Abstract
Lipid droplets (LDs) are unique intracellular organelles that are mainly constituted by neutral lipids (triglycerides, sterol esters). As such they serve as the main site of energy storage in the cell and they are akin to oil emulsions in water. To prevent the direct exposure of the hydrophobic neutral lipids to the aqueous environment of the cytosol, LDs are surrounded by a monolayer of phospholipids that thus behave as a natural surfactant. This interfacial structure is rather unique inside the cell, but a molecular understanding of how the LD structure modulates its functions is still lacking, mainly due to technical challenges in both experimental and computational approaches to investigate oil-in-water emulsions. Recently, we have investigated the structure of LDs using a combination of existing and newly developed computational approaches that are optimized to study oil-water interfaces.1 Our simulations provide a comprehensive molecular characterization of the unique surface properties of LDs, suggesting structure-function relationship in several LD-related metabolic processes.
Keywords
Originally considered as inert lipid globules, lipid droplets (LDs) have recently triggered a revival of interest, mostly due to their crucial involvement in metabolic diseases, and for their role in the infectious cycle of pathogens, which target fat metabolic cells and/or lipid metabolic pathways for their propagation. 2 From a cell biology perspective, in the past few years, LDs have emerged as a crucial player in a multitude of intracellular trafficking pathways that are intimately connected with lipid metabolism, and it has been shown that they can establish functional membrane contact sites with multiple organelles. 3 Thus, it is not surprising that several proteins that were previously associated with other organelles, mainly the endoplasmic reticulum (ER) and the Golgi apparatus, also bind to LDs.2,4,5 These observations raise the question of how these proteins are able to bind membranes with such different molecular structures (Figure 1). This is especially puzzling considering that the lipid composition of the surface of LDs has been reported to be strikingly similar to that of the ER, 6 and that the 2 compartments might even remain in contact throughout the life cycle of LDs. 7
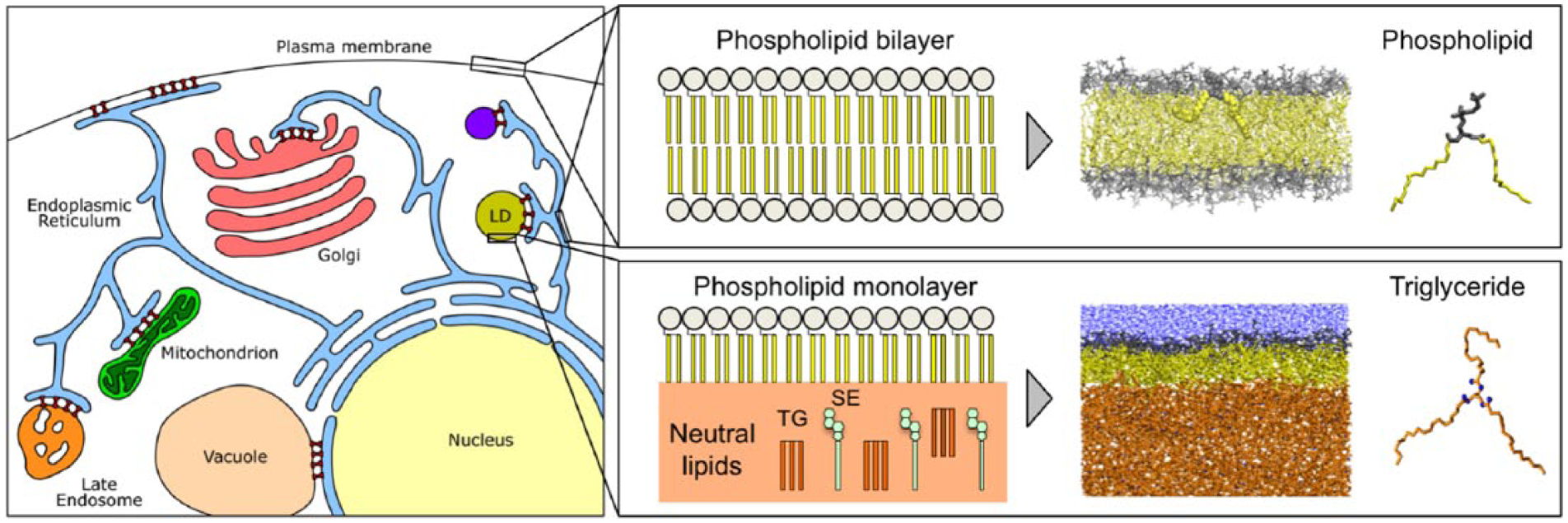
Left: Schematic representation of intracellular organelles and contact sites between them. Right: Schematic comparison between a lipid bilayer (top) and the lipid monolayer of a lipid droplet (bottom). Corresponding representative snapshots from molecular dynamics simulations are also shown.
In the past few years, however, several physicochemical properties of membranes (including curvature, electrostatics, surface tension [ST], etc), have been shown to regulate cellular processes via their interaction with proteins. Recently, this has also been shown to be the case for LDs, where several phenomena, including LD budding from the ER or protein targeting, appear to be driven by physicochemical properties.4,8 Wet-lab methodologies, however, are often unable to characterize membrane-mediated molecular processes with subnanometer resolution, and computational modeling, namely, molecular dynamics (MD) simulations, has emerged as a powerful complementary methodology to investigate the behavior of membranes and membrane-interacting systems directly at the molecular level.
