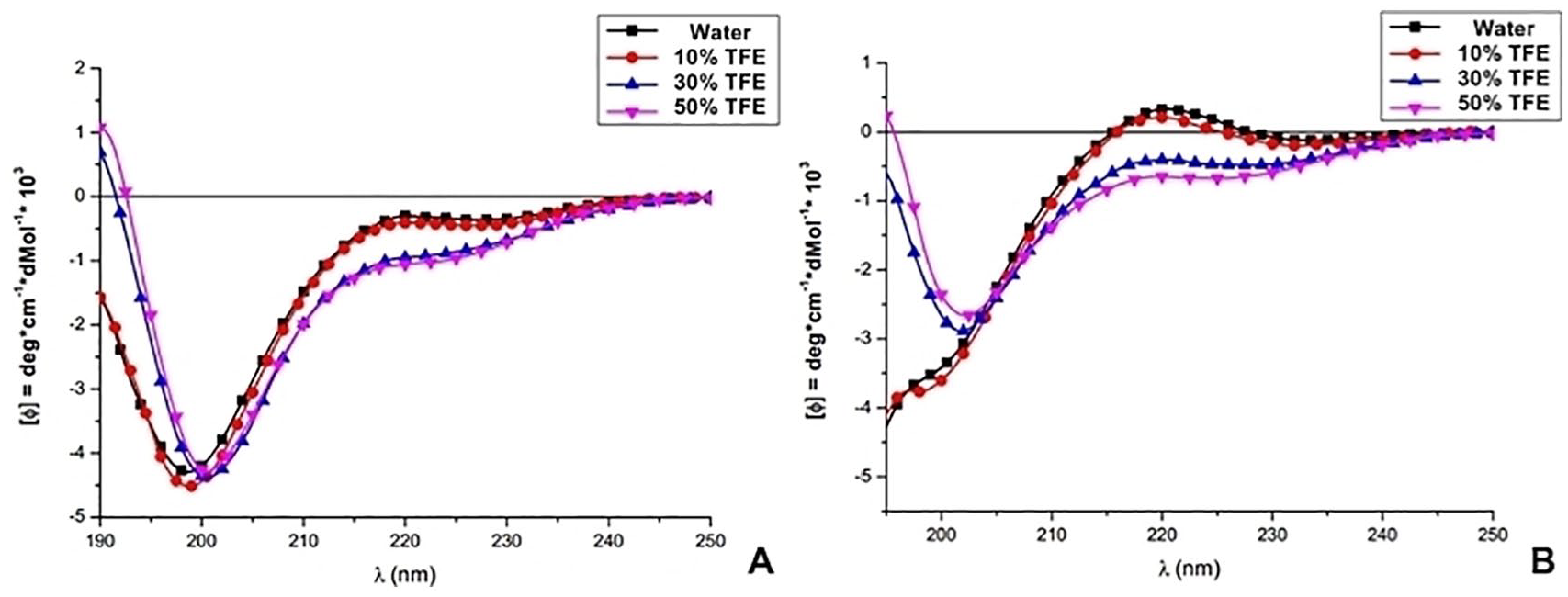
TRP1-TINF and TRP2-TINF CD spectra: (A) Diffraction pattern of polarized light of TRP1-TINF, corresponding to a random structure. (B) Diffraction pattern of polarized light of TRP2-TINF, corresponding to a PPII structure.
TRP1-TINF showed a random secondary structure because of 2 main factors. It is formed of less than 10 amino acid residues and nearly half of these residues have small and neutral side chains (3 glycine residues and 2 proline residues), lowering the possibility of forming a stable secondary structure.
Stable PPII structures presents turns of approximately 120° (3 amino acid residues per turn) and contain 10 atoms in each turn with a hydrogen bridge/bond.83,84 As TRP2-TINF has a PPII conformation and is composed of 10 amino acid residues, it represents a structure with 3 turns.
Tachykinin-related peptides are present in different insect tissues because of hemolymph circulation, 41 thus it is crucial to determine their degradation rate in the plasma to develop improve the understanding of the peptide bioavailability. The percentages of available peptides in the plasma after different incubation times are shown in Figure 8.
