Abstract
Transposable elements (TEs) are mobile genetic elements present in almost all eukaryotic genomes. Due to their typical patterns of repetition, discovery, and characterization, they demand analysis by various bioinformatics software. Probably, as a result of the need for a complex analysis, many genomes publicly available do not have these elements annotated yet. In this study, a de novo and homology-based identification of TEs and microsatellites was performed using genomic data from 3 palm species: Elaeis oleifera (American oil palm, v.1, Embrapa, unpublished; v.8, Malaysian Palm Oil Board [MPOB], public), Elaeis guineensis (African oil palm, v.5, MPOB, public), and Phoenix dactylifera (date palm). The estimated total coverage of TEs was 50.96% (523 572 kb) and 42.31% (593 463 kb), 39.41% (605 015 kb), and 33.67% (187 361 kb), respectively. A total of 155 726 microsatellite loci were identified in the genomes of oil and date palms. This is the first detailed description of repeats in the genomes of oil and date palms. A relatively high diversity and abundance of TEs were found in the genomes, opening a range of further opportunities for applied research in these genera. The development of molecular markers (mainly simple sequence repeat), which may be immediately applied in breeding programs of those species to support the selection of superior genotypes and to enhance knowledge of the genetic structure of the breeding and natural populations, is the most notable opportunity.
Introduction
Eukaryotic genomes are known to be densely made up of repetitive elements, mainly microsatellites and transposable elements (TEs). These repetitive elements, when characterized in a plant species, generate information that can be applied for different purposes in a plant breeding program. For instance, microsatellites can be applied as molecular markers for mapping quantitative trait loci (QTL) for paternity tests, 1 and in the case of transposons for gene regulation, epigenetic studies, genetic engineering, and gene therapy. 2
Transposable elements are classified into 2 main classes, based on the molecular mechanism that mediates their transposition. The elements that use a “copy-and-paste” mechanism belong to class I, and those that use a “cut-and-paste” belong to class II. 3 The increasing diversity of TEs identified in different taxa, mainly in plants, unleashed the unified TE classification system. 4
Transposable elements may respond to more than 50% of the total content of some genomes. 5 This amount can be even higher, up to 70%, in the genomes of some grasses. 6 Although most TEs groups are ancestral and present in basically all the kingdoms, these elements differ significantly from each other, reaching to thousands of different families, only in the plant kingdom. 7 It is known that the expansion and contraction waves in TE numbers can result in dramatic differences between genomes. 8
The repetitive pattern and structural signatures typically found in TEs make them natural candidates for a large-scale bioinformatics analysis. There are 2 computational approaches for the identification and annotation of TEs; the first method is based on structural features (de novo), and the second is the search for similarities in databases (homology based). 9 Although there are many tools for annotation of TEs, 10 this is still an open field of research in the area of bioinformatics. 11
A detailed description of repeats can be useful in refining genome assembling and annotation (especially in complex genomes like those of plants). Moreover, it provides information on genome variability and how they diversified over the evolutionary process. Recent insertion of TE families can help to better understand the evolutionary mechanisms involved in species differentiation. 12 Besides, the epigenetic silencing mechanism may help in understanding the regulation of the transposition activity in plants. 13
The Elaeis genus consists of 2 species, Elaeis guineensis (or African oil palm) and Elaeis oleifera (or American oil palm). The African oil palm is a perennial monocot species that produces high amounts of edible oil in its fruits and seeds. Altogether, this oil crop is responsible for about 35% of all vegetable oil produced worldwide. The American oil palm is similar to the African one in so many aspects. Despite having lower yields, the American oil palm has higher unsaturated fatty acid content, lower height, and tolerance to some important diseases, 14 such as bud rot. African and American oil palms have an estimated genome size of approximately 2 Gb. 15 It has been estimated that a large proportion of repeats is present in the genome of E guineensis14,16; however, there is no public detailed description of the composition and distribution of TEs, as well as microsatellites, in the genomes of these 2 species.
