Abstract
Avian coccidiosis, caused by Eimeria protozoa, presents a significant threat to poultry, with Eimeria tenella being particularly harmful due to its impact on the chicken cecum. Growing resistance to current treatments necessitates alternative therapeutic approaches. Consequently, this study employed an immunoinformatics approach to design a multiepitope vaccine targeting E tenella. Key proteins, including the sporulated oocyst TA4 antigen, alkylglycerone-phosphate synthase, and apical membrane antigen-1, were analysed for epitope prediction. Further comprehensive downstream analysis identified 13 MHC class I, 6 MHC class II, and 7 B-cell epitopes, which were linked with suitable linkers. Also, cholera toxin subunit B was incorporated as an adjuvant, creating a 531-amino-acid construct. The vaccine demonstrated favourable predicted antigenicity, non-allergenicity, and stability properties. Molecular docking predicted interaction with toll-like receptor 15, while immune response simulation showed potential induction of various immunocytes, including helper and cytotoxic T-cells, natural killer cells, and immunoglobulins. The vaccine was predicted to promote antigen clearance after the second dose, suggesting strong memory response potential. These findings indicate the designed vaccine could stimulate a potent protective immune response against E tenella infection. However, further in vitro and in vivo validation studies are necessary to confirm the vaccine’s efficacy before clinical application in poultry immunization programmes.
Introduction
Coccidiosis, caused by protozoan parasites of the genus Eimeria, exerts a significant impact on global poultry production, which leads to significant economic losses within the poultry industry. 1 Historically, prophylactic drugs and attenuated and non-attenuated vaccines are utilized for combating the infection; however, the efficacy of these traditional interventions has been compromised by the relentless evolution of resistance mechanisms by Eimeria species, rendering them increasingly ineffective in mitigating the deleterious effects of coccidiosis. 2 Among the several Eimeria species affecting poultry, Eimeria tenella is notorious for its devastating impact on the chicken cecum, leading to severe intestinal lesions, impaired nutrient absorption, and a significant decline in poultry productivity. 1 Consequently, prophylactic approaches are commonly utilized to control the spread of the infection in poultry. These approaches include the administration of synthetic drugs such as amprolium, nicarbazin, diclazuril, and toltrazuril. 3 However, the spread of drug-resistant E tenella is rapidly spreading, hence necessitating the development of newer and more effective alternatives. 4
The sporulated oocyst TA4 antigen of E tenella, also known as E tenella SAG1, is crucial to the infection of the host, as it facilitates the initial attachment of sporozoites to the host cell surface during invasion, enabling the parasite to establish infection within the intestinal epithelium. 5 Similarly, the apical membrane antigen-1 (AMA-1), which is a type I transmembrane protein, plays critical roles during host cell penetration and growth, and its inhibition has been reported to prevent E tenella’s invasion of host cells. 6 Also, alkylglycerone-phosphate synthase (AGPS) is an enzyme that plays a crucial role in the production of ether lipids, which are components of the cellular membranes. 7 During the sporozoite stage of the life cycle of the parasite, alkylglycerone-phosphate synthase participates in the biosynthesis of ether lipids, which are necessary for the formation and maintenance of E tenella plasma membrane, and these functions are also important during the merogony and gametogony phases, as well as oocyst formation. 8 Consequently, the aforementioned proteins are considered good targets for the development of therapies against E tenella and diseases associated with its infection. In comparing the selected proteins to other proteins expressed by E tenella, the functional roles played by the selected proteins make them attractive for vaccine interventions. For example, the presence of Sporulated Oocyst TA4 Antigen in the sporulated oocyst phase makes it an ideal target for early intervention, potentially preventing the establishment of infection. 9 Similarly, the AMA-1 plays a crucial role in the invasion process of Eimeria tenella. Studies have demonstrated that antibodies targeting recombinant E tenella AMA-1 effectively inhibit sporozoite invasion of host cells. 10 In addition, AMA-1 has been identified as a protective antigen capable of eliciting significant immunoprotection against Eimeria infections. 11 Furthermore, targeting AGPS can disrupt the metabolic processes of the parasite, impacting its survival and replication. While other proteins, such as the immune-mapped protein-1, are recognized for their conservation and protective efficacy across Eimeria species, 12 the targeted selection of proteins in this study aims not only to confer protection but also to prevent infection at critical junctures in the parasite’s lifecycle. Furthermore, targeting AGPS can disrupt the metabolic processes of the parasite, impacting its survival and replication. While other proteins, such as the immune-mapped protein-1, are recognized for their conservation and protective efficacy across Eimeria species, 12 the targeted selection of proteins in this study aims not only to confer protection but also to prevent infection at critical junctures in the parasite’s lifecycle.
