Abstract
Rhoptry protein 18 (ROP18) is a key virulence factor secreted into host cells during the invasion of Toxoplasma gondii (T. gondii) and plays an important role in the pathogenesis of infection. Due to its potential as a vaccine candidate, this study aimed to characterize several properties of the T. gondii ROP18 (TgROP18) protein to support its inclusion in vaccine formulations. Using a range of bioinformatics tools, we investigated its T-cell and B-cell epitopes, physicochemical properties, subcellular localization, transmembrane domains, and tertiary and secondary structures. Our analysis revealed 48 post-translational modification sites in TgROP18. The secondary structure was composed of 4.87% beta-turns, 38.45% random coils, 42.42% alpha helices, and 14.26% extended strands. Several potential T- and B-cell epitopes were identified on ROP18. The Ramachandran plot of both crude and refined models showed that 85.8% and 95.3% of the amino acid residues, respectively, fell within favored regions, indicating energetically stable conformations. Allergenicity and antigenicity assessments indicated that TgROP18 is a nonallergenic, immunogenic protein. Predictions using the C-ImmSim server suggest that TgROP18 can stimulate humoral and cell-mediated immune responses, based on antibody titers and cytokine profiles following antigen administration. These findings provide baseline data for future investigations focused on the potential of TgROP18 in developing therapeutic strategies against toxoplasmosis.
Introduction
Toxoplasmosis, caused by Toxoplasma gondii (T. gondii), is a widespread parasitic infection affecting more than one-third of the global population.1-3 The parasite undergoes 3 main developmental stages: sporozoites in oocysts, bradyzoites in tissue cysts, and tachyzoites. Humans can acquire the infection through various routes, including consuming raw or undercooked meat containing tissue cysts, ingesting of raw or unwashed vegetables or drinking water contaminated with oocysts, organ transplantation, vertical transmission from mother to fetus, and possibly blood transfusions.4-6 Although bradyzoites are associated with the long-term stage of infection, tachyzoites drive acute infection. 7 Although many T. gondii infections are asymptomatic, they can lead to severe complications, particularly in neonates with congenital infection and immunocompromised individuals.3,8,9
Current treatments for toxoplasmosis target only the proliferating tachyzoites during the early stages of infection and often associated with significant side effects.10-12 Importantly, no existing treatments can eliminate the tissue cysts of T. gondii, which can persist in the host. 13 Therefore, vaccination represents a promising strategy for controlling toxoplasmosis.14-16 Given the substantial impact of T. gondii on both public health and the livestock industry, developing an effective vaccine is a compelling priority.14,16-18 Several antigens, including microneme proteins (MICs), surface antigens (SAGs), rhoptry proteins (ROPs), and dense granule proteins (GRAs), have been proposed as potential vaccine targets.14-17,19-21 However, most of these antigens offer only partial protection, 22 underscoring the need to carefully select antigens that can yield a safe and effective vaccine.
Rhoptry proteins play a key role in the parasite’s survival within host cells and are crucial during the invasion process.23,24 Members of the ROP family, including ROP5, ROP18, ROP17, and ROP19, have shown promise as DNA vaccine candidates against T. gondii.25-28 Specifically, several rhoptry antigens, such as ROP18, ROP2, ROP5, and ROP16, are involved in the parasite’s virulence and pathogenesis. Rhoptry protein 18, a highly polymorphic serine-threonine kinase, is secreted into the host cell during invasion and is critical in regulating host-parasite interactions.24,29-31 Its immunogenic potential is well-supported, as ROP18 can trigger a mixed Th1 to Th2 immune response, with a bias toward Th1, evidenced by a higher immunoglobulin (IgG)2a/IgG1 ratio and elevated levels of interferon-γ (IFN-γ) and interleukin-2 (IL-2).32-38 These cytokines are associated with protective immunity, prolonged survival, and reduced formation of tissue cysts in infected mice.
Bioinformatics tools have emerged as powerful resources for identifying B- and T-cell epitopes with high antigenic potential, making them valuable for selecting candidates for vaccine development. Compared with traditional methods, these computational approaches are time-efficient, cost-effective, and provide a high degree of accuracy and precision.39,40 By leveraging bioinformatics, researchers can effectively analyze protein characteristics relevant to both vaccination and diagnostic applications.40,41 In this study, a suite of bioinformatics tools was used to predict the structure, immunogenic epitopes, and other features of the TgROP18 protein. The findings offer new insights into the immunogenic properties of TgROP18, reinforcing its potential as a vaccine candidate against T. gondii infection.
Methods
Amino acid sequence
The amino acid sequence of the TgROP18 protein was obtained in FASTA format from the ToxoDB database, 42 with the following accession number: TGME49_205250 (ME49 strain). The links to all the bioinformatics tools used in this study are provided in Supplemental Table 1.
Physical-chemical characteristics
Key physicochemical properties of the TgROP18 protein were predicted using the ExPASy ProtParam server. The analysis included the number of amino acids, molecular weight (MW), isoelectric point (pI), estimated half-life (in vivo and in vitro), the total number of positively and negatively charged residues, instability index, extinction coefficient, aliphatic index, and the grand average of hydropathicity (GRAVY). 43
Post-translational modification sites
The potential post-translational modification sites of the ROP18 protein were predicted using several bioinformatics tools. N-glycosylation sites were identified with the NetNGlyc 1.0 server, N-acetylation sites with the NetAcet 1.0 server, and phosphorylation sites with the NetPhos 3.1 server.44,45
Subcellular localization and transmembrane domains
The subcellular localization of TgROP18 was predicted using the PSORT II prediction tool. In addition, the TMHMM v. 2.0 server was employed to assess the transmembrane domains of the protein using default parameters. 45
Secondary and tertiary structure prediction
The secondary structure of the ROP18 protein was predicted using the SOPMA server, categorizing the structure into 4 conformational states: helix, sheet, turn, and coil. 46 Disulfide bond prediction and protein solubility (SOLpro) were predicted using the Scratch Protein Predictor tool. 47 The tertiary structure was modeled using SWISS-MODEL, employing a homology modeling approach to generate 3-dimensional (3D) structures based on sequence similarity. 48
Tertiary structure refinement and validation
The 3D model was refined using the GalaxyRefine server, which reconstructed and repacked the side chains and relaxed the overall structure.49-51 The refined 3D model was validated using a Ramachandran plot through the PROCHECK tool, available on the SAVES v6.0 server.52,53 In addition, the overall quality of the model was evaluated using the ProSA-web tool by analyzing the 3D structure in protein data bank (PDB) format.54,55
B-cell epitope prediction
To identify B-cell epitopes, the amino acid sequence of TgROP18 was analyzed using several bioinformatics servers. For linear B-cell epitopes, SVMTriP was used to predict continuous epitopes with a length of 20 amino acids. 56 The Bcepred server was also employed, combining parameters such as flexibility, polarity, surface accessibility, and hydrophilicity at a threshold of 2.38. 57 The ABCpred server, based on a recurrent neural network, was used to predict linear epitopes with an accuracy of 65.93%, using a length of 16 amino acids and a threshold of 0.75. 58 The IEDB (Immune Epitope Database) was further used to assess various properties such as hydrophilicity, flexibility, beta-turns, surface accessibility, and antigenicity.59-64
For conformational (discontinuous) B-cell epitopes, the PDB file of the 3D structure was analyzed using the DiscoTope 2.0 server, which calculated surface accessibility and epitope propensity scores. 65 In addition, the ElliPro tool from the IEDB was used to predict conformational epitopes with default settings (minimum score: 0.5 and maximum distance [Angstrom]: 6). 66
Cytotoxic T lymphocyte epitope prediction
Cytotoxic T lymphocyte (CTL) epitopes were predicted using the CTLpred server, which employs a combination of methods, including support vector machine (SVM) and artificial neural network (ANN). 67 The cutoff scores for SVM and ANN were set at 0.36 and 0.51, respectively, resulting in a predictive accuracy of 75.8%.
