Abstract
Aims:
The emergence of antibiotic resistance is one of the most significant issues today. Modifying antimicrobial peptides (AMPs) can improve their effects. In this study, the active region of Listeriolysin S (LLS) as a peptidic toxin has been recognized, and its antibacterial properties have been evaluated by modifying that region.
Methods:
After extracting the sequence, the structure of LLS was predicted by PEP-FOLD3. AntiBP and AMPA servers identified its antimicrobial active site. It was modified by adding arginine residue to its 3- and N-terminal regions. Its antimicrobial properties on Staphylococcus aureus, Escherichia coli, and Lactobacillus Plantarum were estimated.
Findings:
The results of AntiBP and AntiBP servers demonstrated that a region of 15 amino acids has the most antimicrobial properties (score = 1.696). After adding arginine to the chosen region, the physicochemical evaluation and antimicrobial properties revealed that the designed peptide is a stable AMP with a positive charge of 4, which is not toxic to human erythrocyte cells and has antigenic properties. The results of in vitro and colony counting indicated that at different hours, it caused a significant reduction in the count of S aureus, E coli, and L Plantarum compared with the control sample.
Conclusions:
Upcoming research implies that identifying and enhancing the active sites of natural peptides can help combat bacteria.
Introduction
Bacteria are a group of microbes that play an essential role in the life of organisms. Pathogenic bacteria are the leading cause of death in infections. Antibiotics are the primary weapon of humankind to fight against these tiny but deadly organisms, and they were successful in most cases.1-4 Interestingly, most antibiotics were obtained from these same microorganisms fighting for survival, and by manipulating or modeling their structure, semi-synthetic or synthetic antibiotics were designed.3,5 With the increase in the use of antibiotics, bacteria, as intelligent organisms, have learned during evolution how to resist the deadly effects of antibiotics, and today, they have mainly been successful in this field, so most of the researchers’ investigation is about the discovery of new antimicrobial substances and the resistance to antibiotics is the big challenge of the present age.1,2,4,5 Antimicrobial peptides (AMPs) are a class of antimicrobial substances that have received attention recently.6-8 Antimicrobial peptides are a group of small peptides that exist widely in nature and are an essential part of the innate immune system of various organisms. Antimicrobial peptides have many inhibitory effects against bacteria, fungi, parasites, and viruses.6-9
The cationic peptides polymyxin B and gramicidin S have a long history of clinical use and are commonly found in topical over-the-counter medications. In addition, the cationic lantibiotic nisin is used as an antimicrobial food additive. In contrast, despite multiple series of clinical trials, only one of the newly developed cationic AMPs has shown efficacy in phase 3a clinical trials. However, due to their remarkable broad-spectrum activity, encompassing activity against Gram-negative and Gram-positive bacteria, fungi, viruses, and parasites within a single peptide, there remains significant interest in exploring the therapeutic potential of these molecules.10,11
In general, AMPs kill bacteria by disrupting their membranes or entering bacterial cells to interact with intracellular components.8,12,13 They commonly target and attach to bacterial membranes through peptide-lipid interactions and are of great interest due to their physicochemical nature, broad spectrum, and different mechanisms from antibiotics.8,12,13
Peptide-lipid interactions play critical roles in biological processes such as membrane fusion, cell signaling, and antimicrobial activity.14,15 These interactions involve various mechanisms, including membrane disruption, lipid binding and insertion, pore formation, lipid clustering, lipid flip-flop, and membrane curvature modulation. Cationic AMPs interact with bacterial membranes to disrupt their integrity and induce cell death.14,15 Some peptides can insert into lipid bilayers, leading to changes in membrane structure and function. Overall, the diverse modes of action of peptide-lipid interactions contribute to the regulation of cellular processes and the defense against pathogens.14,15
As mentioned, antibiotics were first obtained from the microorganisms themselves. A peptidic toxin or bacteriocin that has recently received attention is Listeriolysin S (LLS), which is a thiazole/oxazole-modified microcin (TOMM) produced by hypervirulent clones of Listeria monocytogenes and probably when the bacterium enters the human body.16-19 It destroys the intestinal microbiome, facilitating colonization and finally internalizing L monocytogenes bacteria.16-19 The production of this toxin by serotype 4b may be one of the main reasons for the pathogenicity and spread of this serotype of L monocytogenes. In this study, different software has identified the region of LLS with the most antimicrobial properties, and its antimicrobial properties have been strengthened by adding arginine moiety, and its antimicrobial effects on different bacteria have been evaluated.