Traditional vaccine development approaches, such as in vivo experimentation and empirical screening, are often time-consuming, resource-intensive, and may not always yield effective or safe candidates. 13 These challenges are further compounded when dealing with pathogens that exhibit complex immune evasion mechanisms or rapid mutation rates, such as E. tenella, the causative agent of chicken coccidiosis. Moreover, these conventional methods require extensive testing phases, increasing the duration and cost of developing an efficacious vaccine. 14 To overcome these limitations, immunoinformatics has emerged as a powerful tool in modern vaccine design. By leveraging computational methods, immunoinformatics enables the rapid identification and assessment of potential vaccine candidates through epitope prediction, structural modelling, and immune response simulations. These approaches not only accelerate the vaccine development process but also allow for the in silico assessment of immunogenicity and safety before experimental validation.12,15
In this study, we employed a combination of computational tools to design a multiepitope vaccine targeting E. tenella. We hypothesize that a multiepitope vaccine incorporating epitopes from Sporulated Oocyst TA4 Antigen, AMA-1, and Alkylglycerone-Phosphate Synthase will elicit a robust and protective immune response against E. tenella infection in poultry.
Key tools such as the Immune Epitope Database (IEDB) were used to predict immunogenic epitopes, while molecular docking was conducted to evaluate the vaccine’s interaction with Toll-like receptor 15 (TLR15), a key player in avian immune responses. In addition, immune simulations were performed to predict the long-term efficacy of the vaccine in stimulating protective immunity. Given the accuracy of the web servers utilized, the chances of false-positive predictions were minimized. Ultimately, this study seeks to provide a cost-effective and timely alternative to conventional vaccine development methods. 16
Materials and Methods
Protein sequence retrieval
The sequences of the sporulated oocyst TA4 antigen, alkylglycerone-phosphate synthase, and AMA-1 of Eimeria tenella were obtained from the UniProt database (http://www.uniprot.org/) in FASTA format. 17
Prediction of T-cell and B-cell epitopes
T-cell and B-cell epitope predictions were conducted using protein sequences retrieved and analysed through the Immune Epitope Database (IEDB) (https://www.iedb.org/). 18 The NetMHCpan EL 4.0 prediction method (http://tools.iedb.org/mhci/) was employed to predict MHC class I restricted CD8+ cytotoxic T-lymphocyte (CTL) epitopes, while the IEDB-recommended 2.22 prediction method (http://tools.iedb.org/mhcii/) was utilized for predicting 15-mer epitopes specific to MHC class II restricted CD4+ helper T-lymphocytes (HTLs) across common HLA alleles. Due to the unavailability of chicken MHC alleles on the webserver, human HLA alleles were used as a substitute for both MHC-I and MHC-II epitope predictions. 12 In addition, the BepiPred-2.0 linear epitope prediction method (http://tools.iedb.org/bcell/) was utilized to identify potential linear B-cell epitopes (BCLs) of the proteins. 19
Antigenicity and allergenicity prediction
The antigenicity of the selected epitopes was assessed using VaxiJen v2.0 (http://www.ddg-pharmfac.net/vaxijen/), with the accuracy value set to 0.5 and the target organism specified as ‘parasite’. 20 Subsequently, the allergenicity of the predicted epitopes was evaluated using AllerTOP v2.0 (https://www.ddg-pharmfac.net/AllerTOP/), which offers a prediction accuracy of 88.7%. 21
Epitope conjugation
The optimal epitopes selected for vaccine design were those meeting the criteria of being antigenic and non-allergenic. The CTL and HTL epitopes were conjugated using the ‘AAY’ and ‘GPGPG’ linkers, respectively, while the B-cell epitopes were conjugated using the bi-lysine linker. Notably, the cholera toxin subunit B (CTB) (accession number ABV74245.1) was utilized as the adjuvant, and its sequence was conjugated using the ‘EAAK’ linker. 12
Vaccine properties analysis
The antigenicity of the vaccine was assessed using the VaxiJen v2.0 webserver, while its allergenicity was predicted using the AllerTOP webserver. Subsequently, the physicochemical properties of the vaccine were computed using the ProtParam server (https://web.expasy.org/protparam/). 22
Vaccine tertiary prediction and refinement
The I-TASSER webserver (https://zhanggroup.org/I-TASSER/) was employed to predict the 3-dimensional (3D) structure of the vaccine construct. 23 Subsequently, the best model from the predicted structures was refined using the ModRefiner module of the ZhangLab Webserver (https://zhanggroup.org/ModRefiner/), 24 after which a quality assessment of the refined structure was done using the PROCHECK (https://saves.mbi.ucla.edu/) 25 and ProSA webservers 16 based on the Ramachandran plot and z-score analyses.
