Abstract
Background:
The ability to predict and comprehend molecular interactions offers significant insights into the biological functions of proteins. The interaction between surface protein 47 of
Methods:
Public sequences of the
Results:
The examination of motifs reveals a significant degree of conservation in
Conclusions:
The study identifies coevolutionary patterns between P47Rec and Pfs47 related to the speciation and geographic dispersion of
Background
Recent studies have provided an in-depth understanding of the relationship between the protein Pfs47 of
Pfs47 is a member of the 6-cysteine (6-Cys) protein family.
11
The gene
In a recent study, Molina-Cruz et al employed a comprehensive biochemical approach to identify and elucidate the characteristics of the Pfs47 receptor (P47Rec). Their findings revealed that this is a highly conserved protein consisting of 2 natterin-like domains present in the midgut cells of anopheline mosquitoes.
8
The
This study aimed to investigate the interaction between Pfs47 and P47Rec proteins to gain insights into the immune evasion mechanisms employed by
Methods
Sequencing of p47rec gene of Anopheles albimanus and Anopheles darlingi from Honduras
Several sets of primers were designed to amplify partial fragments of the
Sequences, amplicon size and annealing temperature of the primers used to amplify the

Pipeline
A pipeline to generate predictions at multiple levels of functional and structural annotations from nucleotide sequences of the genes that encode the proteins of the Pfs47-P47Rec complex was developed. The putative molecular interactions and evolutionary processes involved in the emergence and speciation of the 2 proteins of the complex were predicted (Figure 1). As an initial strategy, public sequences of

Pipeline used to generate functional and structural annotation predictions of the Pfs47-P47Rec complex. (A) Data mining and development of data sets; (B) data filtering; (C) selection of unique sequences; (D) evolutionary and phylogenetic analysis; (E) pattern recognition and primary protein structure prediction; (F) protein secondary structure prediction and mutations; (G) protein domain search; (H) identification of antigenic sites; (I) prediction of folding proteins; and (J) protein-protein interaction models.
Data mining and filtering
A data mining approach based on global multiple alignments of reference sequences was used. Reference sequences were used as ‘baits’ to identify and select sequences with a high level of similarity (Table 2). The sequences were selected and stored in 2 separate files, 1 for each of the 2 proteins of the complex. These sequences formed the initial data sets that were used in subsequent analyses.