MHC-II and MHC-I binding epitope prediction
Major histocompatibility complex (MHC) class I and II binding epitopes were predicted using the IEDB web tool, which calculates the half-maximal inhibitory concentration (IC50) values. For MHC class II, alleles H2-IAd, H2-IEd, and H2-IAb were selected, and 15-mer peptides were predicted based on the IEDB-recommended method. For MHC class I, alleles H2-Kk, H2-Dd, H2-Db, H2-Kd, H2-Ld, and H2-Kb were analyzed, using a 10-mer peptide length and percentile rank for prediction. 45
Allergenicity, solubility, and antigenicity
Allergenicity of TgROP18 was predicted using the AlgPred server, which integrates 6 distinct methods, with a hybrid approach yielding an 85% efficiency at a threshold of −0.4. 68 Protein solubility was assessed using the SOLpro server. 47 Antigenicity was evaluated using VaxiJen v. 2.0 69 and ANTIGENpro, 70 with the VaxiJen server offering accuracy between 70% and 89%, depending on the target organism. In this study, the “Parasite” target organism was selected with a threshold of 0.5.
Immune simulation
The immune response to TgROP18 was simulated using the C-ImmSim web server. Following a standard clinical protocol, 3 doses of TgROP18 (LPS-free) were administered at 4-week intervals (time points 1, 84, and 168, where each time point equals 8 h). Other parameters included a simulation volume of 10, 1050 simulation steps, a random seed of 12 345, 1000 antigens to inject, and default host Human Leukocyte Antigen (HLA) selection.71,72
Results
General properties of ROP18
The ROP18 protein consists of 554 amino acid residues with a MW of 62 665.51 Da. Its calculated pI is 8.96, indicating that it is more likely to be positively charged under physiological conditions. The protein contains 70 positively charged residues (Arg + Lys) and 63 negatively charged residues (Asp + Glu). The extinction coefficient, measured at 280 nm in water, was 62 255 M–¹ cm–¹. The estimated half-lives of the protein are >30 h in mammalian reticulocytes (in vitro), >10 h in Escherichia coli (in vivo), and 20 h in yeast (in vivo). Based on the ExPASy ProtParam tool, ROP18 was classified as unstable, with an instability index (II) of 42.36. The protein’s hydropathicity, reflected by the GRAVY value, was −0.354, indicating it is relatively hydrophilic, whereas its aliphatic index was calculated to be 83.43, suggesting a moderate level of thermostability.
Prediction of post-translational modification sites in ROP18
Post-translational modification sites in the ROP18 protein were predicted using various online servers. The NetAcet 1.0 tool identified 1 acetylation site, whereas the NetPhos 3.1 tool predicted 47 phosphorylation sites, including 2 on tyrosine (Tyr), 20 on threonine (Thr), and 25 on serine (Ser) residues. The NetNGlyc 1.0 server did not detect any N-glycosylation sites. In total, ROP18 has 48 potential post-translational modification sites (Figure 1A and B).
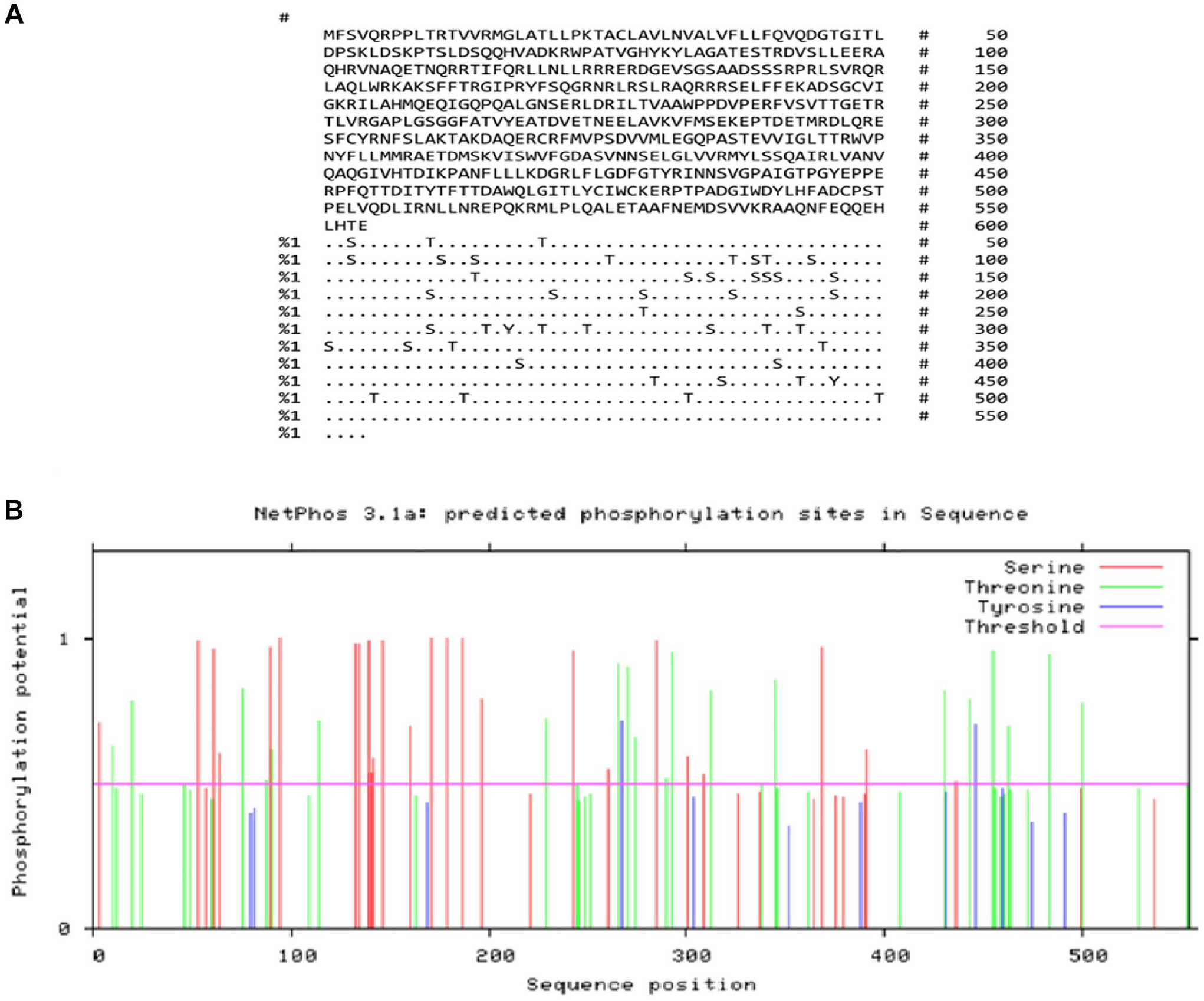
Output from the NetPhos server for predicting phosphorylation sites in TgROP18: (A) Number of predicted sites based on serine (S), threonine (T), and tyrosine (Y) and (B) graphical representation of the predicted phosphorylation sites.