Vaccine and toll-like receptor docking
The 3D structure of the vaccine was subjected to molecular docking against the chicken toll-like receptor (TLR)-15 toll/interleukin-1 receptor (TIR) domain. Notably, the structures of the TLRs were retrieved from the Protein Data Bank (PDB) using the PDB IDs ‘3J0A’ and ‘7YLF’, respectively.26,27 Subsequently, the structures were subjected to docking preparatory steps using UCSF Chimera software, 28 and ClusPro Server (https://cluspro.org/home.php) was utilized for the docking. 29 The 2-dimensional view of the interactions of the vaccine with the TLRs was visualized using Ligplot+ v.2.2.5, while PyMol v3.0 was employed for the 3D visualization. 16
Molecular dynamics simulation
The stability of the complex formed from the docking simulation was assessed using various parameters available via the online server iMODS (https://imods.iqfr.csic.es/). The parameters evaluated included the B-factor, deformability, eigenvalues, variance, covariance map, and elastic network.30-32
Immune simulation
The immune system’s profile following the vaccine administration was simulated using the C-ImmSim server (https://kraken.iac.rm.cnr.it/C-IMMSIM/). 33 The simulation time steps, which correspond to the days of vaccine administration, were set to 1, 84, and 170, while the simulation time step was set to 1050. Time step 1 corresponds to the first injection time, while time steps 84 and 170 correspond to the second and last days of administration, respectively. 16
Results
T-cell and B-cell epitopes identified from Eimeria tenella’s proteins
The UniProt IDs of the retrieved sequences of E tenella’s sporulated oocyst TA4 antigen, alkylglycerone-phosphate synthase, and AMA-1 are P13399, U6KPA3, and F8V414, respectively. Several HTL and CTL epitopes were identified from these proteins, and the epitopes were predicted to be potential binders of several alleles. Some of the identified epitopes are presented in Table 1.
Selected MHC classes I and II epitopes with their antigenicity and allergenicity potential.

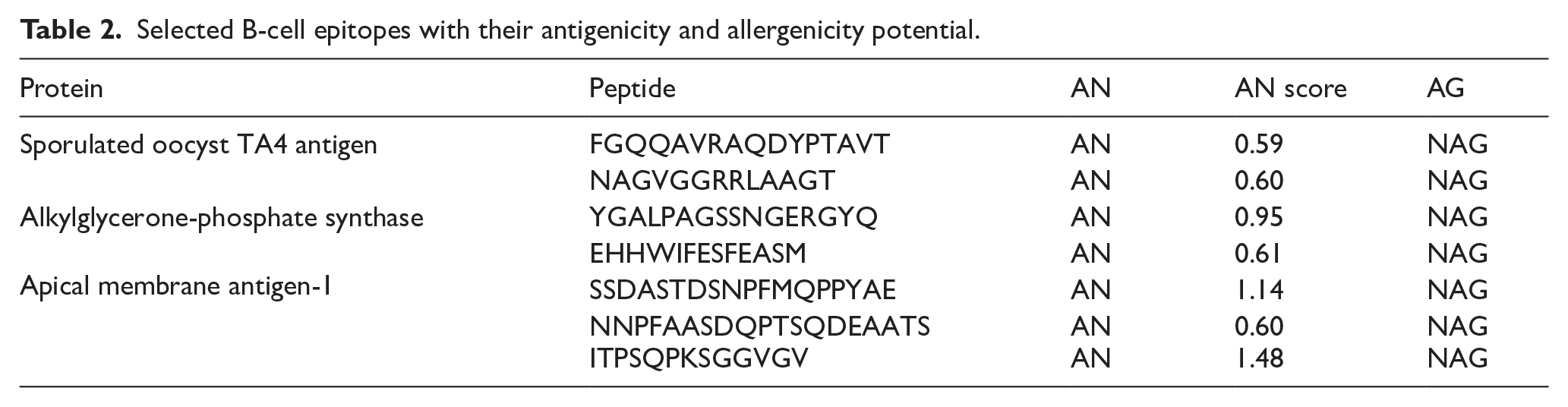
B-cell epitopes present in the proteins were also predicted, and the results are presented in Table 2. Notably, the length of the sequence of the epitopes selected in this study was between 2 and 20.
Selected B-cell epitopes with their antigenicity and allergenicity potential.

