Abstract
Transposable elements (TEs) are known to play a role in genome evolution, gene regulation, and epigenetics, representing potential tools for genetics research in and breeding of conifers. Recently, thanks to the development of high-throughput sequencing, more conifer genomes have been reported. Using bioinformatics tools, the TEs of 3 important conifers (
Transposable Element Insertions Influence Plant Traits
Transposons are DNA sequences that are autonomously duplicated and can “jump” out from one location on the genome and be reinserted at another place. Transposons randomly shift to different loci in the genome, which can alter the original gene structure and lead to gene silencing or inactivation, consequently changing the traits under the regulation of the functional genes.
1
For example, Zhang et al
2
found a 27.2 kb transposable element insertion in the promoter of the
Recent studies have also suggested that plant transposons have important biological functions in gene expression regulation through the generation of noncoding RNAs.4,5 Most long non-coding RNA (lncRNAs) originated from intergenic regions where transposons were abundantly distributed, and some microRNA (miRNAs) and short interfering RNAs (siRNAs) were directly or indirectly derived from transposons. These noncoding RNAs are known to regulate genes in plants. Because TE insertions could influence gene expression and function and eventually cause phenotypic variances between individuals, they have received attention for their potential value in the study of genetics.
Transposable Elements Participate in Genome Evolution
In addition to genetic association studies, TEs have also been used the study of evolutionary processes. For example, miniature inverted-repeat elements (MITEs) were considered the driving force of genomic evolution, and the number of MITEs was significantly and positively correlated with genome size. Because of their abundant variety of copy numbers and insert sites and highly species-specific characteristics, transposons have great potential in the study of gene function and genome evolution.
6
Tang et al identified an active MITE “
Interestingly, when plants experience biotic and abiotic stresses, many TEs in the plant genome are specifically activated and probably transposed under the regulation of methylation or transposons themselves, which has been found in tobacco, tomato, and other plants. 8 Some researchers have suggested that the activation of TEs under stress may accelerate genome recombination, which may also reflect the environmental adaptation process of plants during the particular lengthy evolutionary history of species.
Potential Application of TEs in Conifer Breeding
Conifers (Coniferales) are the largest and most important group of gymnosperms, as well as the most archaic branch. They have a total of 6 families, 69 genera, and 605 species. Although the number of species is limited, most of the important tree species belonging to terrestrial ecosystems evolved from conifers, placing conifers in an important taxonomic status and were usually used for the study of plant evolution history.
9