Date palm (Phoenix dactylifera) is a very well-known palm species, with high economic importance due to its nutritious fruits, as well as due to its ornamental use and wood quality (great tensile strength). 17 This palm has high genomic and phylogenetic similarities with oil palm, 14 has been taxonomically the closest species to the genus Elaeis, and has publicly available genomic data.
The Brazilian breeding program on Elaeis spp., coordinated by Embrapa, has the development of interspecific hybrids between the African and the American oil palm as one of its main goals. A deep characterization of the genomes of these 2 oil palm species is fundamental to further optimize the breeding strategies in use, and this is the main motivation behind this study. The use of publicly available genomic data, from a taxonomically closer species such as date palm, to compare with the genomes of the 2 oil palm species, is understood as a way to strengthen the understanding of the evolution of the repetitive component of these genomes.
This study provides a characterization and comparison of the TEs and microsatellites present in the genomes of the American and African oil palms, as well as the date palm. This analysis can provide insights into the repetitive content of these species and the application of these regions to explore the genetic variability within and among palm species. A comparative analysis based on a scaffold assembly of these genomes was performed, allowing the distribution of TEs on the chromosomes of E guineensis to be unequivocally obtained and highlighting differences with other genome. A full evaluation of African oil palm chromosomes was also included.
Materials and Methods
A pipeline for the analysis of repetitive elements (repeats), which includes some free software typically used in repeats analysis, such as Tandem Repeats Finder (TRF), RepeatModeler, and RepeatMasker, was developed and is detailed below. Local scripts, using programming languages Perl and Python, were developed to automate the data transformation between steps of the scrutiny. This pipeline is under performance enhancement to improve speed through parallelism techniques (Fork, Perl), as well as normalization of software multithread parameters (L.S. Brito et al, 2016 unpublished data).
DNA sequence data
The chromosomes and/or scaffolds from 4 genome drafts were used in this study: (1) E oleifera (EO8) MPOB genome (GenBank accession [gb ac] ASIR00000000), (2) E guineensis (EG5) scaffold (gb ac ASJS00000000) and chromosome (gb ac CM002081.1-CM002096) assemblies from Singh et al, 14 (3) P dactylifera genome (gb ac ATBV00000000) from Al-Mssallem et al, 18 and (4) a local preliminary assembly (version 1.0) of the genome of E oleifera access from the Amazon rainforest, Manicoré, belonging to the E oleifera Germplasm Bank of Embrapa (Illumina Hiseq2000 sequences, assembled with ALLPATHS-LG, unpublished data), resulting in 85 612 scaffold sequences and an N50 of 27 kb.
Identification and classification of microsatellites
The content of microsatellites in oil and date palm genomes was studied. The TRF software was applied to identify microsatellite repeats, 19 using the following parameters: match 2, mismatch 7, delta 7, PM 80, PI 10, minscore 50, maxperiod 500, -f (flanking sequence), -d (data file), and -m (masked sequence file). To summarize the results obtained, the Tandem Repeats Analysis Program (TRAP) software 20 was applied, using the following parameters: -id = 70 (minimum match percentage), -tbf = html + csv (table format), -sort = size (sort field), -rr (flag—create redundancy report), and -trf (flag—create trf-like file).
Identification of repetitive elements
The first step was preformatted with the RepeatModeler software (default settings) that makes up a pipeline with RECON software, 21 RepeatScout, 22 RepeatMasker, TRF, 19 and RMBlast, for the de novo identification of TEs. The types of long terminal repeat (LTR) retrotransposons were identified using the LTR_FINDER software, 23 applying default parameters. All the repeats greater than 100 bp were included in the TE library.
Classification of repetitive elements
The resulting TE library was classified using Blastn (e-value ≤ 1e−5, identity ≥ 70%, and minimum size alignment ≥ 80 bp) against Repbase and the public database MIPS Repeat database, which integrates other databases (TRansposable Elements Platform [TREP], TIRG repeats, PlantSat and GenBank). All TEs identified, but unclassified, were assigned as “retrotransposon not rated” or “DNA transposon not rated.”
