Abstract
Lung cancer is one of the leading causes of cancer-related deaths in the world among both men and women. Several studies in the literature report that overexpression and mutation of the epidermal growth factor receptor (EGFR) are implicated in the pathogenesis of some lung cancers. Nimotuzumab is a humanized monoclonal antibody (mAb) that inhibits EGF binding because it binds to the extracellular domain of the EGFR. Nimotuzumab requires bivalent binding for stable attachment to cellular surface, which leads to nimotuzumab selectively binding to cells that express mAbs of moderate to high EGFR levels, and this could explain its low toxicity. This property has an advantage for development of nimotuzumab as a therapeutic and diagnostic agent. Monoclonal antibodies are large in size (150 kDa), thus penetrating slowly and residing in the blood for extended periods of time (from days to weeks); their use in imaging studies can result in low signal-to-background ratios and poor image quality. A reduction in the size of the immunoglobulin molecule has also been proposed as a means for increasing tumor penetration by mAbs. Nevertheless, it is known that the penetration of mAb into tumor cell is slow, due to its high molecular weight. Therefore, mAb is not very attractive to be used for imaging diagnostic purpose because of its kinetics and potential to elicit antibody response. The objective of this research was to study the homology modeling of a simpler functional molecule based on nimotuzumab, which consists of 2 antigen-binding fragments (Fab), namely, F(ab′)2, using MODELER. The crystal structure of Fab of nimotuzumab from protein data bank was used as a template to construct the model of F(ab′)2. Molecular dynamic simulation was performed to evaluate the stability of F(ab′)2 and conformational changes of F(ab′)2 in simulation. The result showed the dynamic behavior of antigen-binding site region of F(ab′)2 throughout simulation. This result is expected to be useful in the further development of F(ab′)2 fragment nimotuzumab as a lung cancer diagnostic.
Introduction
Cancer is a group of diseases characterized by uncontrolled growth and spread of abnormal cells. 1 Lung cancer is one of the leading causes of cancer-related deaths in the world among both men and women. In the case of lung cancer, non–small cell lung cancer (NSCLC), which accounts for approximately 85% of lung cancer cases, is a malignancy with high mortality. 2 There have been several studies in the literature reporting that overexpression and mutation of the epidermal growth factor receptor (EGFR) are implicated in the pathogenesis of some lung cancers and associated with reduced survival.2-4 The EGFR is a part of the ErbB family of receptor tyrosine kinases (RTKs). These trans-membrane proteins are activated following binding of peptide growth factors of the EGF family of proteins. 5 Its overexpression is not only present in lung cancer, but the corrupted EGFR signaling pathways contribute to a number of processes such as tumor proliferation, apoptosis resistance, angiogenesis, invasion, and metastasis. 6 This phenomenon is important for therapeutic and diagnostic approaches, because the response to anti-EGFR agents such as monoclonal antibodies (mAbs) might depend on the total level of expression of EGFR and ligands in cancer cells.
