Abstract
Background:
Variants of the matrix metalloproteinase (MMP) gene have been associated with multiple malignancies, including stomach, bladder, lung, breast, melanoma, squamous cell skin, and colorectal cancers. Genetic factors often slightly to moderately increase the risk of breast cancer.
Objective:
This study was conducted to ascertain the correlation between breast cancer patients in Bangladesh and the MMP2 rs243865 and MMP8 rs11225395 polymorphisms.
Design:
A case-control study
Methods:
The tetra-primer amplification refractory mutation system polymerase chain reaction (ARMS-PCR) was used for genotyping of 324 healthy individuals and 315 individuals with breast cancer. In addition, we performed in silico expression analysis of MMP2 and MMP8 genes using different databases, such as GEPIA, UALCAN, and GTEx.
Results:
In the case of MMP2 rs243865 polymorphism, additive model 1 (OR = 1.90, 95% CI = 1.03-3.52), dominant model (OR = 2.07, 95% CI = 1.15-3.72), overdominant model (OR = 1.83, 95% CI = 0.99-3.38), and allelic (OR = 1.93, 95% CI = 1.17-3.17) model were significantly associated with enhanced breast cancer risk. For the MMP8 rs11225395 polymorphism, none of the genetic models showed a significant association with increased breast cancer risk. The logistic regression analyses were adjusted for response status (age, BMI, and marital status). In silico analysis showed that MMP2 was more highly expressed in normal tissues, whereas MMP8 was more highly expressed in breast tissues.
Conclusion:
Our findings imply that MMP2 (rs243865) polymorphisms correlate with higher breast carcinoma risk but have no relationship with MMP8 (rs11225395) polymorphisms in the Bangladeshi female population.
Introduction
Breast lobules or glandular milk duct epithelial cells can give rise to a tumor with malignant cells, which is known as breast cancer (BC). 1 About 2,261,419 new diagnoses and 684,996 fatalities have been linked to BC in 2020. 2 In South Asian nations, the prevalence of BC is rising, and both incidence and mortality rates are rising rapidly. In Bangladesh, 13,028 new BC cases were recorded in 2020, accounting for 19% of all cancer cases in women. With an estimated 627,000 BC deaths among women in 2018, Bangladesh is estimated to account for more than 15% of all female cancer mortality in 2018. 3 Three-quarters of female cancer cases in Bangladesh are BC instances, second only to cervical cancer in terms of prevalence. However, the frequency of younger women who get BC has increased significantly in recent years, even though older women over the age of 50 are typically more frequently affected by this disease. 4
The following are the risk factors for BC: advancing age, ethnic background, history of menarche, breast characteristics, reproductive profiles, usage pattern of hormones, alcohol and cigarette consumption, diet, physical activity, and body habitus. Breast cancer risk is often increased slightly to moderately by genetic factors. 5 Recent studies have shown that genetic variations in the germline may alter the likelihood of metastasis and are responsible for variances in the likelihood of BC across individuals. However, identified genes are responsible for just 20% of occurrences of breast cancer., indicating that a large portion of the variation related to the likelihood and apparition of breast carcinoma is caused by numerous unidentified genes, many of which have only modest individual effects.6,7
During routine remodeling and repair processes, MMPs, a crucial class of zinc-based enzymes, hydrolyze fibronectin, interstitial collagen, basement membrane collagen, and other proteoglycans. Six categories of MMPs were identified according to their homology and substrate selectivity. Recent research has revealed that MMPs have an impact on both apoptosis and the enlargement of tumor cells., as well as the immune system of the host against cancer, and thus play a role in tumorigenesis. 8 The extracellular matrix is influenced by MMPs in a way that affects cell growth, MMP cleavage efficiency, and the functionality of growth factors, cytokines, and chemokines. 9 According to several studies, connections have been found between MMP gene variations and a range of malignancies, such as colorectal carcinoma, bladder, stomach, pulmonary, melanoma, breast, and squamous cell skin carcinoma.10 -17
