Abstract
Aims:
Antibiotic resistance is currently a major challenge to scientists. Thus, attempts have been made to develop new compounds with antimicrobial activity. In this research, a new antimicrobial peptide with antibacterial activity was isolated from the plant Urtica dioica.
Methods:
A new antimicrobial peptide, named cliotide U1, was purified through precipitation with ammonium sulfate and reverse-phase high-performance liquid chromatography. In silico methods analyzed the physicochemical properties of cliotide U1. The properties of the peptide, including antibacterial activity, pH stability, heat stability, cytotoxicity, and hemolytic activity, were also examined.
Findings:
The purified peptide was composed of 35 amino acids with a hydrophobicity ratio of 63% and a net charge of + 5. The antibacterial activity of cliotide U1 was observed against gram-negative and gram-positive bacteria with a minimum inhibitory concentration (MIC) of 1 to 4 µM. Cliotide U1 had less than 2% cytotoxic activity at the MIC range against the human embryonic kidney cell line 293 with no clear hemolytic activity. The stability of cliotide U1 was preserved at various temperatures (10-60°C) and pH (6-9).
Conclusion:
Our results demonstrated that cliotide U1 had potent antibacterial potential against gram-negative and gram-positive bacteria. Considering its properties, cliotide U1 can be introduced as a novel antibacterial candidate for expanding new therapeutic drugs.
Introduction
Bacterial infections caused by the misuse of antimicrobial drugs have resulted in many deaths in recent years, thereby becoming a critical public health concern in the 21st century. 1 Around 4.5 million deaths were reported due to bacterial diseases. More than 90% of this death rate was caused by infections due to Escherichia coli, Staphylococcus aureus, Acinetobacter baumannii, and Pseudomonas aeruginosa. 2 The rapid increase in antibiotic resistance has led to the need for developing alternative and effective antimicrobial compounds to treat diseases caused by these pathogens.3,4 Researchers attempt to find substitute procedures or new classes of antimicrobial compounds to combat the pathogens. 5 Over the past few decades, scientists have focused on natural sources to function as antimicrobial compounds, such as oligosaccharides, peptides, and plant extracts. 6 Among the various types of natural compounds (plant extracts, essential oils, and antimicrobial peptides), antimicrobial peptides (AMPs), which are produced in various organisms including vertebrates, invertebrates, marines,7,8 and plants, have attracted much attention as promising bioactive compounds. They have broad-spectrum antimicrobial effect against many microorganisms, including bacteria, protozoans, and fungi.9,10
AMPs are recognized as important peptides of innate immune systems found in most living cells. 11 AMPs are amphipathic and positively charged (+ 2 to + 11) and have different amino acid compositions with lengths of about 10 to 50 amino acids.12,13 Most AMPs are very small in size with a molecular weight of 2 to 10 kDa. 14 Based on the experimental studies, AMPs are involved in the leakage of cell contents and cell death by increasing permeability and disrupting microbial membranes.15,16 Both plasma membrane and intracellular components can be the targets for these peptides. 8 AMPs rapidly eliminate multidrug-resistant microorganisms at low concentrations. During the past 2 decades, different types of AMPs have been recognized. 17
According to the Antimicrobial Peptide Database (APD) (https://aps.unmc.edu/), a total of 3146 AMPs are natural and 250 of them have been isolated from plants. Plant AMPs are very similar in function and structure to human and animal AMPs. 18 Purothionin was the first plant AMP reported to be produced in wheat flour (Triticum aestivum). 19 Plants can secrete AMPs from various organs, such as roots, flowers, leaves, stems, and seeds. 8 Plant AMPs have received notable attention due to properties, such as suppression of DNA, protein and RNA synthesis; suppression of protein folding and enzyme activity; inhibition of mitochondrial function and cell division; and stimulation of reactive oxygen species (ROS). 20 Plant peptides have strong antimicrobial activity against plant and mammalian pathogens and are a promising source of AMPs. 21
Urtica L., known as Stinging nettle, is an annual and perennial plant, known for its stinging hairs, which belongs to the Urticaceae family and is one of the main sources of medicinal compounds and polyphenols. 22 Among these plants, Urtica dioica and Urtica urens have been used as medicinal herbs worldwide. This plant is cultivated in Morocco, Turkey, Brazil, Jordan, and Iran. 23 U. dioica plants are used to treat rheumatic pains, stomachache, skin diseases, respiratory diseases, urinary disorders, anemia, bone and joint pain, colds, and cough. 24 The use of nettle tea is common in traditional medicine for treating allergies, diabetes, and prostatic hyperplasia. 25 According to previous research, different crude extracts of U. dioica showed antimicrobial activity against gram-negative and gram-positive bacteria. 24 To our knowledge, however, AMPs derived from U. dioica have not been thoroughly investigated. We attempted to characterize and isolate a novel antimicrobial peptide from U. dioica.
Materials and Method
Test bacteria and plant collection
E. coli ATCC 25922, S. aureus ATCC 25923, A. baumannii ATCC 19606, and P. aeruginosa ATCC 27853 underwent aerobic incubation for 24 hours at 37°C. Fresh and mature leaves of U. dioica were obtained in May, 2022, from Currab Village in Marand County, East Azerbaijan, Iran. The plants were washed using clean water.