Nimotuzumab is a humanized mAb that inhibits EGF binding because it binds to the extracellular domain of EGFR. 6 In the preclinical studies, nimotuzumab demonstrated antiproliferative, proapoptotic, and antiangiogenic activities, 7 and nimotuzumab displayed a longer half-life and elevated area under the curve than cetuximab at the same dose level. 8 Researchers have studied the clinical efficacy of nimotuzumab in some lung cancers. In the clinical trials, we showed that nimotuzumab combined with chemotherapy significantly improved the objective response rate (ORR) and extended the duration of progression-free survival (PFS) and was generally well tolerated in patients with NSCLC.9,10
One striking outcome from the use of nimotuzumab in the clinic, compared with most of the other drugs targeting the EGFR such as cetuximab and panitumumab, is the absence of adverse effects such as severe skin, renal, and gastrointestinal mucosa. 11 The experimental observations demonstrate that the reason for the low toxicity of nimotuzumab is that the intrinsic properties of nimotuzumab require bivalent binding (binding with both antibody arms to 2 targets simultaneously) for stable attachment to the cellular surface, which leads to nimotuzumab selectively binding to cells that express moderate to high EGFR levels. Previous publications reported a KD of the order of 10−8 M for the nimotuzumab Fab fragment, a 10-fold lower affinity when compared with the cetuximab Fab fragment.12-14 It shows that it has intermediate affinity against the EGFR and hence binds with less avidity, and the target binding interaction is transient. Thus, it spares healthy tissues and avoids severe dose-limiting toxicities. Different from nimotuzumab, cetuximab and panitumumab have higher affinity constants, when EGFR expression is low as in healthy tissues; cetuximab and panitumumab still have high-avidity target binding, although stronger binding clearly leads to higher toxicities. 15 Structural computational studies were identified by Talavera et al, 12 which suggested the mechanism of action of F(ab) fragment; nimotuzumab blocks EGF binding and binds to domain III of the extracellular region of the EGFR, as is the case for several other known anti-EGFR antibodies, but allows active conformation and a basal level of ligand-independent EGFR signaling, whereas cetuximab inhibits also the active conformation efficiently, thereby blocking downstream signaling, which could explain its low toxicity. This property has a very essential advantage for nimotuzumab over other anti-EGFR mAbs including cetuximab and panitumumab, whose high toxicity is well-documented.
In diagnostic areas, researchers have developed a targeted CT scan contrast agent Nanoparticle-Poly Amido Amine Generation 4-Nimotuzumab at the molecular imaging level for the purpose of diagnosing lung cancer. 16 However, mAbs have a large size (150 kDa), thus penetrating slowly and residing in the blood for extended periods of time (from days to weeks); their use in imaging studies can result in low signal-to-background ratio and poor image quality. 17 A reduction in the size of the immunoglobulin molecule has also been proposed as a means of increasing tumor penetration by mAbs such as smaller fragments including F(ab′)2 fragment (110 kDa), Fab fragment (60 kDa), single-chain variable fragments (scFvs; 27 kDa), and minibodies (80 kDa). 18 Previous studies have suggested that only high-affinity Fab fragments bound effectively, independent of the antigen density. In contrast, low-affinity and intermediate-affinity antibodies require the avidity conferred by bivalent binding of F(ab′)2 fragment for effective attachment, which can occur only if the antigen density is elevated. A considerable amount of literature has been shown that the intrinsic properties of nimotuzumab require bivalent binding for stable attachment, which leads to nimotuzumab selectively binding to EGFR-overexpressing cells. 14 There have been several experimental studies in the literature reporting that the optimized digestion process used pepsin enzyme to digest nimotuzumab into its F(ab′)2 fragment and investigate its potential as a therapeutic radioimmunoconjugate. 19
In the past years, how an mAb interacts with antigen has attracted much attention as therapeutic and diagnostic agents. In many such cases, molecular dynamic (MD) simulations may be used. MD simulations have successfully provided information about the interaction between antibody and antigen. Numerous studies have attempted to explain the role of MD simulation to gain a better understanding of the structural stability and dynamics, such as the role of individual amino acids in the internal dynamics of a fully humanized IgG1 trastuzumab and fragments of it or of the domains of the Fc fragment of human IgG1.20,21 Previous computational studies have reported docking simulations and MD simulations performed for the nimotuzumab Fv fragment and a fragment of the region of the EGFR (eEGFR). According to the obtained model, nimotuzumab bound to domain III of the extracellular region of the EGFR, within an area that overlaps with both the surface patch recognized by cetuximab and the binding site for EGF. Most of the contacts of nimotuzumab with eEGFR domain III are accounted for by the heavy chain. 12
In this study, we describe the construction of a model of a monoclonal antibody (nimotuzumab) fragment F(ab′)2 truncated by pepsin from the crystal structures of fragment F(ab′) for lung cancer diagnostics using MODELER 9.19. 22 We use a combination of MD simulation to explore the antibody behavior in aqueous solution and evaluated the conformation, especially the residues on the complementarity-determining regions (CDRs) that play a role in maintaining the stability of the structure of F(ab′)2 fragments of nimotuzumab.