Transmembrane domain and subcellular localization
The TMHMM server v. 2.0 predicted that the ROP18 protein contains a single transmembrane domain (Figure 2A and B). To further explore the subcellular localization, the PSORT II tool was used, suggesting that ROP18 is distributed across several cellular compartments: 11.1% in the mitochondria, 11.1% in the nucleus, 33.3% in the Golgi apparatus, and 44.4% in the endoplasmic reticulum.

Predicted transmembrane domains in the TgROP18 protein: (A) Statistics and a list of the predicted locations of transmembrane helices and intervening loop regions. Length: total length of the protein sequence; number of predicted TMHs: Total number of the predicted transmembrane helices; expected number of AAs in TMHs: The expected number of amino acids in the transmembrane helices. If this number exceeds 18, it is highly likely that the protein is a transmembrane protein or possesses a signal peptide; expected number of AAs in the First 60 AAs: The expected number of amino acids in transmembrane helices within the first 60 amino acids of the protein. If this number is greater than a few, the predicted transmembrane helix at the N-terminus may serve as a signal peptide; total probability of N-terminus orientation: The total probability that the N-terminus is located on the cytoplasmic side of the membrane (source: www.cbs.dtu.dk/services/TMHMM-2.0/TMHMM2.0.guide.html#output). (B) Analysis of the transmembrane domains of TgROP18.
Prediction of the secondary and tertiary structures
The secondary structure of TgROP18 was predicted using the SOPMA tool, which identifies key structural elements such as random coils, extended strands, and alpha helices in proteins. For the 554 amino acid sequence of ROP18, the predicted secondary structure was composed of 42.42% alpha helix (235 residues), 38.45% random coil (213 residues), 14.26% extended strand (79 residues), and 4.87% beta turn (27 residues) (Figure 3A and B).The solvent accessibility of the protein was evaluated using the SCRATCH server, which assessed the hydrophobicity and polarity of the amino acid residues to determine the relative solvent accessibility. Results indicated that accessible residues were distributed across all domains of the ROP18 protein, suggesting favorable solvent-accessibility properties. In addition, the presence of disulfide bonds was predicted using the Scratch Protein Predictor tool. This analysis identified a total of 7 cysteine residues, with 2 disulfide bonds predicted between cysteines at positions 303 to 478 and 320 to 497.
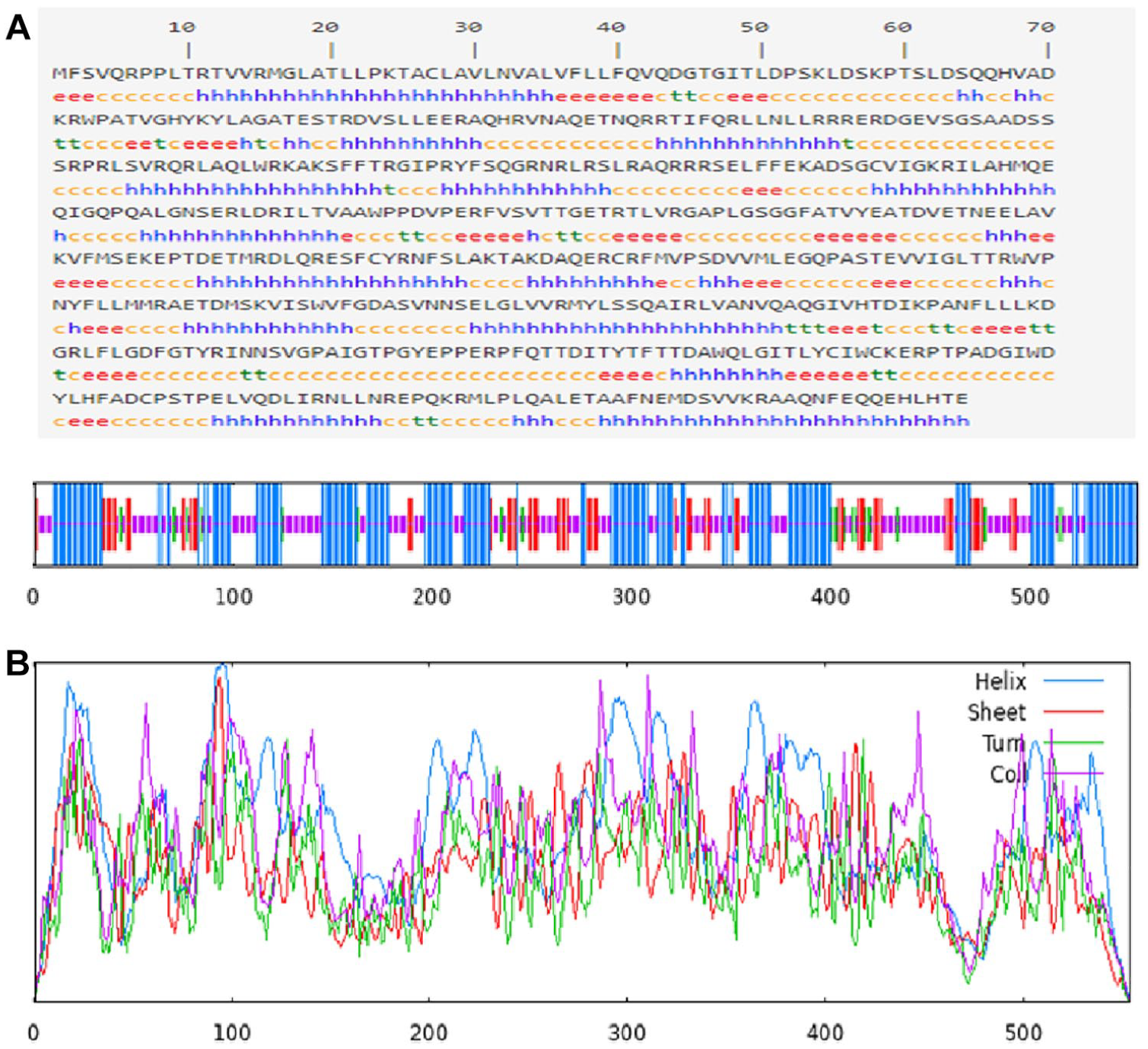
(A) Prediction of the secondary structure of TgROP18 using the SOPMA web server (https://npsa-prabi.ibcp.fr/cgi-bin/npsa_automat.pl?page=/NPSA/npsa_sopma.html). The predicted structural elements include e = extended strand, h = helix, c = coil, and t = turn; (B) Graphical representation of the predicted secondary structure of TgROP18 as generated using the SOPMA server.