Molecular dynamics simulations of ternary systems mimicking LDs, however, have been scarce so far.1,9,10 One possible reason is the numerous technical difficulties that are associated with the investigation of these systems. In comparison with lipid bilayers, for example, more computational resources are generally needed, as the systems are larger (because of the presence of the oil layer) and as full equilibration between oil and phospholipids needs to be achieved. As a consequence, most of the MD studies on LDs had been performed using coarse-grain (CG) approaches,9,10 namely, with the MARTINI force field, 11 where several atoms forming a specific chemical moiety are grouped into individual beads to decrease the number of degrees of freedom and hence speed up the calculations. In our recent work, 1 we also identified a subtler challenge: due to the natural friction and interdigitation between the surface layer of phospholipids and the underlying oil core, we observed important size effects (ie, the results may vary significantly depending on the chosen size of the simulated system, thus indicating that MD simulations on a small system may lead to incorrect results) on both dynamical and structural properties of the LD interface.
The major challenge of investigating LDs, however, is to accurately describe interfacial tensions12,13. To do so, we have developed new parameters for triglycerides that are compatible with the Shinoda-DeVane-Klein CG force field 14 , since this force field was explicitly developed to correctly reproduce surface and interfacial tensions of surfactants and phospholipids. 14 Using this approach, we have been able to investigate LDs and to infer possible structure-function relationship between the unique structure of the LDs and various crucial biological processes taking place on their surface, and especially for what concerns peripheral protein targeting, triglyceride breakdown, and LD biogenesis.
Selective Targeting of LDs by Peripheral Proteins
Our work was originally motivated by the puzzling observation that several proteins that are traditionally localized to the ER and/or the Golgi apparatus can also bind to LDs. All these proteins (Arf1, ArfGAP1, GBF1, etc) possess an amphipathic helix (AH). We had recently proposed a model suggesting that membrane-packing defects, as measured in MD simulations, can be used as a proxy to predict the binding of AH-containing peripheral proteins in vitro and in vivo. 15 By simulating model LDs, we have been able to measure lipid-packing defects on the LD interface. We found that most molecular properties of the surface of model LDs, including lipid-packing defects, resemble those of model ER and Golgi membranes, 16 thus providing a structural explanation of why AH-containing proteins can bind to LDs and other organelles alike.
It must be noted that all these simulations were performed at zero ST and zero curvature, as is commonly the case in MD simulations of membranes. In the case of LDs, curvature is generally considered negligible because LDs have typical sizes in the micrometer range and are hence flat on a molecular scale. However, although ST is generally very low in lipid bilayers (with values typically well below 1 mN/m), this is obviously not necessarily the case for oil emulsions.
Interestingly, the picture starts to change when factoring in the role of ST on surface properties of model LDs. As expected, lipid-packing defects increase with increasing STs. However, this increase is slow when ST is relatively low (<10 mN/m), and it becomes significant when ST is high (>10 mN/m). This behavior suggests an intriguing and novel scenario: when ST is low, only proteins that possess helices “well adapted” to LDs and to membrane binding can target selectively LDs; when ST is high, several proteins, including those that should not bind to LDs physiologically, may also be recruited to LDs. This last scenario is likely to correspond to nonphysiological conditions that are promoted to observe protein binding to LDs using fluorescence microscopy approaches, such as oleate loading and LD induction.
A direct consequence of the previous observation is that we could make the prediction that the ST of intracellular LDs should not go beyond ≈10 mN/m in an unregulated manner; otherwise, the consequences on intracellular trafficking would be potentially devastating. Such high ST levels could lead to trapping on LDs of AH-containing proteins destined for other organelles, possibly leading to aberrant trafficking and spontaneous LD fusion. In collaboration with the Thiam Lab, we have recently reported that the ST of purified LDs is in the 2- to 4-mN/m range, 8 thus confirming our prediction solely based on computational modeling.
Lipid Remodeling and Triglyceride Breakdown
We also observed that interdigitation between triglycerides and phospholipids remained constant for ST < 10 mN/m, whereas it increased dramatically for ST > 10 mN/m. This was quite surprising and has potentially important implications in triglyceride metabolism. Lipid remodeling enzymes, and specifically those that breakdown triglycerides (such as adipose triglyceride lipase [ATGL]), are found at the surface of LDs. Even though their exact mechanism of action remains unknown, increasing substrate availability, as in the case of increased interdigitation, should probably enhance their activity and hence triglyceride breakdown. Thus, our simulations suggest only marginal increased activity of ATGL as a function of ST for values <10 mN/m and a substantial increase for values >10 mN/m. Of note, breakdown of triglycerides would not only decrease the number of oil molecules in the core but also increase the number of surfactant molecules (breakdown of one triglyceride results in the formation of one free fatty acid and one diacylglycerol). As those molecules would reduce ST, our observations suggest the possible presence of a feedback mechanism, whereby on reaching ST values ≈10 mN/m, lipase activity would substantially increase or, conversely, ST increase above 10 mN/m would be strongly prevented in the presence of lipases.
LD Biogenesis
The process of LD biogenesis is generally believed to start from the accumulation of triglyceride molecules between the 2 leaflets of the ER. 17 In such a scenario, a transition from a lipid bilayer devoid of triglycerides to a fully formed LD would involve structural changes similar to those we observed when we move from a lipid bilayer to a model LD. As our MD simulations suggest that an increase in LD size is coupled to a decrease in lipid-packing defects, 1 this may lead to a decrease in the free energy of binding for AH-containing peripheral proteins and hence to their possible unbinding from the membrane surface on LD biogenesis. Thus, our data suggest an interesting molecular mechanism, whereby on LD formation, proteins that were initially recruited to ER membrane are then expelled (possibly to make room for LD-specific proteins) precisely at the site of LD formation and simply as a consequence of changes in the underlying membrane properties.
In summary, although our results represent only a preliminary step toward a structural understanding of LD biology, MD simulations appear primed to provide a significant contribution toward the understanding of complex membrane-mediated processes, including those involving unconventional and fascinating interfaces such as those of LDs.
Footnotes
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Swiss National Science Foundation (#163966).
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
SV wrote this Commentary.