Epitopes antigenicity and allergenicity evaluation
The antigenicity and allergenicity potentials of the T-cell and B-cell epitopes identified from the proteins are presented in Tables 1 and 2, respectively. As evident in the tables, the epitopes possessed high antigenicity potential while also lacking the potential to induce allergenic reactions. Predicted T-cell and B-cell epitopes that did not possess suitable antigenicity and allergenicity potentials are presented in the Supplementary Material. Supplementary Table 1 presents non-viable MHC class I epitopes identified from the Sporulated oocyst TA4 antigen protein; Supplementary Table 2 presents non-viable MHC class II epitopes identified from the Sporulated oocyst TA4 antigen protein; Supplementary Table 3 presents non-viable B-cell epitopes identified from the Sporulated oocyst TA4 antigen protein; Supplementary Table 4 presents non-viable MHC class I epitopes identified from the Alkylglycerone-phosphate synthase; Supplementary Table 5 presents non-viable MHC class II epitopes identified from the alkylglycerone-phosphate synthase; Supplementary Table 6 presents non-viable B-cell epitopes identified from the alkylglycerone-phosphate synthase; Supplementary Table 7 presents non-viable MHC class I epitopes identified from the AMA-1; Supplementary Table 8 presents non-viable MHC class II epitopes identified from the AMA-1; Supplementary Table 9 presents non-viable MHC class II epitopes identified from the AMA-1.
Vaccine designing
The conjugation of the epitopes presented in Tables 1 and 2 using the sequences of the aforementioned linkers and adjuvant resulted in a final vaccine construct containing 531 amino acid sequences. The designed Eimeria tenella vaccine (ETV) contains 13 CTL epitopes, 6 HTL class II epitopes, and 7 B-cell epitopes.
Eimeria tenella vaccine antigenicity, allergenicity, and physicochemical property analyses
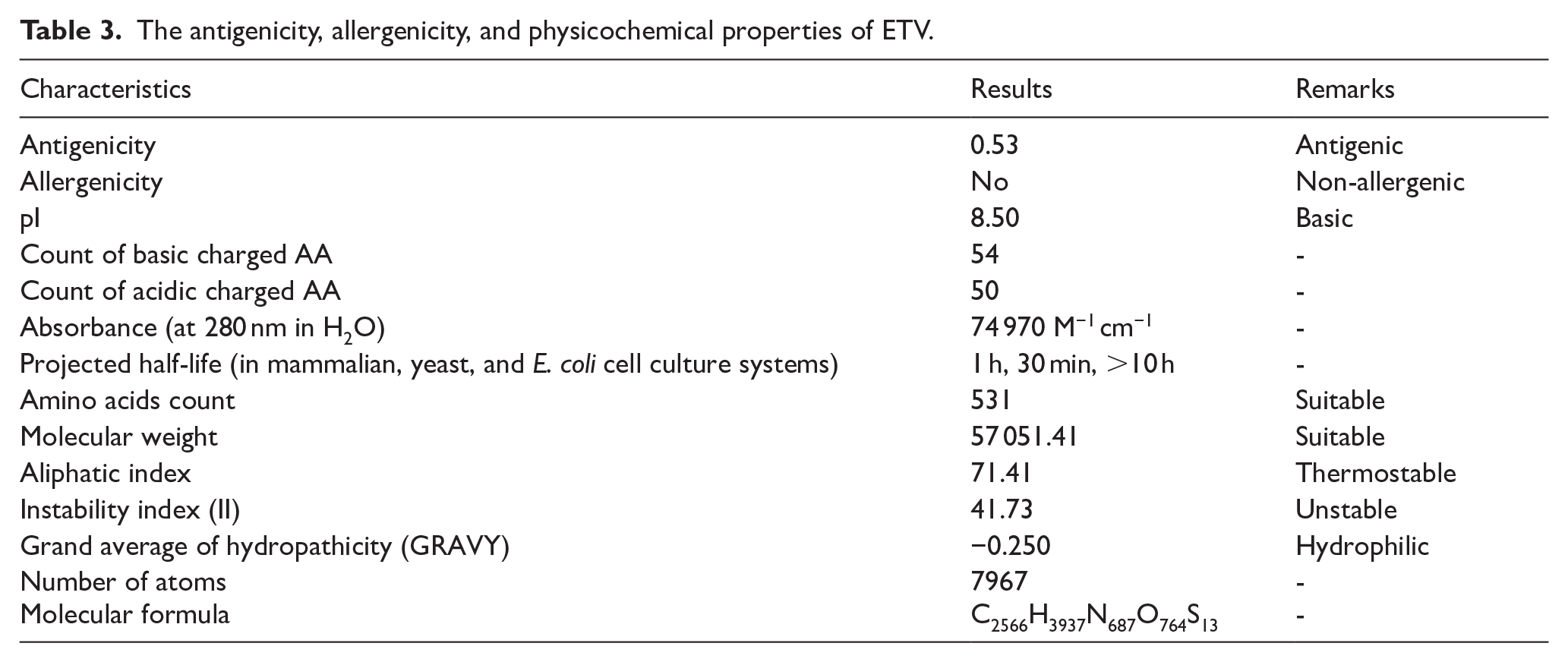
As presented in Table 3, the designed ETV vaccine was found to be antigenic and non-allergenic. The ETV contains 531 amino acid residues and has a chemical formula of C2566H3937N687O764S13. Other parameters include the hydropathicity value, instability index, and isoelectric point, which revealed ETV to be hydrophilic, thermostable, and basic, while it also has a molecular weight of 57051.41 daltons.
