Abstract
WNT proteins constitute a highly conserved family of signaling molecules that play an important role in regulating embryonic development and maintaining adult tissue homeostasis. Their diverse biological functions are mediated through multiple signaling pathways, including both canonical β-catenin-dependent and several non-canonical mechanisms. The regulatory activity of WNT proteins is closely linked to their unique structural organization, the presence of N-terminal signal peptides, and posttranslational modifications. In this study, in silico methods were used to analyze the structural features of WNT proteins. Specifically, the isoelectric points, GRAVY scores, aliphatic indices, and instability indices were determined, and correlation analysis was performed to examine relationships between the latter three parameters. In addition, the characteristics of N-terminal signal peptides in WNT family proteins were investigated, with a particular focus on the bioinformatic prediction of N-terminal peptide lengths in the WNT2B protein isoforms. Furthermore, in silico modeling and molecular dynamics simulations were employed to study the tertiary structure of WNT2B and to assess the significance of O-acylation at serine for the behavior of the mature protein in an aqueous environment. Thus, using computational approaches, new data were obtained on the structural features and dynamic properties of this group of regulatory proteins.
Introduction
WNT proteins are a family of secreted, lipid-modified glycoproteins that regulate ontogenesis and morphogenesis,1,2 participate in stem cell differentiation processes, 3 and contribute to oncogenesis. 4 These proteins function as paracrine factors: under canonical WNT/β-catenin signaling conditions, they are secreted into the extracellular space and diffuse to neighboring cells, where they bind to transmembrane receptor proteins of the Frizzled (FZD) family and the co-receptors LRP5/6. This interaction initiates an intracellular signaling cascade that stabilizes β-catenin, leading to its accumulation and subsequent transcriptional activation of target genes. 5 WNT proteins are also involved in non-canonical signaling pathways, such as the WNT/planar cell polarity (PCP) pathway and the WNT/calcium (Ca2+) pathway. 6 In general, WNTs are known to interact with more than 15 receptors and co-receptors across 7 protein families, 7 highlighting their role as essential bioregulators.
WNT proteins are characterized by a distinct 2-domain structure, in which the N-terminal domain (NTD) resembles a “thumb,” while the C-terminal domain (CTD) is analogous to an “index finger.” 8 Following translation, each WNT protein possesses a signal peptide that directs the synthesized protein into the lumen of the endoplasmic reticulum. 9 - 11 However, due to significant sequence variability among the N-terminal peptides of different WNTs and limited data on the cleavage of these signal peptides during the maturation of WNT proteins, uncertainties remain regarding the shared properties of signal peptides that ensure their functional unity.
Posttranslational modifications also play a critical role in WNT function. Before secretion into the extracellular space, WNT proteins undergo O-acylation with fatty acids in the endoplasmic reticulum and glycosylation in the Golgi complex, to which they are transported in a complex with Wntless (WLS). 12 Unlike glycosylation, acylation is required for the apical secretion of WNT proteins. 8 For instance, in vitro studies have shown that non-glycosylated WNT3A and WNT5A can still trigger intracellular signaling when added to cells. 13 O-acylation renders WNTs hydrophobic, raising questions about the mechanisms of their transport through the extracellular space to target cells. 14 Both the distinct 2-domain structure of WNT proteins and the presence of an acylated fatty acid chain are important for their signaling function, likely influencing their ability to interact with cellular receptors. However, structural data at the tertiary level are available only for a limited number of WNT family members. Moreover, the processes of signal peptide processing, the role of posttranslational modifications, and the impact of hydrophobicity on WNT function require further investigation.
In this study, in silico analysis methods were employed to investigate WNT proteins, aiming to enhance the understanding of their unique structural and functional characteristics.
Methods
The amino acid sequence data of 19 WNT family proteins in Homo sapiens and their corresponding orthologs in other species were obtained from Ensembl 15 and UniProt. 16 For all orthologous protein sequences in the dataset, the aliphatic index, 17 instability index, 18 and grand average of hydropathicity (GRAVY) 19 were calculated, while the isoelectric points20,21 were also determined for the Homo sapiens protein sequences.
The aliphatic index of a protein was calculated using the formula: 17
where X(Ala), X(Val), X(Ile), X(Leu) represent the mole percentages of alanine, valine, isoleucine, and leucine in the protein sequence, respectively; a and b, equal to 2.9 and 3.9, respectively, reflect the relative volumes of these amino acid side chains compared with that of alanine.
The instability index was calculated using the formula: 18
where L is the length of the protein sequence, xiyi + 1 represents the dipeptide starting at position i, and DIWL is the instability weight value of one of the 400 possible dipeptides, as described in Guruprasad et al. 18
The GRAVY was determined as the sum of hydropathy values for each amino acid residue, according to Kyte and Doolittle, 19 divided by the length of the amino acid sequence.