To generate the tertiary structure of the ROP18 protein, the SWISS-MODEL server was employed. Based on this prediction, 6 different 3D models were produced, and the model with the highest sequence identity (99.64%) and maximum coverage was selected. The chosen model covered 100% of the protein, from amino acid positions 1 to 554. The final 3D model is presented in Figure 4.
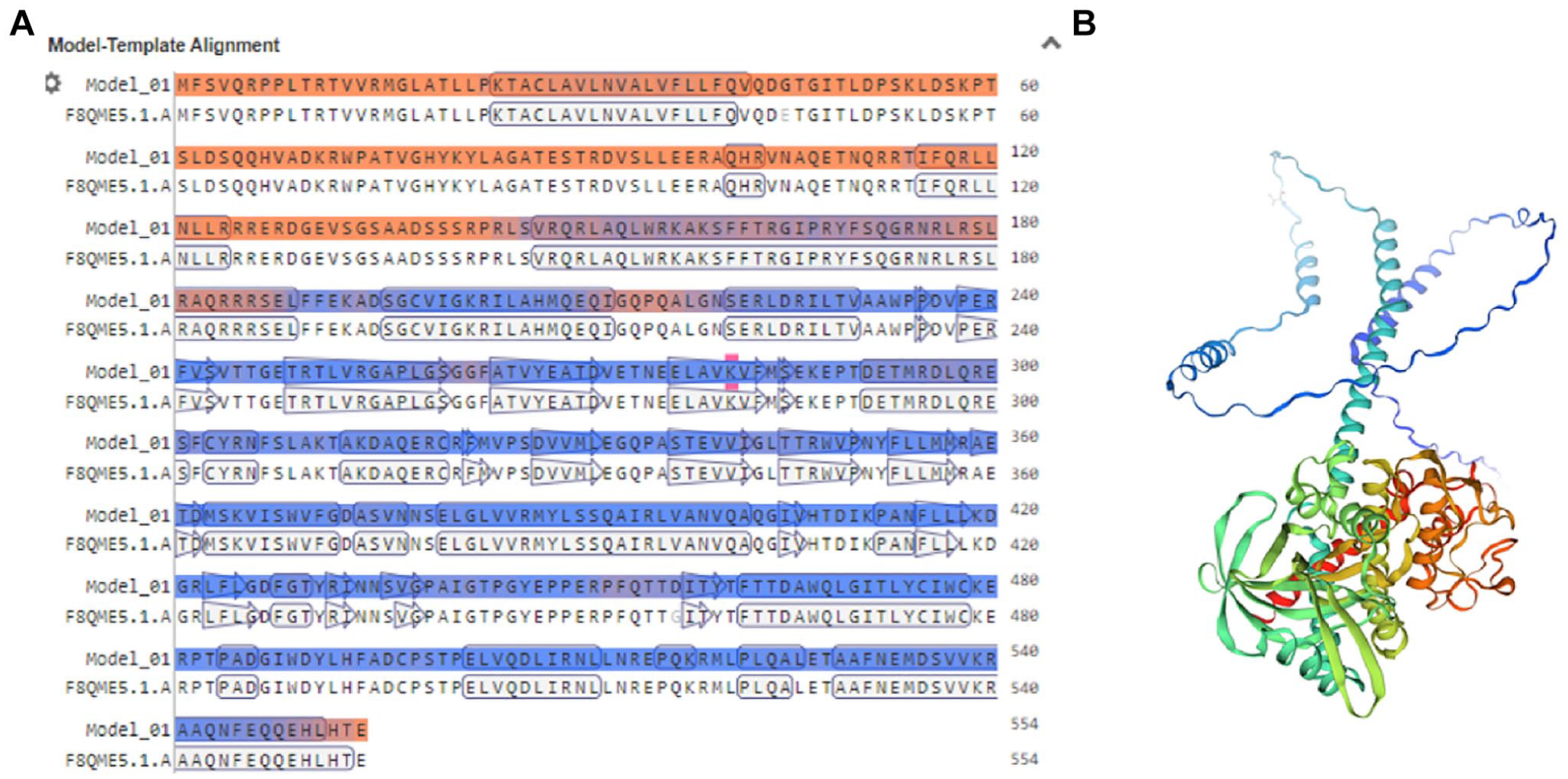
SWISS-MODEL server (https://swissmodel.expasy.org/) output: (A) Model-template alignment and (B) the 3-dimensional structure of TgROP18 protein.
Refinement and validation of the tertiary structure
The 3D structure of TgROP18 was refined using the GalaxyRefine web server to enhance the quality of the predicted model. Postrefinement, the structure was validated using a Ramachandran plot and the ProSA-web tool. These tools assessed the geometry and overall quality of the protein model before and after refinement, with improvements noted in the structural accuracy. The results of the Ramachandran plot and ProSA-web analysis are shown in Figure 5A to D.

Validation of TgROP18 protein tertiary structure. Analysis of the Ramachandran plot using PROCHECK revealed that (A) 85.8% of residues in the crude model and (B) 95.3% in the refined model were located in favored regions. The Z-scores were calculated to be −6.99 for the crude model (C) and −7.25 for the refined model (D), indicating an improvement in the quality of the 3D structure following refinement.
Prediction of B-cell epitopes
Continuous B-cell epitopes of the TgROP18 protein were predicted using the SVMTrip tool. The identified epitopes are ranked by their predicted binding affinity and specificity, with higher threshold scores indicating stronger affinity. These predictions are detailed in Table 1. In addition, the Bcepred server was used to assess several characteristics of continuous B-cell epitopes, including surface exposure, turns, flexibility, hydrophilicity, accessibility, polarity, and antigenic potential. The results are presented in Table 2. These characteristics are critical in defining the antigenic properties of TgROP18. Furthermore, Table 3 provides the predictions from the ABCpred server, which identified B-cell epitopes of 16 amino acids (mer) in length. The server uses a recurrent neural network to assign scores to each epitope, with higher scores (above 0.75) indicating a greater likelihood of the peptide acting as a B-cell epitope.
Linear B-cell epitopes derived from full-length protein using SVMTrip.

The recommended epitopes.
Linear B-cell epitopes predicted using various parameters with the Bcepred server.

Linear B-cell epitopes predicted using the ABCpred online server.

Using the IEDB web tool, the mean threshold scores for several epitope properties were also calculated. These include linear epitope prediction (0.350), hydrophilicity (1.399), surface accessibility (1.000), beta-turns (0.957), flexibility (1.003), and antigenicity (1.027), as shown in Figure 6A to F.
