Abstract
Background:
Zeste White 10 (ZW10) is a key component of the spindle assembly checkpoint (SAC) that maintains chromosomal stability during mitosis. Dysregulation of ZW10 can cause chromosomal instability and aneuploidy—hallmarks of many cancers, including breast cancer, particularly triple-negative breast cancer (TNBC). However, its prognostic and therapeutic relevance in breast cancer remains unclear.
Objectives:
This study aimed to systematically investigate the expression pattern, prognostic significance, mutational profile, and immune associations of ZW10 in breast cancer using integrated omics data.
Design:
A computational, cross-cohort bioinformatics analysis combining transcriptomic, proteomic, mutational, and clinical data from publicly available databases.
Methods:
The ZW10 expression levels were assessed across normal and cancerous tissues using The Cancer Genome Atlas (TCGA), Genotype-Tissue Expression (GTEx), and the Human Protein Atlas data sets. Immune infiltration correlations were analyzed using TIMER2.0, Gene Set Cancer Analysis (GSCA), and TNMplot. Survival analyses were performed using Kaplan-Meier Plotter and TCGA clinical data sets. Protein-protein interaction (PPI) and functional enrichment analyses were conducted using STRING and EnrichR, and mutation data were retrieved from COSMIC and cBioPortal.
Results:
The ZW10 expression was markedly upregulated across multiple cancers, with the highest expression in TNBC. Elevated ZW10 levels correlated with immune cell infiltration and adverse overall survival, while lower ZW10 expression predicted improved relapse-free survival. Protein-protein interaction and enrichment analyses revealed ZW10’s close interaction with key mitotic regulators and its involvement in spindle checkpoint and vesicular trafficking pathways. Mutation analysis identified predominant A > G and A > T substitutions and frequent gene amplifications across malignancies.
Conclusion:
The ZW10 acts as a potential prognostic biomarker and therapeutic target in breast cancer, particularly TNBC, through its dual roles in mitotic regulation and immune modulation. Further experimental validation is warranted to confirm its mechanistic role and therapeutic potential.
Plain Language Summary
Background: Zeste White 10 (ZW10) is a key component of the spindle assembly checkpoint (SAC) that maintains chromosomal stability during mitosis. Dysregulation of ZW10 can cause chromosomal instability and aneuploidy—hallmarks of many cancers, including breast cancer, particularly triple-negative breast cancer (TNBC). However, its prognostic and therapeutic relevance in breast cancer remains unclear. Objectives: This study aimed to systematically investigate the expression pattern, prognostic significance, mutational profile, and immune associations of ZW10 in breast cancer using integrated omics data. Design: A computational, cross-cohort bioinformatics analysis combining transcriptomic, proteomic, mutational, and clinical data from publicly available databases. Methods: The ZW10 expression levels were assessed across normal and cancerous tissues using TCGA, GTEx, and the Human Protein Atlas datasets. Immune infiltration correlations were analyzed using TIMER2.0, GSCA, and TNMplot. Survival analyses were performed using Kaplan-Meier Plotter and TCGA clinical datasets. Protein-protein interaction (PPI) and functional enrichment analyses were conducted using STRING and EnrichR, and mutation data were retrieved from COSMIC and cBioPortal. Results: The ZW10 expression was markedly upregulated across multiple cancers, with the highest expression in TNBC. Elevated ZW10 levels correlated with immune cell infiltration and adverse overall survival, while lower ZW10 expression predicted improved relapse-free survival. The PPI and enrichment analyses revealed ZW10’s close interaction with key mitotic regulators and its involvement in spindle checkpoint and vesicular trafficking pathways. Mutation analysis identified predominant A>G and A>T substitutions and frequent gene amplifications across malignancies. Conclusion: The ZW10 acts as a potential prognostic biomarker and therapeutic target in breast cancer, mainly TNBC, through its dual roles in mitotic regulation and immune modulation.
Introduction
Breast cancer is one of the most prevalent and leading causes of cancer-related deaths in women worldwide. 1 In 2022, female breast cancer ranked as the second most common cancer worldwide, with an estimated 2.3 million new diagnoses, accounting for approximately 11.6% of all cancer cases globally. 2 In 2020, it became the most frequently diagnosed cancer, with an estimated 2 261 419 new cases (11.7% of all cancers) and 684 966 deaths (6.9% of all cancers) globally. 3 It is a heterogeneous disease with various subtypes that differ in their molecular and clinical characteristics, including luminal A, luminal B, HER2-positive, and triple-negative breast cancer (TNBC).4 -6 The classification of breast cancer based on its molecular characteristics has significantly influenced the development of targeted therapies. However, despite advances in treatment, breast cancer remains a significant clinical challenge due to the development of drug resistance and metastatic spread, particularly in aggressive subtypes like TNBC. 7 The pathogenesis of breast cancer is multifactorial, involving complex interactions between genetic predispositions (eg, breast invasive carcinoma [BRCA]1/2 mutations), environmental factors (such as hormonal exposure and lifestyle influences), and dysregulated cellular signaling pathways. 8 These factors collectively contribute to the heterogeneity and aggressive behavior of breast cancer subtypes.9,10
Aberrant mitosis and defective chromosome segregation are fundamental hallmarks of cancer and are particularly prominent in aggressive malignancies such as breast cancer, including TNBC. 11 Disruption of mitotic control mechanisms frequently leads to chromosomal instability (CIN) and aneuploidy, which promote tumor heterogeneity, disease progression, and therapeutic resistance. 12 The spindle assembly checkpoint (SAC) is a critical surveillance mechanism that preserves genomic integrity by ensuring accurate chromosome alignment prior to anaphase onset.13,14 Dysregulation of proteins involved in mitosis can result in genomic instability, a hallmark of cancer. 15 ZW10 (Zeste White 10) is a mitotic checkpoint protein that plays a pivotal role in maintaining chromosomal stability during cell division.16,17 It is a critical member of the complex machinery responsible for regulating the spindle assembly checkpoint (SAC), ensuring that the chromosomes are properly aligned and segregated during mitosis.18,19 The correct functioning of the SAC is vital for preventing chromosomal missegregation, which can lead to aneuploidy—a condition where cells have an abnormal number of chromosomes, often seen in cancer cells.18,20,21 Several multifunctional adaptor proteins, including dynactin, lissencephaly 1, nuclear distribution protein E (NUDE), NUDE-like, Bicaudal D, the Rod-ZW10-Zwilch (RZZ) complex, and Spindly, play pivotal roles in modulating dynein function and its localization. 22 The core structure of the corona comprises ROD, Zwilch, ZW10, and the dynein-dynactin (DD) adaptor Spindly, collectively forming the RZZ-Spindly (RZZS) complex. The kinesin motor protein CENP-E coordinates the loading of the DD complex onto kinetochores, facilitating bidirectional transport of chromosomal cargo during mitosis. 23
