Abstract
Background:
Emergence of human immunodeficiency virus (HIV) drug resistance mutations prior to highly active antiretroviral therapy is a serious problem in clinical management of HIV/AIDS. Risk factors for appearance of drug resistance mutations are not known. We hypothesize that
Methods:
A total of 115 patients were recruited in this study of which 75 were HIV+TB+ coinfected (group 1) and 40 were HIV+TB− (group 2). Blood samples from all the patients were collected and CD4+ cell counts; HIV-1 plasma viral load and sequencing of protease and two-third region of reverse transcriptase of HIV-1 was performed and analyzed for drug resistance pattern.
Results:
For patients with HIV+TB+, 10.6% (8/75) had mutations to non-nucleoside reverse transcriptase inhibitors (NNRTIs), 4% (3/75) to nucleoside reverse transcriptase inhibitors, and only 2.6% (2/75) patients had mutations to protease inhibitors. Interestingly, for group 2 (HIV+TB−), there were only NNRTI mutations found among these patients, and only 3 patients (7.5%) had these drug-resistant mutations. Clade typing and phylogenetic tree analysis showed HIV-1 subtype C predominance in these patients.
Conclusions:
Our study showed that higher percentage of HIV drug resistance mutations was found among HIV+TB+ individuals compared with tuberculosis-uninfected patients. Tuberculosis coinfection may be a risk factor for emergence of high frequency of drug resistance mutations. Studies with a larger sample size will help to confirm these findings from the Indian population.
Introduction
Approximately 40 million people are living with human immunodeficiency virus (HIV) infection worldwide, and thousands of individuals are newly infected each year. 1 In the absence of an effective vaccine, the introduction of antiretroviral therapy (ART) in the early 1990s reduced both mortality and morbidity in these patients. In recent years, however, there are increasing concerns due to the emergence of pretreatment drug resistance mutations (DRMs). 2
As per the latest report by the Joint United Nations Programme on HIV/AIDS (UNAIDS), the prevalence of HIV in India is approximately 0.31%, which in absolute numbers translates to approximately 4.1 million people living with HIV/AIDS.2,3 All newly diagnosed patients in India are currently offered ART consisting of 2 nucleoside reverse transcriptase inhibitors (NRTIs) and 1 non-nucleoside reverse transcriptase inhibitor (NNRTI). Regimens with protease inhibitors (PIs) are reserved as second-line treatment options on failure of the first-line ART. 4 With decreasing costs and the universal use of ART irrespective of CD4 count, emerging data revealed an increased prevalence of drug-resistant (DR) HIV strains ranging from 10% to 20% among ART-naïve patients.5–7
As every virus fights to survive, the emergence of this drug resistance is inevitable. The DRM results from the selective pressure during viral replication, especially in the presence of subtherapeutic levels of ART.8,9A study done in Mumbai in western India showed the prevalence of DR strains among ART-naïve patients to be 9.6%. 10 Similar studies from other parts of India have reported a lower prevalence (<5%) of DRM.11–14
There are limited data on the risk factors for DRM among treatment-naïve individuals. Tuberculosis (TB) is a very common opportunistic infection in patients with HIV. No systematic studies have been performed that identify the pattern of DR mutations among patients coinfected with HIV and TB.
In this study, we explored the prevalence of HIV-1 DR mutations among HIV-TB–coinfected individuals.
Materials and Methods
Patient population
All treatment-naïve patients with HIV visiting the ART clinic of the hospital from July 2012 to January 2016 were screened for eligibility. HIV-1 infection was confirmed using 3 sets of enzyme-linked immunosorbent assay according to NACO (National Aids Control Organization) guidelines.15,16 A detailed treatment history was taken from all patients and spouses (in case they were on highly active antiretroviral therapy [HAART]). Those who reported no prior exposure to antiretroviral drugs were considered ART naïve. Patients with clinical, radiological, or microbiological evidence of TB and not currently on anti-TB therapy were recruited as HIV-TB coinfected. Exclusion criteria included age <18 years; serological evidence of acute hepatitis A, B, C, or E; HBsAg positive; anti-hepatitis C virus antibody positive; and pregnancy. The institute ethics committee approved the study and written informed consent was taken from all patients prior to participation.
Specimen collection
A 10 mL sample of whole blood was taken from each patient. About 3 mL was used for CD4+ T-cell count estimation, and the remaining was centrifuged within 6 hours of collection at 400
Viral load testing and CD4+ T-cell estimation
Viral load testing was performed using the standard protocol of Amplicor HIV-1 Monitor Test, version 1.5 (Roche Molecular Systems Inc., Branchburg, NJ, USA). CD4/CD8+ T-cell counts were determined by flow cytometry using BD FACSCalibur (BD Biosciences, San Diego, CA, USA).