Gelatinase A or type IV collagenase is another name for MMP2, and it is located on chromosome 16q13–q21. 18 This gene consist of 13 exons and 12 introns. 19 A 72 kDa protein is generated during MMP2 gene translation and This zinc-dependent molecule is capable of disrupting extracellular matrix ingredients. 18 MMP2 has been shown to regulate neoplasm cell proliferation, differentiation, invasion, and metastasis in addition to regulating malignancy and immune system monitoring. 20 Numerous tumors types have been found to overexpress MMP2, where it stimulates the growth, mobility, and propagation of malignant tumor cells.21 -24 However, downregulating MMP2 expression can increase tumor cell apoptosis while lowering the development of clones, metastasis, and proliferation of cancerous cells.25 -27 Despite being present in tumors and stromal cells surrounding the tumor, MMP2 was infrequently found in benign breast lesions and normal breast tissue in patients.28 -32 Additionally, it was discovered that breast cancer patients’ circulating MMP2 instances were noticeably greater than the controls. 33 MMP2 contains a polymorphism rs243865 within its promoter region, and this SNP (C > T) occurs at position -1306, which is upstream of the first exon of the MMP2 gene 34 Few research has looked at the connection of MMP2 genetic variation to the susceptibility of BC and they have all focused on a single polymorphism (rs243865).35 -37 Case-control studies conducted in women from Iraq, Mexico, and Saudi demonstrated that a greater risk of breast cancer was linked to MMP2 rs243865 polymorphism.35,38,39 However, case-control studies on Tunisia and Han Chinese women showed that MMP2 rs243865 polymorphism linked to a significantly lower incidence of breast cancer.37,40
MMP8 (also known as collagenase-2 and neutrophil collagenase) once thought that only neutrophils could produce, however, it has now been discovered that various kinds of cells release it. 41 The MMP8 of a human gene contains 12 exons and one of the 9 MMP genes noticed on 11q22.2-q22.3 of chromosome. 42 MMP8 expression has been detected in both BC cells and squamous cell carcinomas.43,44 The promoter region may have a single-nucleotide C to T mutation due to the polymorphism of MMP8 rs11225395 (C-799T). 45 When it comes to promoter activity and protein expression, this polymorphism’s T allele seems to be substantially greater than the C allele.46,47 Studies on prognosis have shown that the T allele is connected to improved overall survival of patients with the initial stages of BC. 48 There is conflicting evidence for the MMP8 gene’s role as an oncogene, even though numerous prior studies have reported that it inhibits carcinogenesis and metastasis. 49 Clarifying the role of the MMP8 gene and the possible prognostic impact of the rs11225395 polymorphism is therefore crucial. Numerous studies have linked polymorphisms in MMP8 to an increased likelihood of BC. For instance, recent research in the Chinese Han community discovered that some haplotypes of various polymorphisms of MMP8 gene including variant rs11225395 enhanced the breast cancerous risk. 50 According to a Belgian study, differences in the MMP8 gene may impact the prognosis of BC and provide evidence for the theory that MMP8 inhibits the spread of cancer. 47 The MMP8 -799C/T (rs11225395) polymorphism may have an indirect influence on the risk factors for BC in Taiwan. 51 A meta-analysis reveals that the MMP8 rs11225395 polymorphism may be connected to a higher risk of cancer in non-Asian populations. 52 However, a Lithuanian case-control study could not discover any correlation between the MMP8 SNP rs11225395 and BC risk. 53
We selected to investigate MMP2 and MMP8 together because, although they belong to different metalloproteinase subfamilies—gelatinases and collagenases—they perform complementary roles in extracellular matrix remodeling. MMP2 is primarily involved in basement membrane degradation, driving tumor cell invasion, while MMP8 contributes to cancer behavior through collagen breakdown and its interactions with immune pathways. By examining variants in both genes, we aimed to capture a more complete picture of how distinct matrix-related processes may shape breast cancer risk.
Based on these findings, MMP2 (rs243865) and MMP8 (rs11225395) may contribute to BC in Bangladeshi individuals. No genetic linkage studies have been conducted on Bangladeshi women to assess and confirm the link between MMP2 (rs243865), MMP8 (rs11225395), and BC.
Methods and Materials
Design of study and participants
Blood samples were collected from 315 BC individuals recruited as cases and 324 healthy participants conscripted from various locations in Chittagong and Dhaka, Bangladesh. The case and control groups in the study ranged in age from 25 to 70 years and from 21 to 74 years, respectively. Each participant was made aware of the intention of the study prior to the collection of blood samples and written consent was obtained from them and/or their families. This study excluded healthy individuals under the age of 18 who were unable to supply the required data. This study also excluded participants with chronic conditions like kidney, liver, or lung disease. We only included Bangladeshi breast cancer patients identified by physicians. The Noakhali Science and Technology University ethics committee approved this project to address ethical concerns (ethics code: NSTU/SCI/EC/2023/058).