Material and Methods
F(ab′)2 construction
Computational experiments were carried out using the following hardware specifications: optional graphics processing unit (GPU) 32 GB, Seagate SATA II Internal Hard Disk 320 GB, and central processing unit (CPU) Core 2 Duo 2.4 GHz. The F(ab′)2 structure of nimotuzumab was prepared using the homology modeling method with MODELER 9.19. 22 The sequence of a target F(ab′)2 fragment is the crystal structure of Fab of nimotuzumab taken from protein data bank (PDB code: 3GKW), 12 and was prepared using MODELER 9.19 to fill in these missing residues by aligning with the full sequence of Fab nimotuzumab from International Immunogenetics Information System (IMGT), 23 as a template. In the F(ab′)2 structure, we added hinge sequence and a few CH2 sequence. 14 Two modified Fab structures were connected by disulfide bridges. Once a target-template alignment is constructed, MODELER 9.19 calculates a 3-dimensional (3D) model of the target completely automatically, and the “best” model can be selected by picking the model with the lowest value of the MODELER 9.19 objective function. 24 The model structures were evaluated by Ramachandran Plot Assessment using PROCHECK server (https://servicesn.mbi.ucla.edu/PROCHECK/).
MD simulation
The simulations were performed on the structure F(ab′)2 solvated in a TIP3P 25 water box using the tleap module of AMBER package. 26 For MD simulation, we ran a 20-ns MD simulation. The size of the water box was chosen to ensure that the systems have at least 10 Å solvation shell in all directions. The total system contains 158 392 atoms. The GAFF 27 is used to describe the intermolecular interaction for F(ab′)2. The solvated structures were then subjected to 1000 steps of steepest descent minimization of the potential energy; this allowed the water molecules to reorganize to eliminate bad contacts with the F(ab′)2 molecule. During this minimization, the F(ab′)2 molecule was kept fixed in its starting conformation using a harmonic constraint, followed by 2000 steps of conjugate gradient minimization and loop part restraint, 2000 steps of conjugate gradient minimization and backbone restraint, and 2000 steps of conjugate gradient minimization. The minimized structure was initially subjected to 60 ps of MD, and the system was gradually heated from 0 to 310 K using weak 5 kcal/mole/Ǻ 2 harmonic constraints on the solute with its starting structure. This allows for slow relaxation of the built F(ab′)2 structure. Equations of motion were solved using the Verlet leapfrog algorithm 28 with an integration step of 1 fs. In addition, SHAKE constraint 29 was imposed on all covalent bonds involving hydrogen atoms.
Once the system is heated, continue to equilibrate the system at constant pressure to allow the density to equilibrate. The subsequent simulation was performed at 1 ns of the backbone constrained structure to equilibrate the system at 310 K. Finally, 20-ns-long constant-pressure constant-temperature (NPT) equilibrium dynamics was carried out, with temperature regulation achieved using Langevin Thermostat. 30 Particle mesh Ewald summations 31 with a real space cutoff of 10 Ǻ were used to calculate electrostatic interactions beyond the nonbonded cutoff distance in the system during all MD runs. The GPU-accelerated PMEMD program written in CUDA 32 was used for production runs. System properties (energies, temperature, pressure, density) and atom coordinates were recorded every 4 ps during the simulations. Every 100 frames of the trajectory were used for analysis, resulting in data points every 8 ps. Analysis of root-mean-square deviations (RSMDs) was performed using CPPTRAJ. 33 The RMSD of backbone atoms (—C–C–N—), using the crystal structure as a reference, was mass-weighted and averaged over each frame considered for analysis. To obtain domain RMSD values, only root-mean-square fitting and RMSD calculations for the domains of interest were averaged. MD simulations were visualized in a virtual environment using VMD 34 and Biovia Discovery Studio Visualizer.