HIV-1 genotyping and DRM
HIV-1 genotyping and mutation analysis was performed using the ViroSeq HIV-1 Genotyping Systems (Abbott Diagnostics, Wiesbaden, Germany). A 1.3 kb protease-RT region of HIV-1 pol gene was sequenced as per standard procedure. 18 RNA extraction was performed on 500 μL of plasma using the guanidine-thiocyanate extraction method. A reverse transcription-polymerase chain reaction (PCR) followed by PCR was conducted to generate an amplicon of 1.3 kb. The amplicons were purified using silica spin columns, and PCR products were run on a 1% agarose gel. The PCR product was then sequenced with a set of 6 primers to sequence 1.3 kb covering the protease gene and two-thirds of the reverse transcriptase gene. Drug resistance mutations were defined according to the WHO Surveillance mutation list 2009 proposed by Bennett et al. 19
Clade typing and phylogenetic tree
HIV-1 subtype was defined using the REGA HIV-1 subtyping tool. 20 Worldwide subtype references were obtained from the Los Alamos HIV database. 21 For the phylogenetic study, nucleotide sequences were aligned using the software programs GeneDoc 8 and Clustal X version 1.83 multiple sequence alignment.
Statistical methods
Mean and standard deviation were computed for data following parametric distribution. Median with range was computed for data following nonparametric distribution. The unpaired
Results
Baseline characteristics
Out of 115 patients recruited in the study, 75 individuals were in the HIV-TB group (group 1) and 40 in the HIV+TB− group (group 2). In group 1, >50% of patients had extrapulmonary TB. None of the patients had any partners on ART. Most of the patients were men in both groups (84% and 67.5%, respectively) with their average age approximately 35 years. Basic demographic and clinical parameters are summarized in Table 1. Group 1 patients had a significantly lower CD4 count and body mass index and a higher viral load (Table 1).
Baseline demographic and clinical characteristics.
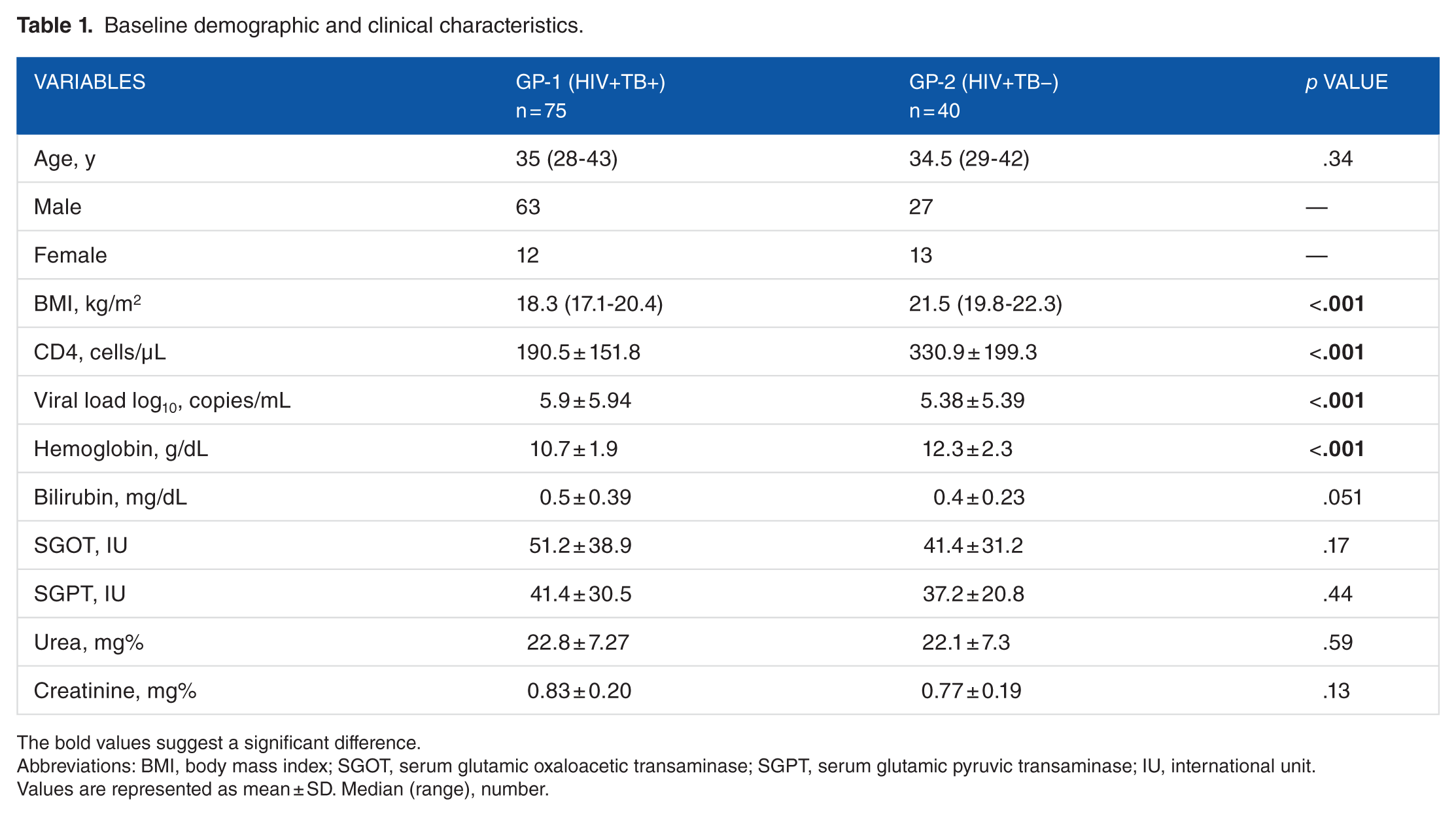
The bold values suggest a significant difference.