Blood collection, isolation of DNA, and quantification of genomic DNA
Each case-control study participant received approximately 3 mL of blood via a plastic syringe which was in 3 mL in amount, which was quickly added to a plastic tube that contained Na2-EDTA. Before DNA was extracted, blood samples were collected and kept at −80°C. DNA isolation was carried out by using the FavorPrep genomic DNA purification kit and the DNA extraction mini kit in accordance with the protocol book that came with the kit (TIANamp Blood DNA Kit, TIANGEN). Using a Micro-volume Spectrophotometer (Genova Nano, Jenway), 2 microliters of the sample were exposed to an absorbance ratio between 260 and 280 nm to determine the concentration and purity of the extracted DNA.
Primer configuration
Two SNPs (rs243865 and rs11225395) were effectively genotyped using the tetra primers ARMS-PCR technique. The tetra primers ARMS-PCR design and method validation process accounted for a variety of factors. Four primers were applied to each gene to amplify the target allele. Primer sequences have been designed by an online software named Primer blast described in Table S1.
Method validation and genotyping
The concentrations of primers, salt, and buffer were used to confirm and choose the annealing temperatures. The PCR premix was made by combining the EmeraldAmp GoTaq G2 green PCR master Mix, DD water, MgCl2, and primers in the appropriate proportions. To create 120 µL of PCR master mix solution (12 samples; 10 µL/reaction), each primer had a volume of 3 µL. The PCR was then started by combining the DNA sample with the premix (10 µL) at the proper concentration. First, gradient PCR was carried out between 54°C and 65°C using different primers and Magnesium chloride quantities to validate the process and parameter selection. Based on these results, the annealing temperatures for rs243865 and rs11225395 were chosen as 58°C for 45 seconds and 61°C for 1 minute, respectively. For rs243865, the PCR sample was heated at 94°C for 5 minutes to denature the DNA and for 1 minute at the same temperature to initiate the PCR. After the annealing stage, the elongation stage was carried out at 72°C for 45 seconds. For rs11225395, the DNA denaturation, PCR initiation and elongation were run for 95°C for 5 minutes, 95°C for 1 minute and 72°C for 1 minute (Table S2). After the tetra-primer ARMS-PCR was completed, amplification bands of various sizes appeared.
For the rs243865 SNP, we found fragments of 211, 379, and 542 base pairs, whereas for the rs11225395 SNP, we observed fragments of 189, 283, and 416 base pairs. The products were examined using 1% agarose gel electrophoresis after the completion of PCR. DNA bands of the genotypes were observed using ethidium bromide (EtBr). Conditions applied to amplify rs243865 and rs11225395 and the length of the PCR product with their respective genotypes under these SNPs are presented in Table S2.
Analysis of gene expression through in-silico study
For in-silico expression analysis, we used 3 publicly available databases. we used 3 publicly available databases. GEPIA (http://gepia.cancer-pku.cn/, accessed 24/09/2024) integrates RNA-seq data from TCGA and GTEx to compare transcriptional levels of MMP2 and MMP8 between breast tumor and normal tissues. UALCAN (http://ualcan.path.uab.edu/analysis.html/, accessed 24/09/2024) provides subgroup analyses of TCGA data based on clinicopathological parameters (e.g., sample type, patient age, cancer stage, race, menopausal status, nodal metastasis, and tumor grade) and reports statistical significance (P values) between comparison groups. GTEx (http://www.gtexportal.org/, accessed 24/09/2024) offers genotype and expression data from normal human tissues, which we used to perform expression quantitative trait loci (eQTL) analysis of MMP2 and MMP8.
Statistical calculation
To determine distributional differences between the frequencies of the cases and controls, the Hardy–Weinberg Equilibrium (HWE) test status was evaluated by using the chi-square test (χ2). Binary logistic regression analysis was used in MedCalc (v19.0.7) to compute the odds ratio (OR) and 95% confidence interval (CI). Software called SPSS 17.0 was used to conduct additional statistical analysis. The statistical significance level, or P value, was established at P < .05.
Results
Distribution of demographic variables
Table 1 provides an analysis of the demographics of both the cases and controls. Women under 45 years of age accounted for 54.91% of BC patients, while the highest proportion of healthy individuals was found in the same age group (57.09%). In contrast, between 22.86% and 25.93% of female patients and controls fell within the 45- to 60-year-old category. The patients had a mean age of 45.65, while the controls averaged 40.37. Average BMI (kg/m2) of patients was 29.39 and for controls it was 22.30. Most patients and controls were married (88.89% and 91.98 %, respectively). The majority of the patients had duct cell carcinoma, which was invasive, comprising 60.95%, while 25.08% of the patients had infiltrating ductal carcinoma cells; however, the remainder of the patients had atypical ductal hyperplasia (1.59%), duct cell carcinoma (3.17%), intraductal carcinoma (1.90%), metastatic duct cell carcinoma (2.86%), and triple-negative breast cancer (2.86%).