The ZW10 plays a pivotal role in mitotic regulation as a core component of the SAC machinery, where it ensures accurate surveillance of chromosome-spindle attachments. Through this function, ZW10 safeguards faithful chromosome segregation, thereby minimizing the risk of aneuploidy and preserving genomic stability during cell division. 13 As such, ZW10 overexpression or mutations may disrupt the SAC, leading to CIN, which is commonly observed in many cancer types, including breast cancer. The association between mitotic checkpoint genes like ZW10 and cancer progression has led to increasing interest in understanding their role as both prognostic markers and therapeutic targets in breast cancer. 24 The ZW10 is frequently overexpressed across multiple cancer types, implicating this protein as a potential contributor to tumorigenesis and malignant progression.25,26 High expression levels of ZW10 have been correlated with poor prognosis in certain cancer types, including breast cancer, particularly in TNBC, a subtype known for its aggressive clinical course and lack of effective targeted therapies. 7 Chromosomal instability resulting from aberrant ZW10 function may contribute to tumor progression and chemoresistance, making it an attractive target for therapeutic interventions. 27 In addition to its oncogenic potential, ZW10’s involvement in cell-cycle regulation and its interactions with other mitotic proteins also present opportunities for identifying new drug targets aimed at restoring proper mitotic function in cancer cells. Furthermore, ZW10’s prognostic value in breast cancer has been a subject of growing interest, with several studies suggesting its potential utility as a biomarker for early detection and monitoring therapeutic responses. The clinical relevance of ZW10 expression, especially in relation to patient survival and treatment outcomes, warrants further investigation to determine its role in guiding personalized treatment strategies. 28 The previously published work on ZW10-interacting kinetochore protein (Zwint-1) in breast cancer demonstrated that Zwint-1 is aberrantly expressed and contributes to breast cancer cell proliferation and poor prognosis through its involvement in kinetochore function and cell-cycle regulation. 28 While that study established a foundational link between a ZW10-associated kinetochore component and breast cancer progression, our current study differs in both focus and scope. Specifically, we directly investigate ZW10 itself rather than its interacting partner, and we extend the analysis to elucidate its independent biological significance, molecular mechanisms, and clinical relevance in the context of cancer. Moreover, our findings provide new insights into how dysregulation of ZW10 contributes to mitotic checkpoint control, genomic instability, and tumor progression, aspects that were not comprehensively addressed in the prior report. By comparing these results, our work builds upon existing knowledge while offering novel mechanistic and translational perspectives that distinguish it from earlier studies and underscore the unique contribution of ZW10 to cancer biology.
Recent progress in bioinformatics has transformed cancer research by enabling the large-scale integration and analysis of multi-omics data sets to identify potential prognostic and therapeutic biomarkers. Open-access resources such as The Cancer Genome Atlas (TCGA) and the Gene Expression Omnibus (GEO) now offer extensive genomic, transcriptomic, and epigenomic data, allowing systematic investigation of cancer-related genes across diverse patient populations. Using computational frameworks that combine differential gene expression profiling, survival analysis, DNA methylation assessment, and immune landscape characterization, researchers can uncover molecular signatures associated with tumor development, clinical outcomes, and therapeutic response.29 -31 These bioinformatics-driven approaches deepen our understanding of cancer mechanisms and accelerate the identification of novel diagnostic and therapeutic targets, thereby advancing precision oncology and personalized medicine.32 -35 Within this framework, comprehensive bioinformatic exploration of ZW10, a kinetochore-associated protein essential for chromosome segregation and mitotic checkpoint control, may yield valuable insights into its molecular role and clinical significance in breast cancer.
This study aims to explore the expression of ZW10 in breast cancer and its potential role in cancer progression and prognosis. By examining the relationship between ZW10 and breast cancer subtypes, we seek to uncover molecular insights into the pathogenesis of breast cancer and highlight the therapeutic potential of targeting ZW10 in the fight against this prevalent disease.
Materials and Methods
Distribution of Zeste White 10 across all major tissues and organs in the human body
The Human Protein Atlas (HPA) aims to create a comprehensive map of all human proteins across various cells, tissues, and organs by integrating multiple advanced technologies. These include antibody-based imaging, mass spectrometry-driven proteomics, transcriptomics, and systems biology approaches. The resource provides open access to its extensive data, enabling both academic researchers and industry professionals to freely explore and analyze the human proteome. The HPA is composed of several specialized resources, each focusing on different aspects of the genome-wide analysis of human proteins, offering a multidimensional view of protein expression and localization across the human body (https://www.proteinatlas.org/about#tissue). Complementary expression data from the Genotype-Tissue Expression (GTEx) project (https://gtexportal.org) were utilized to examine ZW10 transcript levels across a wide range of non-diseased human tissues. 36 The GTEx data, quantified in transcripts per million (TPM), were processed using RNA-SeQC, which applies stringent quality control measures and refined gene annotations to exclude intronic sequences and overlapping transcript artifacts, thereby ensuring accurate and high-confidence expression profiling. 37 To assess the protein expression pattern of ZW10 in breast cancer tissues, immunohistochemistry (IHC) data were retrieved from the HPA database (https://www.proteinatlas.org/). The analysis utilized 3 independent antibodies against ZW10: HPA051253, HPA055410, and CAB011565. The staining intensity and fraction of positive cells were scored by HPA as strong, moderate, weak, or negative, providing a semi-quantitative overview of ZW10 protein expression across multiple breast cancer specimens. Representative IHC images and the distribution of staining intensities for each antibody were obtained directly from the HPA portal. A STROBE checklist from the EQUATOR Network can be found in Supplemental Table S1 for data reporting transparency. 38
Zeste White 10 protein expression in The Cancer Genome Atlas cancer data sets including patients with cancer
The cancer section of the HPA provides comprehensive data on mRNA and protein expression across 31 distinct types of human cancer (available at https://www.proteinatlas.org/about#cancer). This resource is supplemented by millions of in-house generated immunohistochemical images of tissue sections, which are stained to visualize protein localization and expression. In addition, the Kaplan-Meier survival plots are included, illustrating the correlation between the mRNA expression of each protein-coding gene and patient survival across various cancer types. This data set offers valuable insights into how gene expression levels may influence cancer progression and patient outcomes. To ensure consistency in expression analysis, ZW10 expression data from TCGA data sets were normalized using TPM, a widely accepted method for quantifying gene expression levels across samples with varying sequencing depths. The TPM normalization allows for direct comparison of gene expression across samples by accounting for gene length and sequencing depth, thereby providing a standardized measure of transcript abundance. This approach enables accurate assessment of ZW10 expression across different cancer types and facilitates correlation analyses with clinical outcomes.
Expression of protein-coding genes in human cancer cell lines and The Cancer Genome Atlas cancer patients
The cell line resource in the HPA contains information on genome-wide RNA expression profiles of human protein-coding genes in 1206 human cell lines, including 1132 cancer cell lines (https://www.proteinatlas.org/about#cell_line). The transcriptomic analysis involved classification based on the specificity of ZW10 expression across 28 different cancer types. In addition, we conducted distribution and clustering analyses of ZW10 expression across various cell lines and further examined the similarity of these cell lines to their respective cancer types. The expression patterns of ZW10 in TCGA cancer data sets were also explored using the UALCAN web portal, 39 where expression levels of microRNAs (miRNAs) and long non-coding RNAs (lncRNAs) for each tumor subgroup or normal sample were visualized as box-and-whisker plots (https://ualcan.path.uab.edu/cgi-bin/Pan-cancer.pl?genenam=ZW10).