List of
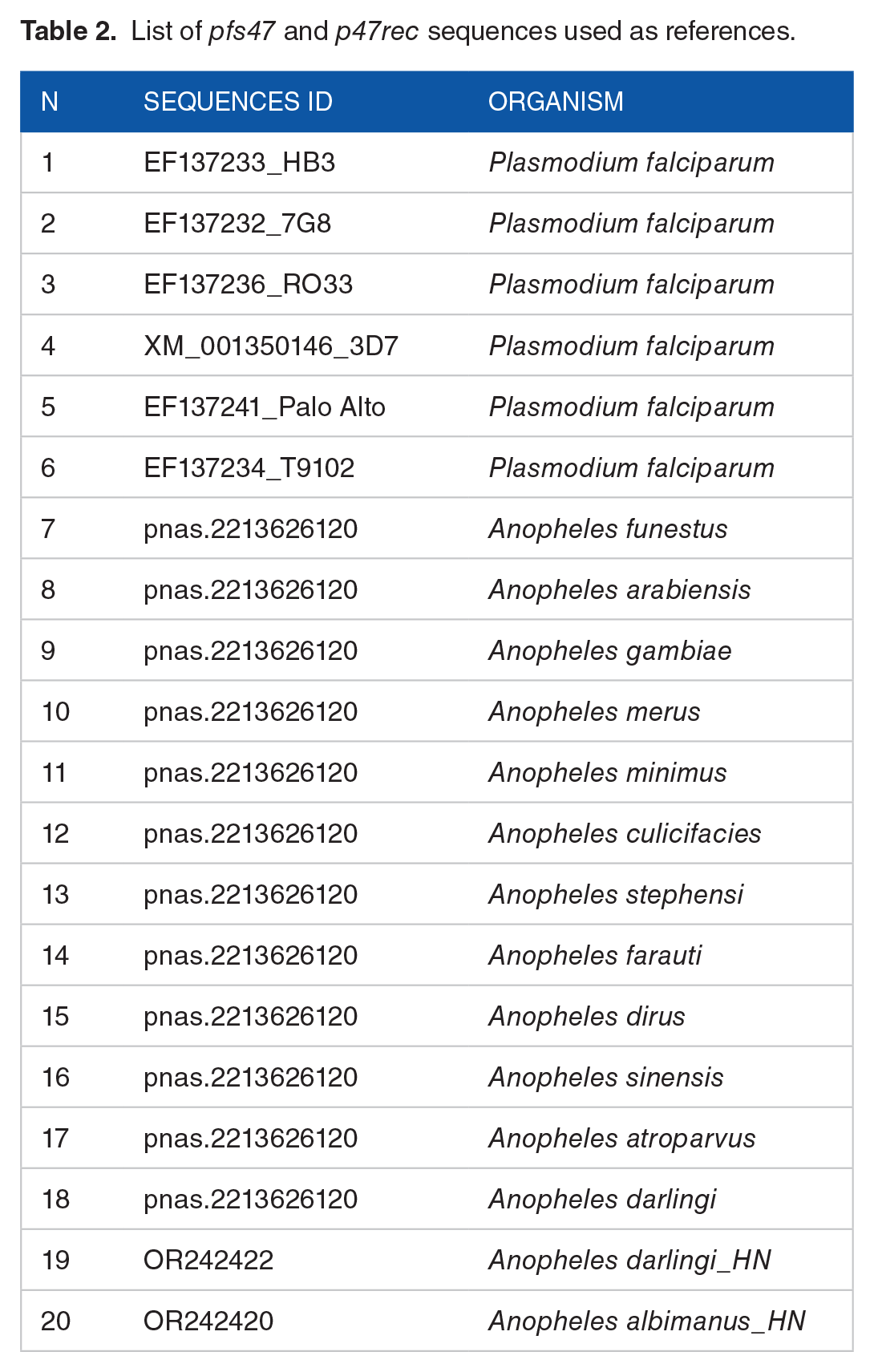
Incomplete, fragmented, redundant or repeated sequences were filtered and deleted from the original data set. Filtered sequences were aligned with the reference sequences in AliView v.1.28 for Linux. 16 The resulting alignments were subjected to haplotype analysis using the R Haplotype package 17 to discard redundant sequences that had not been eliminated by the first filter, keeping unique sequences in the following steps. The resulting sequences were translated into amino acids using the ExPASy Translate Tool 18 and were subsequently aligned to identify and filter haplotypes with synonymous translations. Only reference sequences were retained for subsequent analyses, except for situations where reference sequences existed in haplotypes with multiple sequences. In other cases, a random sequence of each haplotype with multiple sequences was stored. Translated sequences were aligned again, which allowed filtering out the sequences that translated synonymous proteins.
Evolutionary and phylogenetic analysis
Phylogenetic trees were created using the amino acid sequences obtained in the previous analysis through the advanced mode of the Phylogeny.fr server. Trees were generated using sequences aligned with MUSCLE 19 in its full mode. Curation was performed with Gblock 20 (using the analysis model that prevents many contiguous positions from requiring consensus, using the server). For evolutionary inference and distance calculation, the maximum likelihood method with 100 bootstraps and the Jones-Taylor-Thorton (JTT) amino acid substitution model were selected. Finally, the results were visualized and edited in Figtree v.1.3.0 for Linux. Motif pattern identification analyses were performed using the MEME-Suite motif search function v.5.5.3 for Linux, 21 to evaluate the general architecture and infer the possible secondary structures of the proteins.
Pattern recognition and primary protein structure prediction
In addition to the search for motifs, an analysis was carried out to check whether the patterns could be related to the co-adaptation processes of both proteins of the complex, as well as their possible geographical relationship. JalView v. 2.11.2.6 for Linux was used to predict protein secondary structures using the default parameters for the JPred secondary structure predictor. 22 A comparison of the secondary protein structure of the Pfs47 and P47Rec sequences was performed by aligning the proteins in JPred. This analysis sought to determine the likelihood that the mutations could alter the secondary structure of the polypeptide at the places where they are present and thus impact the folding of the proteins.
