Abstract
Hypoxia is an important environmental stressor leading to endocrine disruption and reproductive impairment in fish. Although the hypoxia-inducible factor 1 (HIF-1) is known to regulate the transcription of various genes mediating oxygen homeostasis, its role in modulating steroidogenesis-related gene expression remains poorly understood. In this study, the regulatory effect of HIF-1 on the expression of 9 steroidogenic enzyme genes was investigated in zebrafish embryos using a “gain-of-function and loss-of-function” approach. Eight of the genes,
Introduction
Hypoxia caused by aquatic ecosystem enrichment with nutrients and organic matter is a major global threat that is predicted to worsen as climate change progresses. The adverse impacts of aquatic hypoxia include significant reductions in fisheries production, fish growth, alteration of species composition, and mass fish mortality.1,2 Hypoxia impairs fish reproduction by inhibiting testicular and ovarian development, reducing sperm and egg quality, affecting fertilization, and hatching and influencing the survival of larvae and juvenile fitness. 3 Ovarian and testicular growth, gametogenesis, and oocyte and sperm production were significantly reduced in Atlantic croakers in hypoxic regions of the Mississippi River4,5 and in the northern Gulf of Mexico. 6
In fish, the inhibition of key reproductive processes by hypoxia is associated not only with reduced metabolism but also with the repression of specific hormones and hormone receptors along the hypothalamus-pituitary-gonad (HPG) axis.3,7 Numerous in vivo studies have shown that fish reproduction is affected by hypoxia via alteration of the steroidogenesis pathway.7–11 Steroid hormones biosynthesis is a fundamental process for reproduction in vertebrates, and all steroid hormones are derived from cholesterol. The rate-limiting step in steroid production is the delivery of cholesterol from the outer to the inner mitochondrial membrane, which is mediated first by the steroidogenic acute regulatory (StAR) protein
12
followed by the conversion of cholesterol to pregnenolone by the cytochrome P450 cholesterol side-chain cleavage (P450scc) enzyme encoded by
Hypoxia-inducible factor 1 is a heterodimeric protein consisting of an oxygen-labile HIF-1α subunit and a constitutively expressed HIF-1β subunit.
20
Due to the oxygen-sensitive oxygen-dependent degradation domain (ODD) in the HIF-1α subunit, HIF-1 is rapidly degraded under normoxia.
21
HIF-1α is stabilized under hypoxia and is transported into the nucleus where it dimerizes with HIF-1β
22
to form a heterodimeric complex that binds to the hypoxia-responsive elements (HREs) in the promoters of HIF-target genes.
23
HIF-1 has been shown to negatively regulate
Although more than 100 different genes are now known to be controlled by HIF-1, whether steroidogenic enzyme genes are regulated under hypoxia through HIF-1 remains poorly understood. Zebrafish (
Materials and Methods
Zebrafish maintenance
Wild-type adult zebrafish were maintained under a constant 14 hours:10 hours light:dark cycle at 28°C. Flow through systems were set up in the laboratory to provide a constant temperature and normoxic (7.0 ± 0.2 mg O2 L−1) or hypoxic (1.0 ± 0.2 mg O2 L−1) environments during experimental periods as described previously by Shang et al. 10 Embryos were obtained by natural spawning and cultured in zebrafish E3 embryo medium (5 mM NaCl, 0.17 mM KCl, 0.33 mM CaCl2, 0.33 mM MgSO4, and 5% methylene blue) at 28.5°C and staged according to Kimmel et al. 27
Plasmid construction
The full-length zHIF-1α complementary DNA (cDNA) was amplified by reverse transcription polymerase chain reaction (RT-PCR) from zebrafish embryos and cloned into the pGEM-T Easy Vector (Promega, Madison, WI, USA) to form pGEMT-zHIF-1α. Due to the short half-life of the wild-type HIF-1α protein under normoxia, 28 a zHIF-1α mutant construct (pGEMT-zHIF-1α-ΔODD) was synthesized to increase the stability of the zHIF-1α protein during its overexpression in normoxic embryos. To form pGEMT-zHIF-1α-ΔODD, 2 sites containing proline residues (proline-414 and proline-557) in wild-type ODD (at nucleotide positions 1234-1251: TTG GCA CCT GCA GGC and at nucleotide positions 1659-1680: ATG CTG GCT CCT TAC ATC CCA) were deleted from pGEMT-zHIF-1α using the Gene Tailor Site-Directed Mutagenesis Kit (Invitrogen, Carlsbad, CA USA).