The antigenicity, allergenicity, and physicochemical properties of ETV.
Eimeria tenella vaccine tertiary structure
Evaluation of the tertiary structure of ETV predicted using the I-TASSER webserver revealed 69% of the structure’s residues were in the most favoured region of the Ramachandran plot, as depicted in Figure 1A, while other residues were in the additional allowed regions, generously allowed regions, and disallowed regions with 23.2%, 4.2%, and 3.5%, respectively. Conversely, as depicted in Figure 1B, the refined structure had 85.8% of its residues in the most favoured region, while 10.2%, 1.3%, and 2.7% were in the additional allowed regions, generously allowed regions, and disallowed regions, respectively.

Structural validation of ETV tertiary structure. (A) Unrefined structure. (B) Refined structure.
The refined tertiary structure of ETV is depicted in Figure 2.

The 3-dimensional structure of ETV.
Eimeria tenella vaccine’s interaction with toll-like receptor-15
The ETV was found to possess a binding affinity of −1060.9 kcal/mol and interacted with several residues of the TIR domain of TLR15 as depicted in Figure 3. The ETV residues interacted with the TIR via several hydrogen bonds, as exemplified by the interaction of ASN39 with ARG787. Other hydrophobic interactions are also present, as exemplified by the interaction of GLN213 with PRO830.

Molecular interaction between the Eimeria tenella vaccine (ETV) construct and the TIR domain of TLR15. Amino acids are labelled with (A) for Chain A (TLR15) and (B) for Chain B (ETV construct) to indicate their respective positions within the complex.
Immune simulation of Eimeria tenella vaccine
As evident in Figure 4, immune simulation of ETV revealed the vaccine was capable of inducing an immune response, as evident by the increase in immunocytes and antibody titre values.

The simulation of the generation of immune response induced by ETV. (A) Antigen levels and immunoglobulins. (B) Helper T-lymphocyte count. (C) Cytotoxic T-lymphocyte count. (D) Natural killer cells count. (E) Macrophages count. (F) Concentration of cytokines and interleukins.
Eimeria tenella vaccine toll-like receptor-15 complex stability
As depicted in Figure 5, the results of the molecular dynamics simulation revealed the interaction between ETV and TLR15 as stable. The complex has a low degree of deformability, as evident in Figure 5A, which shows the deformability graph; Figure 5B shows the number of correlated residues of the complex; Figure 5C reveals the B-factor value; and Figure 5D depicts the elastic model.

Stability profile of ETV-TLR15 complex. (A) Deformability graph. (B) Covariance matrix. (C) B-factor. (D) The elastic network model which showcases the pairs of atoms joined together by springs.
Discussion
The global economic burden impacted farmers as a result of coccidiosis, whose etiological organism is E. tenella, is worrisome. Hence, prophylactic measures aimed at preventing the high mortality and reduced productivity associated with its infection are of great importance. In this study, an immunoinformatics approach was employed to design a multiepitope vaccine that can be utilized in combating E tenella infection. One of the challenges of developing an effective vaccine against E tenella is its antigen complexity, and the conjugation of T-cell epitopes from different stages of the organism’s life has been identified as a viable way to surmount this. 2 In parallel with this, the efficacy of a vaccine has been linked to its ability to stimulate both T-cells and B-cells in order to achieve a broad immune response against a pathogen. 12 With this in mind, this study selected both helper T-cell and cytotoxic T-cell epitopes from proteins that play critical roles in the life cycle of the parasite. Noteworthy, the B-cell epitopes selected in this study had lengths less than 20 in order to ensure that their conjugation with other selected epitopes would not result in a vaccine with unsuitable physicochemical properties, hence making the process of experimental production arduous.16,22 To eliminate false-positive epitopes, the identified epitopes were further subjected to antigenicity tests aimed at identifying the epitopes with the potential to be recognized by and interact with the immune system, resulting in the induction of an immune response through the production of antibodies or activation of T-cells against the parasite. Among the epitopes identified, several stand out for their high antigenicity. For instance, the epitope RSQPFDDEQW from the sporulated oocyst TA4 antigen has the highest antigenicity score of 1.73, indicating its potential to induce a robust immune response. The ability of this epitope to bind multiple MHC class I alleles, including HLA-B*58:01 and HLA-A*32:01, enhances its versatility, making it a valuable target for eliciting cytotoxic T-cell responses across diverse populations. Similarly, the epitope FGKNIEFYQPLDSDL from the AMA-1 exhibits a high antigenicity score of 0.93, positioning it as a strong candidate for inclusion in the vaccine. This epitope’s binding affinity to MHC class II alleles such as HLA-DRB1*15:01 suggests its role in eliciting a helper T-cell response, crucial for stimulating antibody production and developing long-term immune memory. Moreover, epitopes such as RAAPVAETLW (antigenicity score of 0.91) and PVVSCVLLQCPNAGV (antigenicity score of 0.91) highlight additional immunogenic targets that contribute to a balanced immune response by engaging both cytotoxic and helper T-cells. The non-allergenic nature of these epitopes further supports their suitability in vaccine development, reducing the risk of adverse allergic reactions.