Secondary and tertiary structure predictions and functional prediction of domains
SignalP v5.0 for Linux 23 was used to analyse the individual sequences of the Pfs47 and P47Rec proteins, to detect signal peptide-like domains. Simultaneously, the sequences were subjected to transmembrane domain identification analysis with TMHMM v2.0 software for Linux. 24 This was carried out to identify the presence of tertiary structures previously described for Pfs47.13,25 In addition, both data sets were subjected to analysis to reveal structures or domains not previously described. Subsequently, the sequences were subjected to tertiary structure prediction using the default settings of AlphaFold software v.2.3.2, 26 which can be accessed through the Colab Research Server of Google.
The tertiary structures were subjected to quality control in the web version of MolProbity software. 27 The predicted angles were evaluated using Ramachandran diagrams, to demonstrate the biological plausibility of the tertiary structures. The structures were modelled using BepiPred v3.0 software in its web version. 28 Antigenic sites with a greater probability of generating epitopes and, therefore, a greater probability of protein-protein interaction were identified.
Protein-protein interaction models
Tertiary prediction results were viewed and edited in UCSF-Chimera X v.1.6.1 for Linux.
29
Finally, 6 Pfs47 structures were subjected to docking analysis alongside 15 P47Rec structures. The
To evaluate the results of the docking simulations, a comparison model was defined based on the following parameters: The final comparisons had to involve at least 2 models as an interaction reference. One of the models was in all cases the consensus protein P47Rec interacting with each Pfs47 receptor. On the contrary, the second reference model used was selected based on experimental evidence of the interactions between the proteins of the complex. This model was called the ‘ideal interaction model’ (IIM). For interaction models of parasite strains originating from Africa (RO33, 3D7 and Palo Alto), the IIM was ‘Pfs47 –
Results
Sequencing of the p47rec gene of Anopheles albimanus and Anopheles darlingi from Honduras
Four sequences (two from each species) of the
Data mining and data set development
The data mining strategy generated a total of 128 and 134 nucleotide sequences for
Evolutionary and phylogenetic analysis
The phylogenetic tree obtained for Pfs47 showed 3 distinct clusters. A first cluster named African

(A) Phylogenetic tree for Pfs47 showing 3 clusters. Cluster 1: African
On the contrary, the phylogenetic tree of P47Rec showed a distribution of clusters that corresponded to the geographical distribution of the anopheline species studied (Figure 2). The analysis identified 5 distinct clusters based on P47Rec: (1) an African cluster, where the consensus sequence of P47Rec was found, (2) an African-Asian cluster and (3) a Eurasian cluster, and 2 clusters (4 and 5) with species from the Americas. Sequences of 3 species of African origin were identified within the African-Asian cluster (
Identification of motif patterns and prediction of primary structures
All Pfs47 sequences displayed the same pattern, which was composed of 11 conserved motifs, except for the EF137234_T9/102 reference sequence, which displayed a distinct pattern at the N-terminus. The missing region in EF137234_T9/102 was later found to be located within the signal peptide domain (Figure 3). Although 11 motifs made up the basic pattern of Pfs47, the analysis identified a total of 14 different motifs, each with at least 1 potential variant (Additional Material 3).

(A) General pattern of amino acid motifs of the Pfs47 protein from
P47Rec, on the contrary, exhibited 10 different motif patterns and 15 conserved motifs. Only 1 motif was different between the number of identified motifs and conserved motifs. Pattern 1 was the most frequent (24/36 sequences) in P47Rec sequences from all continents (Figure 3) (Additional Material 4). Pattern 3 was the second most frequent pattern (3/26), and the 3 sequences included within this pattern presented geographical association. All sequences with pattern 3 belong to American anophelines and included sequences from
Prediction of secondary structures and identification of mutations
The predictions of the secondary conformation of Pfs47 revealed a basic structure made up of 11 alpha helices and 22 beta sheets, arranged as depicted in Figure 4. The N-terminal region of Pfs47 revealed a signal peptide-like domain between amino acids 1 and 18, as well as a cleavage site between amino acids 18 and 19. The secondary structure of the proteins also revealed a putative GPI-type anchoring site and a transmembrane domain close to the C-terminus. Ninety-one mutation sites or 21% of the amino acids that make up all proteins were found. The transmembrane domains and the signal peptide were unaffected, and the mutations were limited to 6 locations in the protein’s central globular domain. Substitutions involving valine (V), leucine (L), phenylalanine (F), methionine (M) or isoleucine (I) occurred in 24 of the 91 mutations (26%) (Additional Figure 1). Furthermore, glutamic acid (E) was replaced by glutamine (Q), aspartic acid (D) or lysine (K) in 15 of the 91 mutations (16%). The remaining Pfs47 mutations were not associated with commonly observed substitution patterns.