Results and Discussion
A construction of F(ab’)2 fragment of nimotuzumab
A complete crystal structure of nimotuzumab, a humanized IgG antibody, does not exist. Previous research has shown that the crystal structure of the nimotuzumab Fab fragment was determined at 2.5 Å resolution with good stereochemistry and no outliers in the Ramachandran plot. 12 Therefore, to generate the starting coordinates for this study, we obtained the sequence of a target F(ab′)2 fragment through the full sequence of Fab nimotuzumab from IMGT 23 and the crystal structure of Fab of nimotuzumab taken from PDB (PDB code: 3GKW) 12 as a template. Based on the study by Vlasak et al that observed a cleavage site in mAbs by pepsin enzyme and the modeling of F(ab′)2 fragment which includes both the Fab domains and the upper, middle, and lower hinge residues as shown in Figure 1, we added the hinge sequence, a few CH2 sequence, and 2 modified Fab structures connected by disulfides bridges. Pepsin can be used to cleave IgG at the C-terminal side of the inter-heavy chain disulfides in the hinge region, producing a bivalent antigen-binding F(ab′)2 fragment and an Fc fragment. A number of studies have found that fragmentation of mAb depends significantly on pH, temperature, and other solvent conditions. 35

Frequently observed cleavage sites in monoclonal antibodies. 35
The 3D structure (Figure 2) was created using MODELER 9.19. 22 The MODELER is a program that can be used for homology modeling based on the alignment of a sequence to be modeled with the template structures and the atomic coordinates of the templates, and automatically calculates a model containing all nonhydrogen atoms.22,24 Studies have found that the structure of an F(ab′)2 fragment forms 2 Fab fragments and crosslinks with the inter-chain disulfide bond. In the pairing of light and heavy chains, the 2 variable domains dimerize to form the Fv fragment which contains the antigen-binding site. The 3D F(ab′)2 structure of nimotuzumab consists of 2 pairs of light chains and 2 pairs of heavy chains, of which the light chain is formed by 2 of these domains, called the variable (VL) and the constant domain (CL). The heavy chains contain a variable domain (VH) and 1 constant domain (CH). CH is formed by 2 β sheets packed face-to-face, linked together by a conserved disulfide bridge. Previous research has shown that the F(ab′)2 fragment contains 2 covalently linked Fab regions while leaving intact some of the hinge region. F(ab′)2 fragments have 2 antigen-binding Fab domains and are divalent with a molecular weight of approximately 110 kDa. 36
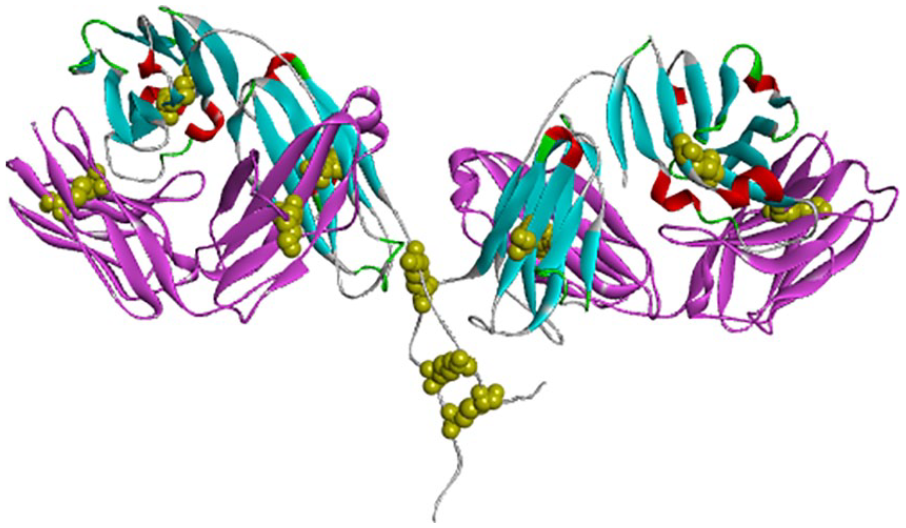
Three-dimensional structure of F(ab′)2 fragments from Discovery Studio 4.1 visualizer: light chain (pink ribbon), heavy chain (blue ribbon), disulfide (yellow), and electrostatic interactions (red).