Annotation of repetitive elements
The RepeatMasker software was applied, with a custom library (combination of repeats of RepBase, MIPs—Munich information center for protein sequence and TE library de novo), to search for TE coordinates. This software was also used to generate a version of a masked sequence with repeat regions. The tool “one code to find them all,” 24 a Perl script to parse the RepeatMasker output file, was used, aiming to organize, summarize, and produce statistics about the RepeatMasker results.
The data generated by “one code to find them all” were used to measure divergence between copies of TEs, by means of the correlation of divergences (in relation to reference), and the proportion of the length of the reconstructed copy compared with the reference element. 24
Results
Large proportions of the 4 genomes studied are repeat sequences: 50.96% of the E oleifera Amazonian genome (EoAG), 42.31% of the E oleifera MPOB genome (EoMG), 33.67% of P dactylifera genome (PdG), and 39.41% of the African oil palm scaffold assembly genome (EgMG) (Table 1). Moreover, 212 722 TE copies were identified in chromosome assembly of African oil palm (Tables 2 and 3). A total of 155 726 microsatellite loci (between mononucleotide and hexanucleotide) were identified in these 4 genomes.
Repeat content in oil and date palm genomes.
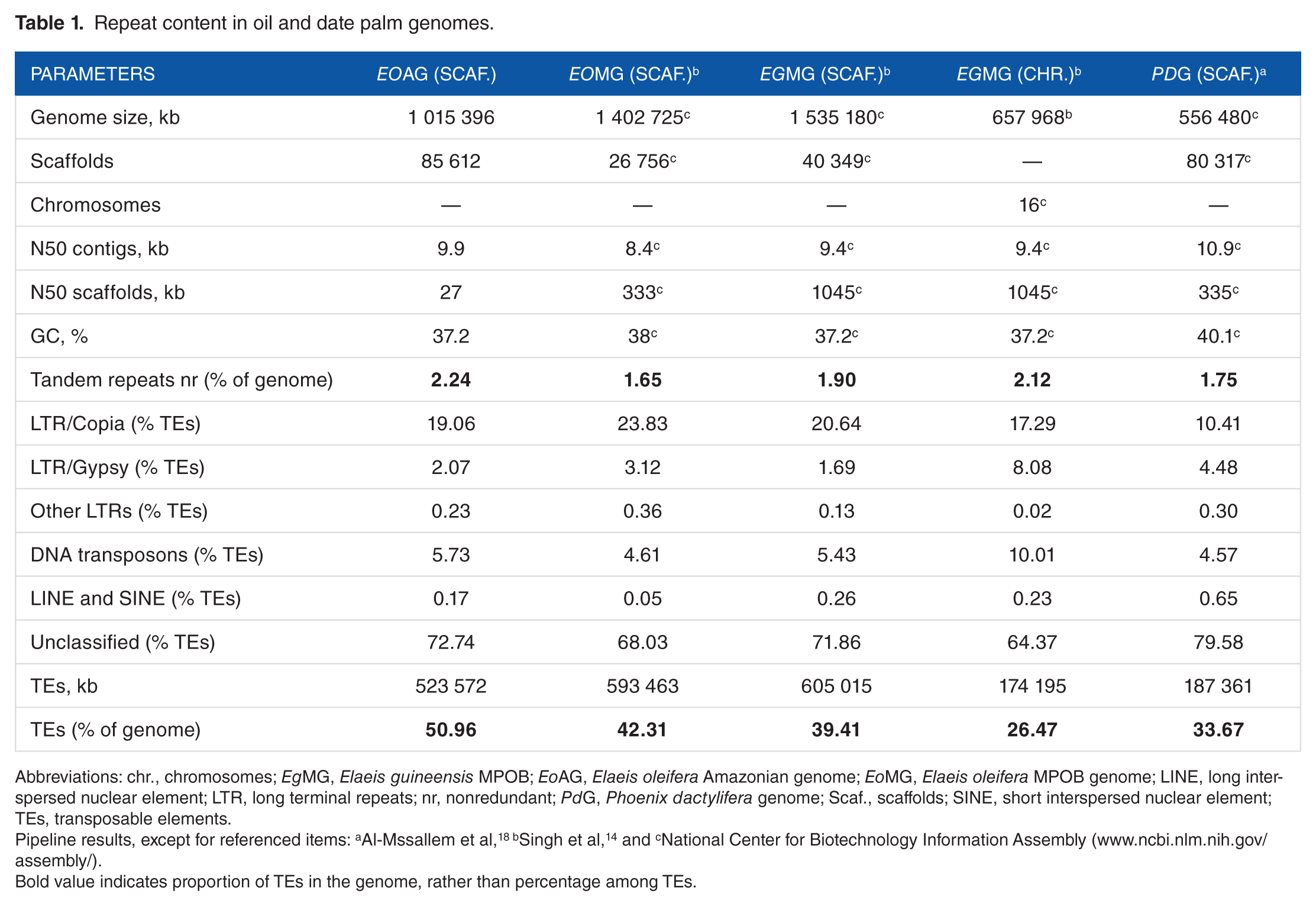
Abbreviations: chr., chromosomes; EgMG, Elaeis guineensis MPOB; EoAG, Elaeis oleifera Amazonian genome; EoMG, Elaeis oleifera MPOB genome; LINE, long interspersed nuclear element; LTR, long terminal repeats; nr, nonredundant; PdG, Phoenix dactylifera genome; Scaf., scaffolds; SINE, short interspersed nuclear element; TEs, transposable elements.
Pipeline results, except for referenced items: aAl-Mssallem et al,18 bSingh et al, 14 and cNational Center for Biotechnology Information Assembly (www.ncbi.nlm.nih.gov/assembly/).
Bold value indicates proportion of TEs in the genome, rather than percentage among TEs.
Total repeat content on Elaeis guineensis chromosomes.

Abbreviation: TE, transposable element.