To calculate the aliphatic index, instability index, and GRAVY for Homo sapiens WNT proteins, the ProtParam tool on the Expasy server22,23 was used. An automated R script was employed to calculate these parameters for orthologous proteins in other species. 24 Pairwise correlations of the aliphatic index, instability index, and GRAVY scores were determined using the Spearman correlation test with a significance level of P < 0.05.
Isoelectric points of Homo sapiens WNT proteins were calculated using amino acid pK values described in Bjellqvist et al20,21 via the ProtParam tool on the Expasy server.22,23
Signal peptide lengths of Homo sapiens WNT proteins were obtained from UniProt. 16 For WNT2B protein isoforms (UniProt: Q93097), where signal peptide lengths were not available in UniProt, 3 predictive tools—PrediSi, 25 SignalP-5.0, 26 and TargetP-2.0 27 —were used, and their results were compared.
Pairwise alignment of WNT2B isoforms was performed using the Needleman-Wunsch algorithm 28 implemented in EMBOSS Needle29,30 with the BLOSUM62 substitution matrix. 31
The WNT2B protein was further used as a model protein for studying tertiary structure of WNT family proteins. Five in silico tools were employed for modeling: SWISS-MODEL Server, 32 based on homology modeling, used structure 6AHY 33 from the RCSB PDB 34 as a template, while AlphaFold3, 35 RoseTTAFold,36,37 D-I-TASSER,38,39 and DMPfold v2 40 , 41 used deep-learning algorithms.
The geometric quality of the modeled tertiary structures was evaluated using MolProbity, 42 ModFOLD9, 43 VoroMQA,44,45 and ProQ3D.46,47 ModFOLD9, VoroMQA, and ProQ3D inherently provide scores on a scale from 0 to 1, where higher values indicate better model geometry. MolProbity has a different scale for model quality assessment 42 and also reports percentiles for each score, with the 100th percentile representing the most accurate model geometry. To enable direct comparison and averaging, MolProbity percentiles were normalized to a 0–1 scale by dividing by 100. This transformation preserved the relative quality ranking and aligned the scale with those of the other tools. The normalized scores from all 4 methods were then averaged for each model, and the model with the highest average score was selected for further analysis.
In silico O-acylation of WNT2B was performed by covalently attaching palmitoleic acid to the hydroxyl group of Ser243. To achieve this, the structure of the modeled WNT2B was structurally aligned in PyMOL 48 with the crystallographic structure 6AHY, which includes a serine residue O-acylated with palmitoleic acid. Following alignment, the coordinates of the palmitoleic acid chain from 6AHY were transferred to Ser243 in WNT2B, and a non-standard residue representing the O-palmitoleoylated serine was created.
Molecular dynamics simulations of the mature WNT2B protein (excluding the signal sequence) were performed using GROMACS 2023.3 software49,50 with a modified CHARMM36 force field. 51 Separate simulations were conducted for the unmodified protein and the protein with Ser243 O-acylated by palmitoleic acid. To construct the topology for the O-acylated residue, the existing CHARMM36 parameters for O-acetylserine (OAS) were used as a structural basis. The OAS topology was extended to include the longer palmitoleic acid chain. Standard CHARMM36 parameters for saturated (alkane) and monounsaturated (alkene) carbon atoms were applied to define the relevant topology parameters.
Simulations were conducted in a dodecahedron box with a 2.4 nm distance from the protein to the box edge, filled with water and 150 mM NaCl. System minimization was followed by equilibration at 310 K and 1 bar in 3 stages: (1) 1 ns NVT with a V-rescale thermostat and 0.1 ps time constant for temperature coupling, (2) 1 ns NPT with a V-rescale thermostat and C-rescale barostat, using 0.1 and 2.0 ps time constants for temperature and pressure coupling, respectively, and (3) 50 ns NPT with a V-rescale thermostat and C-rescale barostat, using 1.0 and 5.0 ps time constants for temperature and pressure coupling, respectively. A 200 ns production run followed.
Trajectory analysis included the calculation of root-mean-square deviation (RMSD), root-mean-square fluctuation (RMSF), radius of gyration (Rg), and solvent-accessible surface area (SASA) for both O-acylated and non-acylated WNT2B.52,53 Backbone atoms were used for 2 trajectory fitting, followed by concatenation. Principal component analysis (PCA) and free energy landscape (FEL) analysis were performed using the C-alpha atoms on the concatenated trajectory. 54
Visualization of WNT2B structures was performed using the open-source PyMOL. 48
Results
Evaluation of stability, hydrophobicity, and isoelectric points of WNT proteins
To gain insights into the molecular characteristics of WNT proteins, the aliphatic indices, GRAVY scores, and instability indices were calculated based on their primary sequences. The dataset included 19 human WNT proteins and their orthologs in other species available in Ensembl, comprising a total of 3911 protein sequences. Distribution and relation between these indices in WNT-related proteins are presented on Figure 1, while the corresponding values for human WNT proteins are summarized in Table 1.