Purification and extraction of peptide
The Seyedjavadi et al’s 26 method was used to isolate AMPs from the U. dioica leaves. First, fresh U. dioica leaves (400 g) were frozen and ground into fine flour in liquid nitrogen. The plant powder was mixed with extraction buffer consisting of 100 mM KCl, 10 mM Na2HPO4, 1.5% EDTA (Sigma-Aldrich, USA), and 15 mM NaH2PO4 and at 4°C on a shaker for 4 hours. The crude extract was then clarified by centrifugation at 4500 r/min at 4°C for 20 min. The obtained supernatant underwent filtration through Whatman No1 filter paper (pore size: 11 µm) and then saturated using 85% ammonium sulfate ([NH4]2SO4) at 4°C for 24 hours. The saturated proteins were subjected to centrifugation at 4°C for 20 minutes at 12 000 r/min. Dissolving of the pellets was done in distilled water (DW). To remove residual ammonium sulfate, the precipitate was dialyzed against DW by a benzoylated membrane (MWCO 2000 Da) (Sigma Aldrich-USA) at 4°C for 12 hours. Insoluble residue was removed from the dialysate by centrifuging the dialyzed suspension at 7500 r/min at 4°C for 10 minutes. The final supernatant was then gathered to purify the peptide. To separate peptides with low-molecular weight, the peptide extract was filtered using a 10 kDa cut-off (Millipore, Bedford, MA, USA) ultra-centrifugal filter, concentrated with a 1 kDa ultrafiltration membrane, and dried via freeze-drying. 27 For the purification of AMPs, the lyophilized extract was dissolved in DW, and was applied to a reversed-phase high-performance liquid chromatography (HPLC) column. The separation was conducted using solution A (0.05% TFA in water) and a gradient of 5% to 65% (v/v) solution B (0.05% TFA in acetonitrile) at 1 mL/min for 85 minutes. Considering the absorbance at 220 nm, the fraction collection was done, followed by lyophilizing by freeze-drying to identify the antimicrobial activity. Fractions with the highest activity were selected for further investigation. To make sure of the purity of the peaks with antimicrobial activity, the fractions were re-injected into the similar column in the similar condition with the same solvent systems. 28
Protein quantification
Total protein estimation after each step of extraction and purification was done according to the method of Bradford. 29
Radial diffusion assay
The antibacterial effectiveness of the fractions was evaluated through the radial diffusion assay (RDA) following the Wang method. 30 A bacterial cell concentration of 1 × 106 CFU/mL was introduced into a sterile petri dish, which was subsequently covered with molten Mueller–Hinton agar (MHA) medium. Wells were created and loaded with the purified peptide, while gentamicin served as the positive control. Following 24 hours of incubation at 37°C, the transparent zones surrounding the wells were gauged. Each experiment was conducted 3 times for accuracy.
Minimum bactericidal concentration and inhibition concentration assay
The minimum inhibitory concentration (MIC) of the active and pure peptide was performed against gram-positive and gram-negative bacteria following Li et al’s 8 method with some changes. In brief, 2-fold peptide serial dilutions were prepared in plates with 96 wells to a final volume of 100 µL. Conventional antibiotics, such as gentamicin, were used as control. The concentration assayed for the purified peptide had a range of 0.25 to 32 μM. Afterward, 100 µL of bacterial suspension (1 × 106 CFU/mL) was transferred to the wells, followed by incubation at 37°C for 24 hours. The pure broth with inoculum suspension and the pure broth alone were considered as the respectively positive and negative controls. MIC was determined as the lowest concentration preventing bacterial development in experimental conditions. Next, 20 µL from each well was cultured on the MHA plate and underwent incubation at 37°C for 24 hours. The minimum bactericidal concentration (MBC) was calculated as the peptide lowest level to kill bacterial cells.
Mass spectrometry and amino acid sequence analysis
The potent and active peptide was lyophilized and subjected to amino acid sequencing. The molecular mass of the isolated peptide was determined with electrospray ionization mass spectrometry (MS) at a mass to charge (m/z). Edman degradation was used for the amino acid sequence of the purified antibacterial peptide with an online connection of an ABI Procise Edman Micro Sequencer (Model 492) with an ABI PTH Amino Acid Analyzer (Model 140 C).
Phylogenetic tree and sequence alignment
A search was performed, and 14 peptides with the highest similarity to the isolated peptide were obtained (https://www.aps.unmc.edu/AP/main.php). The identified peptides and the new peptide were aligned regarding sequencing with the use of the Basic Local Alignment Search Tool (BLAST) (https://www.ncbi.nlm.nih.gov/BLAST). Alignment was adjusted manually, and a phylogenetic tree was designed using CLC Main Workbench software. The phylogenetic tree was evaluated through bootstrap analysis with 100 replications for computing the reproducibility of the tree topology.
Physicochemical and bioinformatics analysis
The physicochemical characteristics of the antibacterial peptide were identified using the ExPASy Proteomics server (http://www.expasy.org/tools/protparam.html) for the Boman index, values of the instability index, molecular weight, net charge, aliphatic index, and the hydrophobic ratio. Its AMP probability was evaluated by 4 machine learning algorithms, such as support vector machine (SVM), random forest (RF), artificial neural network (ANN), and discriminant analysis (DA) from the CAMPR3 server (http://www.camp3.bicnirrh.res.in/). The threshold of each algorithm is between 0.5 and 1. The peptide is AMP if this threshold number is > 0.5. The Protein Structure Prediction (PSIPRED) server (http://bioinf.cs.ucl.ac.uk/psipred/) was used to predict the secondary structure. A helical wheel diagram was drawn using the online software to predict the position of amino acids in the peptide (http://lbqp.unb.br/NetWheels/). The 3-dimensional (3D) structure of the novel peptide was evaluated online using the I-TASSER server (http://zhanglab.ccmb.med.umich.edu/ITASSER/). The model’s quality was investigated using Accelrys, DS Visualizer ver. 1.7. To predict the hemolytic property of the peptide, the HemoPI2 server (https://webs.iiitd.edu.in/raghava/hemopi2/prediction.html) was used.