Abbreviations: BMI, body mass index; SGOT, serum glutamic oxaloacetic transaminase; SGPT, serum glutamic pyruvic transaminase; IU, international unit.
Values are represented as mean ± SD. Median (range), number.
DRM pattern
In group 1 (n = 75), 10 subjects had 14 DRMs; 8, 3, and 2 subjects had mutations to NNRTI, NRTI, and PI, respectively. In group 2 (n = 40), 3 subjects had 4 DRMs; 2 had NNRTI mutations; and 1 had PI mutation; there were no NRTI DRM. There was no statistically significant difference in the prevalence of DRM in the 2 groups (Table 2).
Drug resistance mutation patterns in patients with HIV+TB+ and HIV+TB−.
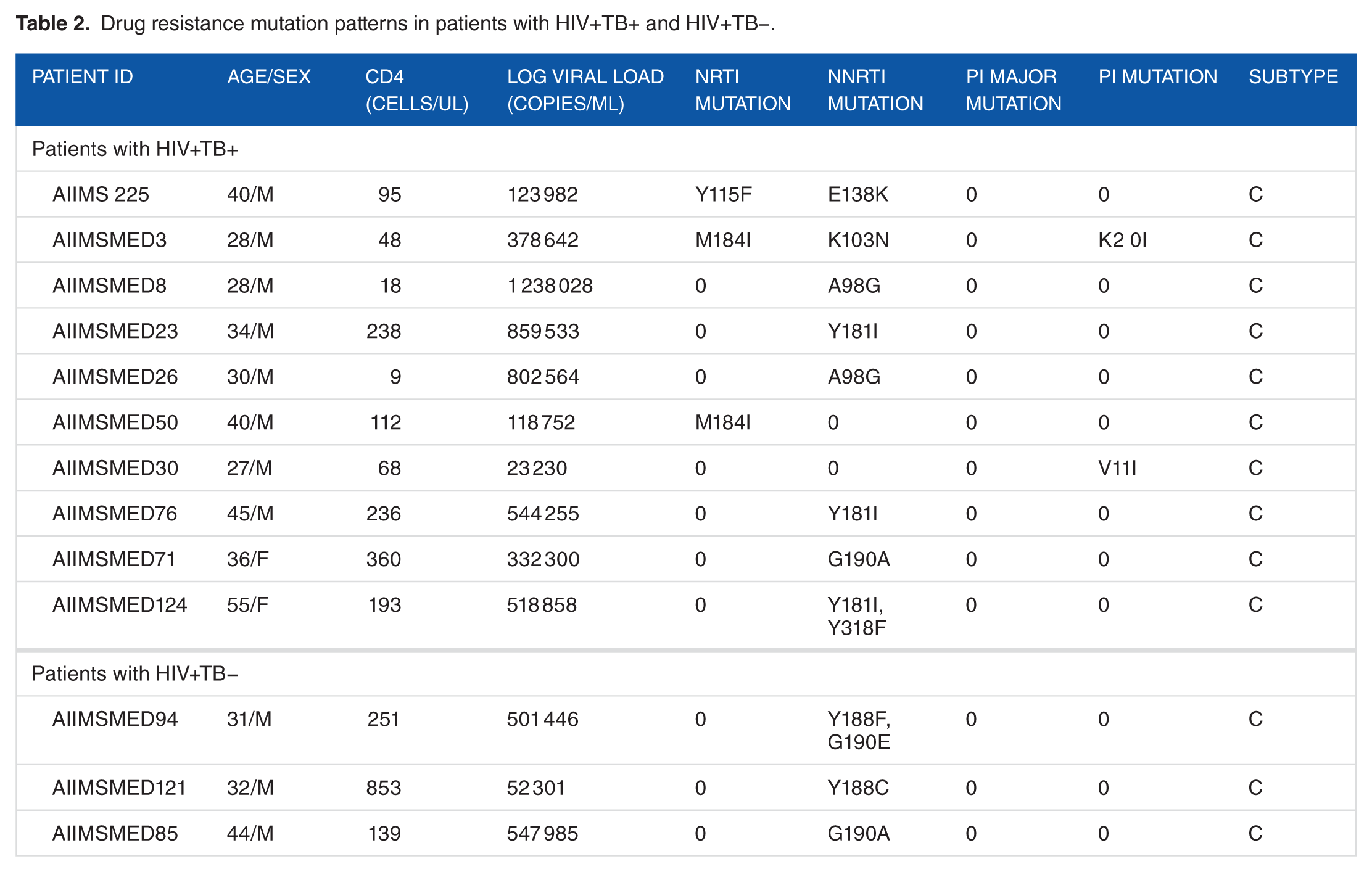
Phylogenetic analysis
Subtype C was predominant in both groups (Figure 1A and B). Variations within the subtype C viruses isolated were also observed with divergence up to 10%. Three sequences from group 1 and 3 sequences from group 2 were found to be of the A subtype. No other subtype was detected from this region.