Distribution of aliphatic index, GRAVY score, and instability index in WNT-related proteins. (A) 3D scatterplot illustrating the relationship between the 3 indices; (B) scatterplot showing the correlation between GRAVY and the aliphatic index; (C) scatterplot showing the correlation between GRAVY and the instability index; (D) scatterplot showing the correlation between the instability index and the aliphatic index. The color gradient represents the length of amino acid sequences in WNT proteins.
Characteristics of Homo sapiens WNT proteins.

The amino acid sequence length in the dataset ranges from 93 to 1004 residues, with an average length of 362. The aliphatic index values span from 30.80 to 110.98, with an average of 67.31. A higher aliphatic index indicates a greater proportion of aliphatic amino acids (alanine, valine, leucine, isoleucine), which enhances thermal stability. Among human WNT proteins, WNT6 and WNT11 exhibit the highest thermal stability, whereas WNT2 and WNT3A have the lowest.
The GRAVY values indicate protein hydrophilicity when negative and hydrophobicity when positive. The GRAVY values in the dataset range from –0.861 to 0.676, with an average of –0.402, suggesting that WNT proteins, in the absence of modifications, are generally hydrophilic. Individual proteins with positive GRAVY values in the dataset represent outliers.
The Spearman correlation coefficient between GRAVY and the aliphatic index is 0.665 (P << .001), indicating a strong positive correlation that is statistically significant (Figure 1B). This suggests that as GRAVY increases (indicating reduced hydrophilicity), the aliphatic index also increases (indicating greater thermal stability).
The instability index values in the dataset range from 16.09 to 96.01, with an average of 47.67. According to established criteria, proteins with an instability index below 40 are considered stable, whereas those above 40 are classified as unstable. Most proteins in the dataset (3207 from 3911) fall into the latter category. Among human WNT proteins, only WNT2, WNT5A, WNT5B, WNT7A, and WNT7B have instability index values below 40, suggesting they are relatively more stable.
The Spearman correlation coefficient between GRAVY and the instability index is –0.272 (P << .001), indicating a weak negative statistically significant correlation (Figure 1C). In contrast, the correlation between the aliphatic index and the instability index is 0.665 (P << .001), representing a strong positive statistically significant relationship (Figure 1D).
The theoretical isoelectric point (pI) values calculated for the 19 human WNT proteins are presented in Table 1. The lowest pI value (7.47) is observed for WNT3, while 7 proteins have pI values between 8 and 9, and the remaining proteins have pI values exceeding 9.
Analysis of signal peptides in WNT family proteins
Table 2 presents data on the N-terminal signal peptides of WNT proteins in Homo sapiens. For the vast majority of WNTs, information on the length of the N-terminal signal peptide is available in the UniProt database. The exception is the WNT2B protein, for which the corresponding information is not available. WNT2B is represented by 2 isoforms (Q93097-1 and Q93097-2). The lengths of the N-terminal peptides for these 2 isoforms were predicted using the bioinformatic tools SignalP 5.0 (Figure 2A), PrediSi, and TargetP-2.0 (Supplemental file).
N-terminal signal peptides of Homo sapiens WNT proteins.

Data on signal peptides in WNT2B isoforms were not available in UniProt and were predicted in this study.

Prediction of the lengths of the N-terminal signal peptide sequences in Q93097-1 (A) and Q93097-2 (B) isoforms of WNT2B using SignalP 5.0 tool.
Analysis of the amino acid sequence of the first isoform of WNT2B (Q93097-1) predicted that its N-terminal peptide spans residues 1 to 58. This is significantly longer than the signal peptides of all other WNT family proteins in Homo sapiens. However, identical results were obtained using 3 bioinformatic tools based on different signal region prediction algorithms.
The second isoform of WNT2B (Q93097-2) is shorter than the first, containing 372 amino acid residues compared with 391 in Q93097-1. The corresponding transcripts are formed through alternative splicing. 55 The Q93097-1 isoform is encoded by 5 exons, whereas the Q93097-2 isoform is encoded by 6 exons, with the last 4 exons being common to both isoforms. Pairwise alignment revealed 83.5% sequence identity between these isoforms. Notably, the N-terminal regions differ, but from position 62 of Q93097-1 and position 43 of Q93097-2 onward, their amino acid sequences are identical.