Molecular docking
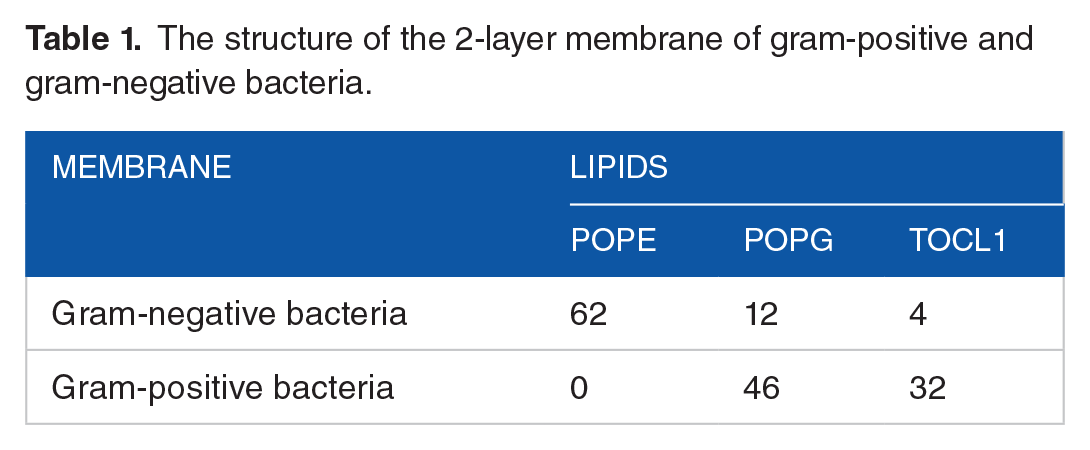
The 3D structure of the peptide was modeled using the I-TASSER online server. SAVES online server and the PROCHECK module were used to assess the quality of this predicted 3D structure. The dual-layer membrane structure for both gram-positive and gram-negative bacteria, as described in the study by Hasannejad-asl et al, 31 was generated via the CHARMM-GUI online server. The specific molecule composition of each dual-layer membrane is detailed in Table 1. For molecular docking, the peptide molecule was prepared using Chimera software version 1.17.1, where Gasteiger charges and nonpolar hydrogen atoms were added, and aromatic carbon rings were defined. Subsequently, the peptide underwent energy minimization using the Amber force field and was saved in pdbqt format after 1000 steps. Docking was performed with Autodock Vina version 1.12.1, designating each dual-layer membrane as a macromolecular receptor and the peptide molecule as the ligand.
The structure of the 2-layer membrane of gram-positive and gram-negative bacteria.
Cytotoxic effects of novel AMP on HEK293 cells
The impact of the antibacterial peptide’s toxicity was examined on the human embryonic kidney cell line 293 (HEK293) cells from the Pasteur Institute in Iran, using the 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) assay. 28 To summarize, Dulbecco’s Modified Eagle’s Medium (DMEM) supplemented with 10% (v/v) fetal bovine serum (FBS) served as the culture medium. The cell lines were cultured in this medium and maintained in an incubator at 37°C with 5% CO2. Seeding of cells was then done in 96-well microtiter plates at a density of 1 × 105 cells per well and exposed to various concentrations of the peptide (0.5-32 μM). Following incubation at 37°C with 5% CO2, 10 µL of MTT solution (0.5 mg/mL in phosphate-buffered saline, PBS Sigma-Aldrich; USA), was introduced to each well and left to incubate for 4 hours under the same conditions. Subsequently, the supernatant from each well was aspirated, and 100 µL of dimethyl sulfoxide was transferred to the wells. The optical density (OD) of each well was evaluated spectrophotometrically at 570 nm using an ELISA reader. For reference, PBS and 1% Triton X-100 were employed respectively as the negative and positive controls. Each test was repeated 3 times for accuracy.
Hemolytic activity
Evaluation of hemolytic activity was performed using Seyedjavadi et al.’s 27 method. At first, we centrifuged the red blood cells (RBCs) of heparinized human whole blood (2400 r/min / 10 min). Washing of the resulting erythrocytes was done 5 times with sterile PBS (pH 7.4), followed by centrifugation at 4500 r/min for 15 minutes. To make a 4% (v/v) solution, the pellets of the RBCs were resuspended in PBS. Next, different serial dilutions of the peptide and RBC solution were mixed and incubated at 37°C for 1 hour. Hemoglobin release was measured via evaluating the supernatant absorption at 570 nm by an ELISA reader. Subsequently, 0.1% Triton X-100 and the RBCs were mixed with PBS and applied, respectively, as the positive and negative controls. The tests were done 3 times. The hemolysis percentage was computed as follows: Hemolysis (%) = ([test OD−negative control OD]/[positive control OD−negative control OD]) × 100.
pH and temperature stability of novel peptide
The effect of pH and temperature was measured on the stability of the peptide, as defined in the work by Khani et al. 11 To assess the impact of pH on the antibacterial efficacy of the new peptide, the peptide specimen was exposed to buffers spanning various pH levels (2-14) at 25°C for 1 hour. Following this incubation, the antibacterial effectiveness was evaluated after adjusting the pH of the solution to 7.2 at room temperature. The peptide was mixed in a 100 µL solution with a pH of 7.2 and served as the standard control. For the examination of thermal stability, the peptide solution underwent incubation at different temperatures (ranging from 10°C to 100°C) for 30 minutes. The peptide that was not subjected to heat served as the reference control. S. aureus ATCC 25923 was used as the indicator to detect the antibacterial activity. The antibacterial activity was measured by RDA as mentioned above. All tests were performed in triplicate.