Among the identified B-cell epitopes, ITPSQPKSGGVGV from the AMA-1 stands out with an antigenicity score of 1.48, indicating its strong potential for triggering an immune response. Similarly, the peptide SSDASTDSNPFMQPPYAE from the AMA-1 has a high antigenicity score of 1.14, further suggesting its importance in activating the immune system. The ability of this peptide to stimulate immune responses without allergenic properties strengthens its role as a critical component in vaccine development. In addition, the peptide YGALPAGSSNGERGYQ from alkylglycerone-phosphate synthase has an antigenicity score of 0.95, positioning it as another strong epitope that could enhance the breadth of immune responses when incorporated into the vaccine. Its non-allergenic status supports its use in a wide population without the risk of triggering allergic reactions.
Lower antigenicity epitopes, such as FGQQAVRAQDYPTAVT and NAGVGGRRLAAGT from the sporulated oocyst TA4 antigen, with scores of 0.59 and 0.60, respectively, still exhibit potential as antigenic targets. These peptides, while less immunogenic, could contribute to a balanced immune response, working in conjunction with highly antigenic epitopes to strengthen the overall vaccine-induced immunity. Interestingly, several of the identified epitopes were predicted to be incapable of stimulating the immune response; hence, they were eliminated. Furthermore, epitopes with the potential to trigger allergic reactions were screened out.
The decision to use human HLA alleles for epitope prediction in this study, while introducing certain biases, ultimately reflects the innovative approach taken in the absence of avian-specific resources. Although immune responses in chickens and humans can differ, there are notable similarities in their adaptive immune systems. Both species rely on MHC molecules to present antigenic peptides to T-cells, engaging similar downstream pathways, including T-cell activation and cytokine production.34,35 This foundational similarity provides a rationale for utilizing human HLA alleles as a surrogate in our immunoinformatics analysis. Current limitations in publicly available webservers for avian-specific MHC prediction necessitated this choice, as several studies have similarly relied on human alleles to inform poultry vaccine design. 36 By leveraging the established functional parallels between the immune systems of chickens and humans, our study effectively harnesses existing knowledge to advance the design of potential vaccines against avian coccidiosis. Importantly, research has shown cross-species reactivity between MHC molecules, reinforcing the validity of our approach. The conserved mechanisms of T-cell-mediated immunity across species lend credence to the predictive power of human HLA alleles in identifying potential immunogenic epitopes for chickens.
The conjugation of the final set of epitopes with suitable properties and the CTB adjuvant resulted in a vaccine construct with 531 amino acid sequences. The selection and conjugation of the CTB adjuvant are to further bolster the immune response that will be elicited upon the administration of the vaccine. Notably, adjuvants from toxins have been reported to improve the immunogenicity of vaccines by inducing the production of immunoglobulins G and A. Specifically, the CTB adjuvant aids in the formation of a GM1-ganglioside receptor-binding pentameric structure in the gut mucosa and modulates the expression of gut cell receptors. 12 Also, studies have documented the efficacy of CTB in avian species. For instance, a study involving oral administration of a recombinant pool of Lactobacillus spp., combined with microencapsulated CTB and flagellin, to broiler chickens demonstrated significant increases in intestinal IgA and serum IgY levels across immunized groups, particularly 3 weeks post-immunization. In addition, the administration of recombinant CTB was associated with a marked elevation in CD8+ T-lymphocyte populations within the cecum, suggesting strong activation of adaptive immune responses in poultry. 37 Furthermore, a separate study underscored the adjuvanticity of CTB in chickens, revealing a significant positive correlation between CTB’s immunogenicity and the specific antibody response elicited against bovine serum albumin when co-administered orally. 38 These findings underscore the potential utility of CTB as an adjuvant in vaccine formulations aimed at poultry. Notwithstanding these encouraging findings, it is recognized that alternative adjuvants may enhance the immune response more effectively and mitigate potential safety concerns.