Given these differences between the WNT2B isoforms, their N-terminal sequences are completely distinct. Using the bioinformatic tools SignalP-5.0 (Figure 2B) and TargetP-2.0 (Supplemental file), the N-terminal peptide of the Q93097-2 isoform was predicted to span residues 1 to 21. The PrediSi tool, however, did not detect a signal motif in this isoform.
Thus, the N-terminal peptide of the Q93097-1 isoform of WNT2B is predicted to be unusually long for WNT proteins (58 amino acids), whereas the N-terminal peptide of the Q93097-2 isoform is predicted to be 21 amino acids in length, a size comparable with other WNT proteins. Consequently, the mature protein of the first isoform consists of 333 amino acid residues, while the mature protein of the second isoform contains 351 residues, making the latter actually larger.
Several common properties can be observed in the signal peptide sequences of Homo sapiens WNT proteins (Table 2). First, they are characterized by leucine-rich motifs. Second, WNT signal peptides are predominantly composed of hydrophobic amino acids. For most WNTs, the proportion of hydrophobic amino acids (I, L, V, C, A, G, M, F, Y, W, H, K, T; shown in one-letter code) in the signal peptides exceeds 75%. The hydrophobic amino acid content is slightly lower only in WNT4, WNT10A, WNT10B, and the Q93097-1 isoform of WNT2B; though even in these cases, hydrophobic residues constitute more than half of the total.
Structure modeling and molecular dynamics of WNT2B
To model the full-size tertiary structure of WNT2B (Q93097-1), several tools were used in parallel, including those based on both deep-learning algorithms (AlphaFold3, RoseTTAFold, D-I-TASSER, and DMPfold2) and homology modeling (SWISS-MODEL server). When using AlphaFold3, RoseTTAFold, and D-I-TASSER, each modeling project submitted to the server yields 5 models, whereas SWISS-MODEL server and DMPfold2 generate a single model each. The modeling results were compared by evaluating the geometry of each obtained model using the MolProbity, ModFOLD9, VoroMQA, and ProQ3D services. The corresponding scores and the mean normalized score for each model (on a scale from 0 to 1), are presented in Table 3. A visualization of the molecule with the highest mean normalized score, generated by each of the 5 services, is shown in Figure 3.
Assessment of structural geometry for different WNT2B models.
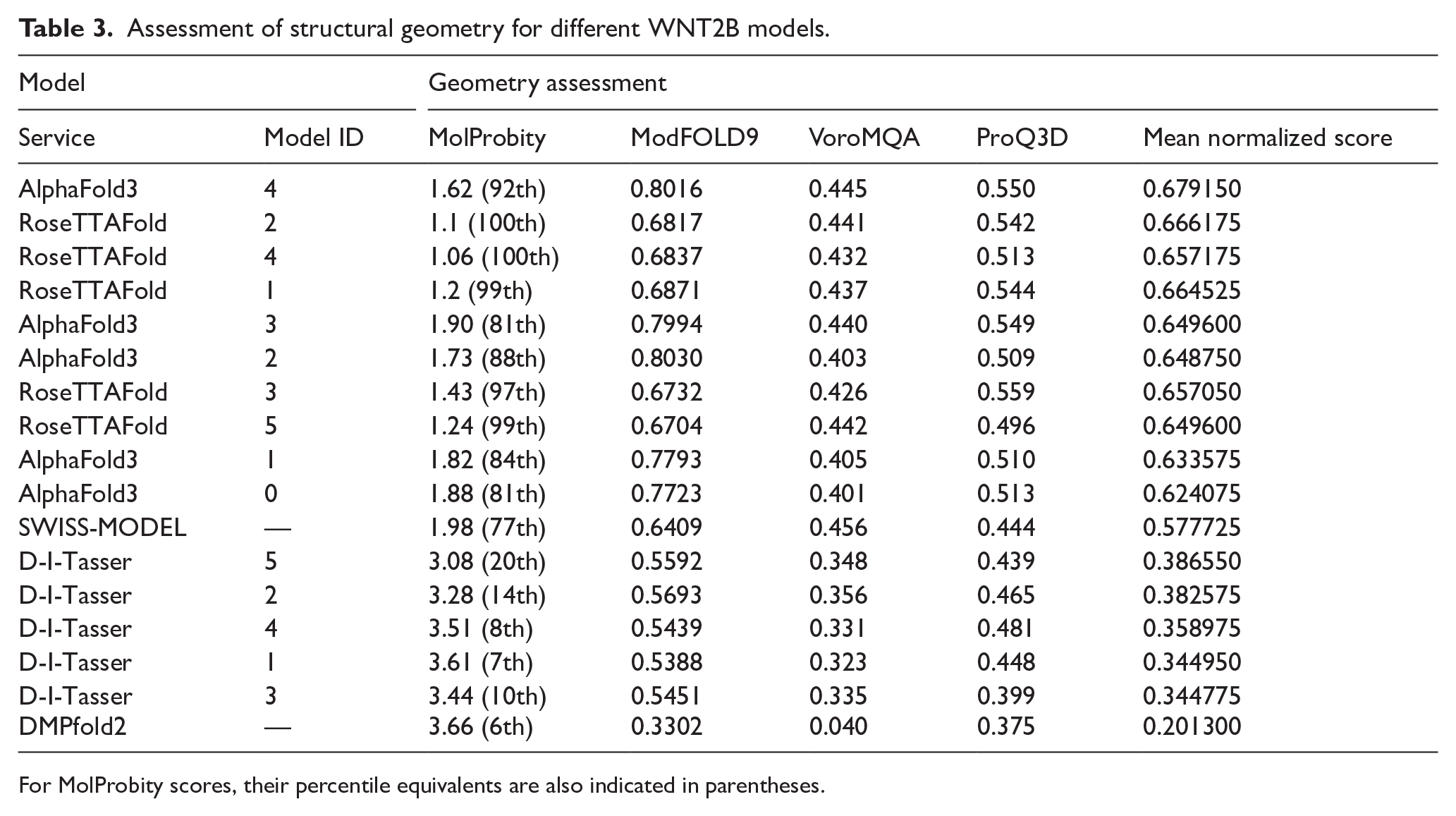
For MolProbity scores, their percentile equivalents are also indicated in parentheses.

WNT2B models generated by different tools: AlphaFold3 (A), RoseTTAFold (B), SWISS-MODEL (C) D-I-TASSER (D), DMPfold2 (E).
The most accurate geometric models of WNT2B were obtained using AlphaFold3 and RoseTTAFold (Figure 3A-B). The model generated by the homology modeling service SWISS-MODEL (Figure 3C) occupies an intermediate position, outperforming those produced by D-I-TASSER (Figure 3D) and DMPfold2 (Figure 3E), despite the latter being based on deep-learning algorithms.