Effects of serum on antimicrobial activity
The stability of peptides in diluted serum was tested following the procedure outlined in the work by Lyu et al. 32 Human serum was first centrifuged at 4500 r/min for 10 minutes, then heat inactivated at 56°C for 30 minutes. S. aureus ATCC 25923 as the indicator in the exponential growth phase was washed 3 times with PBS at 4500 r/min for 10 minutes and subsequently diluted in PBS or a 25% serum solution to achieve a final concentration of 1 × 106 CFU/mL. The bacterial suspensions were then exposed to 2× MIC of the peptide at 37°C for the following intervals: 0, 0.5, 1, 2, 4, and 6 hours. At each time point, 50 μL samples were collected, serially diluted in PBS, and spread on MHA plates. Colony counts were conducted after incubating the plates at 37°C for 24 hours.
Statistical analysis
Data were derived from a minimum of 3 separate experiments. The outcomes were presented as mean ± standard deviation. Analyses were done using GraphPad Prism 5 Statistical software.
Results
Peptide purification and isolation
Following centrifugation, low molecular weight peptides were extracted from U. dioica leaves, lyophilized, and subjected to C18-RP-HPLC. Notably, 17 fractions were isolated, each collected separately for antibacterial activity assessment (Figure 1A). Among these fractions, fraction 9 exhibited the most potent antibacterial activity and was chosen as the target for additional purification. Subsequently, this potent fraction was reintroduced into the same column for re-chromatography under identical elution conditions. The purity of the specific peak was verified by analyzing the eluted fraction at 220 nm (Figure 1B).

(A) Peptide RP-HPLC purification from U. dioica leaves. The arrow displayed the active peak 9. (B) The chromatogram was taken after the re-chromatography process for purification of active peak 9. (C) ESI-mass spectrum of purified antimicrobial peptide. Multiple charged molecular ions are indicated.
Protein quantification
A summary of the purification of antibacterial peptide from U. dioica leaves is presented in Table 2. After Ammonium sulfate precipitation and Ultrafiltration, 8120 and 963 mg of proteins was obtained from 400 g of leaves. The yield of antimicrobial peptide was 5 mg/400 g leaves. The yield of purified peptide is 0.06% of total leaves protein.
Yields of antimicrobial peptide at different stages of purification from 400 g U. Dioica leaves.

Antibacterial activity
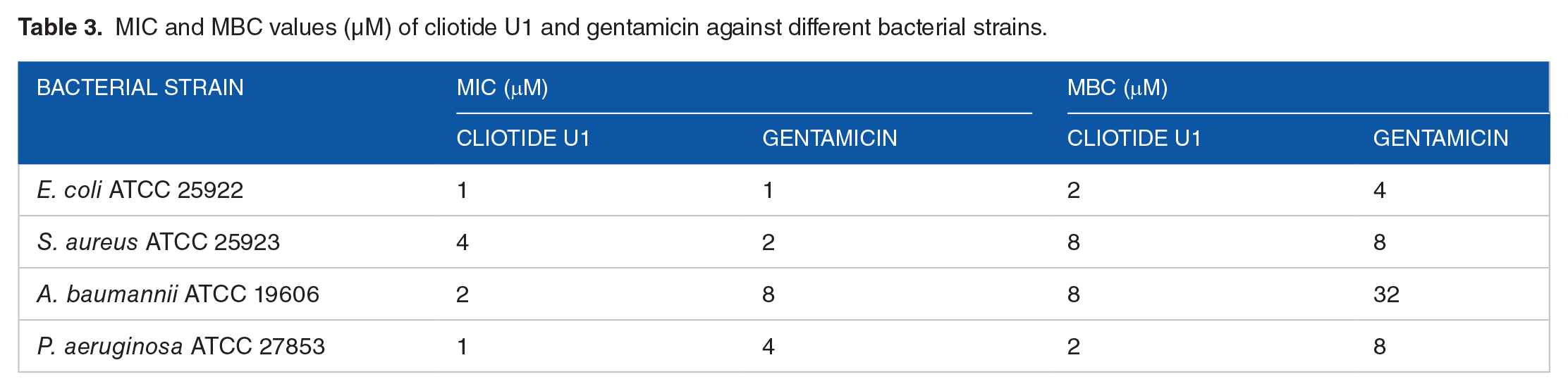
As mentioned earlier, the antibacterial effect of all peaks against gram-negative and gram-positive bacteria was evaluated based on RDA. According to the RDA test results, the peak 9 isolated from U. dioica leaves represented strong antibacterial activity compared with the gentamicin as control. The diameters of the inhibition zones for S. aureus ATCC 25923, E. coli ATCC 25922, A. baumannii ATCC 19606, and P. aeruginosa ATCC 27853 were demonstrated to be 17, 24, 20, and 22 mm for the peptide, respectively. The diameters of the inhibition zones for S. aureus and gram-negative species were 26 and 23 to 27 mm for gentamicin, respectively (Figure 2A and B). The largest inhibition zone diameter was obtained by E. coli and P. aeruginosa, while S. aureus showed the smallest diameter. The MIC values were calculated by the micro-broth dilution approach for further information about the antibacterial activity of the peptide. Gentamicin exhibited an MIC range of 1 and 4 µM. According to Table 3, the extracted peptide has strong activity against the gram-positive and gram-negative bacteria in the MIC of 4 and 1 to 2 µM, respectively. The highest antimicrobial activity was found against E. coli with the significant MIC value (1 µM). Based on the observations, the MBCs of the peptide against bacterial strains are 2 to 4 times superior to its MICs.