Secondary structure predictions of (A) Pfs47 and (B) P47Rec. Red cylinders visually represent Alpha helices, beta sheets are denoted by green rectangles, and loops are depicted as grey lines. Asterisks show the location of the amino acid substitutions that had the greatest effects on secondary structure predictions; while the dotted boxes show the regions that underwent significant structural changes.
The secondary structure of P47Rec was predicted, revealing a minimum of 20 beta sheets (Figure 4). Moreover, it was predicted that P47Rec lacks transmembrane domains, signal peptides and GPI anchor sites. Even so, the alignments showed proteins exhibiting conserved secondary structures and mutations that predominantly impact amino acids possessing chemically similar characteristics (V, L, F, M and I). A total of 47 mutations were identified in the secondary structures of P47Rec, which represent approximately 16% of the amino acids. The remaining mutations did not exhibit discernible substitution patterns, except for those involving hydrophobic amino acids (E, Q, D and K) (Additional Figure 2).
The Pfs47 consensus sequence was subjected to secondary structure prediction, which revealed 3 domains previously described. 25 The composition of this structure consisted of 3 alpha helices and 24 beta sheets (Figure 4). These findings show agreement between the Pfs47 consensus sequence, and the secondary structure prediction calculated by Jpred, whereas the secondary structure prediction of the P47Rec consensus sequence revealed an even distribution of 28 beta sheets throughout 2 protein domains (Figure 4). Moreover, the analysis of P47Rec did not reveal any evidence of signal peptides, transmembrane domains or signalling and membrane anchoring structures.
Prediction of tertiary structures and identification of antigenic sites
To predict the tertiary structure of Pfs47, the removal of the signal peptide from the sequences was performed by the cleavage site. This process was conducted by manually removing the signal peptide sequences from the unique sequences subjected to structural prediction. Tertiary structures unveiled a protein with 2 distinct domains. The first domain was a transmembrane domain, characterized by a helical structure consisting of 20 amino acids. The second domain was globular, taking the form of an equilateral triangle. This domain comprises 3 substructures arranged in a sandwich-like manner, with each substructure composed of an average of 8 antiparallel beta sheets (Figure 5). The globular domain of Pfs47 exhibits sandwich-like structures at both its C- and N-terminal ends. These structures belong to the 6-Cys family and contain 3 disulfide bonds in the central region (Additional Figure 3). On the contrary, the central substructure did not exhibit any discernible functional annotation (Additional Figure 4). Nonetheless, it is notable that the central substructure has 2 cysteines in proximity. However, it appears that these cysteines do not correlate directly with the substructure’s rigidity (Additional Figure 4). According to our substitution models, the region containing the cysteine residues in the core substructure did not exhibit notable changes when substituted with glycine. This suggests that the cysteines may not contribute to the rigidity of the subunit, implying that this region might have a role in the specificity of the interaction with P47rec.

Tertiary structure predictions of (A) Pfs47 and (B) P47Rec. Row 1 of each model presents the cartoon structure. Row 2 shows the volumetric models. Row 3 presents the electrostatic forces. Row 4 shows the hydrophobicity. Columns B, C and D present the models with rotations of 90°C in the
The tertiary structure of the Pfs47 consensus sequence exhibited regions on the upper surface of the globular domain that are less electrostatic towards the apex and more electrostatic towards the base (relative to the triangular structure of the domain) (Figure 5). In contrast, the lower face of the globular domain exhibited fewer electrostatic areas at the ends, while displaying highly electrostatic regions in the central region of the domain. Electrostatic amino acids predominated in 3 of the 4 lateral faces of the protein. Moreover, the transmembrane domain of Pfs47 predominantly comprises lipophilic regions, which is consistent with the predicted characteristics of this domain. In contrast, the globular domain exhibits predominantly hydrophilic regions throughout its surface (Figure 5).