Significant differences were observed between the models in the N-terminal sequences. SWISS-MODEL could not model these regions, as it was absent in the template for homology modeling (Figure 3C). AlphaFold3 predicted these regions as primarily non-globular (disordered), whereas other services generated more structured conformations but with different spatial orientations (Figure 3A). In the mature regions of the WNT2B protein, the models generated by AlphaFold3, RoseTTAFold, SWISS-MODEL, and D-I-TASSER exhibited a similar 2-domain structure characteristic of WNT proteins. However, the WNT2B model produced by DMPfold2 deviated significantly from the others, as it failed to distinguish the 2-domain structure, which includes the “thumb” in the NTD and the “index finger” in the CTD (Figure 3E).
To investigate the dynamics of different domains and structural motifs of WNT2B, as well as to assess the effect of O-acylation on its structure, molecular dynamics simulations were performed. The AlphaFold3 model with the highest mean normalized score was selected for the simulations and truncated by 58 amino acid residues to represent the mature protein.
Figure 4A presents the root-mean-square deviation (RMSD) values for WNT2B models with and without O-acylation at Ser243. During the first 145 ns, the RMSD values remain similar for both models. However, in the O-acylated WNT2B, a transition to an open extended conformation occurs due to the displacement of the CTD with the “index finger” relative to the NTD with the O-acylated “thumb.” This structural transition leads to a significant increase in the RMSD values and their fluctuations during the last 55 ns of the simulation. In contrast, the model without O-acylated Ser243 exhibits smaller RMSD values with less fluctuation throughout the simulation.

Analysis of molecular dynamics trajectories of WNT2B. (A) root-mean-square deviation (RMSD), (B) root-mean-square fluctuation (RMSF), (C) radius of gyration (Rg), (D) solvent-accessible surface area (SASA). Graphs for non-acylated WNT2B are presented in blue, and for O-acylated WNT2B in green.
The root-mean-square fluctuation (RMSF) plot (Figure 4B) reveals differences in the dynamics of residues near Ser243. Specifically, in the O-acylated WNT2B, this region displays higher mobility, likely as a direct consequence of modification with palmitoleic acid. However, more pronounced differences are observed in the “index finger” of the CTD, which exhibits increased mobility in the O-acylated WNT2B due to its transition to an open extended conformation, resulting in higher RMSF values. In contrast, in the non-acylated model, the RMSFs of the C-terminal residues outside the “index finger” are significantly larger.