To monitor the overexpression of zHIF-1α in developing zebrafish embryos, a zHIF-1α-eGFP fusion expression vector was constructed. The open reading frame (ORF) of enhanced green fluorescent protein (eGFP) was amplified from the pEGFP-N1 (Clontech Laboratories, Mountain View, CA, USA) plasmid using the gene-specific primers EGFP-F-Sma: 5′-TCC
Hypoxia exposure experiment
Zebrafish embryos were divided into 2 groups: 1 group was reared in a hypoxic system and the other in a normoxic system. Tanks used for hypoxia exposure were set up as previously described. 10 Dissolved O2 concentrations were monitored continuously using dissolved oxygen meters (Cole-Parmer 01972-00, Vernon Hills, IL, USA) and polarographic probes (Cole-Parmer 5643-00), and adjustment was made using a dissolved oxygen controller (Cole-Parmer 01972-00). Fertilized embryos (4 hours postfertilization [hpf]) were placed in net cages and allowed to develop to desired stages of development (24, 48, or 72 hpf). Each treatment consisted of 5 replicates of 60 embryos each. At the end of the experiments, the embryos were immersed in 200 µL of TRI Reagent (Molecular Research Center, Cincinnati, OH, USA) and stored at −80°C until RNA was extracted. The embryos used for hormone analysis were snap-frozen in liquid nitrogen and stored at −80°C. Animal care and experiments were conducted in accordance with the City University of Hong Kong animal care guidelines.
RNA isolation and first-strand cDNA synthesis
Total RNA was extracted using TRI Reagent (Molecular Research Center) according to the manufacturer’s instructions. First-strand cDNA was synthesized in a 25-µL reaction that contained 0.25 µg (0.5 µg/µL) of random primer, 5 µL of 5 × reaction buffer, 1.25 µL of 10 mM deoxynucleotide triphosphates, 0.65 µL of 40 U ribonuclease (RNase inhibitor) (Invitrogen), 1 µL of 200 U M-MLV Reverse Transcriptase (Promega), and 1 µg of total RNA, which was pretreated with RNase-free deoxyribonuclease I (Invitrogen) to eliminate genomic DNA contamination. cDNA synthesis was performed at 37°C for 1 hour. The cDNAs were kept at −20°C until they were analyzed by real-time PCR.
Real-time PCR
Real-time PCR, which was used to quantify gene expression, was performed using a StepOnePlus Real-Time PCR System (Applied Biosystems, Foster City, CA, USA) as described previously.
11
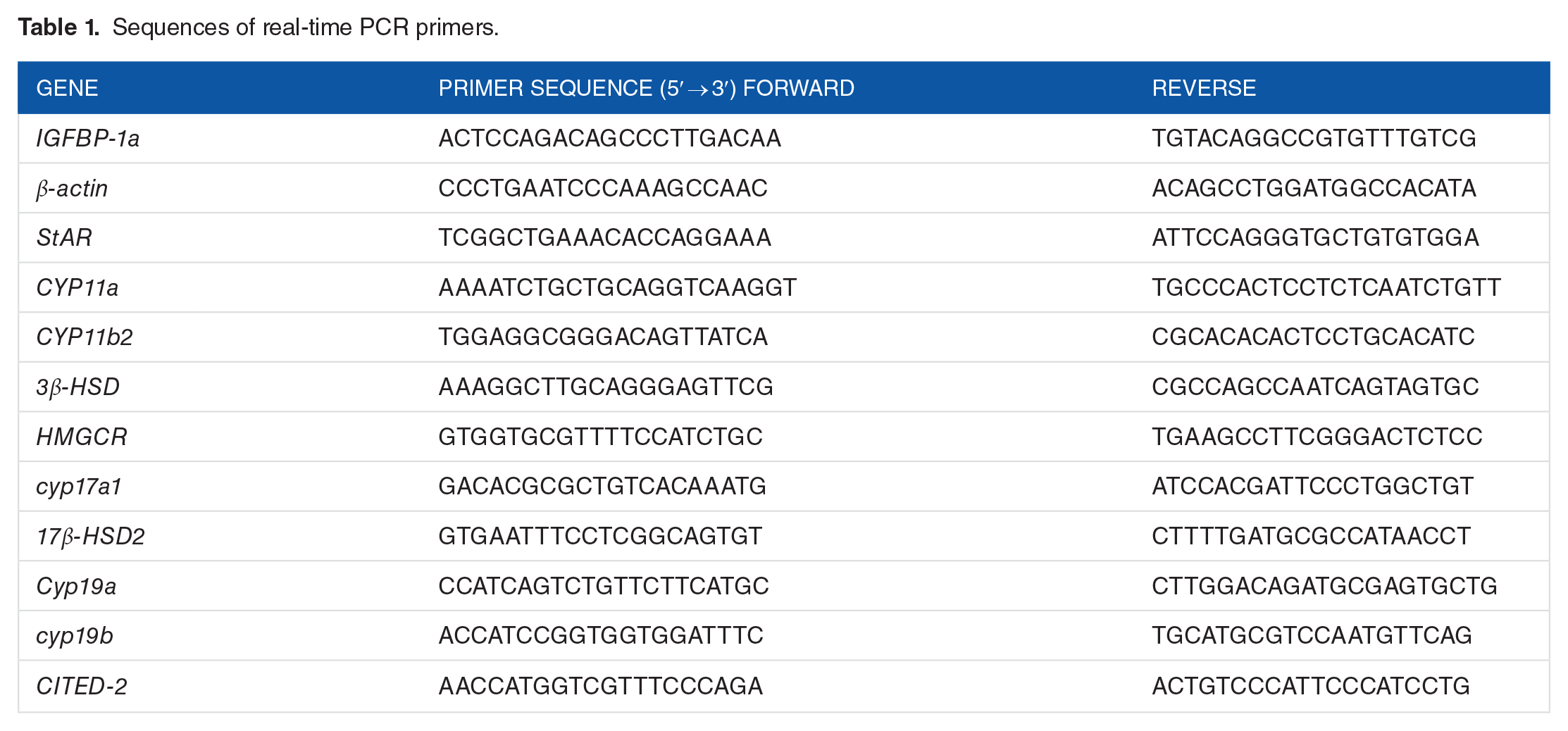