The Ramachandran plot was used for F(ab′)2 fragment structural validation by comparing the structural conformation with the secondary protein structure coordinates that have been modeled. Furthermore, the stereochemical parameters of the proteins such as the main and side chain data of Ramachandran plot were compared. The results of the Ramachandran plot model of F(ab′)2 structure of nimotuzumab have shown, furthermore, the stereochemical parameters of the proteins, such as main and side chain data of Ramachandran plot. It has shown that 94% of residues were located in the most favored regions, whereas 4.3% and 1.7% residues, respectively, fell in the additional allowed and disallowed regions (see Figure 3). That is, >90% of the amino acid residues have allowed conformations. In general, a protein structure with more than 90% of the residues being located in the most favored region of all models is categorized as a good model.
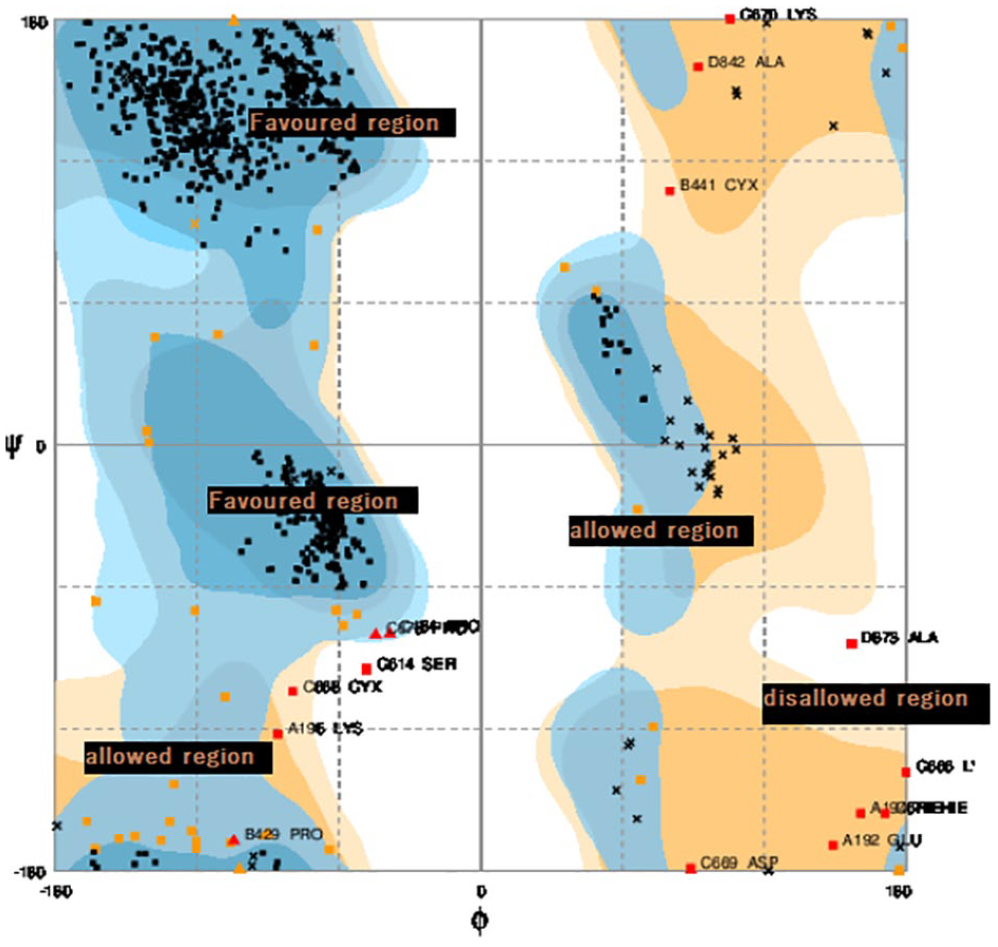
Ramachandran plot assessment of F(ab′)2 nimotuzumab.