Detailed classification of transposable elements identified on Elaeis guineensis chromosomes.

Abbreviations: LINE, long interspersed nuclear element; LTR, long terminal repeats; RC, rolling circle; SINE, short interspersed nuclear element; TEs, transposable elements; tRNA, transfer RNA.
Bold value indicates that RC/Helitron stand out from the other Class II-DNA because they have an exclusive transposition mechanism called rolling circle (RC).
Identification of repeats
Long terminal repeat retrotransposons are the TEs predominantly identified in EoAG—more specifically, those from Copia (19.06%) and Gypsy (2.07%) superfamilies. The non-LTR retrotransposons long interspersed nuclear elements (LINEs) and short interspersed nuclear elements (SINEs) Comprise about 0.17% of the repetitive elements. Other classes of repetitive elements, such as DNA transposons, constitute relatively small proportions of the genome (5.73%). More than half of the repeats in the E oleifera Amazonian genome do not show sequence similarity with other previously identified TEs (Figure 1 and Table 1).

Distribution of transposable elements in the genome of Elaeis oleifera, Amazonian genotype. Transposable elements in silico identified in the draft genome (version 1.0) of a Manicoré genotype from the Germplasm Bank of Caiaué (E oleifera) at Embrapa Amazônia Ocidental. LTR indicates long terminal repeat.
In EoAG, a total of 328 879 loci of tandem repeats (all repeats type) were found, corresponding to 23 056 kb of repeat bases and representing 2.24% of the total bases of the sequence (additional data given in Table S1). The major classes of microsatellites identified were mononucleotides (8.40%), dinucleotides (43.30%), trinucleotides (9.59%), tetranucleotides (10.38%), pentanucleotides (15.80%), and hexanucleotides (12.53%). For each class, the main region repeats found were (T/A)n, 100%; (AT)n, 48%; (TTA)n, 42%; (ACAT)n, 47%; (TATAT)n, 31%, and (TTTTTC)n, 33%, respectively (Figure 2). Among the major classes analyzed, the most abundant are the dinucleotide repeats, with 15 574 identified loci.