PCR assays were conducted using the SYBR Green–based detection method (#4367659; Applied Biosystems, Warrington, UK) according to the manufacturer’s instructions. Sequences of the primers used for real-time PCR are shown in Table 1. Melting curve analysis was performed after the PCR was completed to assess the amplification specificity. The identity of the PCR amplicons was confirmed by DNA sequencing. The cycling conditions were as follows: initial denaturation at 95°C for 10 minutes, followed by 40 cycles performed at 95°C for 15 seconds, and 0°C for 30 seconds. β-Actin was employed as an endogenous control for normalization. All PCR reactions were performed in triplicate. Fold change was calculated according to the following formula:
Sequences of real-time PCR primers.

Estradiol (E2) and testosterone (T) measurement
In total, 60 zebrafish embryos from each replicate were collected and immediately stored at −80°C for hormone extraction (E2 and T) as described by Yu et al. 11 E2 and T levels were measured using a commercial competitive enzyme-linked immunosorbent assay (ELISA) kit (Cayman Chemical, Ann Arbor, MI, USA).
Overexpression of zHIF-1α
Capped eGFP and zHIF1α-ΔODD-eGFP messenger RNAs (mRNAs) were synthesized from linearized pCMV-eGFP and pCMV-zHIF-1α-ΔODD-eGFP, respectively, using the T7 mMESSAGE mMACHINE Kit (Ambion, Austin, TX, USA) according to the manufacturer’s instructions. mRNAs were microinjected at a concentration of 0.5 µg/µL in Hanks buffer containing 0.15% phenol red, and each embryo was injected with 2 nL mRNA at a 1 to 2 cell stage using a FemtoJet microinjection system (Eppendorf AG, Hamburg, Germany). Five replicates of 60 fertilized embryos each were used for both treatments. Injected embryos were incubated under normoxic conditions and sampled at various developmental stages for analyses.
zHIF-1α knockdown
Antisense morpholino oligonucleotides (MOs) were obtained from Gene Tools (Philomath, OR, USA). An MO targeted against the translation initiation region of zHIF-1α (zHIF-1α MO: 5′-CAG TGA CAA CTC CAG TAT CCA TTC C-3′) was used to knockdown the zHIF-1α protein. The standard control MO (5′-CCT CTT ACC TCA GTT ACA ATT TAT A-3′) was used as an injection control. The optimized morpholino amount for zHIF-1α knockdown experiments was 8 ng/embryo. Injected embryos were reared to the desired developmental stages under hypoxic or normoxic conditions as described above.