The findings derived from the analysis of the Ramachandran plots showed few improbable angles, and in most cases, these represented less than 1% of the total angles between amino acids. These angles were generally located near the inclusion zones in the plots in the consensus sequences (Figure 6). The consensus protein P47Rec revealed a tertiary structure that simulates a dumbbell (Figure 6) and has 2 globular domains of the DM9 repeat type (Additional Figure 3), separated by an interdomain loop (Figure 5). Only a small portion of P47Rec showed electrostatic potential, while the rest of the protein surface exhibited regions with low electrostatic potential. A considerable proportion of the protein exhibits regions with hydrophilic properties; however, the hydrophobicity pattern does not reveal notably hydrophobic segments (Figure 5).

(A and C) Location of cysteine residues in the structures of the Pfs47 and P47Rec consensus proteins. (B and D) Ramachandran plots of the consensus proteins of Pfs47 and P47Rec.
Pfs47 showed 18 antigenic sites, among which the sites designated II, III, VI, X, XI, XI and XVI had the greatest abundance of amino acids (Additional Figure 5). The antigenic sites were later identified and defined inside the tertiary structures that were constructed using Alphafold. The visualization of these data was performed using UCSF-ChimeraX. When comparing the Pfs47 consensus structure to the triangle structure, it was observed that the antigenic areas were predominantly situated at the lower and upper right vertices (Additional Figure 6). In contrast, the P47Rec sequences exhibited a total of 10 antigenic regions, which were evenly distributed across the N- and C-terminal domains. Notably, regions I, IV and VII encompassed the highest number of amino acids (Additional Figure 5). As in the case of Pfs47, not all antigenic sites were located on the surface of the folded protein. For the P47Rec consensus, a region close to the interdomain loop with 3 antigenic sites was observed (Additional Figure 6). The N-terminal domain of the P47Rec consensus protein exhibited a greater number of antigenic sites on its front face compared with the C-terminal domain, which had only a small number of antigenic sites.
Pfs47-P47Rec interaction models
A total of 8400 PDB files containing the top 100 best molecular docking models of the Pfs47-P47Rec complex were analysed. All generated models were filtered, selecting only those models that presented RMSD less than 40 Å. Then, the best models of each docking were displayed, and only those interaction models that involved the upper face of Pfs47 were filtered. A total of 686 interactions were chosen for further analysis. In the absence of an existing method to assess this particular analytical approach, we aimed to develop evaluation criteria that would enable an objective assessment of the generated models. The evaluation focused on specific metrics, including interaction distance, similarity with the consensus model and IIM, and the geographical relationships between the Pfs47-P47rec complexes and their relationship with the specificity of each possible model. The overall average RMSD of the selected models was 35.92 Å (Additional Material 5). The models exhibiting the lowest RMSD for each docking process were chosen. These selected models were then visually examined and compared with the remaining models for each Pfs47 receptor (Figure 7). For the Pfs47-RO33 interaction, the reference models showed diametrically different positions, while the consensus model showed P47Rec located in the centre of the upper face of Pfs47 and tilted to the left. P47Rec from