The physicochemical properties of the designed vaccine construct were evaluated to determine its suitability for further development. The construct demonstrated antigenicity (0.53), indicating its potential to elicit an immune response, while its non-allergenic nature suggests a lower risk of hypersensitivity reactions. The molecular weight (57.05 kDa) and theoretical isoelectric point (pI = 8.50) were comparable to other in silico-designed vaccines, such as those developed against breast cancer (47 kDa, pI = 9.35), 39 melanoma (39.8 kDa, pI = 9.61), 40 and SARS-CoV-2 (51.64 kDa, pI = 10). 41 Thermal stability is an essential parameter for vaccine stability and efficacy. The aliphatic index of the construct (71.41) suggests thermostability, although it is slightly lower than that reported in SARS-CoV-2 (79) 41 and breast cancer vaccines (67.45), 39 but it remains within the range observed in previously designed constructs. The hydrophilic nature of the vaccine, indicated by a negative GRAVY value (−0.250), is in line with other studies, such as those targeting TMEM31 (−0.215) 39 and SARS-CoV-2 (−0.354). 41 This hydrophilicity is beneficial for solubility and effective interaction with the immune system.
The instability index of the designed vaccine (41.73) classifies it as unstable, contrasting with previous studies where vaccine constructs were predicted to be stable (instability index < 40).39,41 However, similar values have been reported for other vaccine constructs, and further structural optimizations or in vitro expression studies may help address this limitation. The estimated half-life in mammalian cells (1 hour 30 minutes), yeast (30 hours), and E coli (> 10 hours) aligns with the expected range reported for previous vaccine constructs. Overall, the designed vaccine exhibits physicochemical properties that are comparable to, and in some cases, aligned with, previously developed multiepitope vaccines. Despite a slightly higher instability index, the vaccine’s thermostability, solubility, and antigenicity highlight its potential as a viable candidate for further experimental validation.
The tertiary structure of ETV was generated via ab initio modelling, with the Ramachandran plot revealing the structure as having high quality. A high percentage of the residues in the most favoured regions of the Ramachandran plot indicates a well-structured protein, suggesting that the backbone conformation is energetically stable and consistent with known protein structures.
TLR15 has been reported to be upregulated in chickens during E tenella infection; it is commonly upregulated in immunocytes, including heterophils and macrophages, and it has been suggested to trigger pathways involving other immune system molecules like MyD88 and nuclear factor kappa B (NF-κB) to fight the infection.42,43 The tertiary structure of the ETV was found to interact strongly with the TIR domain of TLR15, as evidenced by the docking score of −1060.9 kcal/mol. The interaction was also found to be stable; the complex had a low degree of deformability, while the eigenvalue showed that a high energy level is required to dissociate the complex. Other parameters, such as the covariance map, showed the complex has a high number of correlated residues, while the elastic model revealed numerous stiff regions in the complex. In the context of E tenella vaccine development, interactions with TLR15 are biologically significant as they enhance the ability of a vaccine to induce a strong immune response. By activating TLR15, the vaccine can stimulate both the innate and adaptive branches of the immune system, leading to the production of antigen-specific T-cells and antibodies.42,43 This makes TLR15 interactions especially valuable in poultry vaccines, where they can potentiate immune responses, improve vaccine efficacy, and potentially provide broader protection against pathogens.
The immune simulation of ETV’s response activation pattern demonstrated a significant increase in IgM and IgG levels following the administration of the final vaccine dose. Importantly, antibody levels remained elevated for a period of 1 year, indicating sustained immune memory. In addition to this humoral response, high populations of natural killer (NK) cells and macrophages persisted throughout the year, suggesting continued immune surveillance. Cytotoxic T-cells (CD8+) and helper T-cells (CD4+) were also present at high levels for the first 100 days, representing strong cellular immunity post-vaccination. A notable decrease in immune cell activity was observed after 50 days, indicating that the vaccine successfully initiated a controlled immune response. The induction of protective immunity against avian coccidiosis in chickens has been linked to the production of interleukins, such as interleukin (IL)-2, tumour necrosis factor (TNF)-α, and transforming growth factor (TGF-β), which aligns with the cytokine responses observed in our immune simulation. This finding supports the potential of ETV to elicit a robust immune response in chickens. While we did not manually set the baseline values used in the immune simulation, the server’s pre-configured parameters enabled us to effectively assess immune dynamics. The significant increases in immunocytes and antibody levels above these default baseline values demonstrate the vaccine’s ability to provoke both humoral and cellular immunity, further emphasizing its promise as a potential vaccine candidate. As evident in Supplementary Figure 1A, a sharp increase in T-helper (TH) cell count was observed shortly after immunization, reaching peak values within the first 50 days. This rapid expansion signifies a robust activation of the adaptive immune response, likely triggered by antigen presentation and immune system priming. Following this peak, non-memory TH cells exhibited a sharp decline, while memory TH cells persisted over an extended period. The retention of memory TH cells suggests that the vaccine construct is capable of inducing long-term immunological memory, which is critical for prolonged immunity and rapid response upon subsequent antigen exposure.