Distribution of demographic variables of breast cancer patients and controls.

Distribution of genotype data between patients and controls
The distribution of genotypes for both MMP2 rs243865 and MMP8 rs11225395 polymorphisms based on HWE is presented in Table 2. The genotype distribution for both genetic variants for both cases and controls was reported to be compatible with HWE (P > .05). The CC genotype was detected in 75.23% of BC patients and 87.96% of healthy individuals for rs243865, whereas 21.5% of cancer patients and 10.8% of controls had the CT genotype. For the TT genotype, 3.17% were patients and 1.23 % were controls. The observed genotypes for MMP2 rs243865 in both patients and controls (Table 2) revealed that the distribution of genotypes was consistent with the HWE value. In these cases, the value was χ2 = 3.625, P = .860, and in the controls, the value was χ2 = 5.325, P = .934. PCR amplification bands of the CC, CT, and TT genotypes for SNP rs243865 and product sizes are shown in Figure 1.
Genotype data distribution and Hardy–Weinberg equilibrium (HWE) test status.


PCR amplification bands of CC, CT & TT genotypes for SNP rs243865 and product sizes were 211 bp for the C allele, 379 bp for the T allele, and 542 bp for the control band. The first lane represents a 100 bp DNA ladder; lanes 1, 3, 4,6, 7, and 10 indicate CC genotype; lane 9 indicates CT genotype; lanes 11 and 12 indicate TT genotype.
In contrast, when looking at the rs11225395 SNP genotypes, 75.87% of patients and 84.88% of controls had the GG genotype. AA and AG genotype frequencies were 20.63 and 3.49% in patients and 13.27 and 1.85% in controls, respectively. No substantial variations in genotype distribution were found between patients and controls. Our observed genotypes for MMP8 rs11225395 in the patient as well as in healthy people depicted in Table 2 revealed that the distribution of genotypes was consistent with the HWE value. In these cases, the value was χ2 = 5.592, P value = 0.860, and in the controls, the value was χ2 = 6.880, P = .920. PCR amplification bands of the GG, AG, and AA genotypes for SNP rs11225395 and product sizes are presented in Figure 2.

PCR amplification bands of GG, AG & AA genotypes for SNP rs11225395 and product sizes were 189 bp for the G allele, 283 bp for the A allele, and 416 bp for the control band. The first lane represents a 100 bp DNA ladder; lanes 2, 5, 8, 9, 10, and 11 indicate GG genotype; lanes 1, 6, 7, and 12 indicate GA genotype; lanes 3 and 4 indicate AA genotype.
Genotypic association of rs243865 and rs11225395 with breast cancer
The relationship between the MMP2 rs243865 polymorphism and BC risk with adjusting response status (adjusted by Age + BMI + Marital Status) is shown in Table 3. In the case of rs243865, women with the CT genotype had a 1.90 times greater probability of developing BC, while those bearing the TT genotype had a 3.90 times elevated risk, CT versus CC: OR = 1.90, 95% CI = 1.03 to 3.52, P = .035; TT versus CC: OR = 3.90, 95% CI = 0.83 to 18.25, P = .072, respectively; additive model 1 showed statistical significance, but additive model 2 demonstrated a statistically non-significant association. It was discovered that having a dominant model, recessive model, overdominant model, or allele model was linked with an elevated breast cancerous risk (CT + TT vs CC: OR = 2.07, 95% CI = 1.15 to 3.72, P = .015; TT vs CC + CT: OR = 3.51, 95% CI = 0.75 to 16.39, P = .110; CT vs CC + TT: OR = 1.83, 95% CI = 0.99 to 3.38, P = .050; T vs C: OR = 1.93, 95% CI = 1.17 to 3.17, P = .010, respectively). The allele, dominant, and overdominant models showed statistically significant associations; however, the recessive model showed no statistically significant associations. Frequencies of the foremost allele (C) in patients (86.03%) and controls (93.36%). However, the minor allele (T) was found in 13.96% and 6.63% of the patients and controls, respectively.
Quantitative risk analysis of rs243865 and rs11225395 polymorphism on breast cancer patients through genotypic models with response status (adjusted by age + BMI + marital status).

In the case of the test of association, bold indicates statistically significant (P < .05).