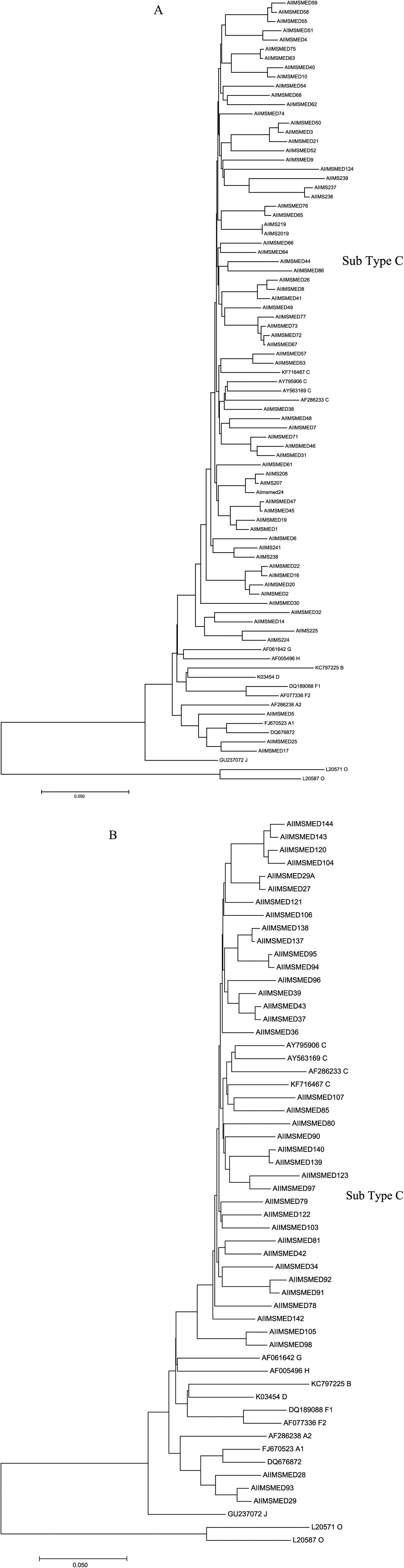
Phylogenetic trees of (A) HIV+TB+ ART-naïve and (B) HIV+TB− ART-naïve samples. 1302-bp DNA sequences of PI and RT regions from HIV-1 were analyzed and aligned with reference sequences from different subtypes. GeneBank accession number and corresponding subtypes of reference sequences are given. KF716467, AY795906, AY563169, and AF286233 are subtype C; AF061642 is subtype G; AF005496 is subtype H; KC797225 is subtype B; K03454 is subtype D; DQ189088 and AF077336 are subtype F. FJ670523 and DQ676872 are subtype A; GU237072 is subtype J and L20571 and L20587 are subtype O. ART indicates antiretroviral therapy; PI, protease inhibitor; RT, reverse transcription.
Discussion
The WHO guidelines classify DRM prevalence in a geographical area as low (<5%), moderate (5%–15%), and high (>15%). 21 This classification signifies the level of drug resistance surveillance programs required for monitoring primary HIV-DR.