Statistical analysis
The Student
Results
Hypoxia alters the expression pattern of steroidogenic enzyme genes
To confirm that HIF-1 signaling was activated in the zebrafish embryos under the experimental hypoxic conditions employed in this study, real-time PCR was performed to measure the expression of 2 HIF-1 target genes,
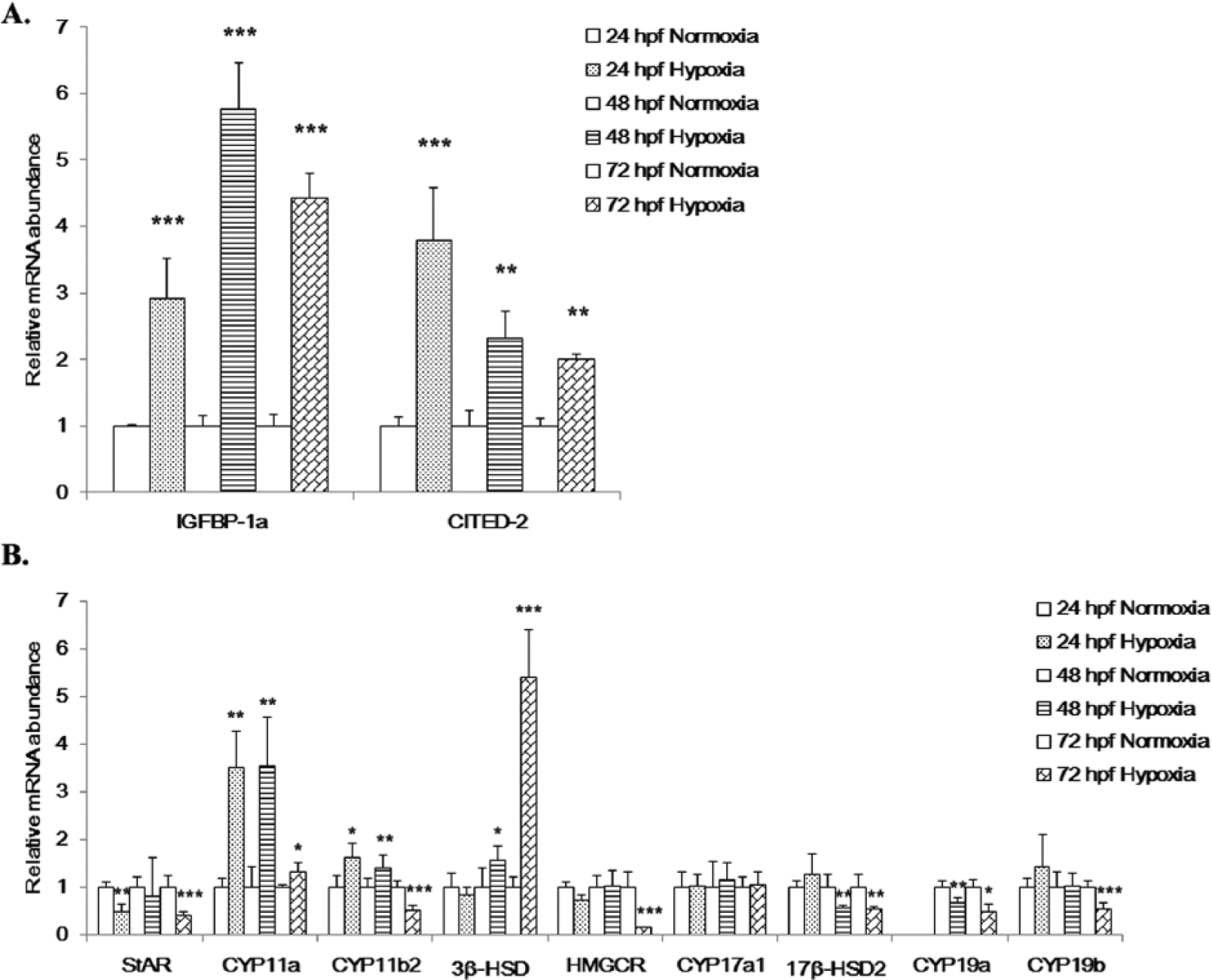
Effects of hypoxia on the expression of hypoxia markers and steroidogenic genes. Effect of hypoxia on the expression of (A) the HIF-1 target genes, IGFBP-1a and CITED-2 and (B) 9 steroidogenic enzyme genes. Fertilized zebrafish embryos were exposed to normoxic (7.0 ± 0.2 mg O2 L−1) or hypoxic (1.0 ± 0.2 mg O2 L−1) conditions for 24, 48, or 72 hours. Gene expression was quantified using real-time PCR and normalized against β-actin mRNA. Data are presented as the mean relative fold change ± SD with respect to the normoxia control (its expression level was arbitrarily set to 1). Expression levels significantly different from the control are indicated by asterisks (
The mRNA expression levels for 9 selected steroidogenic genes in embryos exposed to hypoxia were differentially affected (Figure 1B). Five steroidogenic genes were significantly downregulated.