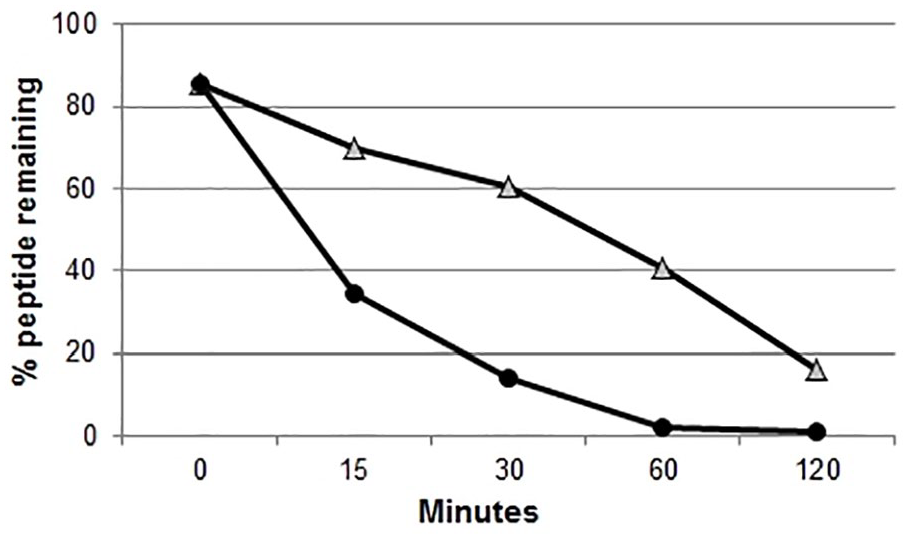
Degradation of TRP1-TINF and TRP2-TINF in the plasma. Percentage of whole peptides per incubation time (0-120 minutes). TRP1-TINF is represented by Δ and 16% of the peptides were not degraded after 120 minutes. TRP2-TINF is represented by (•) and was entirely degraded after 120 minutes.
Within 15 minutes, the percentage of TRP1-TINF decreased to 69% peptide remaining; this process occurs gradually, with only 16% of the whole peptides remaining in solution at 120 minutes.
TRP2-TINF was more susceptible to degradation. In 15 minutes, the peptide concentration decreased to 34% and in 30 minutes decreased to 14%. By 60 minutes, 2.3% of the peptide remained, and no peptide was detected at 120 minutes. The molecular weight of each peptide and the degradation curves were analyzed by mass spectrometry to determine the amino acid loss (Table 4).
Amino acid loss by degradation in plasma.
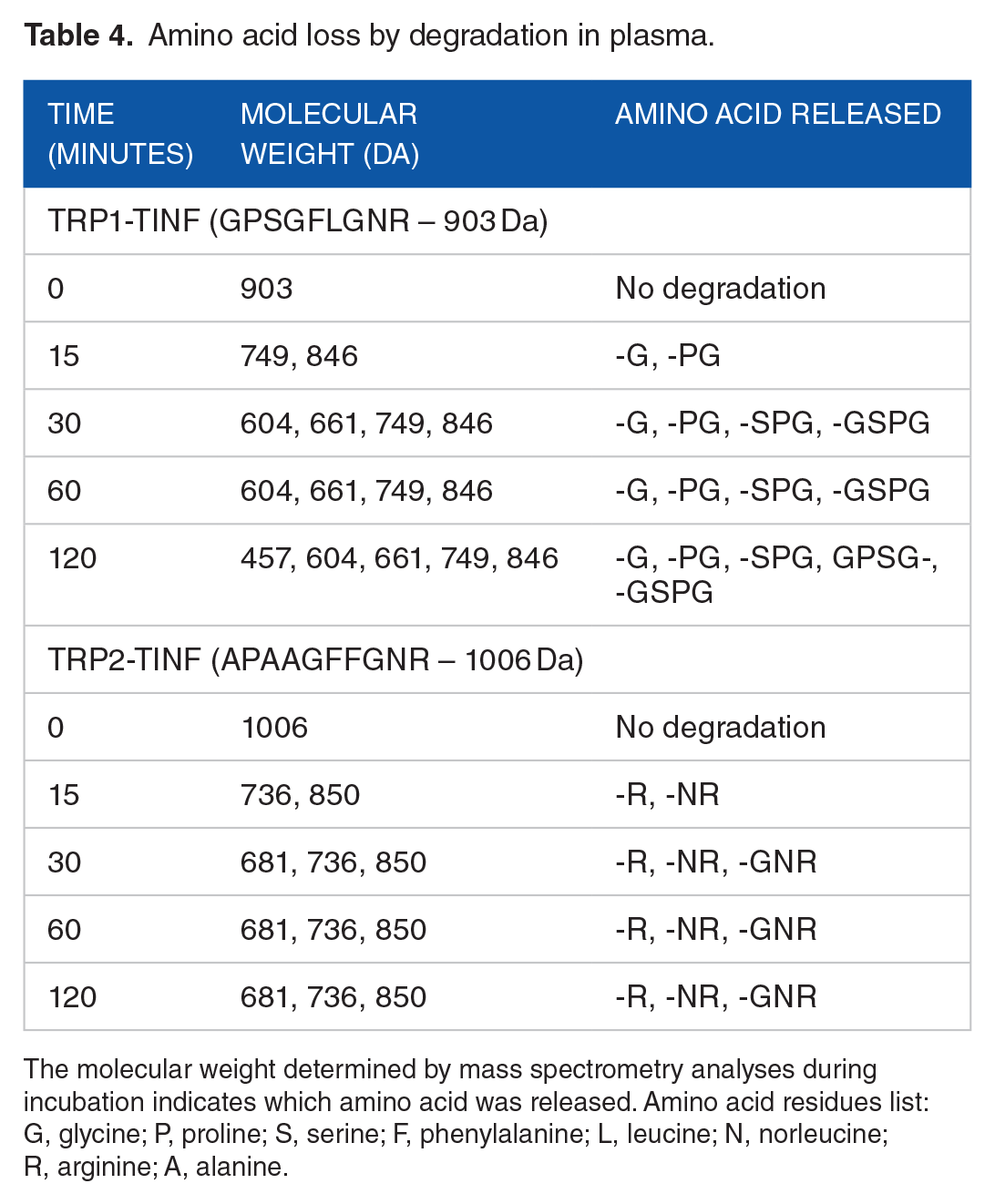
The molecular weight determined by mass spectrometry analyses during incubation indicates which amino acid was released. Amino acid residues list: G, glycine; P, proline; S, serine; F, phenylalanine; L, leucine; N, norleucine; R, arginine; A, alanine.
By observing the amino acid loss at each time point, it is possible to determine if the sequence was degraded by aminopeptidases, carboxypeptidases, or endopeptidases. After 15 minutes of incubation, 3 molecular weights were detected, and 903, 846, and 749 Da were also registered. These molecular weights correspond to the entire sequence and loss of G and G-P, respectively. Additional amino acid losses continually occurred until 120 minutes. At the last time point, 5 molecular weights corresponding to the full sequence without G (846 Da), G-P (749 Da), G-P-S (661 Da), G-P-S-G (604 Da), and G-P-S-G-F (457 Da) were observed. These losses indicate the susceptibility of the peptides to degradation by aminopeptidase enzymes. 85