In modeling, mAb requires precise identification of the residues that have an impact on the interaction or affinity of the antibody with its target antigen. The CDRs of the mAb are assumed to account for the recognition of the antigen. Its contains also the antigen-binding site. Kabat and co-workers were the first to introduce a systematic approach to identify CDRs based on the assumption that CDRs are the most variable regions between mAbs. Kabat Numbering Scheme revealed 3 hypervariable regions in the variable region of the light chain and heavy chain. 37 Figure 4 shows that the active regions or the CDRs are divided into 6 active regions, namely, CDR1, CDR2, and CDR3 in the heavy chain and CDR1, CDR2, and CDR3 in the light chain based on Kabat alignment result. CDR1 light chain regions of F(ab′)2 consist of 16 amino acid residues (residues 24-34), CDR2 region consists of 7 amino acid residues (residues 50-56), and CDR3 (residues 89-97) consists of 9 amino acid residues. In the heavy chain, CDR1 region consists of 5 amino acid residues (residues 26-35), CDR2 consists of 17 amino acid residues (residues 50-65), and CDR3 consists of 14 amino acid residues (residues 95-102) (Figure 4). Comparison of results with previous findings shows that the contacts of Fab fragment nimotuzumab with eEGFR domain III are accounted for by the heavy chain (CDR H1), contributing to binding with 2 residues Tyr H27 and Tyr H32. CDR H3 contributes the majority of the interactions with 7 consecutive residues (from H98 to H100D). In the light chain, 7 residues interact with the EGFR, 2 from CDR L1 (Asn L28 and Tyr L32), 4 from CDR L2 (Tyr L49, Lys L50, Phe L55, and Ser L56), and Leu L46 at the base of CDR L2. 12 In this study, F(ab′)2 structures superimposed the crystal structure of Fab of nimotuzumab taken from PDB (PDB code: 3GKW) 12 using Discovery Studio 4.1 visualizer based on their sequence alignment to confirm that our structure complies with these findings of past studies by Talavera et.al and then the matching residues for superimposition are automatically identified based on the aligned residues in the sequence alignment, especially on their CDRs. The overlay structure of the (Fab′)2 fragment suggests that the CDRs in the heavy chain and light chain share similar residue features with the corresponding CDRs in the heavy chain and light chain in the Fab as a template (figure does not show).

Complementarity-determining regions of nimotuzumab F(ab′)2 light chain and heavy chain based on Kabat alignment result.
MD simulation
To study the molecular behavior, 20 ns of MD simulations was performed in the F(ab′)2 fragment of nimotuzumab structure to investigate the structural stability and conformational evolution. We calculated the time series of the atom-positional RMSD to reflect the impact of the stability on the F(ab′)2 fragment of nimotuzumab structure, and they were subsequently analyzed to evaluate the balance of trajectory in MD simulation. 38 At the duration of the initial, the trajectory system fluctuated between 0.9 and 2.5 Å from the starting time, and this fluctuation continued until the MD simulation. The RMSD value slightly increased when starting the simulation, but there was a sharp rise from 8000 ps until 11 000 ps, which was decreased after 13 000 ps. In comparison, the hinge region appears to be quite flexible. The motions of this region appear to be larger, which is not unexpected, and it is highly exposed to solvent in 17 500 ps (Figure 5). The size of the RMSD values reflects the degree of difference between the 2 structures: the overall RMSD of the bound and free antibody structures. More recent studies have confirmed that the constant domains tend to have larger RMSD values than the variable domains. They hypothesized that a high movement of the constant domain might be correlated with signal propagation.

Root-mean-square deviation profile of F(ab′)2 of nimotuzumab during 20 ns simulation.