Table 3 illustrates the association between MMP8 rs11225395 polymorphism and BC risk with adjusting for response Status (adjusted by age + BMI + marital status). This table recommended that additive model 1 revealed that BC was 1.12 times as high in the AG mutant heterozygote genotype than in the GG normal homozygote genotype (AG vs GG: OR = 1.12, 95% CI = 0.60 to 2.11, P = .160). In contrast to the normal homozygote genotype of GG, the AA mutant homozygote also depicted a 3.51 times elevated likelihood of BC progression, although the correlation was not statistically relevant (GG vs AA: OR = 3.51, 95% CI = 0.95 to 12.94, P = .157). The combination of the AG and AA genotypes in the dominant model showed a 1.37-fold increased risk of BC progression compared with the GG genotype (AG + AA vs GG: OR = 1.37, 95% CI = 0.77 to 2.43, P = .290). The AA genotype in the recessive model had a 3.45-fold higher risk of BC (AA vs GG + AG); however, this difference was not statistically significant (OR = 3.45, 95% CI = 0.94 to 512.69, P = .058). The risk was found to be 1.07 times higher in the overdominant model (AG vs GG + AA), and the relation was not statistically significant (OR = 1.07, 95% CI = 0.57 to 2.00, P = .840). The minor allele (A) exhibited a 1.44 times greater risk with BC progression in the allele model when compared with the major allele G, and the relation was also statistically nonsignificant (A vs G: OR = 1.44, 95% CI = 0.90 to 2.29, P = .120).
In-silico gene expression of MMP2 and MMP8 gene
Box plots in Figure 3 indicated that MMP2 expression is significantly elevated in normal tissues compared with breast cancerous tissues, whereas MMP8 expression is much greater in breast cancerous tissues than in normal tissues. Figure 4 illustrates that elevated expression levels were observed in normal tissue samples (Figure 4I), and individuals aged 41–60 years (Figure 4II) and Stage 1 (Figure 4III), White patients (Figure 4IX), and perimenopausal women (Figure 4X), and N3 status (Figure 4XI), respectively. Similarly, Figure 5 shows box plots indicating elevated expression levels in tumor samples (Figure 5I), and individuals aged 61–80 years (Figure 5II), and stages 1 and 2 (Figure 5III), White patients (Figure 5IX), and perimenopausal patients (Figure 5X), and patients with N0 status (Figure 5XI) separately.