Overexpression of zHIF-1α mRNA delays embryonic development and the expression pattern of steroidogenic genes
To investigate the relationship between zHIF-1α expression and steroidogenesis in zebrafish, overexpression and knockdown of zHIF-1α were conducted by microinjection of the in vitro–transcribed zHIF-1α-ΔODD-eGFP mRNAs and zHIF-1α translation-blocking morpholinos (MO) into embryos, respectively. Five treatment replicates were used in each experiment. The total numbers of embryos injected with the eGFP mRNA (control) and zHIF-1α-ΔODD-eGFP mRNA were 719 and 1081, respectively. Based on the embryo staging criteria described by Westerfield, 31 the development of embryos microinjected with zHIF-1α-ΔODD-eGFP mRNA was found to be delayed by at least 6 hours when compared with that of the eGFP-injected and uninjected controls at 24 hpf (Figure 2A).
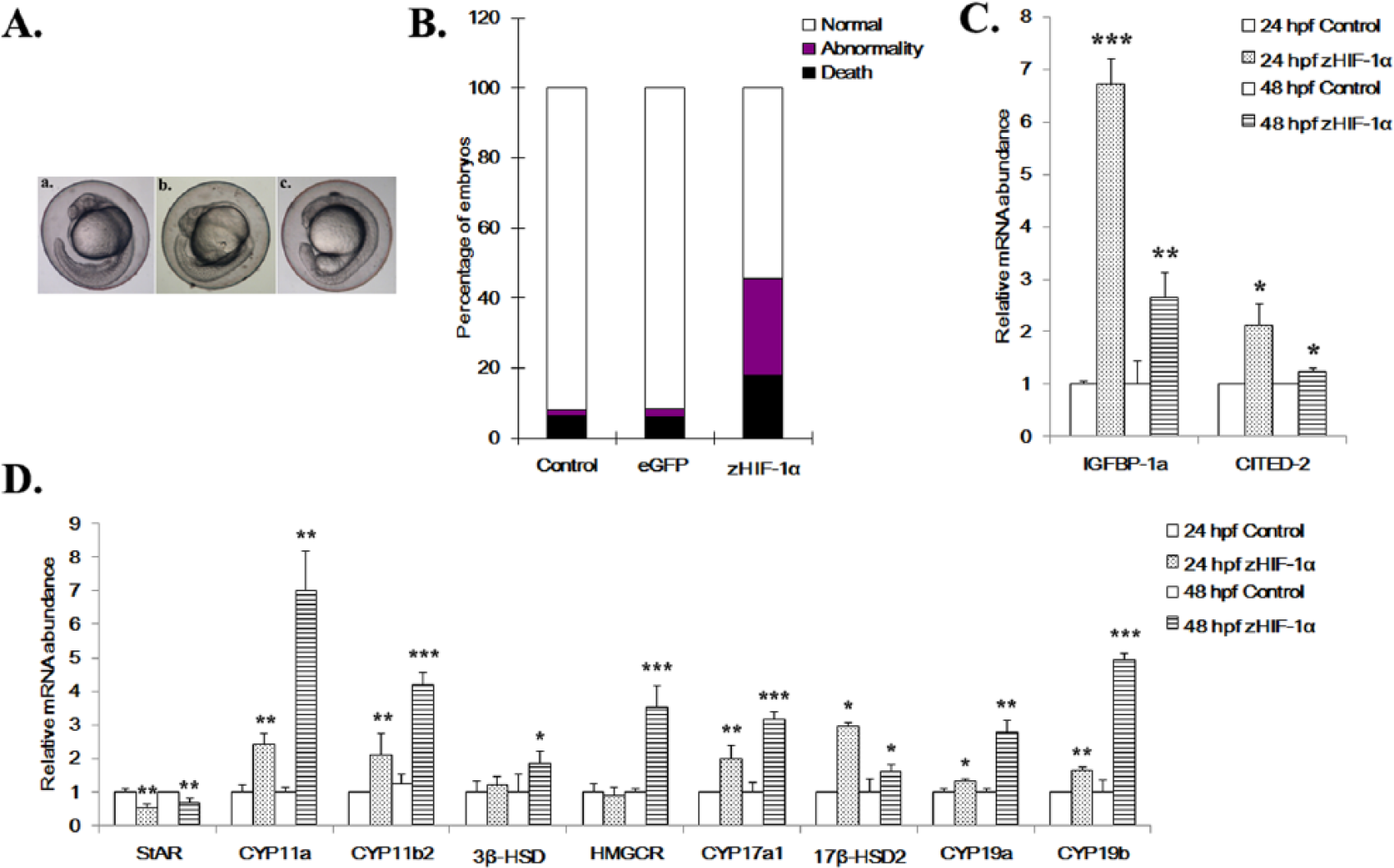
Effects of zHIF-1α overexpression on embryonic development and the expression of hypoxia markers and steroidogenic genes. (A) Development of zebrafish embryos at 24 hpf. a, uninjected control embryo; b, embryo microinjected with eGFP mRNA shows normal development (Prim-6 stage); c, embryo microinjected with zHIF-1α-ΔODD-eGFP mRNA shows developmental retardation (at approximately 18-somite stage). (B) Percentage of mortality and morphological abnormality in zebrafish embryos (24 hpf) microinjected with zHIF-1α mRNA. The effect of zHIF-1α overexpression on steroidogenic gene expression was examined by microinjecting zHIF-1α mRNA into 1- to 2-cell stage zebrafish embryos and maintained under normoxic conditions. Zebrafish embryos microinjected with eGFP mRNA and maintained under normoxia were used as the control. Because no statistically significant difference was observed between the data sets of the microinjected control and the no-injection control (One-way analysis of variance with a