Analysis of Zeste White 10 expression and immune infiltration profile in breast cancer
TIMER2.0 (http://timer.cistrome.org/) is a comprehensive web tool designed to analyze and visualize immune cell infiltration across various cancer types using data from TCGA. It provides multiple analytical modules to explore the relationships between gene expression and immune infiltration. Specifically, the Gene_DE module allows users to compare the expression levels of a selected gene between tumor tissues and their adjacent normal counterparts across all TCGA cancer types. The TCGA data set integrated into TIMER2.0 includes a wide range of cancer types along with corresponding normal tissue data, enabling detailed investigations into gene expression patterns and immune microenvironment dynamics in cancer. 40 To analyze the expression levels of ZW10 in BRCA, we utilized the Gene Set Cancer Analysis (GSCA) (https://guolab.wchscu.cn/GSCA/#/expression). The GSCA was also used for the analysis of ZW10 expression in different stages of breast invasive cancer. 41 For RNA-seq-based comparative analysis, TNMplot (https://tnmplot.com/analysis/) was employed to assess ZW10 expression across all TCGA cancer types as well as normal, tumor, and metastatic tissues using its pan-cancer visualization and gene expression in breast cancer. 42
Survival analysis
The Kaplan-Meier plotter (www.kmplot.com) database, which integrates both clinical and gene expression data, was utilized to evaluate the prognostic significance of ZW10 mRNA expression in breast cancer.10,43,44 This data set contained relapse-free survival (RFS) information for 4929 patients and overall survival (OS) data for 2976 individuals. To investigate the influence of ZW10 expression, patient samples were categorized into high- and low-expression groups based on the median expression level after uploading the gene expression data. The Kaplan-Meier survival curves were subsequently generated, displaying the number of patients at risk beneath the primary plot. The platform also provided statistical outputs, including the log-rank P-value, hazard ratio (HR), and 95% confidence intervals, considering a P-value < .05 as statistically significant. To maintain data reliability, “array quality control” measures were implemented to eliminate biased arrays. In addition, survival data from TCGA were retrieved using the UALCAN platform (http://ualcan.path.uab.edu/), enabling further analysis of survival probability in breast cancer patients based on ZW10 expression levels, with menopause status also taken into account. 39
Gene correlation analyses
To identify genes that interact with or co-express alongside ZW10, we conducted a correlation analysis using its RNA expression data. The Pearson correlation analysis was applied to the expression levels of all protein-coding genes, utilizing the UALCAN platform. Genes that exhibited significant positive or negative correlations with ZW10 were further analyzed. To ensure data accuracy, genes with low-expression levels (median TPM < 0.5) were excluded from the analysis. Gene pairs with a Pearson correlation coefficient of 0.3 or greater were categorized as positively correlated, while pairs with a coefficient of −0.3 or lower were classified as negatively correlated. 39
Protein-protein interaction network construction
The STRING database (http://string-db.org/) is a comprehensive resource for constructing functional protein interaction networks. It integrates experimentally validated and computationally predicted protein-protein interactions (PPIs) across various organisms. In this study, we focused on interactions specific to Homo sapiens and applied a high-confidence threshold of 0.9 to enhance the reliability of the results. This approach facilitated the identification of key PPIs involving ZW10, underscoring its functional significance in cellular processes. 45
Mutation and copy number variation analysis of Zeste White 10
The mutations in ZW10 were analyzed using the Catalog of Somatic Mutations in Cancer (COSMIC, GRCh38 • COSMIC v97) database, which provides an extensive collection of somatic mutations discovered across various human cancer types. This resource allows for the detailed examination of genetic alterations, offering valuable insights into the mutation landscape of ZW10 in cancer.46,47 The distribution of various mutations was included and presented in the form of a pie chart.
Genetic alteration analysis of Zeste White 10 in pan-cancer
Data on ZW10 copy number alterations and mutation profiles across various cancer types were retrieved from the cBioPortal database. This platform integrates genomic data sets from multiple large-scale cancer studies, enabling the comprehensive assessment of genetic alterations, including gene amplifications, deletions, and specific mutations in ZW10 across diverse tumor types.
Functional enrichment analysis
The ZW10-interacting partners were identified using the STRING database with a confidence score threshold of >0.9. The top 10 genes with the strongest association were selected for downstream functional analysis. To elucidate the biological processes, cellular components, molecular functions, and pathways associated with the ZW10-centric network, Gene Ontology (GO) and Reactome pathway enrichment analyses were performed. The gene set, comprising ZW10 and its 10 interactors, was submitted to the EnrichR web server (https://maayanlab.cloud/EnrichR/). 48 The analysis was run against the GO Biological Process 2025, GO Cellular Component 2025, GO Molecular Function 202, and Reactome Pathway 2022 libraries. Terms with an adjusted P-value (Benjamini-Hochberg) of <.05 were considered statistically significant.
Statistical analysis
The significance levels are denoted as follows: ns indicates not significant, * corresponds to a P-value < .05, ** represents a P-value < .01, and *** denotes a P-value < .001. Log-rank P-values were computed alongside HRs and their corresponding 95% confidence intervals to assess statistical significance. A P-value threshold of <.05 was used to determine statistically significant results.
Results
Zeste White 10 expression in human tissues and cancer
To characterize the expression profile of ZW10, we examined its distribution at both the transcript and protein levels across normal human tissues and cancer using data from the HPA, GTEx project, and TCGA. Analysis of GTEx and HPA transcriptomic data revealed that ZW10 exhibits a tissue-specific expression pattern, with relatively high expression observed in several tissues, including the breast (Figure 1A and Supplementary Figure S1).

Overview of the expression of ZW10 in human tissues and organs (https://www.proteinatlas.org/ENSG00000157456-CCNB2/tissue). The ZW10 is expressed at the protein level (A) and RNA level (B) in human tissues and organs, indicating that ZW10 is a tissue-specific protein (https://www.proteinatlas.org/). (C) Representative immunohistochemistry images showing moderate ZW10 protein expression in breast cancer tissues. (D) Bar charts illustrating the distribution of ZW10 staining intensities across analyzed cases.
Protein-level expression of ZW10 in breast cancer tissues was further evaluated using IHC data from the HPA. As shown in Figure 1B to D, ZW10 protein was detectable in most breast cancer samples and was predominantly localized to the cytoplasm of tumor cells. Consistent staining patterns were observed across multiple antibodies. Specifically, antibody HPA051253 showed moderate staining in 10 of 11 cases and weak staining in 1 case; antibody HPA055410 demonstrated moderate to strong staining in 10 of 11 cases; and antibody CAB011565 showed weak staining in 10 of 11 cases and moderate staining in 1 case. Collectively, these results confirm the presence of ZW10 protein in breast cancer tissues, with predominantly weak to moderate expression intensity.
To further assess ZW10 expression in cancer, transcript abundance was examined across cancer-derived cell lines and tumor samples. The RNA specificity and TPM analyses indicated that ZW10 is expressed in multiple cancer cell lines (Figure 2A). Consistent with these findings, TCGA-based analyses demonstrated increased ZW10 mRNA expression in a broad range of tumor types compared with corresponding normal tissues, including breast cancer (Figure 2B). Elevated ZW10 expression was also observed in tumors originating from the adrenal gland, colon, liver, lung, ovary, pancreas, prostate, rectum, skin, stomach, testis, thyroid, and uterus (Figure 2C).