Frequency (%) of the most common simple sequence repeat (SSR) motifs in the genome of Elaeis oleifera, Amazonian genotype. Frequency was estimated for each class of SSRs.
In total, 155 726 loci of microsatellites (between mononucleotide and hexanucleotide) were identified in the genomes of oil and date palms. For EoAG, EoMG, and EgMG assembly scaffolds, there are 35 968, 41 808, and 48 788 microsatellite loci, respectively, and 29 162 loci in PdG (Figure 3 and additional data given in Table S1). In E guineensis assembly chromosomes, the total number of loci identified is 31 179 (additional data given in Table S1).

Comparison of simple sequence repeat (SSR) amount among oil and date palm genomes. Amount of mononucleotide, dinucleotide, trinucleotide, tetranucleotide, pentanucleotide, and hexanucleotide in EoAG (scaf.), EgMG (scaf.), EgMG (Chr.), and PdG (scaf.). EgMG indicates Elaeis guineensis MPOB genome; EoAG, Elaeis oleifera Amazonian genome; EoMG, Elaeis oleifera MPOB genome; PdG, Phoenix dactylifera genome; Scaf., scaffolds; Chr., chromosomes.
The composition of TEs was very similar among the 4 genomes studied. For the 4 sets of scaffolds used (EoAG, EoMG, PdG, and EgMG), the number of TEs/copies was around 50.96% (523 572 kb)/591 808, 42.31% (593 463 kb)/585 241, 33.67% (187 361 kb)/347 513, and 39.41% (605 015 kb)/608 682, respectively (Table 1 and additional data given in Tables S2 to S5).
Distribution and classification of TEs on the chromosomes of African oil palm
A total of 212 722 TE copies were identified, with a total size of 174 195 kb, representing 26.47% of the sequence. Among the 16 chromosomes of the African oil palm, chromosomes 6 and 15 are the ones presenting the highest repeat coverage (Table 2 and additional data given in Table S6); however, the distribution of TE classes was to a certain degree similar in all chromosomes (Figure 4).

Chromosomal distribution of TEs in Elaeis guineensis, the African oil palm. Each chromosome of E guineensis MPOB genotype was analyzed for the proportion of the types of TEs. LINE indicates long interspersed nuclear element, LTR, long terminal repeat; MPOB, Malaysian Palm Oil Board; TE, transposable element.
Figure 5 shows the most representative TE families in each chromosome. The most characterized LINE families are L1 and L1-Tx1, whereas the 2 most represented DNA transposon families are CMC-EnSpm and hAT-Ac. For the LTR retrotransposons, Copia and Gypsy were the most frequent superfamilies. Copia is the most abundant one on all chromosomes. The distribution of the main families of TEs per chromosome was also examined. The repeats have been classified and are described below.

Chromosomal distribution of the most represented transposable elements (TEs) in Elaeis guineensis, the African oil palm. (A) DNA/MuLE-MuDR and DNA/CMC-EnSpm families are the most represented DNA transposons. (B) LINE/RTE-BovB and LINE/L1 families are the 2 most represented LINE superfamilies. (C) LTR/Gypsy and LTR/Copia families are the 2 most represented LTR superfamilies. (D) Unknown and unspecified are the 2 most represented unclassified repeats. LINE indicates long interspersed nuclear element; LTR, long terminal repeat.
Among all the class I retrotransposons identified in the African oil palm chromosomes, 25 558 copies have been classified as LTR elements, totalizing 44 226 kb. The 4 main superfamilies are Caulimovirus, Copia, Gypsy, and ERV1 (Table 3). Chromosomes with the largest representation of these elements were 6 and 9 (28.94% and 27.77%, respectively).
However, only 609 and 148 copies have been classified as belonging to the LINE and SINE families, respectively, totalizing 372 and 20 kb. The 5 main LINE families are L1, L1-Txt1, L2, RTE-BovB, and Tad1 (Table 3). Chromosomes 2 and 5 are the ones with the greatest abundance of this element (0.29% and 0.34%, respectively). The SINE/transfer RNA family responded to 95.95% of the SINE elements found (Table 3).