Changes in the radius of gyration (Rg) during the simulation are shown in Figure 4C. These results align well with the RMSD data: during the first 145 ns, the Rg values remain similar, but toward the end of the simulation, one model transitions to a less compact state, leading to higher Rg values, whereas in the other model, the Rg values remain stable. In contrast, the solvent-accessible surface area (SASA) values (Figure 4D) and their fluctuations remain similar for both proteins throughout the simulation. This can be attributed to the overall structure of the WNT2B molecule, in which the large “thumb” and “index finger” regions remain well exposed to water molecules. As a result, relative changes in the positions of these regions do not significantly affect the accessibility of amino acid residues to solvent, and consequently, the SASA values remain similar in compact and open extended conformations.
Figure 5 shows the structural alignment of the non-acylated and O-acylated WNT2B models at 100 ns and 200 ns, illustrating the transition of one model to the open extended conformation toward the end of the simulation.

Structural alignment of WNT2B models at 100 ns (A) and 200 ns (B). Non-acylated WNT2B is shown in blue, and O-acylated WNT2B in green.
Principal component analysis (PCA) and free energy landscape (FEL) analysis (Figure 6) based on molecular dynamics simulations indicate that the model without O-acylation exhibits a single stable conformational state, in which the molecule remains for the majority of the simulation time. In contrast, the model with O-acylation displays 2 predominant states, which can be attributed to the transition of WNT2B from the compact state to an open extended conformation in the final quarter of the simulation. The primary conformation of WNT2B in both models, with and without O-acylation, is largely similar, as evidenced by the overlap in the PCA and FEL plots, particularly in regions with the lowest Gibbs energy, which correspond to the predominant stable states of the molecules.

Principal component analysis (PCA) and free energy landscape (FEL) analysis for molecular dynamics of WNT2B. (A-C) PCA plots for 2 models of WNT2B simultaneously (A), non-acylated WNT2B (B), and O-acylated WNT2B (C). (D-E) FEL plots for non-acylated (D) and O-acylated (E) WNT2B. Dots corresponding to non-acylated WNT2B are shown in blue on PCA plots, and for O-acylated WNT2B in green.
Discussion
WNT family proteins are widespread regulators of ontogenesis and are found across a broad range of animal species. A previous study 56 examined the presence of paralogous WNT family genes in 12 vertebrate and 11 invertebrate species, demonstrating that vertebrates possess a greater number of paralogous WNT genes than invertebrates (on average, 18 vs 7). 56
In humans, 19 genes are known to belong to the WNT family. In this study, a dataset of 3911 protein sequences available in Ensembl, orthologous to human WNT proteins, was analyzed. For each sequence, the GRAVY, aliphatic index, and instability index were calculated, and pairwise correlations between these parameters were determined. A strong positive correlation was observed between GRAVY and the aliphatic index (Spearman ρ = 0.665). By comparison, a previous study 57 on a different protein dataset reported a very strong correlation (ρ = 0.79), which, according to the authors, supports the notion that highly hydrophobic proteins are more likely to be thermostable.
The correlation analysis between GRAVY and the instability index revealed a weak negative relationship (ρ =–0.272), suggesting a trend toward decreased instability with increasing protein hydrophobicity. However, the most unexpected finding was a strong positive correlation between the aliphatic and instability indices (ρ = 0.665), which appears to indicate that an increase in thermal stability is associated with an increase in overall instability. Given that a previous study questioned the theoretical significance of the instability index, 58 we consider the values of instability index of WNT proteins to be a subject of debate, meriting further investigation.
A defining characteristic of WNT proteins is the presence of N-terminal signal peptides, 8 -10,33,59 which direct newly synthesized proteins into the lumen of the endoplasmic reticulum. However, the processes governing their cleavage and structural organization remain insufficiently explored. Despite the presence of publications in which these peptides are mentioned, relatively few studies have directly addressed their mechanistic or structural roles. One study has demonstrated that the signal peptides found in WNT proteins contain a hydrophobic N-terminal region that is essential for their entry into the secretory pathway. 60 Other study suggests cotranslational cleavage of the signal peptide sequence. 61 In addition, experimental evidence obtained through site-directed mutagenesis revealed that deletion of the signal sequence in WNT1 results in a loss of its transforming ability in mammary epithelial cells. 62 These findings provided the foundation for further research using genetically engineered constructs to explore WNT ligand-receptor interactions, 63 as well as the structural determinants of WNT7A responsible for its packaging into extracellular vesicles. 64
In this study, we predicted the signal peptide lengths in the WNT2B protein isoforms using bioinformatic methods. This protein is of particular interest because WNT2B gene was the first among WNT family genes to be identified as capable of generating multiple protein isoforms due to alternative splicing. 55 The 2 isoforms of WNT2B—Q93097-1 (originally WNT-2B2) and Q93097-2 (originally WNT-2B1)—exhibit distinct expression profiles. 55
Our bioinformatic predictions indicate that the Q93097-1 isoform possesses an N-terminal peptide of 58 amino acid residues, which is longer than that of other WNT proteins, including the alternative isoform Q93097-2. A review of the literature provides a likely explanation for these findings: previous studies65,66 suggest that the Q93097-1 isoform functions as a type II transmembrane protein with an extracellular WNT-type domain. Furthermore, it has been proposed 65 that the Q93097-1 isoform does not possess a conventional signal peptide like other WNTs; instead, residues 35–53 form a sequence that acts as a transmembrane domain.