Materials and Methods
In silico study
Extraction sequence and structure modeling of Listeriolysin S
The LLS amino acid sequence was extracted from the UniProt database, and PEP-FOLD3 software 20 modeled its 3D structure. The secondary structure was predicted by the PSIPRED server (http://bioinf.cs.ucl.ac.uk/psipred/) as well.
Identification of the most active antimicrobial region of Listeriolysin S
The sequence of 49 amino acids of LLS was introduced to AntiBP Server. 21 The algorithm of this software is based on methods Quantitative Matrices (QM), Artificial Neural Network (ANN), and Support Vector Machine (SVM) and the use of binary patterns of peptide sequences. Furthermore, the overall accuracy of this server is ~92.11%. Also, the AMPA server, 22 a theoretical approach to identifying active regions in antimicrobial proteins, was used.
Modifying the antimicrobial properties of the target area in Listeriolysin S and investigating its antimicrobial and physicochemical properties
After identifying the antimicrobial active region of LLS, its antimicrobial properties were strengthened by adding 4 arginine amino acids in the C- and N-terminal regions. Owing to the presence of the guanidine group, arginine amino acid increases the binding affinity of peptides to negatively charged groups of cell membranes, such as carboxyl, phosphate, and sulfate. Then, its antimicrobial properties were checked with APD3, 23 ToxinPred, 24 ClassAMP, 25 and DBAASP servers. 26 The PEP-FOLD3 server modeled the final 3D structure of the peptide in question.
The ultimate peptide construct was synthesized using solid-phase synthesis methodology (GenScript, Piscataway, NJ, USA). Also, the physicochemical characteristics of the designed peptide were analyzed using the ProtParam program provided by Expasy. 27
Visual presentations
Pymol software 28 and LigPlot+ software 29 were employed to visualize the 3D structure of the studied peptides.
In vitro study
The studied strains and culture media
In this study, the standard strains of Staphylococcus aureus ATCC 33591, Escherichia coli ATCC 10536, and Lactobacillus plantarum subsp. Plantarum ATCC 13643 was used. Also, Mueller Hinton broth and brain heart infusion (BHI) agar are used to grow S aureus and E coli. The MRS (De Man-Rogosa-Sharpe) broth was used for L plantarum growth.
Investigating the effect of the designed peptide on bacterial survival
The antimicrobial activity of the desired peptide was tested against the studied bacteria strains by the microbroth dilution method using 96 U-shaped wells microdilution plates. 30 Accordingly, after adding 100 μL of Muller Hinton Broth (MHB) broth (Oxoid, UK) to each well of a 96-well plate, 100 μL of different concentrations of the studied peptide (32, 64, and 68 μg/mL) was added to a well. Finally, 100 μL of the overnight culture containing ∼105 cells was added to each well. Plates were incubated at 37°C, and colony count technique was done at 0, 4, and 28 hour post-inoculation. Each assay was performed in triplicates and repeated at least twice.31-33 Vancomycin (1 μg/mL) and ciprofloxacin (0.008 μg/mL) antibiotics were employed as selective agents for the ATCC strains of S aureus and E coli, respectively, serving as controls in the study.
Statistical analysis
Meaningful results were estimated employing a 1-way variance analysis (ANOVA) obeyed by the Tukey post hoc test and the Dunnett test for multiple comparisons (as demonstrated in figure legends). All data examination and statistical graphing were performed using GraphPad Prism 8.