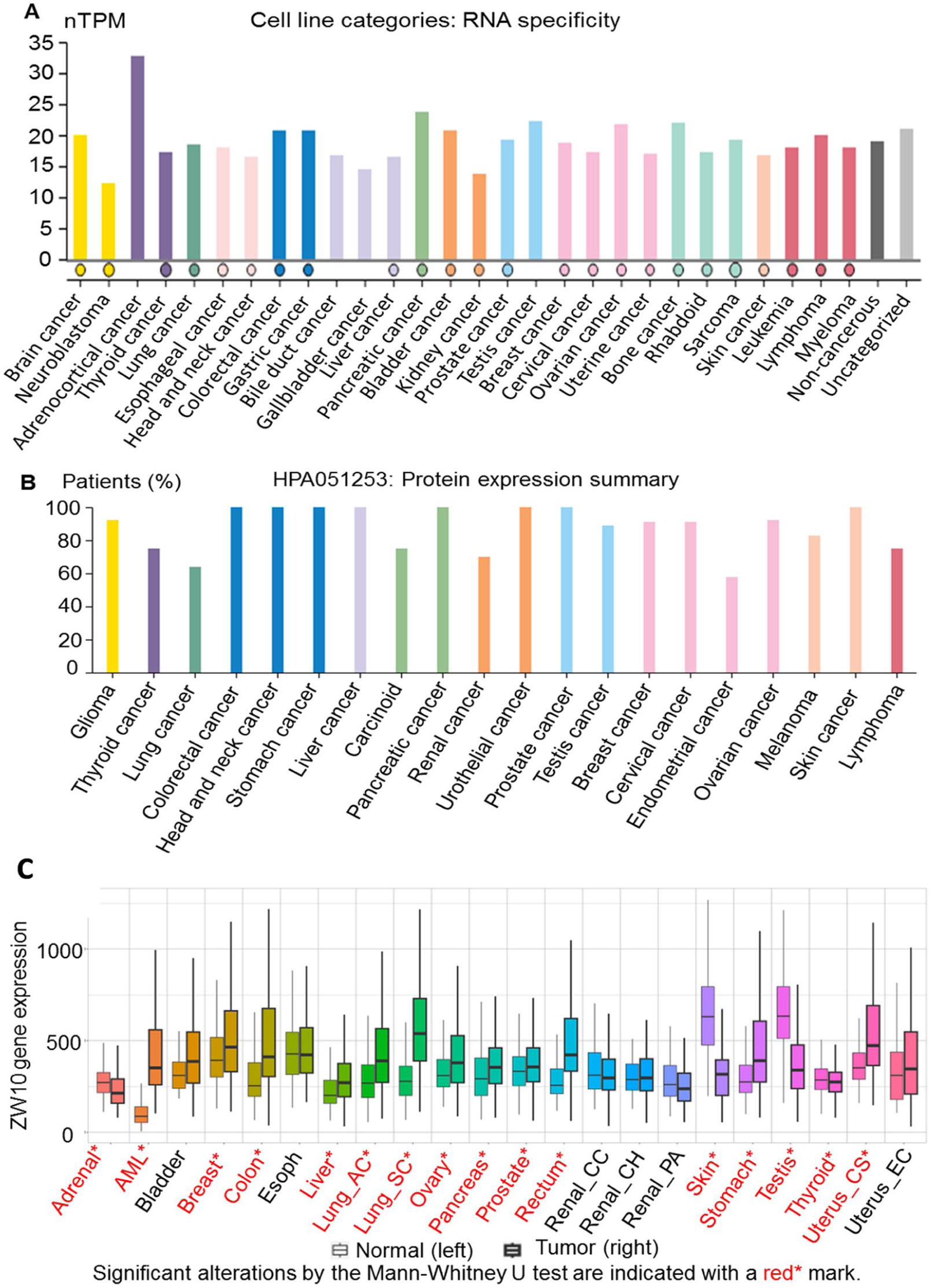
Overview of the expression of ZW10 in human cancer. (A) Expression profile of ZW10 in different cancer cell lines with respect to their RNA specificity (https://www.proteinatlas.org/ENSG00000086827-ZW10/cell+line). (B) ZW10 is expressed at the protein level in different human cancers (https://www.proteinatlas.org/ENSG00000086827-ZW10/cancer). (C) Pan-cancer analysis of ZW10 mRNA expression comparing tumor and corresponding normal tissues using RNA-seq data from The Cancer Genome Atlas (TCGA).
Overall, these results demonstrate that ZW10 is broadly expressed at both the transcript and protein levels in breast cancer and multiple other malignancies, providing a foundation for subsequent analyses of its prognostic relevance and biological associations.
Zeste White 10 expression and its association with breast cancer subtypes
Given the observed overexpression of ZW10 in various cancers, we aimed to explore its potential clinical significance in the progression of breast cancer. Utilizing transcriptomic data sets from TCGA, we analyzed ZW10 expression across various clinical stages and molecular subtypes of breast cancer. The analysis revealed a significant elevation of ZW10 expression across all stages of disease progression, indicating its persistent presence throughout tumor development (Figure 3A to D).

Expression of ZW10 in breast cancer stages and subtypes. (A) HCCS gene signature expression in breast adenocarcinoma, including the normal, tumor, and metastatic, was analyzed using RNA-seq data. (B) HCCS gene expression in breast adenocarcinoma, accounting for the different stages, namely stage I, stage II, stage III, and stage IV, using RNA-seq-based data (https://tnmplot.com/analysis/). Statistical significance was determined using the Kruskal-Wallis test. (C) Boxplots depict the expression levels of ZW10 across different subtypes of breast cancer, as analyzed through the Gene Set Cancer Analysis (GSCA) platform using TCGA transcriptomic data. (D) Boxplots depict the expression levels of ZW10 mRNA across different pathological stages of breast cancer, as analyzed through the GSCA platform using TCGA transcriptomic data. The P-values < .05 were considered statistically significant. (E) Differential expression of ZW10 in breast cancer subtypes, with a focus on triple-negative breast cancer (TNBC) subcategories. Boxplots illustrate ZW10 expression across different breast cancer stages and molecular subtypes.
To refine our understanding of ZW10’s role within breast cancer heterogeneity, expression levels were further stratified according to molecular subtypes. Particular attention was given to TNBC, a subtype characterized by the absence of estrogen receptor, progesterone receptor, and HER2 amplification, and known for its aggressive behavior and limited treatment options. Notably, ZW10 expression was markedly higher in TNBC samples compared with other subtypes, suggesting a possible link between ZW10 upregulation and the aggressive clinical phenotype of TNBC (Figure 3E). The elevated expression of ZW10 in TNBC, in conjunction with its upregulation across broader breast cancer categories, highlights a potential association between ZW10 activity and tumor aggressiveness. These findings suggest that ZW10 may contribute to the oncogenic landscape of breast cancer, particularly in clinically challenging subtypes such as TNBC. Its consistent expression pattern underscores its potential utility not only as a molecular marker for disease progression but also as a candidate for targeted therapeutic intervention.