The WNT2B Q93097-1 isoform is predicted to function as a transmembrane-type ligand for FZD receptors or to be released following cleavage at the C-terminal adjacent position of the transmembrane domain.65,66 This prediction aligns with our findings, which indicate that the cleavage site is located after the 58th amino acid residue. In contrast, our analysis suggests that the Q93097-2 isoform contains a signal peptide of 21 amino acid residues, and according to previous studies, it functions as a secreted protein. Furthermore, given the evolutionary conservation of the Q93097-1 isoform sequence, it is considered the predominant isoform.65,66 Thus, alternative splicing in the WNT2B gene represents a notable case within the WNT family, allowing the generation of isoforms with partially distinct sequences and different functional properties. The present study supports previously proposed ideas65,66 regarding the critical role of N-terminal sequences in determining the functional properties of WNT2B isoforms. However, since in this study the cleavage site positions in both WNT2B isoforms were predicted using computational methods, further in vitro experiments are needed to validate these findings.
The WNT2B protein under investigation belongs to the group of canonical WNT signaling activators.1,66- 68 It plays a critical role during early mammalian embryogenesis and functions as a stem cell factor for neural and retinal progenitor cells. 66 Its significance in ontogenesis also extends to its involvement in the TBX5–WNT2/WNT2B signaling axis, which provides the molecular basis for the developmental connection between lung formation and cardiac anatomical structures essential for pulmonary blood flow, a feature observed in vertebrates with lungs. 1 WNT2B has been implicated in several states, including Alzheimer disease, 69 infiltrative Crohn disease, 70 inflammatory bowel disease, 71 and oncology.72,73
In this study, the WNT2B isoform Q93097-1 was further used as a representative model to analyze several structural characteristics relevant to WNT family proteins. Specifically, the investigation focused on modeling the tertiary structure of WNT2B, including its posttranslational O-acylation at the Ser243 residue.
Molecular dynamics simulations were performed to investigate the structural fluctuations in WNT2B and to assess the impact of O-acylation at Ser243 on protein properties. Although the overall behavior of both acylated and non-acylated forms was similar, only O-acylated WNT2B exhibited a transition from a compact to an open extended conformation. This transition is attributed to the increased mobility of the “index finger” in the CTD. Similar conformational changes have been reported in earlier works which studied WNTs without O-acylation. For instance, investigations of WNT3 and WNT3A 74 demonstrated high mobility in the regions forming the “thumb” and “index finger” structures, with the latter showing the highest mobility, consistent with our findings. A study of WNT4 75 demonstrated that loops in the NTD and CTD of this protein contributed significantly to fluctuations during molecular dynamics simulation, with the loop corresponding to the “index finger” exhibiting the highest peak in the RMSF profile. In contrast, a separate study of WNT1 76 did not report increased mobility in the CTD, while another investigation 77 observed CTD mobility but reported a conformational transition toward pocket closure and a more compact structure of WNT1. These findings collectively suggest that WNT family proteins exhibit diverse dynamic behaviors in the liquid phase, with the capacity for transitions between several conformations, reflecting distinct structural plasticity among family members.
O-acylation, involving the addition of monounsaturated palmitoyl oleate (C16:1Δ9) to a serine residue, is a characteristic modification of WNT family proteins. This process facilitates their accessibility to transporters within the cell and in the intercellular space. 12 O-acylation is catalyzed by the endoplasmic reticulum enzyme Porcupine (PORCN).12,67 Palmitoleic acid plays an important role in WNT transport, secretion, and signaling by mediating interactions between WNT proteins and both the transport protein WLS and the FZD receptor. 10 - 12 To the best of our knowledge, these molecular dynamics simulations of WNT2B are the first to investigate the conformational dynamics not only of this WNT family member, but also of a WNT protein with such a specific posttranslational modification. However, further studies are needed to explore the dynamic behavior of WNT proteins with other posttranslational modifications, such as glycosylation, which may also play critical roles in their structure and function.