The differentiation of TH cells into distinct functional states was further analysed. The majority of TH cells transitioned into an active state early in the immune response, characterized by rapid expansion within the first 50 days. This was followed by a gradual decline as cells transitioned into resting or anergic states. The presence of duplicating TH cells during the early immune response phase suggests efficient proliferation and immune activation. Notably, the proportion of resting TH cells remained stable over time (Supplementary Figure 1B), further supporting the vaccine’s ability to induce long-lasting immune memory. A detailed breakdown of TH cell subtypes provides insights into the polarization of the immune response. During the peak immune response phase, the dominant population was the TH1 cells, which are key in orchestrating cellular immunity. The eventual emergence of regulatory T-cells suggests an intrinsic mechanism for immune regulation, to prevent excessive immune activation and potential autoimmunity. The persistence of TH17 cells, which are involved in inflammatory responses, may also contribute to enhanced immune activation, potentially augmenting vaccine efficacy (Supplementary Figure 1). However, it is worth noting that the simulations provide valuable insights into potential vaccine efficacy by predicting immune activation patterns based on computational algorithms and historical data. For example, while our simulation indicated sustained antibody levels, experimental data could reveal variations influenced by genetic diversity, environmental factors, and other biological complexities influenced by antigen persistence, dosage, and the overall health of the host. Hence, the results that will be obtained from experimental validation of the immune response might differ.
Current vaccines against coccidia primarily consist of first-generation live, wild-type Eimeria tenella oocyst vaccines, used since the 1950s. 13 While effective in inducing protective immunity, these vaccines face challenges such as inconsistent administration and the need for recycling live oocysts, which can lead to high exposure and clinical disease, often necessitating anticoccidial drugs post-vaccination. 44 Newer second-generation vaccines utilize attenuated Eimeria lines with reduced pathogenicity but still rely on in vivo production, making them costly and limiting availability. 44 In contrast, our proposed vaccine offers distinct advantages. It leverages computational approaches to enhance immunogenicity while minimizing pathogenic risks, potentially allowing for more efficient production methods. Our vaccine also demonstrates the ability to induce robust immune responses, characterized by sustained antibody levels and high immune cell populations.
While the immunoinformatics-driven approach employed in this study offers a rapid and cost-effective strategy for designing a potential multiepitope vaccine against E tenella, challenges are anticipated in translating these findings into poultry application. Notably, several of the prediction tools and immune simulation servers utilized, such as those for epitope binding and C-ImmSim, are primarily designed for modelling human immune responses. This may limit the accuracy of the predicted immune dynamics in chickens, which possess a distinct immunological architecture. Furthermore, there may be discrepancies between in silico predictions and actual biological responses, as epitopes identified computationally may not retain their immunogenicity, structural conformation, or stability when expressed in vivo. Issues related to construct expression, solubility, and proper folding in suitable expression systems may also pose hurdles in vaccine development. Moreover, the host-pathogen interactions in avian species, which are not fully recapitulated by current computational models, could significantly influence vaccine efficacy. Therefore, while the vaccine construct shows promise in silico, its poultry-specific utility requires rigorous experimental validation, including codon optimization for avian expression systems, structural and functional assessment, and controlled in vitro and in vivo immunogenicity assays, to substantiate its real-world application and mitigate any concerns in subsequent peer scrutiny.
Conclusions
Summarily, this study utilized immunoinformatics approach to design a multiepitope vaccine against E tenella based on the antigenic epitopes identified from the sporulated oocyst TA4 antigen, alkylglycerone-phosphate synthase, and AMA-1 proteins. The designed vaccine was predicted to be capable of eliciting a long-lasting immune response and possesses the physicochemical properties of an ideal vaccine candidate. Conclusively, in silico evaluation of the designed vaccine revealed its potential to be effective in combating E tenella infection; however, further experimental studies are needed to further validate the results of this study.
Supplemental Material
sj-docx-1-bbi-10.1177_11779322251348307 – Supplemental material for Immunoinformatics-Aided Design of a Multiepitope Vaccine Against Eimeria Tenella-Associated Chicken Coccidiosis
Supplemental material, sj-docx-1-bbi-10.1177_11779322251348307 for Immunoinformatics-Aided Design of a Multiepitope Vaccine Against Eimeria Tenella-Associated Chicken Coccidiosis by Adeyinka I Fadahunsi, Adewale J Atansuyi, Abayomi I Adebayo and Clifford A Chineke in Bioinformatics and Biology Insights
Footnotes
Author Contributions
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Ethical Approval
Ethical approval was not required for this study as it did not involve any human participants, animal subjects, or the use of confidential data.
Human and Animal Rights
This article does not contain any studies with human or animal subjects.
Informed Consent
There are no human subjects in this article and informed consent is not applicable.
Data Availability Statement
All data generated or analysed during this study are included in this published article.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