Expression of MMP2 and MMP3 gene in breast cancerous tissues versus normal tissues (data retrieved from GEPIA database).

Expression levels of MMP2 stratified by (I) sample type (P > .05), (II) patient age [significant difference between 41 and 60 years versus 61–80 years, P < .05], (III) cancer stage [significant differences for Normal vs Stage 2, Stage 1 vs Stage 2, and Stage 2 vs Stage 3, all P < .05], (IV) patient race [significant differences for Normal vs African American and White vs African American, P < .05], (V) menopause status [significant difference between peri- and post-menopause, P < .05], and (VI) nodal metastasis status (all comparisons, P > .05). Data retrieved from the UALCAN database.
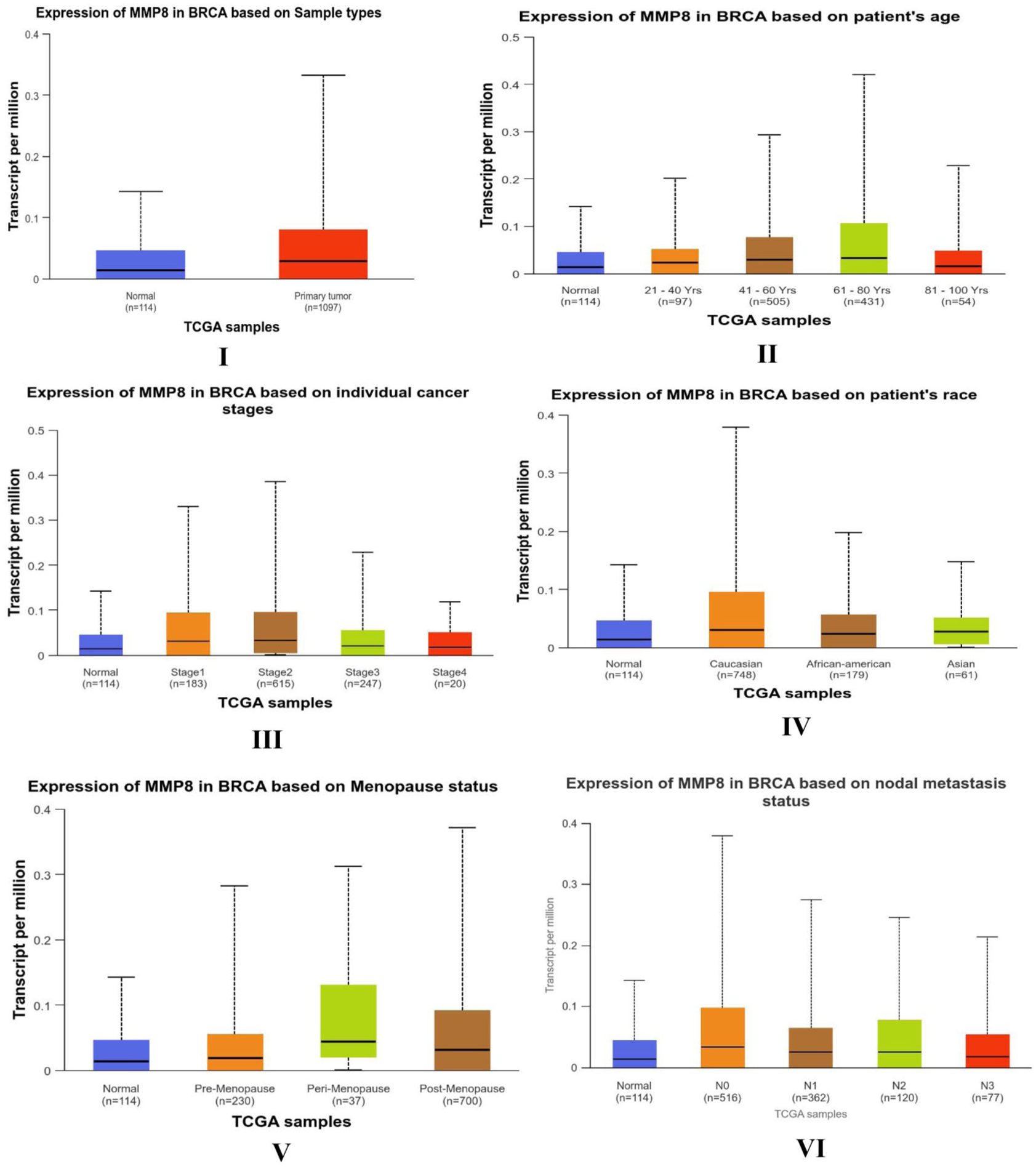
Expression levels of MMP8 stratified by (I) sample type (P < .05), (II) patient age [significant differences for Normal vs 41–60 years, Normal vs 61–80 years, 41–60 years vs 81–100 years, and 61–80 years vs 81–100 years, all P < .05], (III) cancer stage [significant differences for Normal vs Stage 1, Normal vs Stage 2, Normal vs Stage 3, Stage 1 vs Stage 4, Stage 2 vs Stage 4, and Stage 3 vs Stage 4, all P < .05], (IV) patient race [significant differences for Normal vs White, White vs African American, and White vs Asian, P < .05], (V) menopause status [significant differences for Normal vs pre-menopause and Normal vs post-menopause, P < .05], and (VI) nodal metastasis status [significant differences for Normal vs N0, Normal vs N1, and Normal vs N2, all P < .05]. Data and P values were obtained from the UALCAN database.
The genotype-based expression levels for the MMP2 rs243865 (P = .67) and MMP8 rs11225395 (P = .65) polymorphisms in breast mammary tissues did not differ statistically significantly based on their genotypes, according to the eQTL plots. The GTEx study revealed that the mutant allele (T) of MMP2 rs243865 reduces MMP2 mRNA expression, whereas the mutant allele (A) of MMP8 rs11225395 also correlates with reduced MMP8 mRNA expression in breast mammary tissues (Figure 6).

Genotype-based mRNA expression of MMP2 rs243865 and MMP8 rs11225395 variants. The values enclosed in the brackets indicate the frequency of carriers with different genotypes.