Results
Listeriolysin S structure
As studying a protein’s second and third structure can help us understand its function better, the second and third structure of LLS was analyzed by PSIPRED and PEP-FOLD3 server, respectively.
As shown in Figure 1, the second structure of LLS mainly consists of coils structure (47%) and also 2 regions with strand (20%) and helix (16%) structures. Figure 2 also shows the cartoon representation of the third structure resulting from server PEP-FOLD3. The red area is considered the most active part of LLS in this study.

Predicted secondary structure of LLS by PSIPRED.

Predicted 3D structure of LLS by PEP-FOLD3. The red area is where the most antimicrobial activity has been detected.
Identifying the most active part of Listeriolysin S
The most active parts of LLS are shown in Table 1.
Active antimicrobial areas of Listeriolysin S.

These results were obtained by AntiBP and AMPA servers. Finally, the area with the highest score (1.696) was selected for further study.
Modifying the antimicrobial properties of the Listeriolysin S
After selecting the region with the most antimicrobial properties by adding 4 arginines to its N- and C-terminal regions, a 19 amino acid peptide was obtained (RRGCCSCSCSTCTCTCTRR), shown in Figure 3.

The cartoon structure of the designed peptide in this study.
Different servers’ results showed that the peptide has a positive charge of 4 and high antimicrobial properties. It is antigenic, non-toxic, and has no hemolytic activity on human erythrocyte cells. More details are given in Table 2.
The physicochemical properties of the studied peptide.

The results of the antimicrobial effects of the modified peptide on the survival of the studied bacteria
The results of this part showed that the peptide under study has high antimicrobial effects, so as shown in Figure 4, 4 hours after exposure to a concentration of 32 μg/mL of the peptide under study, the number of living cells of S aureus bacterium compared with the control sample has decreased by about 4 logs. This decrease has reached about 9 logs after 24 hours (P < .0001).
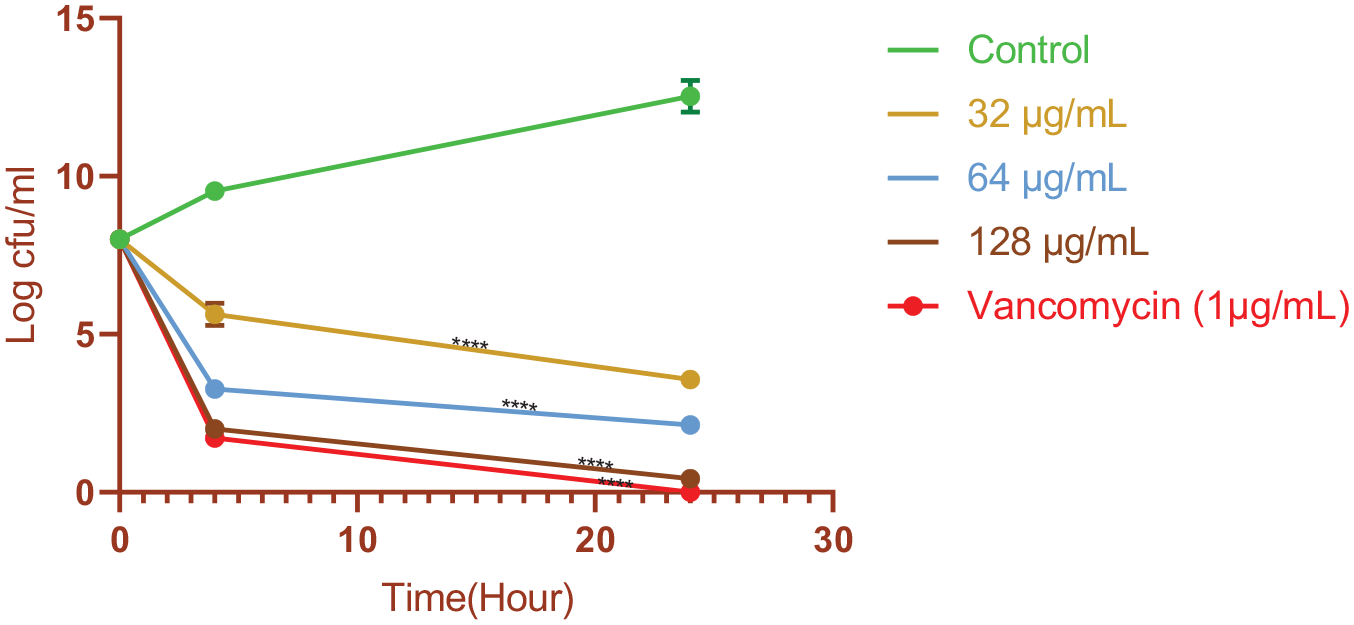
Effect of the designed peptide on S aureus survival. Untreated bacteria were used as controls. The error bars indicate the SD from 3 replicate samples. *P ⩽ .05; **P ⩽ .01; ***P ⩽ .001; ****P ⩽ .0001 using 2-way ANOVA and the Dunnett multiple comparisons test.