(A and B) Antibacterial activity of peak 9 and gentamicin against gram-negative and gram-positive bacteria were measured using the radial diffusion method. (A) Quantitative results of the inhibition zone diameters of peak 9 and gentamicin for tested bacteria with values that are given as the mean ± SD. (B) The inhibition zone of peak 9 and gentamicin against (a) S. aureus ATCC29213, (b) E. coli ATCC 25922, (c) P. aeruginosa ATCC 27853, (d) A. baumannii ATCC 19606.
MIC and MBC values (µM) of cliotide U1 and gentamicin against different bacterial strains.

Identification of peptide
As per the nano-HPLC-ESI-MS/MS spectrometry findings, the peptide’s molecular weight was determined to be 3995.89 Da (Figure 1C). Amino acid sequencing was accomplished using Edman degradation, revealing a fragment consisting of 35 amino acid residues (LIVAASLVYDFYTWIAKKVALLRIAKKVLYLARNA).
Phylogenetic tree and sequence alignment
The peptide sequence did not exhibit complete sequence homology with any previously studied AMPs. Also, the investigation of the BLAST findings approved that the selected peptide is a new and novel antibacterial peptide isolated from U. dioica leaves, which was named cliotide U1. Through the analysis of the phylogenetic tree and sequence alignment, it was determined that the cliotide U1 sequence bore the closest resemblance to cliotide T10, which was isolated from the flowers of Clitoria ternatea (Figure 3A and B).
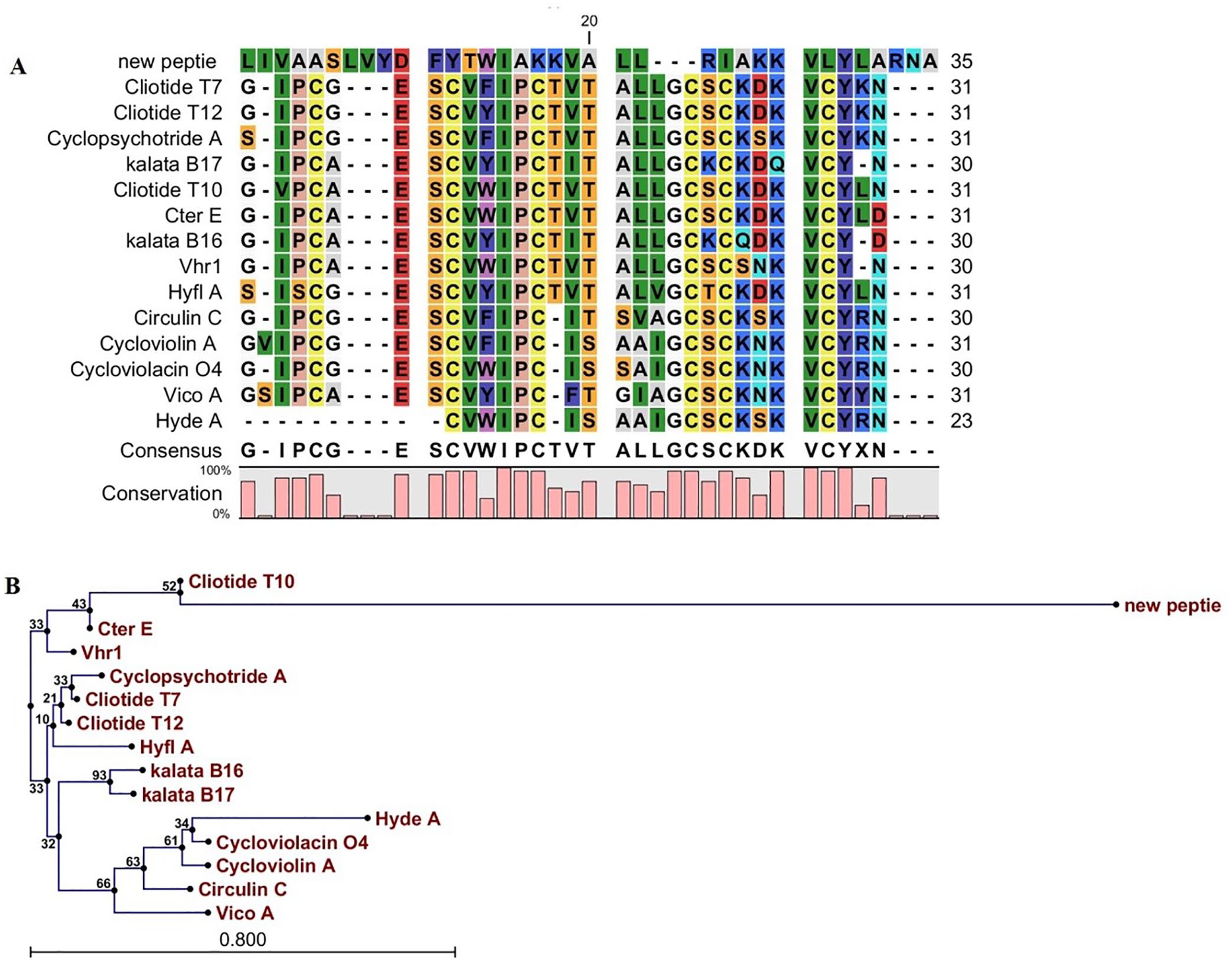
The alignment (A) and phylogenetic tree (B) of cliotide U1. (A) The alignment of cliotide U1 amino acid sequence and the sequences related to other antimicrobial peptides. Color display is settled to emphasize amino acid conservation along all peptides. (B) Phylogenetic tree of cliotide U1. The phylogenetic tree was acquired by CLC main workbench software. Each sequence name is written at the end of the relevant branch. The tree reliability was determined using the bootstrap method with 100 replications.