Together, these results reinforce the hypothesis that ZW10 plays a role in tumorigenesis and provide a compelling rationale for future functional studies to elucidate its mechanistic contribution to breast cancer pathophysiology.
Association between Zeste White 10 expression and immune cell infiltration in breast cancer
Building on the observed upregulation of ZW10 in breast cancer and its subtypes, particularly in triple-negative cases, we further explored the potential immunological role of ZW10 by examining its correlation with immune cell infiltration within the tumor microenvironment. Utilizing transcriptomic profiles from TCGA breast cancer data sets, the Spearman correlation analysis was performed to assess the association between ZW10 expression and the abundance of diverse immune cell populations.
The analysis revealed statistically significant positive correlations between ZW10 expression and multiple immune cell types, including CD8+ cytotoxic T cells, CD4+ helper T cells, regulatory T cells (Tregs), neutrophils, monocytes, macrophages, myeloid dendritic cells, natural killer (NK) cells, mast cells, and common lymphoid progenitor cells (Figure 4A to J). These results suggest that ZW10 expression is closely linked to both pro-inflammatory and immunosuppressive immune components, pointing toward a potential regulatory role in shaping the immune milieu of breast tumors.

Correlation between ZW10 expression and immune cell infiltration in breast cancer. Scatter plots illustrate the Spearman correlation between ZW10 expression and the abundance of various immune cell populations in breast cancer based on TCGA transcriptomic data. (A) CD8+ T cells, (B) CD4+ T cells, (C) regulatory T cells, (D) neutrophils, (E) monocyte, (F) macrophages, (G) myeloid dendritic cell, (H) natural killer (NK) cell, (I) mast cells, and (J) common lymphoid progenitor cell, each demonstrated a statistically significant positive association with ZW10 expression levels.
To determine whether this immune association is conserved across molecular subtypes of breast cancer, further stratified analyses were conducted within basal-like, HER2-enriched, luminal A, and luminal B subgroups. The correlation patterns between ZW10 expression and immune cell infiltration remained consistent across these subtypes (Supplementary Figure S2), reinforcing the hypothesis that ZW10 may serve as a central mediator of immune-tumor interactions in breast cancer.
These findings indicate that ZW10 may play a dual role in breast cancer biology—not only contributing to tumor progression through its elevated expression but also serving as a key regulator of the immune microenvironment. The integration of quantitative immunocytochemistry underscores the utility of ZW10 as a measurable biomarker that could aid in identifying specific immune cell populations within the tumor milieu, offering new avenues for immunotherapeutic targeting in breast cancer.
Clinical outcome of Zeste White 10 expression in breast cancer prognosis
To elucidate the prognostic significance of ZW10 expression in breast cancer, we extended our analysis to assess patient survival outcomes using clinical data obtained from TCGA. Specifically, we examined the association between ZW10 mRNA expression levels, menopausal status, and survival probability in breast cancer patients. Stratification of the cohort indicated that postmenopausal patients with high ZW10 expression exhibited significantly poorer OS compared with those with other groups (Figure 5A). The observed difference in survival probabilities between these groups was statistically significant (P < .05), suggesting that ZW10 expression may have a clinically relevant impact on breast cancer progression in postmenopausal women.

Survival analysis of ZW10 in breast invasive carcinoma. (A) Impact of ZW10 expression and menopausal status on the survival outcomes of patients with breast invasive carcinoma (BRCA). Data were analyzed using the UALCAN platform (http://ualcan.path.uab.edu/) based on TCGA data sets. Values with P < .05 were considered significant. (B, C) Kaplan-Meier survival curves illustrating the association between ZW10 expression and patient survival outcomes in breast invasive carcinoma. (B) Overall survival (OS) analysis (n = 2976) was performed using mRNA-seq data of breast cancer patients, and the right panel shows the significance vs cutoff values between lower and upper quartiles of expression. (C) Relapse-free survival (RFS) analysis (n = 4929) was performed using mRNA gene chip data of breast cancer patients.
To further substantiate these findings, we conducted an independent survival analysis using the Kaplan-Meier plotter, which leverages extensive gene expression data sets to evaluate the prognostic value of ZW10. The Affymetrix probe ID 204812_at, corresponding to ZW10, was utilized to analyze both OS and RFS. Patients were stratified into high- and low-expression groups based on the median ZW10 mRNA expression level.
In the OS analysis, patients with higher ZW10 expression exhibited a significant trend toward reduced survival (n = 2976, HR = 1.71 [1.36-2.15], P = 4.6e-06), while lower ZW10 expression levels were associated with a modest improvement in survival. This trend reached statistical significance (Figure 5B), indicating a potential association between elevated ZW10 expression and adverse survival outcomes in breast cancer patients.
Conversely, a significant association was observed between ZW10 expression and RFS. Patients in the high ZW10 expression group experienced a significantly shorter RFS compared with those with lower expression levels (n = 4929, HR = 0.86 [0.78-0.95], P = .0042) (Figure 5C). These findings suggest that reduced ZW10 expression may serve as a favorable prognostic marker for relapse prevention in breast cancer.
Collectively, these results underscore the clinical relevance of ZW10 expression in breast cancer prognosis. The significant association between lower ZW10 expression and improved RFS highlights its potential as a prognostic biomarker in breast cancer. Furthermore, the observed relationship between lower ZW10 expression and enhanced OS further suggests its role in breast cancer progression, particularly in the context of its elevated expression in TNBC. These findings provide a compelling rationale for further investigation of ZW10 as a therapeutic target in breast cancer.
Zeste White 10-associated gene interaction and protein-protein networks in breast cancer
In light of the established overexpression of ZW10 in various cancers and its prognostic relevance in breast cancer, particularly in TNBC, we further explored the broader molecular framework in which ZW10 operates. Genes implicated in oncogenesis often function within dynamic regulatory networks that influence fundamental cellular processes, including the cell cycle, mitotic fidelity, and tumor progression. As a potential master regulator, ZW10 may orchestrate diverse downstream pathways contributing to tumor biology.
To uncover the molecular interactions associated with ZW10, we performed expression-based correlation analyses to identify genes co-expressed with ZW10 across cancer tissues. This analysis revealed the top 25 genes showing strong positive correlation with ZW10 expression (Figure 6A), indicative of coordinated activity within potentially shared biological processes. For clarity, we have analyzed 1 gene (DLAT gene) that is positively correlated (r ⩾ 0.3) with ZW10 as a representative image of all genes (Figure 6A, right panel). In parallel, the top 25 genes negatively correlated with ZW10 were identified (Figure 6B), suggesting regulatory divergence or involvement in opposing cellular functions. Similarly, we have analyzed 1 gene (PHPT1 gene) that is negatively correlated (r ⩽ 0.3) with ZW10 as a representative image of all genes (Figure 6B, right panel).