Moreover, conifers have been regarded as dominant species in the forest for more than 200 million years, not only for supporting large industries by providing wood, fiber, and energy but also for making great contributions to the global ecosystem. Due to their high ecological significance and economic importance, the genetic breeding of conifers has become an important research subject worldwide. With the rapid development of genome sequencing technology, an increasing number of high-quality plant genomes have been reported, from which researchers could get genetic data that could be used for the study of genetic mechanisms and for identifying key genes associated with important economic traits. These genomic resources could be a great resource when making breeding plans and, in addition, accelerate the process of genetic breeding in plants. Although several plant whole-genome sequences have been reported, the sequences available from conifers are limited due to their extremely large genomes and assembly difficulty. Examples of such conifers include
In a previous study, we reported our work on the identification and classification of TEs in 3 important conifers,
Summary of identified TEs in the
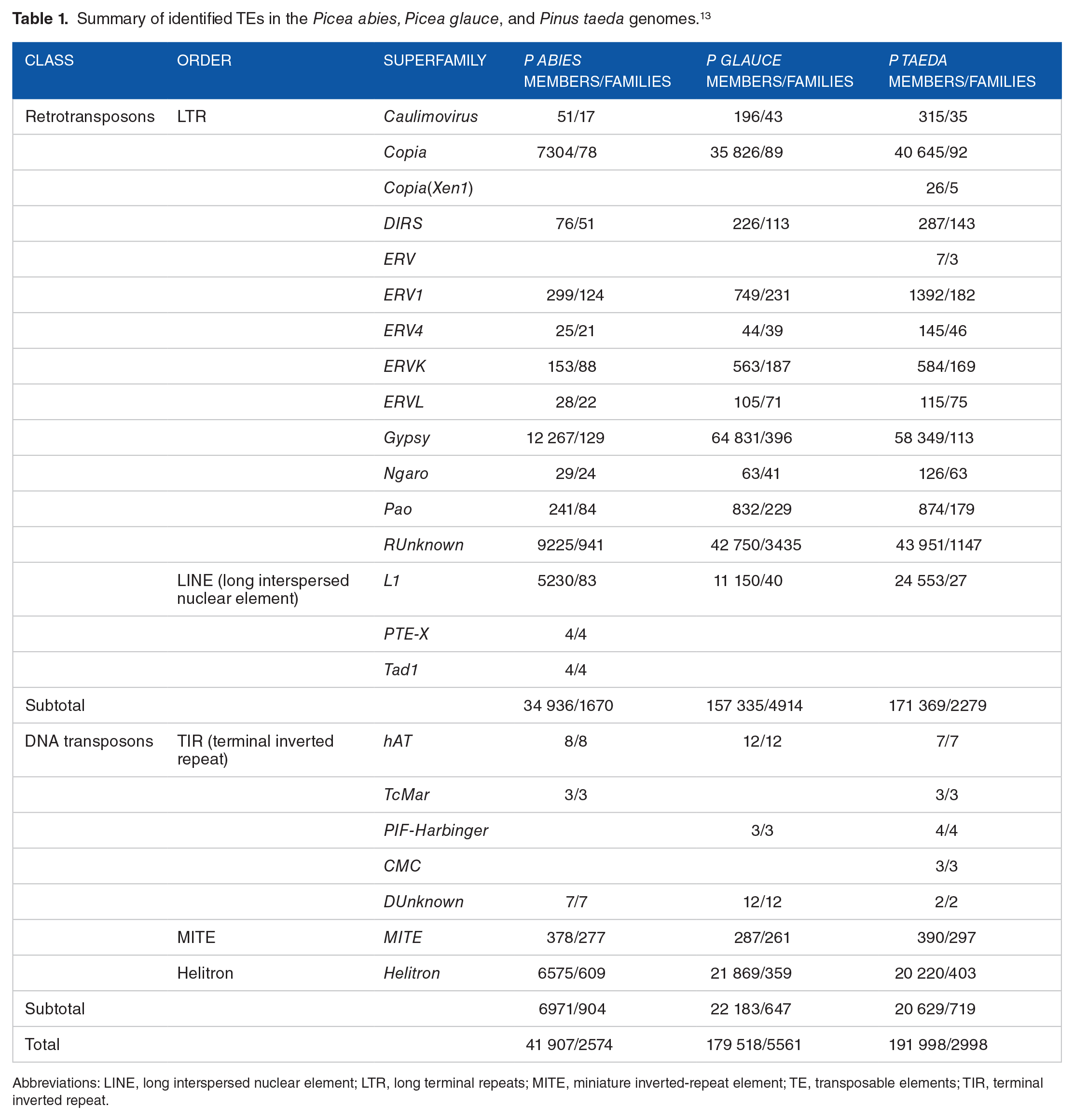
Abbreviations: LINE, long interspersed nuclear element; LTR, long terminal repeats; MITE, miniature inverted-repeat element; TE, transposable elements; TIR, terminal inverted repeat.
Transposon-based mutation libraries are important tools for the study of functional genomics in economic crops, including
Molecular markers are an important tool for genetic studies and breeding, the construction of linkage maps, and population genetics analysis. Transposable elements constitute most of the repetitive sequences and are distributed throughout the genome in plants; thus, TE could have great potential to be explored as molecular markers. Liang et al
27
identified 336 long terminal repeats (LTR) effective primer pairs that could amplify DNA from
Conclusions
TEs have great potential to be used in conifer breeding; however, a limited number of related studies have been reported. In future studies, mutant libraries of conifers could be constructed to identify essential genes associated with target traits, according to the genotype and phenotype variations in the library. Moreover, TE-based molecular markers could be developed for use in molecular breeding in conifers. It is important to mention that the massive genome sizes and relatively long life cycles of conifers make the use of TEs in conifer breeding more challenging than in annual crop breeding and much foundational work is needed.
Footnotes
Acknowledgements
The authors would like to thank Thirteenth Five-Year Plan for Key & Research Project of China (2017YFD0600606-09) for the financial support.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Fundamental Research Funds of Chinese Academy of Forestry (CAFYBB2019ZX001); Thirteenth Five-Year Plan for Key & Research Project of China (2017YFD0600606-09).
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
JW, NL, FY, and YX composed, reviewed, and approved the final article.