One of the most important aspects of WNT modification with palmitoleic acid is the introduction of hydrophobic properties, which complicates the crystallographic study of these proteins. 59 Indeed, our GRAVY calculations for Homo sapiens WNT proteins yielded negative values, indicating their hydrophilic nature in the absence of modifications. A previous study raised the question of how hydrophobic, O-acylated WNT proteins can persist in an uncomplexed state within a predominantly aqueous environment. 8 It has been proposed that WNTs may interact with carrier molecules or the plasma membrane, or that the hydrophobic regions of the “thumb” and “index finger” fold into a compact conformation forming the “fist.” 8 However, our study of free WNT2B does not support the hypothesis of intramolecular folding in an aqueous environment. On the contrary, for O-acylated WNT2B, our findings suggest protein unfolding in solution, leading to an open extended conformation. This observation at least partially supports the hypothesis that carrier molecules are required to maintain hydrophobic WNTs in an aqueous environment. A recent study 64 demonstrated that long-distance transport of WNT proteins can occur via their association with the surface of extracellular vesicles. The same study also discussed alternative potential carriers, including lipoproteins, soluble WNT-binding proteins, and mechanisms involving cytonemes. 64
The present study, although primarily focused on WNT2B, alongside other in vitro and in silico investigations of various WNT family members, contributes to a deeper understanding of the structural and functional properties of this protein family. Several shared features, including the presence of N-terminal signal peptides, overall hydrophobicity, posttranslational O-acylation, and a flexible 2-domain structure, appear to be important for the signaling functions of WNT proteins. For instance, it was shown 78 that human WNT3 and WNT3A, despite their high sequence similarity, undergo substantial conformational changes upon interaction with FZD and WLS.33,78 These changes include the rotation of hairpin-1 and the W-loop (which together form the “thumb”) by approximately 30° and 90°, respectively, and hairpin-2 (which forms the “index finger”) by approximately 60°. 78 Similarly, comparisons between human WNT8A in complex with WLS and Xenopus Wnt8 in complex with FZD revealed significant shifts in hairpin positioning, including alterations in the orientation of the palmitoleic acid attached to the hairpin in the “thumb” region, in order to accommodate specific binding interactions.12,79 Such structural flexibility may be essential for WNT proteins’ signaling activity and may facilitate a broad range of other molecules, such as PORCN, RYK, ROR1, GPC4, and PTK7. 12 Furthermore, a detailed understanding of WNT structural characteristics and their interaction mechanisms holds potential for the development of targeted therapies.59,80 The hydrophobic nature of WNT proteins also presents opportunities for their incorporation into liposomal delivery systems. 81
The computational study presented here provides a foundation for future in vitro experiments aimed at validating the obtained results and advancing our understanding of the structure and function of a broader range of WNT family members. Further research focusing on elucidating the tertiary structure of the N-terminal regions of WNT proteins, the mechanisms underlying the transport of hydrophobic O-acylated WNTs in aqueous environments, and the dynamic structural adaptations of WNTs during interaction with cell surface receptors holds significant promise.
Conclusions
The properties of WNT family proteins and the characteristics of their tertiary structure were investigated. Based on GRAVY index calculations, it was determined that proteins of this family are predominantly hydrophilic, suggesting that they acquire hydrophobic properties as a result of posttranslational lipid modifications. The N-terminal sequences of human WNT proteins, which function as signal peptides, were analyzed. Previously unknown lengths of N-terminal sequences in 2 different isoforms of human WNT2B, which are likely cleaved during the maturation of these proteins, were predicted. Molecular dynamics simulations were performed using both O-acylated (at Ser243) and non-acylated forms of mature WNT2B to explore the impact of lipid modifications on protein behavior. The ability of WNT2B to transition from a compact state to an open extended conformation was observed, offering insights into the structural flexibility of this protein and contributing to a broader understanding of WNT proteins’ behavior in an aqueous environment.
Supplemental Material
sj-docx-1-bbi-10.1177_11779322251353347 – Supplemental material for Bioinformatic Analysis of WNT Family Proteins
Supplemental material, sj-docx-1-bbi-10.1177_11779322251353347 for Bioinformatic Analysis of WNT Family Proteins by Konstantin Midlovets, Natalia Volkova and Mykyta Peka in Bioinformatics and Biology Insights
Footnotes
Ethical Considerations
This article does not contain any studies with human or animal participants.
Consent to Participate
Not applicable.
Consent for Publication
Not applicable.
Author Contributions
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.