Bioinformatics and structural analysis
The peptide’s physicochemical characteristics were determined using the ProtParam tool on the ExPASy server. As outlined in Table 4, cliotide U1 has a molecular weight of 3995.89 Da and consists of 35 amino acids. It carries a positive charge of + 5, a hydrophobic ratio of 63%, and a protein-binding potential (Boman index) of 0.13 kcal/mol. The instability index and aliphatic index of cliotide U1 were calculated to be 16.52 and 153.43, respectively. According to the online prediction servers CAMPR3 (Table 4), cliotide U1 is AMP. The secondary structure of the cliotide U1 peptide was predicted using the PSIPRED online server, indicating an α-helix structure (Figure 4A). Analysis through helical wheel projection (Figure 4B) suggested that the peptide possesses both hydrophobic and hydrophilic regions, indicative of amphipathic α-helix conformations. Nonpolar residues, such as alanine, valine, leucine, phenylalanine, isoleucine, tyrosine, and tryptophan cause the hydrophobicity of the peptide. The data obtained by the I-TASSER server showed the α-helix structure in the cliotide U1 peptide (Figure 4C). Evaluation by I-TASSER also showed that the C-score, a confidence score to calculate the global accuracy of the models (ranging from −5.0 to 2.0), was 1.02 for our new peptide. The peptide was analyzed using the HemoPI server, a tool specifically designed to predict hemolytic activity based on amino acid composition and physicochemical properties. According to the HemoPI prediction, the peptide exhibits a hemolytic activity with an estimated HC50 value (concentration for 50% hemolysis) at 19.8 µM.
The probability of antimicrobial activity and physicochemical features of cliotide U1.

ExPASy server was used to estimate physicochemical parameters (charge, pI, hydrophobicity, Boman index, Aliphatic index, and instability index). In silico tools described in the methodology section were used to predict whether the peptide had antimicrobial properties.
Abbreviations: SVM, support vector machine; RF, random forest; ANN, artificial neural network;

(A) Graphical result from secondary structure prediction of cliotide U1 using PSIPRED. (B) Helical wheel diagram of cliotide U1 showing the amphipathic α-helix conformation. (C) The 3D structure of cliotide U1 accepted the α-helix predicted structure of I-TASSER. The 3D structure model imagined by Accelrys discovery studio visualizer software.
Molecular docking
The quality assessment of the predicted 3D structure was conducted using the Ramachandran plot and the PROCHECK module within the SAVES server. The results indicated that 90.9% of the residues fall within the most favored regions, while 9.1% are located within the allowed regions (Figure 5A).

(A) The predicted structure of the AMPs and corresponding Ramachandran plot. Cliotide U1 exhibits an α-helical 3D structure, further validated by the clustering of amino acids in the most favored region of the α-helix on the Ramachandran plot. (B) The interaction between cliotide U1 and both gram-negative and gram-positive membranes, illustrating the peptide’s capability to penetrate the hydrophobic region of the bacterial membrane.
Molecular docking of the peptide with each of the bilayer membranes of gram-positive and gram-negative bacteria revealed that the minimum binding-free energy of the peptide to the gram-negative and gram-positive membranes was −12.4 and −9.2 kJ/mol, respectively, indicating a stronger binding affinity of the peptide to the gram-negative membrane. Furthermore, visualization of the peptide interactions with each membrane demonstrated that the peptide fully penetrated the hydrophobic region of the gram-positive bacterial membrane, while interaction with the gram-negative membrane displayed a carpet-like mode, with only a short segment at each end of the peptide penetrating the membrane structure (Figure 5B). These findings suggest that the peptide induces cell death in gram-positive bacteria by forming pores in the membrane, whereas for gram-negative bacteria, it disrupts membrane protein functions, leading to bacterial cell death.
Cytotoxicity and hemolytic activity
As shown in Figure 6A, the cytotoxic effects of cliotide U1 were assayed on HEK293 cells. At the concentration of 1 to 4 µM, peptide toxicity was less than 6%. Thus, this peptide had no significant toxic effect at its MIC and MBC concentrations. Cliotide U1 stimulated 18.2% of hemolytic activity at the highest level (32 µM). Analysis of the hemolytic activity of cliotide U1 on the RBCs illustrated that this new peptide induced 8.5% hemolytic activity at 4 µM (Figure 6B).

Cytotoxicity, hemolytic activity, and stability of cliotide U1. (A) Cytotoxicity activity of cliotide U1 on HEK293 mammalian cell line using MTT assay. (B) Hemolytic assay of cliotide U1 against human erythrocytes. The effects of pH (C) and temperature (D) on the antibacterial activity of cliotide U1 against S. aureus ATCC 25923 cells.
pH and temperature stability
As the results in Figure 6C show, the effect of pH was measured on the antimicrobial peptide at different pH values from 2 to 12. The highest pH stability of cliotide U1 was observed at pH 6 to 9. The peptide lost its activity in the acidic condition (2-5). Cliotide U1 was maintained with 35% and 15% activity at pH 11 and 12, respectively. The cliotide U1 activity was investigated at various temperatures (10°C-100°C). Based on the finding in Figure 6D, cliotide U1 was constant at various temperatures (10°C-60°C) for 1 hour. However, the antimicrobial activity decreased significantly with increasing the temperature to 70°C. The new peptide has lost its activity at temperatures above 90°C.