Gene expression correlation analysis of ZW10 in BRCA. (A) Expression pattern of ZW10 in breast invasive carcinoma. Heat map showing the top 25 genes positively correlated with promoter DNA of ZW10 in breast invasive carcinoma. Right panel: representative image of all other genes. Scatter plot highlighting the gene (DLAT gene) with the highest Pearson correlation coefficient (r > 0.3) that is positively correlated with ZW10 across all breast cancer samples. (B) Heat map showing the top 25 genes that are negatively correlated with the promoter DNA of ZW10 in breast invasive carcinoma. Right panel: representative image of all other genes. Scatter plot highlighting the gene (PHPT1 gene) negatively correlated with ZW10 a lower Pearson correlation coefficient (r < 0.3) across all breast cancer samples. Genes with extremely low expression (median TPM < 0.5) are filtered out of the list.
These correlation patterns provide important insights into the functional landscape of ZW10. Genes with positive correlation may act synergistically within oncogenic pathways that support mitotic regulation, proliferation, or metastatic potential. On the contrary, negatively correlated genes could participate in antagonistic pathways or tumor-suppressive mechanisms that counterbalance ZW10-mediated effects. The identification of these co-regulated genes adds a layer of complexity to our understanding of ZW10’s role in cancer and may inform future functional studies aimed at elucidating regulatory hierarchies.
Building upon the gene correlation analysis, we constructed a PPI network for ZW10 using high-confidence data retrieved from the STRING database. Protein interactions are critical in regulating signaling cascades, and mapping these interactions provides a deeper understanding of ZW10’s role within cellular systems. The resulting PPI network revealed a set of top predicted functional partners with high interaction confidence scores (Supplementary Figure S3). Among these were RINT1 (score = 0.999), ZWILCH (score = 0.999), KNTC1 (score = 0.999), STX18 (score = 0.998), NBAS (score = 0.995), Zwint-1 (score = 0.991), DCTN2 (score = 0.991), BUB3 (score = 0.974), USE1 (score = 0.957), NDC80 (score = 0.952), CENP-E (score = 0.910), BUB1 (score = 0.945), BNP1 (score = 0.942), BUB1B (score = 0.906), DCTN1 (score = 0.644), and PLK1 (score = 0.514).
Strikingly, several of these interacting proteins—such as CENP-E, BUB1, BUB1B, PLK1, and DCTN1—are integral to mitotic checkpoint control and chromosomal stability. 18 ,49 -51 Their association with ZW10 points toward its active involvement in processes such as spindle assembly, kinetochore-microtubule attachment, and regulation of anaphase onset. These mitotic regulators are frequently dysregulated in cancer, implicating ZW10 as a potentially critical node within mitosis-related oncogenic pathways.
Together, these findings extend previous observations by mapping both transcriptional and protein-level interactions of ZW10, revealing its centrality in cell-cycle regulation and mitotic checkpoint fidelity. The integration of expression correlation and PPI analysis reinforces ZW10’s functional relevance in breast cancer and highlights new avenues for mechanistic investigation. These networks not only provide a framework for future functional studies but also lay the groundwork for exploring ZW10 and its interactors as therapeutic targets, particularly in aggressive cancer subtypes such as TNBC.
Mutational landscape of Zeste White 10 across human tissues and its oncogenic implications
Following the comprehensive analysis of ZW10 expression, immune cell infiltration, prognostic relevance, and interaction networks, we further investigated the genetic alterations of ZW10 to elucidate its potential oncogenic implications. Genetic mutations in key regulatory genes, particularly oncogenes and tumor suppressors, are critical drivers of malignant transformation and tumor progression. To characterize the mutation spectrum of ZW10 across diverse cancer types, we utilized data from the COSMIC.
Our analysis identified 2 primary categories of ZW10 mutations: missense substitutions and synonymous substitutions (Figure 7A). Missense mutations, which result in amino acid changes, may disrupt the structural integrity or functional domains of ZW10, potentially impairing its role in mitotic checkpoint regulation. Conversely, synonymous mutations do not alter the protein sequence but may influence regulatory processes, including mRNA stability, splicing, or translation efficiency.

ZW10 mutations in all primary tissue types that are curated by COSMIC. Pie charts demonstrating the (A) distribution and (B) percentage of the substitution types of ZW10 in all TCGA cancers based on results from the COSMIC database (https://cancer.sanger.ac.uk/cosmic/gene/analysis?ln=ZW10#distribution).
A more detailed examination of the mutational profile revealed that A > G (10.18%) and A > T (3.51%) transitions were the most prevalent nucleotide changes within the ZW10 coding sequence (Figure 7B). These specific alterations could impact transcript processing or protein synthesis, thereby contributing to ZW10 dysregulation in tumor cells. The localization of these mutations within functional domains suggests a potential link to disrupted mitotic control and CIN, key hallmarks of cancer.
Given the influence of genetic alterations on gene expression, we further explored ZW10 genetic alterations using the cBioPortal database to assess its mutation and amplification profiles across a pan-cancer cohort. Our findings indicated that gene amplification was the most common genetic alteration of ZW10 across multiple cancer types, followed by missense mutations (Supplementary Figure S4). This pattern of amplification suggests that ZW10 dysregulation may contribute to cancer progression by enhancing gene dosage effects.
Notably, ZW10 mutations were detected across various malignancies, including breast, lung, prostate, ovary, liver, skin, testis, and penile cancers. This widespread mutational distribution implies that ZW10 may exhibit tissue-specific roles in oncogenesis, potentially modulated by distinct molecular contexts. These findings align with our previous observations of ZW10 overexpression in multiple cancers and its associations with immune modulation, patient prognosis, and regulatory interaction networks, particularly in breast cancer and TNBC.
The concurrent upregulation of ZW10 expression and occurrence of genetic mutations further underscores its potential contribution to tumorigenesis through transcriptional dysregulation and genomic instability. In addition, the PPI network identified several key mitotic regulators, including those involved in spindle assembly and kinetochore function. The ZW10 mutations could potentially disrupt these interactions, leading to aberrant mitotic checkpoint signaling and CIN, further driving cancer progression.
In summary, the mutational landscape of ZW10 provides critical insights into its potential oncogenic role across diverse cancer types. The identification of recurrent substitution patterns, amplification events, and their distribution across malignancies strengthens the hypothesis that ZW10 may serve as a molecular driver of cancer progression. These findings underscore the need for targeted functional studies to validate ZW10’s regulatory role in mitotic control and its potential as a therapeutic target, particularly in aggressive subtypes such as TNBC.
Functional enrichment of the Zeste White 10 interactome implicates dual roles in mitosis and trafficking
To define the biological functions of the ZW10 interaction network, we performed GO and Reactome pathway analysis. The results reveal that ZW10 and its partners are significantly enriched in 2 core cellular processes. Mitotic Spindle Checkpoint and Chromosome Segregation: The most significantly enriched terms were overwhelmingly related to cell division: The network was highly enriched for biological processes such as “Mitotic Spindle Checkpoint Signaling” (GO:0007094), “Regulation of Attachment of Spindle Microtubules to Kinetochore” (GO:0051988), and “Sister Chromatid Segregation” (GO:0000819) (Figure 8A). Accordingly, the core cellular components identified were the “Kinetochore Microtubule” and “Spindle Microtubule” (Figure 8B). This was further corroborated by Reactome pathways, including “Mitotic Spindle Checkpoint” and “Amplification of Signal from Unattached Kinetochores” (Figure 8D). These findings computationally validate that the ZW10 interactome is a critical component of the machinery that ensures accurate chromosome segregation during mitosis.