A total of 15 254 copies have been classified as class II (DNA transposons) on the African oil palm chromosomes, totalizing 16 983 kb. CMC-EnSpm is the most frequent one, with a total of 7544 copies and 18 165 fragments, totalizing 10 825 kb (Table 3). CMC-EnSpm is widely dispersed among the 16 chromosomes, with the lowest percentage of appearance on chromosome 14 and the highest on 15. Besides this family, 8 other families were identified: Academ (27 kb), Crypton (2 kb), Dada (240 kb), Hat (families Ac, Blackjack, Charlie, Tag1, and Tip100, totalizing 2873 kb), Mule-MuDR (2879 kb), PIF-Harbinger (44 kb), Sola (94 bp), and rolling-circle transposons—Helitron (461 kb) (Table 3).
The majority of TEs copies (80.30%) was grouped as unclassified, being subdivided into 2 groups: unspecified (43.92%) and unknown (36.37%). Altogether, they account for 170 814 copies, totalizing 112 131 kb (Table 3).
Divergence of TEs
Ratios close to “1” (full-length elements) and divergence close to “0” could indicate events of recent insertion of TEs in the genome. Figure 6 shows DNA transposon and LTR retrotransposon superfamilies as potential recent insertions (with some full-length elements), whereas LINE-like elements present low divergence but of different sizes. Each point represents a TE copy.

The plot of the divergence of transposable element (TEs) in Elaeis guineensis, the African oil palm. The divergence has been plotted for the most represented families of DNA transposons (upper panel), LINE-like elements (half panel), and LTR retrotransposons (lower panel). (A) DNA/hAT-AC, (B) DNA/CMC-EnSpm, (C) LINE/L1, (D) LINE/L1-Tx1, (E) LTR/Copia, and (F) LTR/Gypsy. Each point corresponds to a copy. Copies with divergence close to 0 and ratio close to 1 correspond to potentially active and full-length copies. LINE indicates long interspersed nuclear element; LTR, long terminal repeat.
Discussion
Microsatellites and TEs present in the oil and date palm genomes were identified and analyzed using a pipeline for de novo and homology-based identification of repetitive elements. This report is the first with a detailed analysis of repeats in the whole genome of oil palm. Here, the not yet published genome of an Amazonian oil palm genotype belonging to the E oleifera Germplasm Bank of Embrapa and the recently released genomes of African and American oil palms 14 provide an opportunity for the analysis of repeat content with implications for the development of genetic markers, genome assembly, phylogenetic analysis, and epigenetic studies.
Oil and date palm genomes are mainly composed of repeats
A large portion of these 4 genomes available and studied is composed of TEs (50.96%—EoAG, 42.31%—EoMG, 39.41%—EgMG, and 33.67%—DpG). This fact correlates with the C-value paradox in which the genome size in eukaryotes is associated with the number of repetitive regions and not gene content. 25 Small genomes, such as the one of Arabidopsis thaliana, have only 10% of repetitive DNA. This value is much higher in the genomes of other plants, such as poplar (42%), 26 papaya (51.9%), 26 apple (42.4%), 28 and African oil palm (57%). 14
The difference in the repetitive content of the 2 American oil palm genomes reflects the discrepancy in the assembly stage of these genome projects. Elaeis oleifera AG assembly (unpublished data) is a preliminary version based on Illumina HiSeq reads, and E oleifera MG is a finished version based on 454/Roche reads. The Illumina approach has low cost and short reads, whereas Roche/454 approach has higher cost and longer reads. 29
The content of TEs found in the African oil palm genome scaffolds (39.41%) was different from that described by Singh et al 14 (57%). Nonetheless, the amount (in percentage) of LTR retrotransposons found by Beulé et al 16 is very close to the results found in this work. This study shows that, on average, 26.47% of E guineensis chromosome length is made of TEs. These numbers can be explained, in part, by the bias in partial mapping (only ~680 Mb from more than 2 Gb are already mapped), given that repeat regions are typically harder to assemble and tend to form smaller contigs, which makes it more difficult to be included in genetic mapping. Our results show many copies of full-length TEs and possible recent transposition in E guineensis (Figure 6).