Effects of serum on antimicrobial activity
The stability of cliotide U1’s antimicrobial efficacy in serum was assessed. Figure 7 illustrates that, when cliotide U1 is present in PBS, the initial bacterial concentration is approximately 105 CFU/mL. The cliotide U1 peptide without any serum (in PBS, phosphate-buffered saline), there is a rapid and drastic reduction in bacterial count, reaching near zero CFU/mL by 120 minutes. This indicates that cliotide U1 is highly effective at killing S. aureus in the absence of serum. In the presence of 25% serum, there is a significant reduction in bacterial count, though the decline is slower compared with the PBS + cliotide U1 condition. By around 180 minutes, bacteria levels approach zero. This suggests that while the presence of serum slows down the bactericidal effect of cliotide U1, it does not prevent the peptide from eventually killing the bacteria.

Effect of serum on cliotide U1 at 2 × MIC against S. aureus ATCC 25923. Data are mean ± SD of 3 independent experiments.
Discussion
Due to the emergence of multidrug-resistant bacterial strains, many studies have been conducted to discover novel antimicrobial agents. As mentioned earlier, most organisms produce a wide variety of AMPs to protect themselves against pathogens. Previously, U. dioica was shown to have therapeutic properties and is used as a source of traditional medicine, food, and raw ingredients. 33 In our study, the antimicrobial peptide was obtained from U. dioica leaves, and purification, identification and characterization of the newly isolated peptide were carried out. Previous research indicated that the crude U. dioica extract exhibited potent antimicrobial properties against gram-positive and gram-negative bacteria. 22 Another study highlighted the beneficial impact of U. dioica on more than 30 types of gram-negative and gram-positive bacteria. 25 The purification of natural resources for the production of antimicrobial peptides shows considerable potential due to the variety and biocompatibility of these natural AMPs. However, the initial discovery of AMPs was reliant on their isolation from natural sources, which typically required large amounts of raw biological material to extract only small quantities of pure peptides. 34 Our results indicate that the yield of purified peptide accounts for 0.06% of the total protein in the leaves. Today, it is feasible to isolate AMPs on a large scale using recombinant DNA technology or chemical synthesis.
Cliotide U1 showed no similarity with other AMPs reported in the Scientific Database of AMPs successfully proven to be a novel herbal antibacterial peptide against gram-positive and gram-negative bacteria. In silico molecular docking studies revealed that cliotide U1 binds with greater affinity to gram-negative bacterial membranes (−12.4 kJ/mol) compared with gram-positive membranes (−9.2 kJ/mol). As reviewed in the work by Malanovic et al, 35 we hypothesize that additional membrane properties and components may have effects on AMP activity, which could explain the different effects observed with cliotide U1 on gram-positive and gram-negative bacteria. For example, gram-positive bacteria have a thick peptidoglycan-based cell wall containing lipoteichoic acid that may act as an AMP trap, leading to a reduction in the local peptide concentration on the cytoplasmic membrane and ultimately protection against membrane disruption induced by AMPs.36,37 These results explain why cliotide U1, such as cecropins and OMN6,35,38 has a better antimicrobial effect on gram-negative bacteria than on gram-positive bacteria. Supporting this hypothesis, the MIC and MBC data demonstrate that cliotide U1 exhibits stronger antibacterial activity against gram-negative bacteria. For instance, it shows MIC values of 1 μM against E. coli and P. aeruginosa, and 2 μM against A. baumannii. In contrast, the MIC for S. aureus is higher at 4 μM, consistent with the reduced effectiveness against gram-positive bacteria. The highest MBC against E. coli was observed at 2 μM, which likely reflects the influence of the bacterial cell wall. This pattern is in line with similar findings by Tincho et al, 39 who also observed that AMPs exhibited more potent effects against gram-negative bacteria compared with gram-positive ones, further supporting the importance of membrane composition in AMP efficacy. This integrated analysis highlights how the binding affinity of cliotide U1, along with the structural and compositional differences between bacterial membranes, influences its antibacterial activity, especially in gram-negative bacteria.
Furthermore, as shown in Figure 5B, cliotide U1 can disrupt both gram-positive and gram-negative bacterial membranes with some difference in its activity as a function of the nature of the cell wall of each bacteria. Cliotide U1 acts through the carpet-like model, covering the surface in parallel alignments to the gram-negative membrane but, in gram-positive bacteria, this peptide probably destroyed the bacteria by forming pores in the membrane.
Evaluation of predictive and physicochemical properties of in silico cliotide U1 was performed using various online tools. Cliotide U1 has 35 amino acids with a net charge of + 5 and a Boman’s index of 0.13 kcal/mol. The net positive charge of cliotide U1 caused their electrostatic attraction to anionic phospholipids in bacterial membranes. 40 Estimating the efficiency of cliotide U1 to bind to other proteins was provided by Boman’s index. A Boman index value of less than 1 indicates a strong antimicrobial effect of AMPs with few side effects. 41 One of the important parameters to facilitate the interaction of AMPs and lipid bilayers is hydrophobicity. Cliotide U1 consists of 7 hydrophobic residues (63%) that facilitated interactions with bacterial membranes, leading to disruptions in membrane integrity. Research indicates that hydrophobicity is a key parameter influencing the antimicrobial efficacy of AMPs.42,43 This interaction often results in the formation of pores in the bacterial membrane, leading to degradation and ultimately causing the death of the bacteria. Furthermore, the stability and aliphatic index are 16.52 and 153.43, respectively; the aliphatic rate is the relative volume of side chains, and it is also associated with thermostability.41,44
The helical wheel diagram of cliotide U1 illustrates its amphipathic α-helical structure, a characteristic feature of this AMP. Peptides featuring α-helical configurations are recognized as a crucial category of cytolytic AMPs. 45 Research has shown that α-helical antimicrobial peptides (AMPs) with an amphipathic structure can significantly enhance antibacterial effectiveness. 46 These peptides are capable of interacting with bacterial membranes. 47 Typically, they feature a dual composition: a charged portion that aids in membrane interaction and a hydrophobic portion that enables insertion into the lipid bilayer. In general, antimicrobial peptides (AMPs) eliminate bacteria by either disrupting their membranes or by penetrating the bacterial cells to interact with intracellular components.48,49 Models of action for AMPs acting on the cytoplasmic membrane include pore formation (eg, barrel stave or toroidal pores) and carpet mechanism. 50 The findings from the molecular docking analysis highlight the differential interaction of the peptide with the bilayer membranes of gram-positive and gram-negative bacteria. The binding-free energy values of 9.2 kJ/mol for gram-negative membranes and 12.4 kJ/mol for gram-positive membranes suggest a stronger binding affinity of the peptide to the gram-negative membrane. This observation is critical, as it implies that the peptide may exhibit selective cytotoxic effects based on the bacterial cell wall structure.