Gene ontology and reactome pathway enrichment of the ZW10 interactome. The network of ZW10 and its top 10 correlated genes was analyzed for functional enrichment. The bar charts display the most significant terms in the categories of (A) Biological Process, (B) Cellular Component, (C) Molecular Function, and (D) Reactome Pathways.
Golgi-to-ER Retrograde Vesicular Transport: Concurrently, the analysis revealed a second, distinct functional module within the network dedicated to intracellular trafficking. We observed significant enrichment for terms like “Retrograde Vesicle-Mediated Transport, Golgi to ER” (GO:0006890) and “Regulation of ER to Golgi Vesicle-Mediated Transport” (GO:0060628) (Figure 8A). The associated cellular components included various vesicle membranes, such as the “COPI-coated ER to Golgi Transport Vesicle” (GO:0030134) and “ER to Golgi Transport Vesicle Membrane” (GO:0012507) (Figure 8B). The Reactome pathway analysis solidified this with enrichments for “COPI-dependent Golgi-to-ER Retrograde Traffic” and “Intra-Golgi and Retrograde Golgi-to-ER Traffic” (Figure 8D). The molecular function analysis primarily indicated roles in “GTPase Binding” and “Sequence-Specific DNA Binding” (Figure 8C), suggesting mechanisms for regulatory control and potential gene expression influence. In summary, the functional enrichment analysis provides a strong biological rationale for the ZW10 interactome, implicating it directly in the fundamental processes of cell-cycle regulation and intracellular vesicle trafficking.
Discussion
This study provides a comprehensive multiple omics characterization of ZW10, elucidating its expression patterns, prognostic relevance, immune associations, interaction networks, and mutational spectrum across cancers, with an emphasis on breast cancer and TNBC. Our findings indicate that ZW10 is significantly overexpressed in several malignancies and is closely associated with poor survival outcomes, suggesting its oncogenic potential and utility as a prognostic biomarker.
The ZW10’s upregulation across diverse tumors aligns with earlier reports linking mitotic checkpoint dysregulation to CIN and tumor progression.20,52 -54 In breast cancer, high ZW10 expression is particularly pronounced in TNBC, a subtype known for its aggressive phenotype and limited treatment options.7,24 Elevated ZW10 expression was associated with poorer OS and shorter relapse-free survival, indicating that aberrant ZW10 activity may drive tumor aggressiveness and therapy resistance.7,55
Consistent with transcriptomic analyses, IHC data from the HPA confirmed that ZW10 protein is highly expressed in breast cancer tissues but not detected in normal breast tissues. This differential expression pattern implies a potential role of ZW10 in breast cancer progression. As a kinetochore-associated protein involved in chromosome segregation and mitotic checkpoint regulation, the elevated expression of ZW10 may contribute to increased proliferative capacity and CIN in tumor cells. The concordant staining patterns obtained from multiple independent antibodies enhance the reliability of these observations. Together, these results suggest that ZW10 may act as a potential oncogenic factor and diagnostic or therapeutic biomarker in breast cancer, warranting further functional validation in independent cohorts.
While traditional biomarkers like Ki-67 (a proliferation marker) and HER2 (a receptor tyrosine kinase) are well-established in clinical practice, ZW10 offers unique advantages. Unlike HER2, which is relevant only in HER2-enriched subtypes, ZW10’s prognostic power spans multiple breast cancer subtypes, including TNBC, where biomarkers are scarce. 7 In addition, ZW10’s involvement in mitotic regulation and CIN provides mechanistic insights into tumor aggressiveness, a feature not fully captured by Ki-67 alone.54,55 However, further validation in prospective clinical cohorts is needed to determine whether ZW10 outperforms or complements existing markers.
The ZW10 is a key component of the mitotic checkpoint, ensuring proper chromosome segregation by regulating kinetochore-microtubule attachments.54,55 Dysregulation of ZW10 leads to CIN, a hallmark of cancer progression, and may explain its association with poor prognosis. Intriguingly, ZW10 also correlates with immune cell infiltration, including CD8+ T cells and macrophages, suggesting a dual role in both tumor proliferation and immune evasion. 54 This finding aligns with recent studies highlighting the interplay between mitotic errors and antitumor immunity.52,55 Recent pan-cancer analyses further validate ZW10’s upregulation in pancreatic adenocarcinoma (PAAD) and its role in cell-cycle progression via E2-F1 signaling. 54 In addition, CRISPR screening in hepatocellular carcinoma (HCC) identifies ZW10 as a tyrosine kinase inhibitor (TKI)-resistance gene, expanding its relevance beyond breast cancer. 55 These studies underscore ZW10’s broad oncogenic potential and justify further exploration in therapeutic contexts. Given ZW10’s involvement in TKI resistance, 55 targeting its interactors (eg, ZWINT, BUB1) or downstream pathways (eg, CDK1-mediated cell-cycle progression) could offer novel treatment strategies.24,54 Small-molecule inhibitors disrupting ZW10’s mitotic functions or immunomodulatory effects may be particularly relevant for TNBC, where current options are limited.
Immunohistochemistry is a vital laboratory staining technique employed to detect specific immune cell populations within mouse and human tissues, including tumor specimens. 56 By implementing quantitative immunocytochemistry with established positive controls, IHC enables precise characterization of immune cell infiltration, thereby facilitating the identification of novel biomarkers, such as ZW10. In the context of breast cancer, assessing the tumor immune microenvironment, including lymphovascular invasion in primary tumors, skin recurrences, and visceral metastases, provides critical insights into the temporal and spatial dynamics of tumor-infiltrating lymphocytes (TILs). This approach allows for the evaluation of immune cell populations, such as CD8+ cytotoxic T cells, CD4+ helper T cells, macrophages, and Tregs, across distinct tumor sites and stages of disease progression. By establishing ZW10 as a potential biomarker through quantitative immunocytochemistry, we can systematically evaluate its expression in conjunction with immune cell infiltration patterns. This combined analysis not only aids in delineating the immune landscape of breast tumors but also helps to identify specific immune subsets that may correlate with clinical outcomes and therapeutic responsiveness. 57 To better contextualize the observed association between ZW10 expression and immune infiltration, a plausible mechanistic framework can be proposed based on ZW10’s established roles in mitotic checkpoint regulation and intracellular trafficking. Dysregulation of mitotic checkpoint components has been shown to promote CIN, leading to micronuclei formation and the accumulation of cytosolic DNA, which can activate innate immune pathways such as the cyclic GMP-AMP synthase-stimulator of interferon genes (cGAS-STING) axis.58 -60 Activation of this pathway is known to induce type I interferon signaling and shape immune cell recruitment within the tumor microenvironment. 61 In parallel, ZW10’s role in Golgi-to-ER retrograde vesicular transport suggests a potential influence on antigen processing and presentation, processes that are critical for effective tumor immune surveillance. Disruption of vesicular trafficking has been linked to altered major histocompatibility complex (MHC) presentation and impaired cytotoxic T cell recognition. 62 Therefore, the positive correlation between ZW10 expression and multiple immune cell populations observed in this study may reflect a dual functional role, whereby ZW10-driven mitotic stress enhances immunogenic signaling while concomitant alterations in intracellular trafficking contribute to immune evasion. Collectively, these mechanisms provide a biologically coherent explanation linking ZW10 overexpression to immune infiltration patterns in breast cancer and support its relevance as an active regulator of tumor-immune interactions rather than a passive biomarker. Thus, the integration of immunocytochemistry with immune profiling underscores ZW10’s role as a quantifiable marker of immune-tumor interactions, offering a compelling framework for future immunotherapy studies in breast cancer.