Elaeis oleifera and E guineensis have high similar repetitive content, and the main difference was found in the percentage of DNA transposons in E guineensis chromosome assembly (Table 1). In what concerns the total DNA transposon value for each genome (EoAG—31 kb, EoMG—29 kb, EgMG scaf.—33 kb, and EgMG chr.—17 kb), there is some consensus among the 3 assemblages in scaffolds. The different patterns from E guineensis chromosome assembly reflect the difficulty in assembling the repeat regions. When comparing our data with content of TEs in date palm 18 (Table 1), it is possible to see a great similarity, reinforcing the close phylogenetic relationship between oil palm and date palm.
The TE effects have great influence on gene expression and genome evolution in plants. 30 Considering that exactly the same analysis was applied to these 4 different data sets, one can observe quantitative and qualitative differences in TE profiles of the African and the American oil palm genome sequences, which may be evidence of different mechanisms of transposition and regulation of such elements in the 2 species.
Diversity of microsatellites
This study has identified 155 726 microsatellite loci, which are potential molecular markers of E guineensis, E oleifera, and P dactylifera. Microsatellite markers stand out for being multiallelic, codominant, and highly reproducible. 31 A few development work and application of microsatellite markers are available for the American oil palm.32,33
It was found that dinucleotide repeats are the most frequent in the genomes studied, corroborating what is found in other plant species (48%-67%), in different data sets. 34 Within the dinucleotide class, the most frequently identified was AT. Due to the lower instability of A/T bonds, probably the mutation rate in this genome is high, 35 which ultimately increases the level of polymorphism. These observations are consistent with studies in apple, 36 Arabidopsis, 37 soybean, 38 and papaya, 39 among other plants, and show that AT-rich motifs are much more prevalent in the genomes of higher plants.
There was a clear difference in dinucleotide content between EoAG and EoMG (Figure 3), a fact that can be explained partially by the 2 different sequencing technologies (Illumina HiSeq and Roche 454) used and a fact that the assemblies are still in early versions. However, considering that several classes of TEs present similar proportions, this can also indicate a greater polymorphism in the EoMG genotype, which needs to be confirmed through population genotyping.
Our result corroborates those found in E oleifera by Zaki et al, 32 which also presents genomic microsatellites for this species. Those authors also found a high percentage of dinucleotides (63.6%) and tested 20 simple sequence repeats (SSRs) to evaluate the genetic diversity in germplasm accessions of Elaeis spp. 32 Although such analysis proved to be efficient in revealing diversity patterns, one needs to consider its limitations regarding the relationship between individuals due to reduced number of markers. Hence, our analysis demonstrated the existence of a large number of repetitive elements, including SSRs; we can now develop and validate a larger number of markers to be further used in genetic analysis. The availability of SSRs, with known genomic positions and others features, will represent an outstanding genomic resource for basic and applied research in Elaeis spp.
Regarding the microsatellite content in the evaluated genomes, there is considerable variation (between 1.65% and 2.24%) among them. This level of variation is expected to be found within species that are phylogenetically close, such as oil palm (Elaeis spp.) and date palm, 14 mainly due to 3 reasons: (1) highly polymorphic genomic microsatellites, (2) study performed on partial versions of the genomes, and (3) bias in the pipeline applied in the study. However, due to the high number of microsatellite loci identified, many should exhibit polymorphism when genotyped in vitro.
Our results on the characterization of microsatellites in the genome of P dactylifera (Table 1) are in accordance with those found by Al-Mssallem et al, 18 who identified 1.94% of SSRs in the genome (our result was 1.75%). In relation to the total number of SSRs identified, P dactylifera was the one with the lowest content among the 3 species studied (EoAG—32 947, EoMG—40 984, EgMG scaf.—48 007, EgMG chr.—30 641, and DpG—28 867). Based on this fact, we can suggest that the oil palm genome is more polymorphic than the date palm genome.