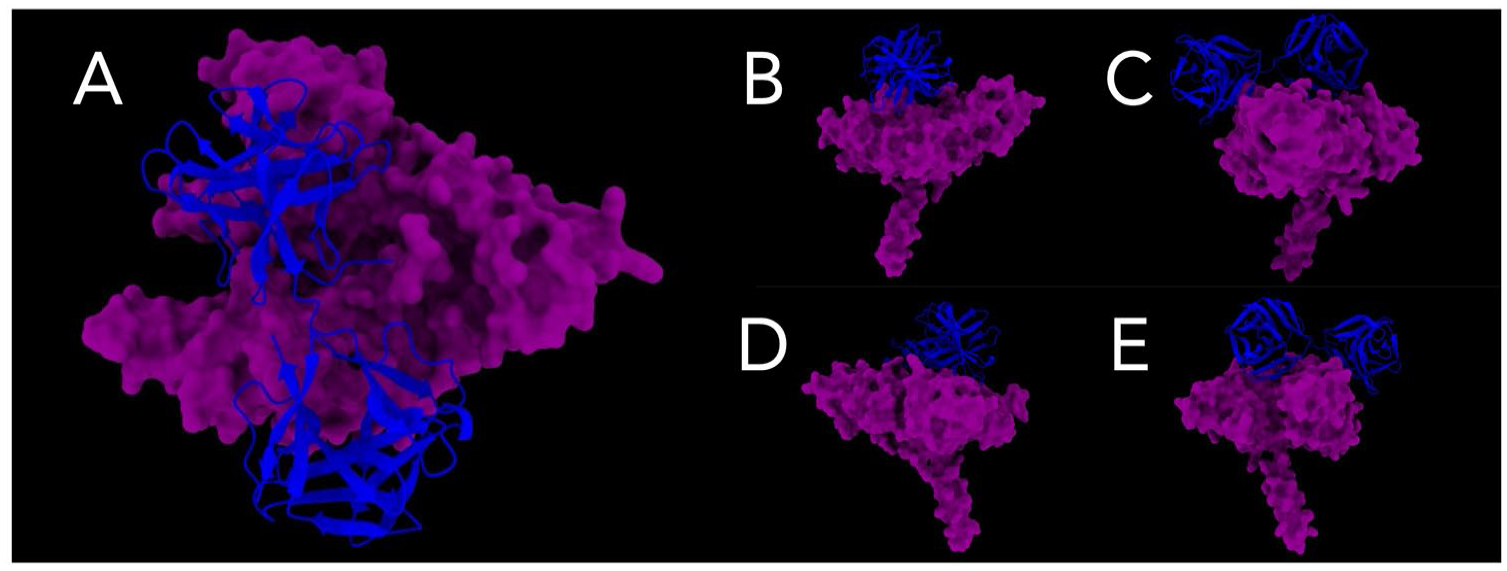
Interaction models between Pfs47-P47Rec. (A) The upper face of the interaction site between the consensuses is depicted. (B) to (E) provide lateral views of the consensus interaction site.

Optimal docking models for the complexes formed by the interaction of Pfs47 and P47Rec. (A) Pfs47-RO33, (B) Pfs47-3D7, (C) Pfs47-Palo Alto, (D) Pfs47-7G8, (E) Pfs47-HB3 and (F) Pfs47-T9/102.
Regarding Pfs47-3D7, it was observed that both the consensus and the IIM were situated in similar locations and had identical orientations. With the exclusion of models Pfs47-3D7-
In the Pfs47-HB3 model, it was observed that the P47Rec consensus and the IIM were not consistent in position or orientation. Using the Pfs47-HB3-
Discussion
The findings of our study indicate that the Pfs47 derived from African strains of the parasite, as well as those from nonhuman primate parasites, have the highest similarity to the Pfs47 consensus sequence. Although the consensus is not a prediction of ancestry, this sequence was located equidistant from all the clusters, so its structure is representative of all other proteins. However, it is advisable to expand the databases to determine more precise evolutionary structures. This outcome could be attributed to the African origin of
On the contrary, the AAA cluster exhibited a greater relationship with the African cluster
The anophelines of the Old and New World exhibit significant evolutionary divergence, occurring approximately 95 million years ago. This divergence coincides with the separation of the South American continent from its present-day African continent.
34
These results could indicate a sequential dispersion of P47Rec from Africa to other geographic regions. However, to verify this hypothesis, it is still necessary to expand the proteins analysed, including those from other anopheline species.
35
The phylogenetic tree based on the P47Rec sequences shows sequential clusters that go from Africa to Asia, Eurasia, South America and Central America, suggesting that the speciation of South American and Central American anophelines is more recent.8,36 In contrast, the intermediate cluster positioned between the Asian cluster and the American subcluster comprised the reference sequence of
The structure of the phylogenetic tree of P47Rec demonstrates the presence of diversity across the anopheline species that were analysed. This observation may be associated with the geographical spread of the genus, as well as with vector speciation. According to the motif patterns of Pfs47, a highly conserved protein structure is observed, as described by other authors.13,25 Considering the role of Pfs47 in evading the immune system of its vectors, it is reasonable to expect that a significant part of its structure exhibits a notable level of conservation.