Data from this study fit in the WHO moderate zone (11.3%; 13/115), suggesting the need for more surveillance data on primary DR in the treatment of naïve patients.
In this study, we observed an overall mutation of 13.3% DRM among the patients with HIV-TB and 7.5% among the patients with HIV alone. A previously published study reported a prevalence of 3% in treatment-naïve HIV-infected patients of whom none had TB coinfection. 12 The results from the present and previous studies suggest that HIV-TB–coinfected patients have higher DRM. However, because the CD4 count was lower and the viral load was higher in the HIV-TB group, it can also be argued that those with DR mutations had higher immunosuppression and therefore had more active TB infection.
Interestingly, we found that among the TB-coinfected patients, the prevalence of NRTI mutations was higher. Most of the NRTI mutations result in a less-fit virus, but it is possible that because of altered immunity due to TB infection, these mutant viruses are able to replicate more efficiently. Because only 2 patients in group 1 had exclusive NRTI or PI mutation, we could not compare the viral load of these patients with the rest of the group.
In our study, M184V, an NRTI mutation that confers resistance to lamivudine, was isolated in only 2 cases. It is seen as a transmitted drug resistance mutation in ART-naïve patients and is most commonly found in patients failing ART. This may be because the mutation decreases the replicative capacity of the virus, which thus reverts back to the wild type. 22 A similar low prevalence was reported by 2 earlier studies from India (1.6% and 2.5%).23,24
We found 2 participants in group 1 with PI mutations (K20I and V11I) in the protease gene. This mutation is not part of the surveillance list and was not known to confer therapeutic drug resistance until now. Because PI-based regimens are used less frequently in the country in which the study 2 was performed, the prevalence of major PI mutation may be assumed to be <5%, as also corroborated by this study and previously published literature. 25
Most of the patients within both groups of our study were infected with subtype C of HIV-1. Subtype C is the common subtype circulating in all regions of India, followed by subtype B, which is common in northeastern states, and subtype A, which is seen in the western and northern regions of India.26–30
The emergence of HIV DRM is a global concern, and HAART is the only option to prevent AIDS progression among the infected individuals. Risk factors for the emergence of DRMs are not yet very clear.
There is evidence of a reduction in CD4 cell count in patients with HIV associated with decreased interferon gamma production leading to an increased risk of reactivation or infection with TB. In addition, pro-inflammatory cytokine production by TB granulomas (especially interferon alpha) has been associated with increased immune activation that increases HIV viremia. 31 It has been known for years that HIV-1 virus infection may act as an amplifier and make patients more susceptible to TB. However, this study helps understand the bidirectional mechanism between HIV and TB, suggesting that an opportunistic infection like TB, in the case of HIV-1–infected patients, might exacerbate the condition of patients with HIV and subsequently lead to HIV DRM.
This study shows that there is a higher prevalence of DRM in TB-coinfected patients. The statistical strength of this conclusion is limited because of the small sample size. Whether TB itself leads to a higher prevalence of mutations needs to be further explored. We did not follow-up patients for virological responses and therefore cannot say with certainty that these detected mutations would lead to treatment failure. As we only studied treatment-naïve patients, it would be interesting to see whether concomitant anti-TB therapy leads to the emergence of DRM in HIV-TB–coinfected patients. As HIV and TB run hand in hand in many endemic countries, this topic is of huge public health interest.
Continued surveillance and systematic prevalence studies in India and around the world are required to assess primary drug resistance patterns and possible risk factors.
Conclusions
There is a higher percentage of HIV DRMs among HIV-TB–coinfected patients compared with those with HIV-1 only. Tuberculosis coinfection may be a risk factor for the emergence of DRMs. Similar studies, with a larger sample size, are required to confirm these findings.
Footnotes
Acknowledgements
The support of ART, DOTS staff; NACO and ICMR is much appreciated. The authors especially thank Anita and Pooja, support staff for their contributions.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship and/or publication of this article: This work is supported by National AIDS Control Program, Ministry of Health and Family Welfare, Government of India. The authors thank National AIDS control organization for funding the project.
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
SS provided inputs to the study design, helped in data analysis and interpretation, wrote the manuscript, and did the final editing. KG, MK and DM helped to write the manuscript. BKD, DM, and NHK conducted laboratory tests for CD4, plasma viral load, and HIV drug resistance. RMP and KG performed data analyses. All authors read and approved the final manuscript.
Ethical Approval and Consent to Participate
Prior approval of the Institute Ethics Committee and informed written consent were taken from all participants before the start of the study.