Discussion
With an estimated 23 lakh new cases in 2020, BC surpassed lung cancer as the primary cause of cancer incidence among women globally. 54 BC ranks second place among the most persistent cancers and is the most prevalent malignancy in women. 55 Affluent, poor, and middle-class countries all have higher cancer burden due to population increase, geriatric population, lifestyle changes, and environmental exposure. 56 Genetic factors like familial records or gene mutations, are major contributing factor to developing breast carcinoma. 57
The MMP family consists of over 25 members, which are categorized based on their structural conformation and reaction molecules. 58 Upregulation of MMPs has been found in various types of cancer. Therefore, they have been identified as effective predictive indicators in different stage of cancer. 59 They contribute to the breast cancer by modulating immune responses. By cleaving a broad range of cellular materials, including extracellular matrix constituents, adhesion molecules, tyrosine kinases, proteinases, and growth factor components (precursor, binding protein), MMPs contribute to an increased cancer risk. 9 MMP2, also called gelatinase A, is an enzyme connected to the growth of cells, inflammation, angiogenesis, invasion of tumors, metastasis, and the carcinogenesis of BC. 60 A binding site in the MMP2 gene’s promoter region is compromised by a C > T transition at nucleotide -1306 for the transcription factor Sp1 (Specificity Protein 1). The C > T polymorphism (rs243865) diminishes the binding affinity of Sp1, making it a significant candidate for association studies across various pathologies linked to MMP2, including metastasis and tumor invasion. 61 In addition, MMP8 has also been identified as an associated biomarker for BC development. 51 MMP8, also termed as a neutrophil collagenase due to its production by neutrophils, has been demonstrated to impact BC cells, especially when it pertains to metastatic behavior. 47 Plasma concentrations of MMP8 were inversely related to the probability of distant metastasis through protection against lymph node metastasis. 62 In a 2017 study, elevated expression of MMP8 was found in the serum of BC patients. 63
The results of this research suggest a relationship between MMP2 rs243865 polymorphism and a raised likelihood of breast carcinoma. Our research also demonstrated similar outcomes, women carrying CT and TT genotypes had 1.90 times and 3.90 times substantially increased chance (P = .035 and P = .072, respectively) of originating breast carcinoma in comparison to the CC genotype. Additionally, we discovered that a higher risk of BC was linked with the minor T allele for the rs243865 polymorphism. The present study also revealed that the MMP8 rs11225395 polymorphism is not related to the extended possibility of developing BC. In our analysis, women with the GG genotype had a considerably higher chance of getting BC than those with the AG and AA genotypes, by 1.12 and 3.51 times, respectively (P = .160 and P = .157). Our allelic model proved that individuals carrying the A allele of the MMP8 gene rs11225395 polymorphism are not significantly prone to an enhanced susceptibility to progressing breast carcinoma (P = .120). Additionally, our research revealed that an elevated risk was linked to both the dominant model (P = .290) and the recessive model (P = .058), non-significantly.
Similar to our findings, MMP2 rs243865 polymorphism was substantially correlated with an elevated breast carcinoma risk in the Saudi population. 39 According to a prior study, homozygotes for the rare or minor allele rs243865 (-1306C/T) were more likely to develop BC. 64 Mexican women with BC had a considerably higher frequency of the MMP2 -1306 CC genotype compared with controls, 35 but in our study, the MMP2 -1306 CC genotype was significantly less common in BC patients than in controls in Bangladeshi women. Furthermore, we found an increased risk for the allele model (P = .010), dominant model (P = .015), and overdominant model (P = .050), all of which met the statistical significance threshold, except for the recessive model. Thus, our findings reconcile with several research projects involving other ethnic groups. In case of MMP8 rs11225395, the association was found with a higher risk of malignant melanoma, according to a Polish study, with OR of 1.69 (P = .040) for the A/A genotype and 1.49 (P = .035) for the A/G genotype compared with the G/G genotype. 64 Polish women carrying the rs11225395 polymorphism of the MMP8 gene was linked to an increased risk of malignant melanoma (OR = 1.54, P = .017). 64 However, in Belgium, there was a correlation between the MMP8 gene variant rs11225395 T allele and decreased metastasis of BC (OR = 0.7, P = .02). 47 A correlation between rs11225395 of MMP8 and breast cancer was also discovered in a study in the Taiwanese population. 51
In addition, The GEPIA and UALCAN databases indicate that normal tissues have substantially higher levels of MMP2 in comparison to breast malignant tissues, and breast malignant tissues have significantly higher MMP8 mRNA levels than normal tissues.65,66 Our study found that in silico expression is elevated in normal tissue among individuals aged 41 to 60 years, at stage 1, White descent, perimenopausal status, and N3 category for MMP2. Conversely, for breast tumors, expression is heightened in individuals aged 61 to 80 years, at stages 1 and 2, White descent, perimenopausal status, and N0 category for MMP8. 65 We completed gene expression analysis via eQTL in the GTEx portal, which indicated that the mutant alleles of both MMP2 rs243865 and MMP8 rs11225395 polymorphisms result in reduced mRNA expression in breast mammary tissues. 67