The atom-positional root-mean-square fluctuations (RMSFs) of F(ab′)2 of nimotuzumab are shown in Figure 6. The RMSF illustrates the deviation from the mean structure over a dynamic ensemble (flexibility or fluctuation changes) of amino acid residues. The highest fluctuations were observed in atom numbers 14 071 and 17 367. However, the residue does not play an important role in the binding site. Numerous studies have attempted to explain the dependence of the RMSF values on the position of amino acid residues, with one of the most significant fluctuations being on the C-terminal end. The lowest mobility in the light chain was revealed in the regions that interacted with the heavy chain. The heterogeneity of Fab fragments was revealed in the heavy and light chains. 39 According to an investigation by Wang et al, 39 amino acids within the variable domains have generally fluctuated more than in the constant domains. This could be attributed to the tendency of antibodies to have flexible variable domain, which allows them to better search for and ultimately bind their target. 33

Root-mean-square fluctuation profile of F(ab′)2 of nimotuzumab during 20 ns simulation.
One important parameter that correlated with the conjugation reaction of F(ab′)2 of nimotuzumab with lysine residues is the solvent-accessible surface area (SASA). The SASA of a protein is the protein surface in contact with the solvent molecule. The solvation effect plays an important role in the maintenance of protein stability and folding involving hydrophobic interaction. 40 Hydrophobic interactions established among nonpolar amino acids ensure the stability of globular proteins in solution by shielding the nonpolar amino acids in hydrophobic cores, away from the aqueous environment. 41 The analysis shows that the lysine residues display SASA of ~ 284-350.1 Å 2 in the simulation period, whereas there are several lysine residues that have a high exposure to solvents in the light chain—Lys 55, 199, 259, 627, and 840 residues (Figure 7)—and in the heavy chain—18, 249, 252, 257, and 401 residues (Figure 8). Among the lysine residues, only one appears in the CDRs, lys 55. The reduced solvent-accessible area in the proteins indicates that the proteins may decrease the probability of their interaction with other molecules.

Solvent-accessible surface area in lysine residues in the light chain using VMD.

Solvent-accessible surface area in lysine residues in the heavy chain using VMD.
Conclusion
Modeling and MD simulations were performed on F(ab′)2 nimotuzumab for a total simulation time of 20 ns. In homology modeling, the 3D F(ab′)2 structure of nimotuzumab consists of 2 pairs of light chains and 2 pairs of heavy chains; the heavy chains contain a VH and a CH. The structural validation was studied used Ramachandran plot, with more than 90% of the residues being located in the most favored region of all models being categorized as a good model. Identify CDRs based on the assumption that CDRs are the most variable regions between mAbs. Kabat Numbering Scheme and the studies have shown that the active regions or the CDRs are divided into 6 active regions. The result showed that the dynamic behavior of F(ab′)2 fragment nimotuzumab at 20 ns is conformally stable. The results of this study could inform the design of better F(ab′)2 fragments. For further research, it is advisable to perform MD simulations of the binding affinity explored with the receptor.
Footnotes
Acknowledgements
The authors are grateful to the Directorate of Research and Community Service Universitas Padjadjaran; Dean of the Faculty of Mathematics and Natural Sciences, Head of the Department of Chemistry, and Head and the entire staff of Biochemistry Laboratory. We are also grateful to the laboratory facilities provided by Department of Chemistry, Faculty of Mathematics and Natural Sciences Universitas Padjadjaran and to thank to all those who have given advice, motivation, and material support.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The authors are grateful to the Directorate of Research and Community Service Universitas Padjadjaran.
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
Y.S. conceived and designed the experiment, performed the experiment, analyzed and interpreted the data, and wrote the manuscript. A.R.R.F. conceived and designed the experiments, and analyzed and interpreted the data. Z.N. performed analysis of the data. A.H. analyzed and interpreted the data. M.Y. conceived and designed the experiments, performed the experiment, and analyzed and interpreted the data. M.R. performed the experiment, and analyzed and interpreted the data. A.M. conceived and designed the experiments, performed the experiment, and analyzed and interpreted the data. U.M.S.S. conceived and designed the experiments, analyzed and interpreted the data, wrote the manuscript, and compiled and revised the manuscript.