The Pfs47-T9/102 sequence exhibited a distinct motif pattern in the signal peptide region, which differed somewhat from the patterns observed in other evaluated polypeptides. Based on the available evidence, it can be argued that the observed variation in the Pfs47-T9/102 pattern is unlikely to have any significant effect on the protein’s interaction capacity. Our analysis of the annotations indicated that both the motif pattern of Pfs47-T9/102 and that of the rest of the proteins possess a cleavage site that would remove the signal peptide from all proteins.
On the contrary, P47Rec has a higher abundance of motif patterns compared with Pfs47. Only pattern 3 encompasses species exclusively from a single continent. The remaining patterns encompassed an assortment of anopheline species originating from various geographical locations.
The primary variations in the configurations of P47Rec occur mainly in the C-terminal region, indicating that this segment may play a crucial role in the specific interaction with Pfs47. This finding lends support to the hypothesis proposing coevolutionary associations between Pfs47 and P47Rec, dependent on their geographical locations.9,25,37
The mutations found in the primary structures of Pfs47 and P47Rec have the potential to impact protein folding, particularly when these mutations involve amino acids with different charges. The structure of proteins can be modified by changes in charges and the lengths of the side chains of amino acids, which can lead them to reject one other due to their polarities or the volume they occupy. A greater number of mutations would generate a greater number of structural modifications. Nevertheless, the secondary and tertiary structures of the examined sequences, along with the consensus sequence, exhibited similar basic structures. The observed similarities may indicate that slight structural differences in the complex account for the selectivity observed between
Concerning tertiary structures, it was shown that Pfs47 exhibits 3 sandwich-shaped structures within its globular domain, which are likely responsible for its interaction with P47Rec. 13 A few cysteine residues in the globular domain of Pfs47 at its C and N-terminal ends imply a potential association with the protein’s rigidity. Conversely, the central structure of Pfs47 contains only 2 cysteines, indicating a possible involvement of this region in the induced adjustments during the formation of the complex 13,37,38 confirming the prediction of a loop structure at this site, likely involved in binding to P47Rec. 8
The lack of membrane anchoring structures is noteworthy, as observed in P47Rec. This result suggests that P47Rec could be a subunit of a larger protein complex that provides the necessary structures to anchor to the membrane, but a different hypothesis supports the findings of Molina-Cruz et al, proposing that P47Rec may exhibit cytoplasmic localization and have a function in the arrangement of the cytoskeleton within the midgut cells of mosquitoes. 8
The prediction of interactions between the consensus sequences of both proteins shows more precise interactions between the 6-Cys structures of Pfs47 and the N-terminal domain of P47Rec. Interactions between the central substructure of the globular domain of Pfs47 and some models of P47Rec were observed, suggesting a potential role for the N- and C-terminal regions of Pfs47 and the N-terminal end of P47Rec in mediating the primary interaction within the complex. Nevertheless, it seems likely that the central structure of Pfs47 and the C-terminal end of P47Rec exhibit distinct interaction regions, potentially accounting for their respective specificities.9,13,37 Some interaction models of the complex, such as those observed in the Pfs47 receptors of American strains of the parasite and the P47Rec variants of American anophelines, support the hypothesis of more recent coevolution.
25
It is noteworthy that a majority of the assessed docks of the Pfs47-P47rec complex with geographical proximity exhibited similarities to their respective IIMs or consensus models. This finding implies that the coevolution of Pfs47 and P47rec may have led to particular interactions between
Overall, the study’s novelty lies in its integrative approach combining bioinformatics, phylogenetics, structural biology and region-specific data collection to provide a comprehensive understanding of the coevolution and interaction between Pfs47 and P47Rec. This research not only fills gaps in current knowledge but also paves the way for future studies to explore these complex biological interactions further.