As a TARMS-PCR technique was designed and validated for the first time in Bangladesh, instead of conventional PCR-RFLP methods for detecting these SNP, some constraints were identified in our current study, such as the selection of identified SNPs from a public database, which did not include novel SNPs, and small sample size. In addition, complete and thorough sociodemographic, clinicopathological, and gene-environment interaction data are lacking. Furthermore, the relatively small sample size did not represent the whole scenario of all the BC patients in Bangladesh. Besides, clinicopathological features were also insufficient to describe the actual scenario of BC patients and their linkage with the genotype distribution of the respective gene. We intend to extend this study to a larger population in the future to validate these findings. Research with a large sample size may reveal more significant facts that help us to detect the vital genetic events responsible for developing breast carcinoma. This study identified these SNPs as contributing factors to BC development in Bangladeshi women. We recommend conducting functional analysis at the molecular and cellular levels to check whether these SNPs influence the higher or lower activity of these enzymes. In addition, it is essential to investigate whether variations in enzyme levels relate to breast cancer in the Bangladeshi population. Such studies could help to determine whether these SNPs can serve as biomarkers for breast cancer in Bangladeshi women. Personalized medicine approaches may be developed based on enzyme activity levels. Furthermore, future studies should explore how MMP2 and MMP8 polymorphisms affect breast cancer susceptibility in various ethnic groups to validate their broader involvement.
Conclusion
Our findings revealed that the T allele of MMP2 rs243865 SNP is notably linked with the chance of developing BC, but the A allele of MMP8 rs11225395 is not significantly linked with the risk of BC in Bangladeshi women. This study clarifies that MMP2 rs243865 polymorphisms are strongly correlated with elevated breast cancer risk, but MMP8 rs11225395 polymorphisms are not strongly correlated with breast cancer in Bangladeshi females. Future research attempting to ascertain the influence of these genetic variations on medication responses in patients with breast cancer would greatly benefit from our findings. They may also aid in the creation of personalized risk assessment techniques and possible treatment plans for Bangladeshi women with breast cancer.
Supplemental Material
sj-docx-1-bcb-10.1177_11782234251392714 – Supplemental material for MMP2 rs243865 and MMP8 rs11225395 Gene Polymorphisms Are Correlated With Breast Cancer Risk in Bangladeshi Women: Evidence From a Case-Control Study
Supplemental material, sj-docx-1-bcb-10.1177_11782234251392714 for MMP2 rs243865 and MMP8 rs11225395 Gene Polymorphisms Are Correlated With Breast Cancer Risk in Bangladeshi Women: Evidence From a Case-Control Study by Anamika Datta, Sayma Binte Mohammad, Khokon Kanti Bhowmik, Md Abdul Barek, Arafat Miah, Md. Shafiul Hossen, Md. Saddam Hussain and Mohammad Safiqul Islam in Breast Cancer: Basic and Clinical Research
Footnotes
Acknowledgements
This case-control study was conducted at the Laboratory of Pharmacogenomics and Molecular Biology, Department of Pharmacy, Noakhali Science and Technology University. So, we would like to acknowledge them for their help during the study. We thank the authority of the National Institute of Cancer Research and Hospital, Dhaka, Bangladesh, for their cooperation and help during the collection of blood samples.
Ethical considerations
This study was conducted in a manner compliant with the tenets of the Declaration of Helsinki. The Ethics Committee of Noakhali Science and Technology University gave its permission to our investigation (ethics code: NSTU/SCI/EC/2023/058).
Author contributions
Anamika Datta: Conceptualization, Data curation, Investigation, Methodology, Resources, Validation, Visualization, Writing—original draft.
Sayma Binte Mohammad: Conceptualization, Data curation, Investigation, Methodology, Resources, Validation, Writing—original draft.
Khokon Kanti Bhowmik: Formal analysis, Validation, Writing—review & editing.
Md Abdul Barek: Data curation, Validation, Writing—review & editing.
Arafat Miah: Formal analysis, Validation.
Md. Shafiul Hossen: Formal analysis, Validation.
Md. Saddam Hussain: Conceptualization, Software, Supervision, Writing—review & editing.
Mohammad Safiqul Islam: Conceptualization, Writing—review & editing
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was partly funded by the Research Cell of Noakhali Science and Technology University.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability statement
All the data presented here have been added to the manuscript. Additional data will be available from the corresponding author on a valid request.
Transparency statement
The corresponding author affirms that this manuscript is an honest, accurate, and transparent account of the study being reported; and that no important aspects of the study have been omitted.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