At 24 hpf, 17.8% of the embryos microinjected with zHIF-1α-ΔODD-eGFP mRNA were dead compared with the 6.1% dead embryos observed with eGFP mRNA microinjection. The percentage of mortality of the embryos microinjected with zHIF-1α mRNAs was approximately 3-fold greater than that of the eGFP control (Figure 2B). In addition, the percentage of abnormal embryos observed in the zHIF-1α microinjection experiments was 27.5% (297/1081), whereas only 2.2% (16/719) of abnormal embryos were observed in the eGFP control group. Following microinjection with the zHIF-1α-ΔODD-eGFP mRNAs,
Following microinjection of the zHIF-1α mRNAs, the expression patterns of all 9 steroidogenic genes were found to be affected in zebrafish embryos at 24 and 48 hpf (Figure 2D). With the exception of
Knockdown of zHIF-1α alters expression of steroidogenic genes
The experiment in which zHIF-1α was knocked down with a morpholino was conducted to complement the study in which zHIF-1α was overexpressed. Following zHIF-1α knockdown, morphological abnormalities were observed in morphant embryos at 48 hpf under normoxia (Figure 3A). The observed deformities included small head circumference, bent notochords, deformed hearts, distorted abdomens, and short curved tails. Knocking down zHIF-1α also resulted in significantly shorter zebrafish embryo body length at the same developmental stage (Figure 3A). In morphant embryos at 24 hpf, the expression levels of
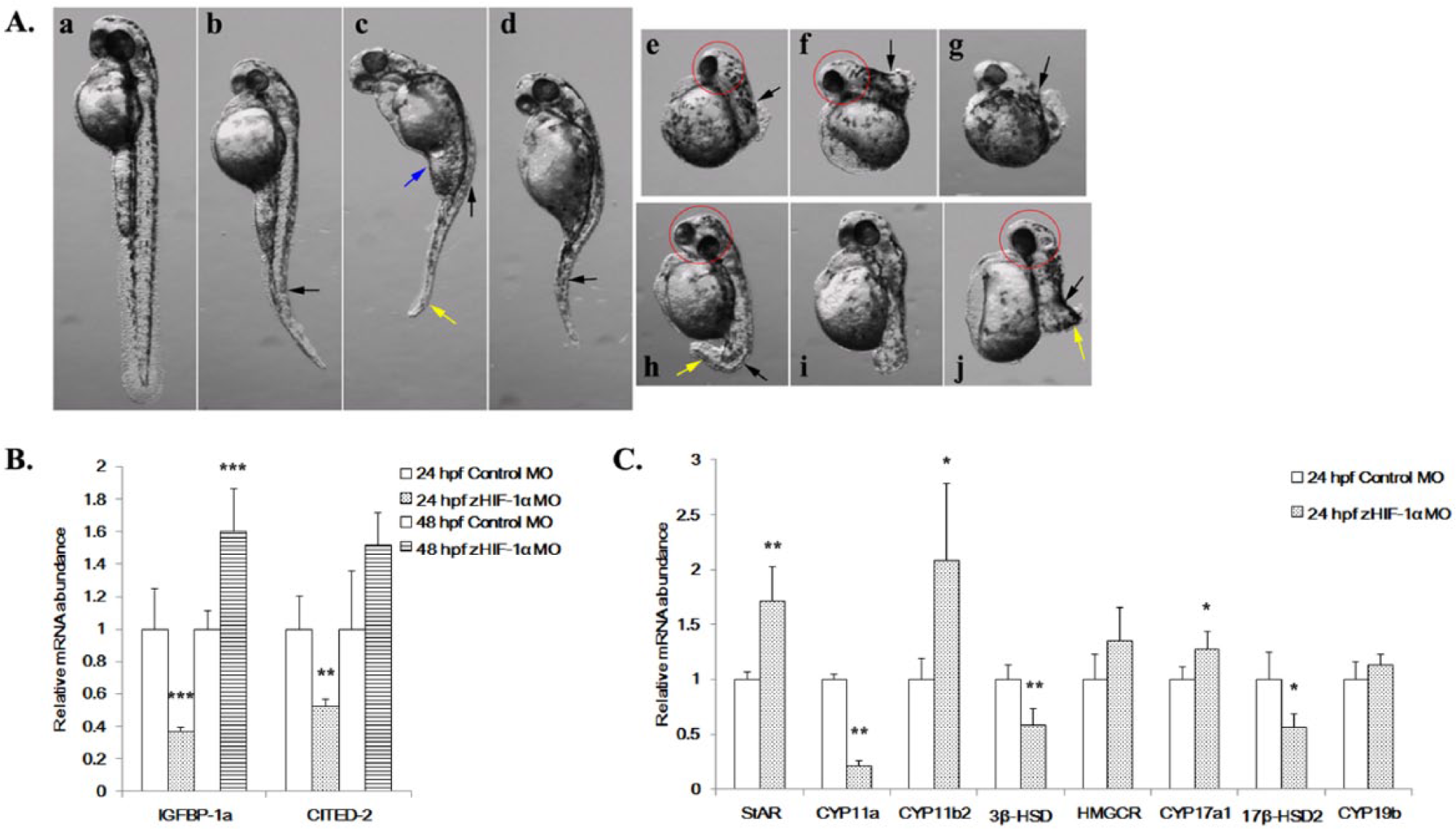