The survival analyses revealed that high ZW10 expression was associated with improved but correlated with poorer OS in breast cancer patients. The apparently divergent associations of ZW10 expression with RFS and OS reflect endpoint-specific and context-dependent tumor biology rather than an unresolved inconsistency. In this study, RFS was evaluated using the Kaplan-Meier Plotter data set, which predominantly captures recurrence dynamics in earlier-stage disease, whereas OS analysis incorporated TCGA cohorts characterized by longer follow-up and a higher representation of advanced, treatment-refractory tumors. Consistent with this distinction, our subtype-stratified analyses demonstrate that ZW10 expression is the highest in aggressive breast cancer subtypes, particularly TNBC, which disproportionately influence OS outcomes. Moreover, immune infiltration analyses reveal that high ZW10 expression correlates simultaneously with cytotoxic immune cells (eg, CD8+ T cells) and immunosuppressive populations (eg, Tregs and macrophages), suggesting a dual immune context that may transiently restrain recurrence while promoting long-term immune dysfunction and tumor progression. Finally, functional enrichment and protein-protein interaction analyses show that ZW10 participates in both mitotic checkpoint control and vesicular trafficking pathways, indicating pleiotropic, stage-dependent roles that are well recognized to produce differential prognostic effects for mitotic regulators. Collectively, these data support a biologically coherent, data-driven explanation for the differential RFS and OS associations observed for ZW10.
Protein-protein interaction analysis revealed that ZW10 interacts with key mitotic checkpoint regulators, including RINT1, ZWILCH, KNTC1, and BUB3. These proteins are essential for spindle checkpoint function, chromosome alignment, and kinetochore-microtubule attachment, processes that are critical for genomic stability. 13 The involvement of ZW10 in small GTPase-mediated signal transduction and chromosome segregation further supports its role in mitotic regulation and tumor development. Notably, overexpression of mitotic regulators such as BUB1 and PLK1 has been associated with chemoresistance and poor prognosis in breast cancer, highlighting the potential of ZW10 and its interacting partners as therapeutic targets. 27 The functional enrichment of the ZW10 interactome provides a compelling biological rationale for our findings, implicating it in both mitotic fidelity and vesicular trafficking. We hypothesize that this duality explains its complex role in breast cancer: dysregulation of the spindle checkpoint could drive genomic instability and tumor progression, while its role in retrograde transport may alter antigen presentation and contribute to immune evasion. This model positions ZW10 as a critical integrator of 2 core cancer hallmarks.
Mutation analysis using the COSMIC database identified missense and synonymous substitutions as the predominant alterations in ZW10, with A > G (10.18%) and A > T (3.51%) being the most common mutations. These mutations could potentially disrupt ZW10 function, leading to mitotic errors and tumor progression. Prior studies have demonstrated that mutations in mitotic checkpoint genes contribute to aneuploidy and cancer aggressiveness, reinforcing the significance of ZW10’s mutational landscape. 28
Although the present study does not provide direct mechanistic or pharmacological evidence supporting ZW10 as a therapeutic target, our findings suggest that ZW10 may represent a biologically relevant candidate for further investigation in breast cancer. The ZW10 is consistently overexpressed in aggressive tumor subtypes, exhibits subtype- and endpoint-dependent prognostic associations, and occupies a central position within networks related to mitotic checkpoint regulation and vesicular trafficking. In addition, its significant correlations with multiple immune cell populations indicate a potential role in shaping the tumor microenvironment. Importantly, these observations are derived from integrative transcriptomic, survival, immune infiltration, and network analyses and should be interpreted as hypothesis-generating rather than confirmatory. Future studies employing functional perturbation, pathway-specific assays, and therapeutic sensitivity models will be necessary to determine whether ZW10 plays a causal role in tumor progression or represents a viable therapeutic vulnerability. Until such experimental validation is available, ZW10 should be considered a candidate biomarker and biological regulator rather than an established therapeutic target.
Limitations and future directions
This study is based exclusively on integrative bioinformatics analyses of publicly available data sets, including TCGA, the HPA, COSMIC, STRING, and the Kaplan-Meier Plotter. While these resources provide valuable large-scale insights into ZW10 expression patterns, prognostic associations, immune infiltration, and mutational profiles, the lack of experimental validation represents an important limitation of the present work. Consequently, the observed associations should be interpreted as hypothesis-generating rather than mechanistically definitive.
Future studies should incorporate functional validation to clarify the biological role of ZW10 in breast cancer. In particular, genetic perturbation approaches, such as siRNA-mediated knockdown or overexpression of ZW10 in relevant breast cancer models, including TNBC cell lines, followed by assessment of cell proliferation, apoptosis, and cell-cycle regulation, would help establish causal relationships. In addition, validation of ZW10 expression at the mRNA and protein levels in independent breast cancer tissues or cell lines using techniques such as quantitative reverse transcription polymerase chain reaction (qRT-PCR) and Western blotting would strengthen its potential utility as a biomarker. Furthermore, pathway-specific functional assays and therapeutic sensitivity analyses will be required to determine whether ZW10 contributes directly to tumor progression or treatment response. Addressing these limitations through experimental and clinical validation will be essential to translate the current in silico findings into biologically and clinically meaningful conclusions.
Conclusion
In conclusion, this study provides a comprehensive integrative analysis of ZW10 expression, prognostic relevance, and immune associations in breast cancer. Our results demonstrate that ZW10 exhibits subtype- and endpoint-dependent prognostic patterns and is closely associated with key components of the tumor immune microenvironment. These observations suggest that ZW10 may serve as a context-dependent biomarker reflecting tumor progression and immune regulation. However, the present findings are derived from bioinformatics analyses and do not establish causal or mechanistic roles. Further experimental and clinical studies will be required to elucidate the functional significance of ZW10 and to determine its potential relevance in therapeutic or translational settings.
Supplemental Material
sj-docx-1-bcb-10.1177_11782234261428784 – Supplemental material for Zeste White 10 May Serve as a Prognostic Biomarker and Therapeutic Target for Human Breast Cancer
Supplemental material, sj-docx-1-bcb-10.1177_11782234261428784 for Zeste White 10 May Serve as a Prognostic Biomarker and Therapeutic Target for Human Breast Cancer by Sm Faysal Bellah, Md Alim Hossen, SM Saker Billah, Olanrewaju Ayodeji Durojaye and Md Obayed Raihan in Breast Cancer: Basic and Clinical Research
Footnotes
Acknowledgements
Ethical Considerations
This study is a computational biology research study and is exempt from institutional review board (IRB) approval.
Consent to Participate
Not applicable.
Consent for Publication
Not applicable.
Author Contributions
Funding
The authors received no financial support for the research; authorship, and/or publication of this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
All data generated or analyzed during this study are included in this published article and the Supplementary Materials.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