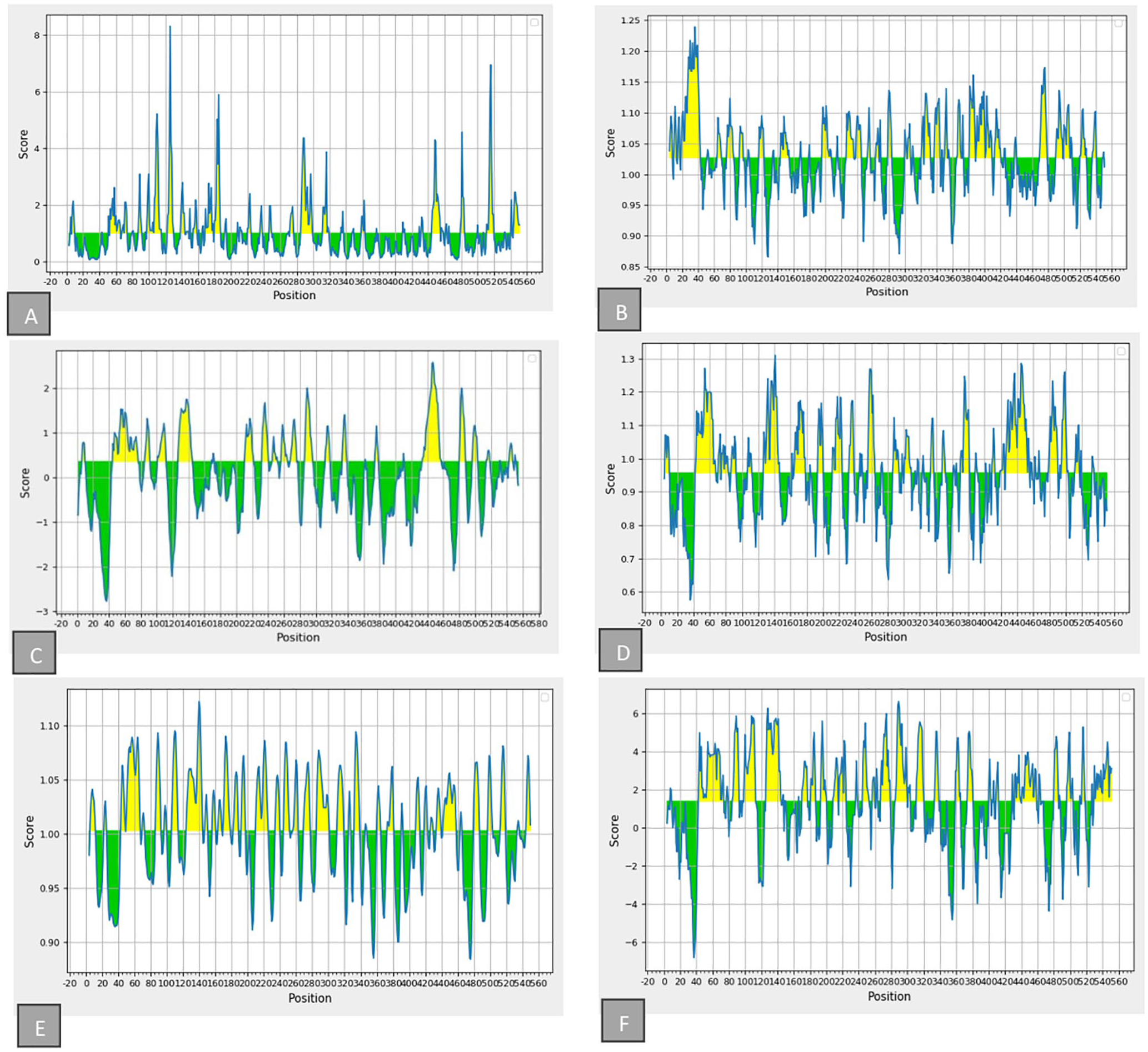
Propensity scale plots of TgROP18: (A) Surface accessibility; (B) antigenicity; (C) bepipred linear epitope; (D) beta-turn; (E) flexibility, and (F) hydrophilicity. The x-axis represents the position, whereas the y-axis indicates the score. Yellow regions (above the threshold) denote favorable areas related to the properties of interest, whereas green regions (below the threshold) indicate unfavorable areas concerning these properties.
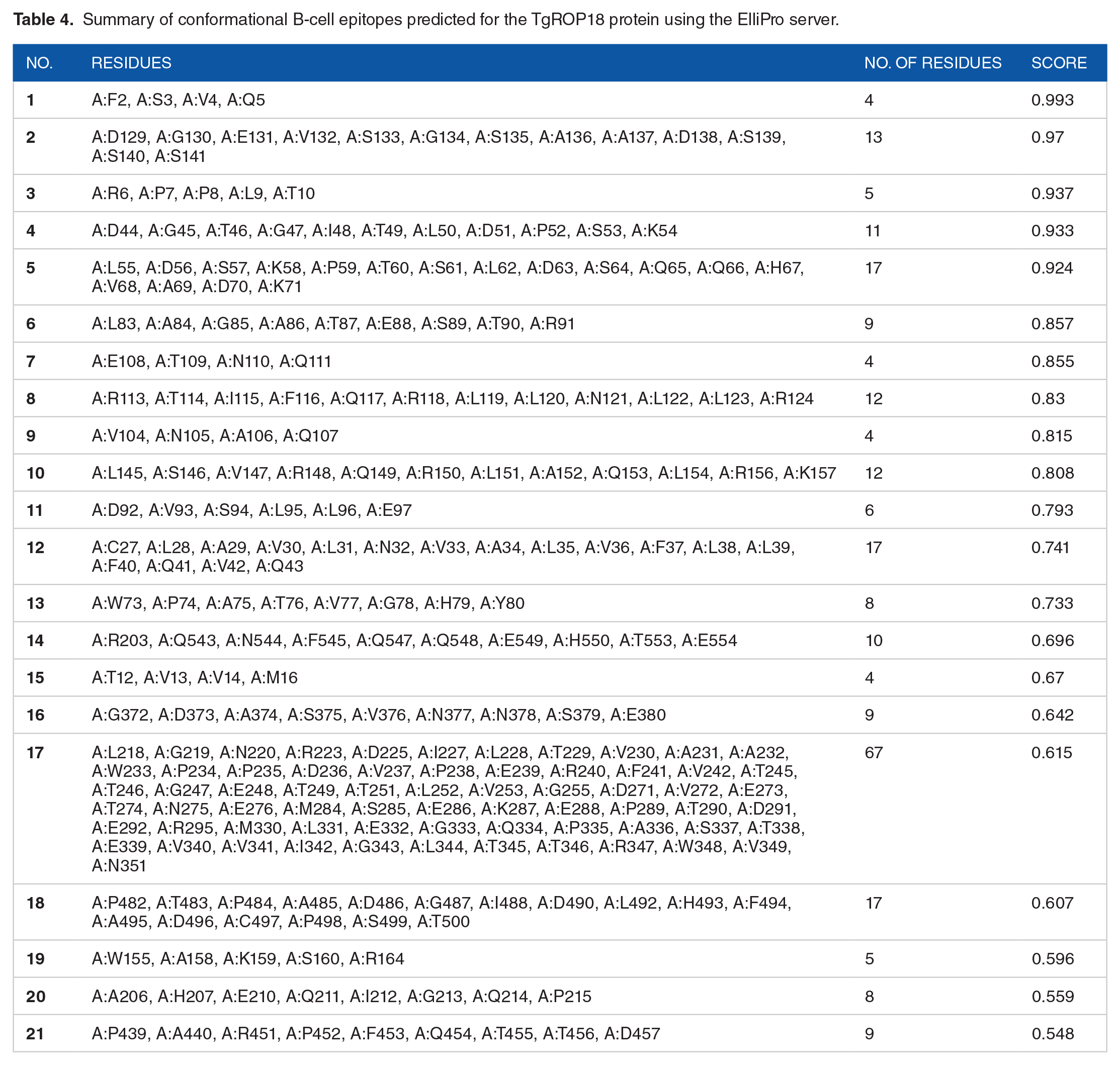
The DiscoTope 2.0 server identified 152 residues as potential B-cell epitope residues out of the 554 amino acids in TgROP18. In addition, 21 discontinuous B-cell epitopes were predicted in the 3D model of TgROP18 using the ElliPro tool (Figure 7 and Table 4).