Effects of zHIF-1α knockdown on zebrafish embryonic development and the expression of hypoxia markers and steroidogenic genes under normoxia. (A) Morphological abnormalities of the zHIF-1α knockdown embryos at 48 hpf under normoxia. The effect of zHIF-1α knockdown on steroidogenic gene expression was examined by microinjecting zHIF-1α MOs into 1- to 2-cell stage zebrafish embryos and maintained under either normoxia or hypoxia. Zebrafish embryos microinjected with a standard control MO were used as controls. All morphants were imaged at the left lateral view. a, Control: embryo injected with control MO; b-j, embryos microinjected with zHIF-1α MO; b, c, and d, morphants with decreased body length, bended notochords (black arrows), and distorted abdomens (blue arrows); e-j, morphants with bended notochords, small head circles (red circles), and short curved tail (yellow arrows). (B) Effects of zHIF-1α knockdown on IGFBP-1a and CITED-2 expression under normoxia. (C) Effects of zHIF-1α knockdown on steroidogenic gene expression under normoxia. Gene expression was quantified by real-time PCR and normalized against β-actin mRNA. Data are presented as the mean relative fold change ± SD with respect to the gene expression level in the control (its expression level was arbitrarily set to 1) for each experiment. Expression levels significantly different from the control are indicated by asterisks (
The expression patterns of the steroidogenic genes in the zHIF-1α morphants were different from those of the control embryos (microinjected with control MO) under normoxia (Figure 3C). Three steroidogenic genes were specifically upregulated in the zHIF-1α morphants.