At the concentration of 128 μg/mL, after 24 hours, the number of bacteria decreased by about 12 logs. This part showed that the studied peptide is effective on S aureus, and its antimicrobial effects increase with increasing concentration and have a direct relationship.
It is notable that the antimicrobial efficacy of the peptide at a concentration of 128 μg/mL closely approached the effects of vancomycin, a selective antibiotic. Nevertheless, vancomycin at a concentration of 1 μg/mL was able to eradicate all bacteria within 24 hours, even in this susceptible strain under investigation (Figure 4).
Also, as shown in Figure 5, 4 hours after exposure to a concentration of 32 μg/mL of the desired peptide, the number of living cells of E coli bacterium compared with the control sample has decreased by about 4 logs. This decrease has reached about 7 logs after 24 hours (P < .0001).

Effect of the designed peptide ion E coli survival. Untreated bacteria were used as controls. The error bars indicate the SD from 3 replicate samples. *P ⩽ .05; **P ⩽ .01; ***P ⩽ .001; ****P ⩽ .0001 using 2-way ANOVA and the Dunnett multiple comparisons test.
At a concentration of 128 µg/mL, after 24 hours, the number of bacteria decreased by about 12 logs. Also, as depicted in Figure 5, the antimicrobial activity of the antibiotic ciprofloxacin at a concentration of 0.008 μg/mL surpassed that of the investigated peptide. It is important to note that the studied strain exhibits complete susceptibility to the specific antibiotic under investigation, which has been designated as the preferred antibiotic. It is also acknowledged that in cases of antibiotic resistance, the observed results may vary.
This section showed that the studied peptide has a bactericidal effect against E coli and that its antibacterial effect increases with increasing concentration, and there is a direct relationship. In general, the results of this part showed that the studied peptide has similar killing properties for these 2 bacteria.
The antimicrobial results of the studied peptide on bacteria L plantarum also showed that the number of live bacteria decreased significantly in the sample treated with the peptide compared with the control sample.
It is shown in Figure 6; for example, after 24 hours of treatment with the studied compound, the number of live cells of L plantarum bacteria decreased by about 11 logs compared with the control sample (P < .0001). Interestingly, researchers believe that LLS inactivates the antilisterial effects of probiotics.

Effect of the designed peptide on L plantarum survival. Untreated bacteria were used as controls. The error bars indicate the SD from 3 replicate samples. *P ⩽ .05; **P ⩽ .01; ***P ⩽ .001; ****P ⩽ .0001 using 2-way ANOVA and the Dunnett multiple comparisons test.