Gram-positive bacteria possess a thick peptidoglycan layer, which primarily contributes to their structural integrity and resistance against environmental stresses. The penetration of the peptide into the hydrophobic region of the gram-positive membrane suggests that it may form pores, leading to cell lysis. Such mechanisms are consistent with previous studies that have shown peptides targeting the membrane integrity of gram-positive bacteria through pore formation, resulting in leakage of intracellular components and subsequent cell death. 51
In contrast, the interaction of the peptide with gram-negative membranes, characterized by a carpet-like mode, indicates a different mechanism of action. The limited penetration of the peptide into the membrane suggests that instead of forming pores, the peptide may disrupt membrane protein functions. This mode of action is supported by the unique structure of gram-negative bacteria, which includes an outer membrane that protects the cell from various antimicrobial agents. The interaction with the membrane proteins may lead to alterations in permeability or interfere with essential membrane-associated processes, ultimately resulting in cell death. 52
The differential binding and interaction modes of the peptide with gram-positive and gram-negative membranes not only underscore the complexity of bacterial cell wall structures but also open avenues for the development of novel antimicrobial peptides. Understanding the specific interactions and mechanisms can guide the design of more effective peptides with enhanced selectivity and potency against resistant bacterial strains.
To use this peptide in the medical industry, it should not have side effects, such as cytotoxicity and hemolytic effects on mammalian cells. Therefore, it is necessary to evaluate this parameter. 53 As shown in the results, cliotide U1 showed slight hemolytic activity against human erythrocytes and cytotoxicity on HEK293 at effective concentrations in the MIC and MBC range. Although computational tools such as HemoPI provide valuable insights, they rely on trained models that generalize from existing datasets. These models may not fully capture complex biological interactions specific to our peptide. 54 The predicted HC50 value of 19.8 µM is based on sequence-derived features and statistical patterns but does not account for experimental conditions, such as peptide folding, membrane composition, or interaction dynamics, in the test environment. Experimental variability (eg, pH, temperature, or RBC membrane composition) can significantly affect hemolytic outcomes. Since HemoPI provided a useful starting point for evaluating hemolytic potential, the experimental results confirm that the peptide’s hemolytic activity is weak, emphasizing the importance of validating computational predictions through laboratory experiments. This highlights the complementary roles of in silico and in vitro approaches in peptide characterization.
After hemolytic activity and cytotoxicity, pH and temperature stability are other important factors that affect antimicrobial peptide activity and their interaction with cell membranes. 55 According to the results, cliotide U1 was stable in the pH range of 6.0 to 9.0 but lost its activity in acidic pH. Furthermore, cliotide U1 was indeed stable at different temperatures from 10°C to 70°C. According to APD database and aliphatic indices, the peptide chemical and how the structure plays a role in the thermal stability of AMP are described.
In the presence of serum, the bactericidal effect of cliotide U1 was reduced, probably due to hydrolysis of the peptide by proteolytic enzymes present in serum. These observations highlight the challenge of maintaining peptide stability under physiological conditions. Several strategies, including lipidation and other structural modifications, have been proposed to enhance peptide stability and activity in serum. Recent studies have shown that lipidation and N-glycosylation can significantly improve the stability and efficacy of antimicrobial peptides.56,57 Future investigations into these strategies could help optimize the clinical use of cliotide U1.
Limitations of this study include the preliminary nature of our findings and the need for additional experiments. To fully validate the proposed mechanism of action of cliotide U1, confirmatory in vitro assays are essential. Techniques such as fluorescence microscopy or fluorescence-based assays 58 could provide direct visualization of the peptide’s interaction with bacterial membranes and help identify whether pore formation or membrane disruption is occurring. These assays are important next steps in confirming the findings of our in silico studies and further elucidating the peptide’s antimicrobial mechanism. Pending tasks involve in vivo studies, which are essential for understanding the pharmacokinetics and interactions of cliotide U1 in complex biological environments, and exploring potential synergistic effects with existing antimicrobial agents. These tasks are key to advancing development of cliotide U1 as a clinical antimicrobial agent.
Conclusion
In this study, a new antimicrobial peptide isolated from U. dioica leaves, called cliotide U1 with an α-helical structure, was investigated. It had a potent antimicrobial effect on gram-negative and gram-positive bacteria without significant cytotoxicity and hemolytic activity. Overall, the results indicated that cliotide U1 could be an efficient agent to fight pathogenic bacteria.
Footnotes
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The Infectious Diseases and Tropical Medicine Research Center, Shahid Beheshti University of Medical Sciences, Tehran, Iran, provided funding for this study as part of Mahnaz Nasre Taheri PhD thesis (grant no. 43005149).
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
MG and GE conceived and designed the study; MNT, MG, SSS, GE, and AH performed experiments and collected data; MG, MNT, and SSS analyzed and interrelated results; MG supervised, directed, and managed the study; MNT, MG, SSS, GE, and AH approved the final version to be published.