Predicted discontinuous B-cell epitopes on the 3D structure of TgROP18 protein by ElliPro. The lengths and scores of the epitopes are indicated. The white rods represent the ROP18 protein, whereas the yellow domains highlight the conformational B-cell epitopes.
Summary of conformational B-cell epitopes predicted for the TgROP18 protein using the ElliPro server.
Identification of CTL epitopes
Cytotoxic T lymphocyte epitopes of TgROP18 were predicted using the CTLpred web server, based on SVM and ANN models with cutoff values of 0.36 and 0.51, respectively. The top ten 9-mer CTL epitopes, including CLAVLNVAL27-35, SVRQRLAQL146-154, VLNVALVFL30-38, SELGLVVRM379-387, TRTVVRMGL10-18, YLSSQAIRL388-396, WVPNYFLLM348-356, QRLLNLLRR117-125, WPATVGHYK73-81, and QRRTIFQRL111-119, are listed in Table 5.
Predicted ROP18 epitopes identified by the CTLpred a server.

CTLpred, available at http://www.imtech.res.in/raghava/ctlpred/index.html.
Nine amino acids were included in the analysis.
The default artificial neural network (ANN) and support vector machine (SVM) cutoff scores were set to 0.51 and 0.36, respectively.
Prediction of MHC class I and MHC class II binding epitopes
T-cell epitopes in TgROP18 were also predicted based on their binding affinity to MHC class I and class II molecules, estimated through the IC50 values calculated using the IEDB software. The analysis identified TgROP18 peptides that are predicted to strongly bind MHC molecules, which are essential for antigen presentation. The results, including the minimum percentile ranks for each MHC allele, are summarized in Tables 6 and 7.
Details of the selected MHC-I T-cell epitopes from the TgROP18 protein sequence, as obtained using the IEDB a server.
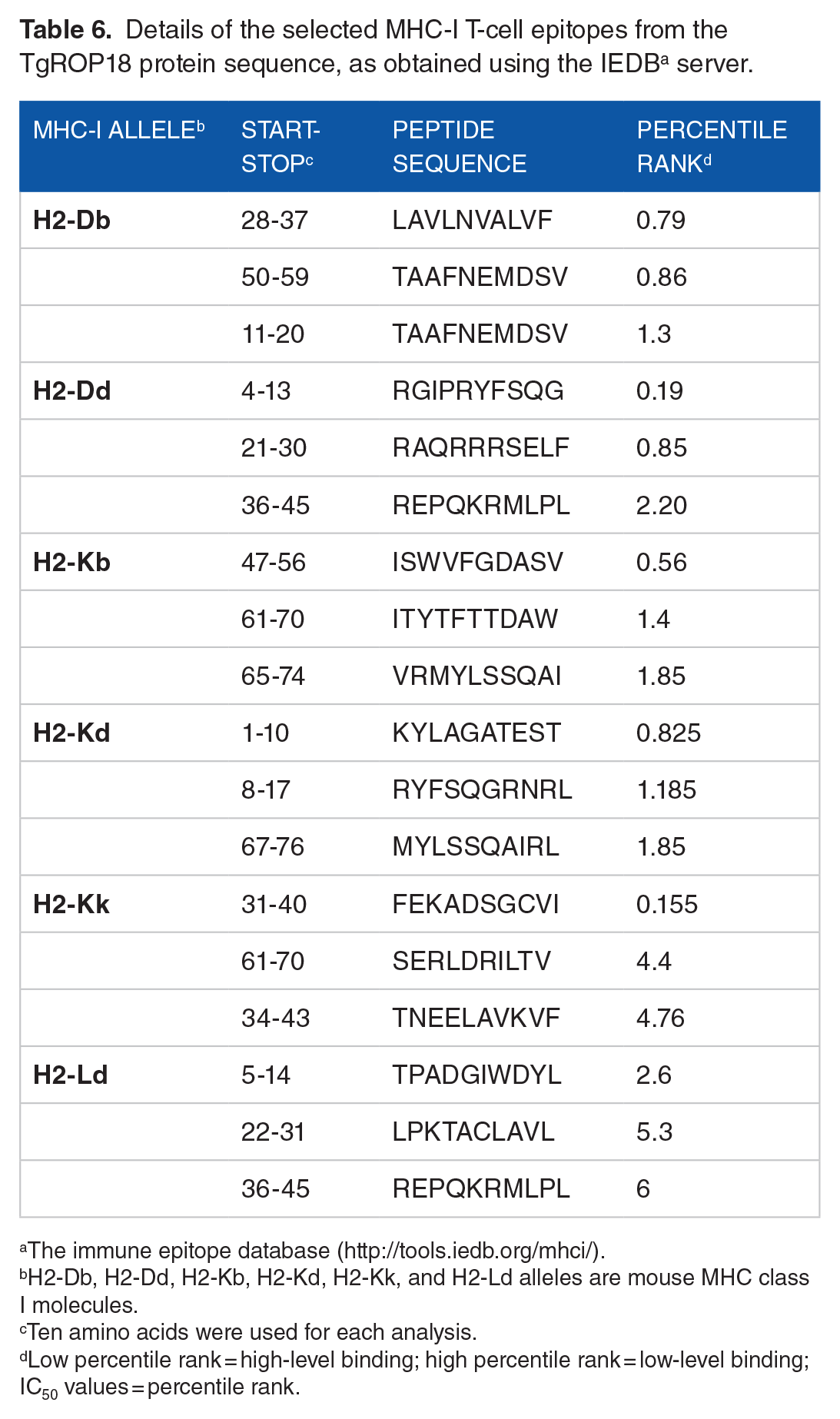
The immune epitope database (http://tools.iedb.org/mhci/).
H2-Db, H2-Dd, H2-Kb, H2-Kd, H2-Kk, and H2-Ld alleles are mouse MHC class I molecules.
Ten amino acids were used for each analysis.
Low percentile rank = high-level binding; high percentile rank = low-level binding; IC50 values = percentile rank.
Details of the selected MHC-II T-cell epitopes from the TgROP18 protein sequence, as obtained using the IEDB a server.

The immune epitope database (http://tools.immuneepitope.org/mhcii).
H2-IAb, H2-IAd, and H2-IEd alleles are mouse MHC class II molecules.
Fifteen amino acids were used for analysis each time.
Low percentile rank = high-level binding; high percentile rank = low-level binding; IC50 values = percentile rank.