Conclusions
The findings of our study indicate that there are coevolutionary patterns between P47Rec and Pfs47 associated with the speciation and geographic dispersion of
Supplemental Material
sj-png-1-bbi-10.1177_11779322241284223 – Supplemental material for Coevolutionary Analysis of the Pfs47-P47Rec Complex: A Bioinformatics Approach
Supplemental material, sj-png-1-bbi-10.1177_11779322241284223 for Coevolutionary Analysis of the Pfs47-P47Rec Complex: A Bioinformatics Approach by Andrés S Ortiz-Morazán, Marcela María Moncada, Denis Escobar, Leonardo A Cabrera-Moreno and Gustavo Fontecha in Bioinformatics and Biology Insights
Supplemental Material
sj-png-2-bbi-10.1177_11779322241284223 – Supplemental material for Coevolutionary Analysis of the Pfs47-P47Rec Complex: A Bioinformatics Approach
Supplemental material, sj-png-2-bbi-10.1177_11779322241284223 for Coevolutionary Analysis of the Pfs47-P47Rec Complex: A Bioinformatics Approach by Andrés S Ortiz-Morazán, Marcela María Moncada, Denis Escobar, Leonardo A Cabrera-Moreno and Gustavo Fontecha in Bioinformatics and Biology Insights
Supplemental Material
sj-png-3-bbi-10.1177_11779322241284223 – Supplemental material for Coevolutionary Analysis of the Pfs47-P47Rec Complex: A Bioinformatics Approach
Supplemental material, sj-png-3-bbi-10.1177_11779322241284223 for Coevolutionary Analysis of the Pfs47-P47Rec Complex: A Bioinformatics Approach by Andrés S Ortiz-Morazán, Marcela María Moncada, Denis Escobar, Leonardo A Cabrera-Moreno and Gustavo Fontecha in Bioinformatics and Biology Insights
Supplemental Material
sj-png-4-bbi-10.1177_11779322241284223 – Supplemental material for Coevolutionary Analysis of the Pfs47-P47Rec Complex: A Bioinformatics Approach
Supplemental material, sj-png-4-bbi-10.1177_11779322241284223 for Coevolutionary Analysis of the Pfs47-P47Rec Complex: A Bioinformatics Approach by Andrés S Ortiz-Morazán, Marcela María Moncada, Denis Escobar, Leonardo A Cabrera-Moreno and Gustavo Fontecha in Bioinformatics and Biology Insights
Supplemental Material
sj-png-5-bbi-10.1177_11779322241284223 – Supplemental material for Coevolutionary Analysis of the Pfs47-P47Rec Complex: A Bioinformatics Approach
Supplemental material, sj-png-5-bbi-10.1177_11779322241284223 for Coevolutionary Analysis of the Pfs47-P47Rec Complex: A Bioinformatics Approach by Andrés S Ortiz-Morazán, Marcela María Moncada, Denis Escobar, Leonardo A Cabrera-Moreno and Gustavo Fontecha in Bioinformatics and Biology Insights
Supplemental Material
sj-png-6-bbi-10.1177_11779322241284223 – Supplemental material for Coevolutionary Analysis of the Pfs47-P47Rec Complex: A Bioinformatics Approach
Supplemental material, sj-png-6-bbi-10.1177_11779322241284223 for Coevolutionary Analysis of the Pfs47-P47Rec Complex: A Bioinformatics Approach by Andrés S Ortiz-Morazán, Marcela María Moncada, Denis Escobar, Leonardo A Cabrera-Moreno and Gustavo Fontecha in Bioinformatics and Biology Insights
Footnotes
Acknowledgements
Not applicable.
Author Contributions
GF and ASO-M involved in conceptualization. DE, GF, ASO-M, LAC-M and MMM involved in methodology. ASO-M, GF and DE involved in validation. ASO-M and GF involved in formal analysis. DE, GF, ASO-M and LAC-M involved in investigation. GF involved in resources. ASO-M, GF and DE involved in data curation. ASO-M and GF involved in writing – original draft preparation. DE, GF, ASO-M, LAC-M and MMM involved in writing – review and editing. GF involved in supervision. DE involved in project administration. GF involved in funding acquisition.
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship and/or publication of this article.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship and/or publication of this article: The National Autonomous University of Honduras’ Genetic Research Center provided funding for this project.
Ethics Approval and Consent to Participate
Not applicable.
Consent for Publication
Not applicable.
Availability of Data and Materials
The data sets used and/or analysed during the current study are available from the corresponding author on reasonable request.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