Discussion
Antimicrobial resistance (AMR) occurs when microorganisms such as bacteria, viruses, fungi, and parasites develop resistance to antimicrobial drugs that were previously effective in treating infections. 34 This means that the drug can no longer kill or prevent the growth of microorganisms, leading to persistent infections, increased morbidity and mortality, and health care costs. 34 Antimicrobial resistance is a global public health threat, affecting both humans and animals.34,35
It is caused by a combination of factors, including overuse and misuse of antibiotics in humans and animals, poor infection prevention and control practices, and a lack of emerging new antibacterial drugs. 35 The emergence of microbial resistance against antibiotics is one of the biggest challenges of the current century, so a significant part of researchers’ efforts is in this direction. 35
One of the antimicrobial classes beyond antibiotics is AMPs with various effects and suitable physicochemical properties for therapeutic use. 36 In this study, the active region of the LLS peptide toxin was identified by different bioinformatics servers, and its antimicrobial properties were strengthened by adding arginine bases on both sides of the studied peptide. Transport and binding of peptides to cells is often enhanced by increasing the content of arginine or other guanidinium groups. The guanidine group shows a great affinity for binding to the negatively charged groups of the membrane, such as phosphate, carboxyl, and sulfate. This study showed that the modified peptide has high and similar antimicrobial effects on Gram-positive S aureus and Gram-negative E coli bacteria. This study also indicated that cationic groups increase the antimicrobial properties of peptides. There needs to be more information about LLS.
Notably, it has been reported that in response to oral Listeria infection in mice, the gut microbiota are upregulated due to the bactericidal properties of LLS. Accordingly, LLS may target prokaryotic cells, leading to the colonization of L monocytogenes into the intestine.
In a study by Mohammadzadeha et al in 2019, they demonstrated that L plantarum bacteria in cell culture inhibit the growth and invasion of Listeria strains lacking the gene encoding LLS, whereas strains with LLS are not only killed by L plantarum but also they caused the death of some of these bacteria. They concluded that LLS probably neutralizes the antilisterial properties of probiotics such as L plantarum and can cause listeriosis in the human body by destroying the host’s flora. 17
Changing the structure of antimicrobial substances or conjugating them with different carriers, such as liposomes and nanoparticles, can be an excellent solution to increase efficiency and reduce the possibility of AMR. We observed that the modified peptide has elevated and comparable antimicrobial effects on Gram-positive S aureus and Gram-negative E coli bacteria.
A study by Chen et al in 2020 showed that modifying the N-terminal cholesterol of PMAP-37 (F34-R) could improve its antibacterial activity. They acknowledged that Chol-37 (F34-R) showed excellent antibacterial activity and remarkable therapeutic effects in vivo. Therefore, Chol-37 (F34-R) may be a candidate for antimicrobial agents against microbial infection in the clinic. 37
In another study by Wenya Zhou et al, adding lipoic acid (LA) to a peptide with the sequence RIWVIWRR-NH2 (Bac8c), a natural AMP, significantly increased its potency. The LA-Bac8c showed a strong depolarization of the membrane and the ability to penetrate the outer membrane. The cell membrane treated with LA-Bac8c was destroyed due to the leakage of the bacterial cell components, and its antibiofilm property also showed a significant increase. In that study, the LA-Bac8c exhibited lower Minimum Inhibitory Concentration (MIC) Minimum Bactericidal Concentration (MBC) values against S aureus and methicillin-resistant S aureus (MRSA) than Bac8c. 38
Conclusions
The forthcoming study showed that identifying and strengthening the active areas of natural peptides can be suitable for fighting bacteria.
Footnotes
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was financially supported by a research grant from Deputy of research, School of Medicine, Shahid Beheshti University of Medical Sciences, Iran [Grant No. 43008821]. The funding agency has no role in the design of the project, work execution, analysis, interpretation of the data, and manuscript writing or submission.
Author Contributions
MG, and AS conceived and designed the study; MG, AS, MSN, KB, MSGL and MM performed experiments and collected data; MG, AS and MSN analyzed and Interrelated Results; MG Supervised, directed and managed the study; MG, AS, MSN, KB, MSGL and MM Final approved of the version to be published.
Consent for Publication
Not applicable.
Ethics Approval and Consent to Participate
Not applicable.
Availability of Data and Materials
Not applicable.