Evaluation of solubility, allergenicity, and antigenicity
The solubility of the ROP18 protein after overexpression in E coli was evaluated using the SOLpro server, which predicted a solubility score of 0.8171. In addition, allergenicity predictions based on a hybrid model showed that TgROP18 is nonallergenic. The antigenicity of the ROP18 protein was assessed using 2 different tools: VaxiJen v. 2.0 calculated an antigenicity score of 0.4466 (below the threshold of 0.5), whereas ANTIGENpro gave a higher score of 0.562483, indicating moderate antigenic potential.
Immune simulation profile
The immune response simulation performed using the C-ImmSim server demonstrated that the TgROP18 protein, when administered as a vaccine candidate, elicited strong humoral responses, particularly after the third injection. Notably, high titers of IgM (~120 000), IgG1 (~90 000), and a combination of IgG and IgM (~210 000) were predicted (Figure 8A). In addition, a robust response of Th-associated cytokine IFN-γ (~420 000 ng/mL) was triggered by the TgROP18 protein (Figure 8L).

In silico immune simulation: (A) immunoglobulin production in response to TgROP18; (B, C) B lymphocytes population; (D) plasma B lymphocytes count subdivided per isotype (IgM, IgG1, and IgG2); (E, F) TH cell (CD4+) population; (G, H) TC cell (CD8+) population; (I) dendritic cells per state; (J) macrophage population per state; (K) NK cell population; and (L) cytokine production levels (ng/mL) in response to TgROP18.
Following the administration of TgROP18, the simulation predicted that Th cells began proliferating approximately 5 days postexposure, with active T-CD4+ cells persisting for around 350 days (Figure 8E and F). Similarly, T-CD8+ (cytotoxic T cells) showed an increase in activity for several weeks after exposure (Figure 8G and H). The total number of natural killer (NK) cells increased for approximately 10 days after antigen exposure, contributing to IFN-γ secretion and the elimination of tachyzoite-infected cells (Figure 8K and L). Further details are illustrated in Figures 8A to L.
Discussion
Infection with T. gondii poses a significant threat to public health.1,3,6 To combat toxoplasmosis, several preventive measures have been proposed, with vaccination being one of the most effective solutions. 73 Over the past few decades, numerous studies have focused on developing vaccines to protect against infectious diseases. The initial approach to vaccine production, based on Pasteur’s principles, involved isolating, inactivating, and injecting the pathogen. 74 However, these conventional methods are often labor-intensive, unsafe, expensive, and frequently ineffective.
Advancements in biotechnology have enabled the development of protein and DNA vaccine formulations generated in modified vector organisms, each with unique advantages and disadvantages.75,76 Today, it is possible to accurately identify the immunodominant epitopes of a specific molecule using advanced computer systems, software, or web-based tools. A notable example of this is the successful application of reverse vaccinology techniques for Neisseria meningitidis serogroup B in 2000, which led to the licensure of the MenB vaccine. 77 Since then, various in silico vaccination initiatives have emerged to address cancer and other diseases caused by protozoan agents.78-82
Toxoplasma ROP18 is a highly polymorphic serine-threonine kinase that is secreted into host cells during parasite invasion, regulating the intracellular proliferation of T. gondii.29-31 This protein forms a complex with ROP5, ROP17, and GRA7, facilitating the inactivation of immunity-related GTPases (IRGs) through phosphorylation. In addition, ROP18 can inhibit NF-κB signaling and parasite control pathways that are dependent on tumor necrosis factor (TNF) and CD-40, in association with the parasite’s excretory-secretory antigens. 83
In this study, we employed various bioinformatics techniques to characterize the specific features of the ROP18 protein. The amino acid sequence of ROP18 comprises 554 residues, with an average MW of 62 665.51 Da, indicating favorable immunogenic properties. 84 Using the ExPASy ProtParam tool, we determined that the GRAVY and aliphatic index of the ROP18 sequence are −0.354 and 83.43, respectively. A negative GRAVY value signifies that the protein is hydrophilic, which allows for greater interaction with surrounding water molecules. A high aliphatic index suggests that the target protein is stable across a broad temperature range. According to El Hajj et al., 24 ROP proteins feature an N-terminal signal sequence containing a transmembrane domain and a C-terminal hydrophobic motif. In this study, we predicted the presence of 1 transmembrane domain in the ROP18 sequence.
Proteins undergo various enzymatic modifications, such as glycosylation, acylation, and phosphorylation, collectively known as post-translational modifications (PTMs). These modifications are essential for multiple functions, including membrane anchoring, signal transduction, altering the protein’s half-life, and other regulatory roles.85,86 Predicting PTM sites is important for selecting the appropriate host to ensure effective expression of recombinant proteins. 87 We used the NetAcet 1.0 and NetPhos 3.1 servers to predict the N-acetylation and phosphorylation sites of the ROP18 protein, respectively. Our findings indicated that TgROP18 contains 48 putative PTM sites, comprising 1 acetylation site and 47 phosphorylation sites, suggesting that these modifications may regulate specific protein activities.
We employed the SOPMA tool to estimate the secondary structure of the ROP18 protein. The results showed that TgROP18 comprises 4.8% beta-turns, 14.26% extended strands, 42.42% alpha helices, and 38.45% random coils. The internal alpha and beta helices of proteins, characterized by high hydrogen bond energies, are crucial for maintaining structural integrity and establishing strong interactions with antibodies. 45 The spatial structure of proteins is widely recognized as a determinant of their biological activity. Therefore, analyzing protein structure is vital for predicting tertiary structures and understanding the connections between structure and function.41,45 We used the online SWISS-MODEL tool to predict and generate the 3D structure of the TgROP18 protein. To evaluate the generated 3D model, we used PROCHECK to create a Ramachandran plot. Following the selection of the best model through SWISS-MODEL, we refined it using the GalaxyRefine server. Initially, 85.8% of amino acid residues were located in favored regions of the Ramachandran plot, indicating energetically favorable conformations for the protein structure. After refinement, this percentage improved to 95.3%.
During T. gondii infection, strong cell-mediated and humoral immune responses are induced.88,89 Specific IgG antibodies must be produced to restrict parasite attachment to host cell receptors. These antibodies also facilitate the rapid clearance of T. gondii by immune cells, such as macrophages, helping to prevent reactivation of the infection. 89 In addition, the production of IFN-γ by T cells is a crucial factor limiting both acute and long-term infections. This important cytokine prevents the reactivation of tissue cysts. Notably, both T-CD4+ and T-CD8+ cells play a role in limiting infection, although IFN-γ and T-CD8+ cells are more critical in controlling disease.88,90 Determining protein antigenicity and specificity through epitope analysis is essential for understanding target antigen structures and activities. Furthermore, epitope analysis can shed light on the pathogenesis and immune mechanisms of pathogens, aiding in epitope-based vaccine development via reverse vaccinology.41,77,91
The prediction of epitopes primarily relies on several protein structural features. Analyzing a single index alone does not provide a comprehensive understanding of the target epitope; therefore, various important indices are available for epitope prediction, including antigenicity, flexibility, hydrophilicity, and surface accessibility.59-63 A peptide that performs well in any of these indices is likely to interact with antibodies and act as an epitope. Consequently, we used several web-based servers to predict T- and B-cell epitopes in this study. Linear B-cell epitope analysis using the Bcepred, SVMTrip, ABCpred, and Bepipred servers revealed that the ROP18 protein possesses positive epitopes with acceptable indices. We also used the CTLpred server to predict CTL epitopes and selected the top 10 epitopes for TgROP18. CTLpred employs sophisticated machine learning algorithms, such as SVM and ANN, to predict CTL epitopes that are critical for vaccine design. Moreover, the IEDB online server was used to estimate the IC50 values of the peptides binding to both MHC-I and MHC-II. The results indicated that T lymphocyte epitopes on TgROP18 exhibit strong binding to MHC molecules. Importantly, lower IC50 values (or percentile ranks) correspond to better T-cell epitopes, suggesting higher affinity. T-cell-mediated cellular immunity is vital for restricting toxoplasmosis, as T. gondii is an obligatory intracellular parasite. 88 Therefore, identifying the type of T-cell-mediated immune response involved is essential for developing effective vaccine candidates. 88 Previous studies have shown that in immunized mice, ROPs can induce long-lasting immunity along with both humoral and cellular immune responses, leading to prolonged survival.15,17,22 Notably, the allergenicity and antigenicity analyses indicated that ROP18 is a non-allergen and an antigenic protein.