The degradation profile of TRP2-TINF is compatible with degradation by a carboxypeptidase. Within 15 minutes, fragments with molecular weights of 850 and 736 Da were detected, corresponding to the entire sequence without R and RN. In agreement with this, at 30, 60, and 120 minutes RNG loss was observed (681 Da). This degradation is indicative of carboxypeptidase activity. 86
Conclusions
These results increase the understanding of triatomines physiology and represent a pharmacological relevant study object, providing the first description of TKRP molecules produced by T. infestans and demonstrating the antimicrobial potency and physicochemical features of the isolated molecules.
Footnotes
Acknowledgements
The authors are thankful to the referees for critical review of the manuscript, and to colleagues of the laboratory who helped during the procedures, mainly to Dr Ronaldo Zucatelli Mendonça and Juliana Cuoco Badari for their help with the cell culture experiments and Ivan Ferreira for helping with English editing.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by Fundação de Amparo a Pesquisa do Estado de São Paulo (FAPESP/CeTICS; Grant No. 2013/07467-1) and CNPq Conselho Nacional de Desenvolvimento Cientifico e Tecnologico (Grant Nos 130409/2014-6, 142224/2016-2, and 472744/2012-7).
Declaration of Conflicting Interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
LCLD was responsible for the development of all experiments and writing the manuscript; PIdS Jr participated during the purification experiments, mainly HPLC, and all of the mass spectrometry analysis; AdM participated during peptide synthesis, in plasma degradation, circular dichroism, and mass spectrometry analysis; and FLA participated during circular dichroism experiments. All the authors approve the entire manuscript content and ensure the accuracy and integrity of any part of the work.