Recent studies have implemented the genome-wide strategy for the development of microsatellite markers in plants.40,41 The advantage of this approach is to get a large number of markers distributed evenly throughout the genome, which is ideal for genetic mapping studies. The construction and deployment of a microsatellite database for the scientific community would have a high impact on the genetic studies of oil palm due to the fact that this type of marker is highly informative and has a wide range of applications.
Using the tools of TRF and TRAP software, included in our pipeline, oil palm genome was systematically searched for microsatellites to develop genetic markers. This approach saves both cost and time. This result showed that in addition to SSRs developed from traditional genetic library screening 42 and other methods, oil palm genome sequence is a rich resource for the rapid identification and development of microsatellites.
Abundance of the different classes of TEs
Little differences in TE classes were found among the 4 genomes used in this study. Retrotransposons are the most abundant TEs in Elaeis spp. genomes analyzed here. This result was expected because large differences in size of plant genomes are usually associated with the presence of different amounts of retrotransposons. The larger the plant genome, the greater the chance it contains a lot of retroelements. For example, large genomes, such as barley, comprise up to 70% of these elements, 43 whereas in small genomes, such as rice, these elements represent only 17% of the genome composition. 44
In class I, there was a much greater presence of LTR compared with LINE and SINE families. The 2 superfamilies that stood out among the LTR families were Copia and Gypsy—what appears to be typical of monocot genome. 45 The LINE and SINE ratio was low because such elements appear to be more abundant in animal genome than in plant genome. 4
Class II of TEs is poorly represented in oil palm genomes, and the most present superfamilies of DNA transposons in American and African oil palms, as well as date palm, are the CMC-EnSpm and hAT elements. Members of the hAT superfamily are found in many monocotyledonous, such as those of the Ac-Ds family in maize. 46
An interesting fact was the high proportion of elements not classified in Elaeis spp. genomes. This fact can be explained by fact that the databases of repeats in monocotyledonous closely related species are not yet well described. One could overcome this limitation with a greater focus on the annotation and storage of TEs in genome projects of plants and other organisms.
In conclusion, to the best of the authors’ knowledge, this is the first detailed description of all genome repeats for American and African oil palms, as well as date palm. In the genomes analyzed, there are high diversity and abundance of TEs and microsatellites . The identified repeats are potential genetic markers for these species and will be used for assembly and genome full annotation of these complex plant genomes. Moreover, the SSRs which are being developed and validated will be used as framework markers to allow the bridging of other marker types, such as SNPs, and relevant information (eg, structure) between breeding populations. In addition, the complexity of this analysis stimulated us to produce a pipeline to improve efficiency in full TEs and tandem repeat analyses, under optimization and documentation (LS Brito et al, unpublished).
Footnotes
Acknowledgements
The authors acknowledge funding to JAFF by the Coordination for the Improvement of Higher Education Personnel (CAPES), a Foundation within the Ministry of Education in Brazil, via the Graduate Program in Plant Biotechnology, Federal University of Lavras (UFLA).
Peer review:
Five peer reviewers contributed to the peer review report. Reviewers’ reports totaled 2144 words, excluding any confidential comments to the academic editor.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The grant (01.09.0073.04—ProDendê Project) for this study was awarded by the Brazilian Ministry of Science, Technology, and Innovation (MCTI) via the Brazilian Innovation Agency—FINEP. The authors confirm that the funder had no influence over the study design, the content of article, or selection of this journal.
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
MTSJ, JAFF and EFF conceived and designed the study. JAFF, LSB and EFF developed and tested the pipeline. JAFF, LSB, APL, AAA, MTSJ and EFF wrote the manuscript. JAFF and EFF produced the figures. All authors read and approved the final manuscript.
Internet Resources
The short-read data will be available publicly through the NCBI SRA database under the accession numbers SRR3545584, SRR3545585, SRR3545586, SRR3545587, SRR3545588, and SRR3545589. The BioProject is available under accession number PRJNA319554 and the BioSample under accession number SAMN04893731.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