In addition, B-cell epitopes of the antigen were predicted using the ABCpred web server, and the first server developed based on a recurrent neural network with fixed-length patterns. This tool can predict epitopes with 65.93% accuracy using a recurrent neural network. 58 Identifying conformational B-cell epitopes essential for antibody-antigen interactions is another critical step in in silico analysis. Based on the findings from the DiscoTope 2.0 and ElliPro servers, we identified 152 and 21 discontinuous epitopes for B lymphocytes, respectively. Our findings indicate that several epitopes can simultaneously elicit both humoral and cellular immune responses. These include VPNYFL349-354, RAQRRRSELF181-190, AQLWRKAKS152-160, RGIPRYFSQG164-173, DGIWDYL486-492, ISWVFGDASV367-376, TAAFNEMDSV528-537, and GLVVRMYLSSQAIR382-395. These epitopes may serve as focal points for future multi-epitope vaccine design studies.
Our results suggest that the ROP18 protein can elicit robust humoral and cell-mediated immune responses, as evidenced by the antibody titers and cytokine levels induced by the administration of TgROP18, which were predicted using the C-ImmSim web server. C-ImmSim functions as a collection of models within a single program, incorporating “key facts” from current immunologic knowledge, including the diversity of specific elements, MHC restriction, and antibody hypermutation. This tool was developed to create a digital system capable of conducting experiments analogous to real-life laboratory tests, assisting biologists in testing theories and ideas about immune system functioning both in vivo and in a controlled environment. 71
Although our study was conducted solely through in silico methods, previous research on this gene has indicated that ROPs can enhance the survival time or rate in mice. For example, a DNA vaccine encoding pROP18-MIC3 and ROP18 demonstrated that mice immunized with pROP18-MIC3 exhibited stronger humoral and Th1-mediated responses, as well as prolonged survival against toxoplasmosis. 37 Another study involving GRA7 and ROP1 of T. gondii found higher IgG2a levels, increased production of IFN-γ and TNF-α, longer survival time, and a greater reduction in cysts. 92 Numerous articles have explored ROP-based vaccines using a combination of bioinformatics platforms and various programs to predict potential B- and T-cell epitopes, offering promising vaccine options for preventing toxoplasmosis. However, evidence supporting the protective efficacy of some of these vaccines in animal studies is still lacking.28,45,93-101 Past studies have often faced limitations in their methodologies such as lack of comprehensive bioinformatics analyses for the target protein.38,102 This study comprehensively examined nearly all essential indicators, including physicochemical properties, antigenicity, solubility, allergenicity, subcellular localization, transmembrane domains, tertiary and secondary structures, potential T- and B-cell epitopes, and virtual immune simulation. This thorough analysis can aid in developing an effective vaccine against T. gondii. T. gondii employs a wide array of virulence factors to evade the host’s immune response and survive within the host. Consequently, creating a vaccine candidate targeting a specific antigenic protein may not provide robust protection due to the parasite’s immune evasion strategies, including the inactivation of IRGs, GTPase-dependent antimicrobial systems, resistance to oxidative stress via peroxiredoxins (TgPRXs), and regulation of host gene expression. 83 Ultimately, the B- and T-cell-based epitopes predicted in this study can be considered immunodominant regions for future multi-epitope vaccine constructs.
In this study, we used various bioinformatics online servers and tools to predict the structure, immunogenic epitopes, and other characteristics of the TgROP18 protein. Our research is centered on computational predictions, and we recognize that environmental factors can influence the expression and functionality of proteins. Future studies could investigate these aspects through in vitro and/or in vivo experiments to provide a more comprehensive understanding of ROP18. Besides, this study is primarily focused on the in silico prediction of immunodominant epitopes of TgROP18. As such, it does not involve the engineering of a multi-epitope vaccine candidate or subsequent validations through immune sensitization. Therefore, assessments of immune pathway sensitization, both in silico and in vivo, were not conducted. We acknowledge the importance of these validations for a comprehensive understanding of immune responses and suggest that future work should address this aspect.
Conclusions
This study provides a comprehensive analysis of the subcellular localization, transmembrane domain, physicochemical parameters, secondary and tertiary structures, and potential T-cell and B-cell epitopes of ROP18. TgROP18 has been shown to possess a nonallergenic and immunogenic nature. In addition, the results from epitope prediction using various bioinformatics resources indicated that the TgROP18 protein contains several promising T- and B-cell epitopes, suggesting its potential utility in developing an effective T. gondii vaccine. Given the challenges associated with treating toxoplasmosis in endemic communities, the findings of this study could offer a solution to the limitations of the current toxoplasmosis medications.
Limitations
This study only involved in silico analysis of TgROP18. Future research should incorporate wet lab experiments to validate these findings and establish a more comprehensive understanding by correlating the identified epitopes with immune responses both in vitro and in vivo. Furthermore, combining immunodominant B- and T-cell epitopes of TgROP18 with appropriate genetic adjuvants would be advantageous.
Supplemental Material
sj-docx-1-bbi-10.1177_11779322251315924 – Supplemental material for In Silico Analysis and Characterization of the Immunogenicity of Toxoplasma gondii Rhoptry Protein 18
Supplemental material, sj-docx-1-bbi-10.1177_11779322251315924 for In Silico Analysis and Characterization of the Immunogenicity of Toxoplasma gondii Rhoptry Protein 18 by Masoud Foroutan, Amir Karimipour-Saryazdi, Ali Dalir Ghaffari, Hamidreza Majidiani, Arezo Arzani Birgani, Elaheh Karimzadeh-Soureshjani, Shahrzad Soltani and Hany M Elsheikha in Bioinformatics and Biology Insights
Footnotes
Declaration of conflicting interest:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Abadan University of Medical Sciences, Abadan, Iran, Grant/Award Number: 1607.
Author contributions
Ethics approval and consent to participate
This study received ethical approval from the Abadan University of Medical Sciences Research Ethics Committee (
Data availability
The data that support the findings of this study are available in the supplemental material of this article.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