To confirm whether changes in the expression of steroidogenic genes under hypoxia (Figure 1B) were due to regulation by zHIF-1α, embryos in which zHIF-1α had been knocked down were exposed to hypoxia. Comparing embryos in which zHIF-1α had been knocked down with the control MO-injected embryos under hypoxia,
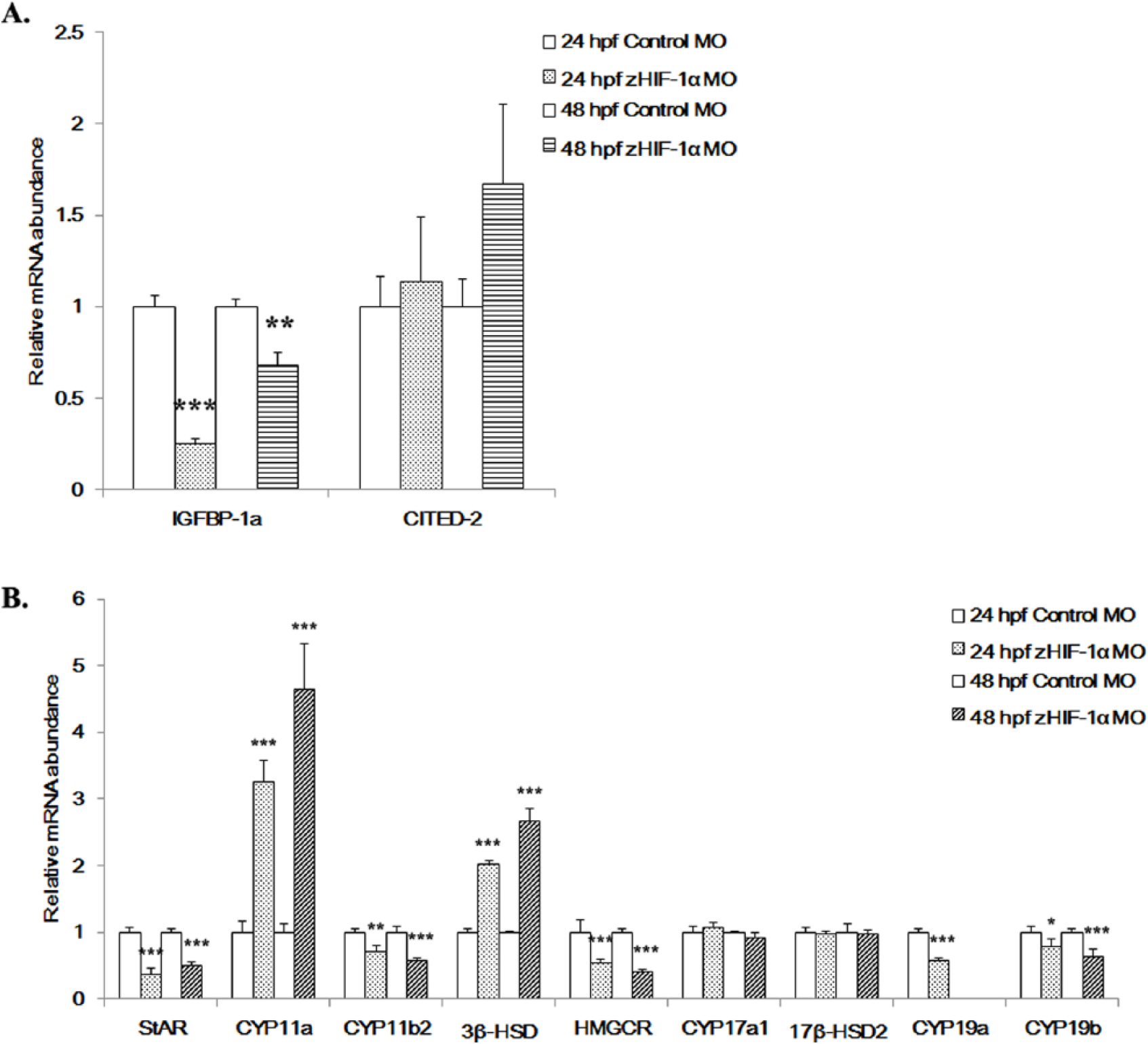
Effects of zHIF-1α knockdown on the expression of hypoxia markers and steroidogenic genes under hypoxia. (A) Effects of zHIF-1α knockdown on the hypoxia marker genes IGFBP-1a and CITED-2 under hypoxia. Methods for zHIF-1α knockdown are detailed in Figure 3. (B) Effects of zHIF-1α knockdown on steroidogenic gene expression under hypoxia. Gene expression was quantified by real-time PCR and normalized against β-actin mRNA. Data are presented as the mean relative fold change ± SD with respect to the gene expression level in the hypoxia control (its expression level was arbitrarily set to 1) for each experiment. Expression levels significantly different from the control are indicated by asterisks (
In the zHIF-1α morphant embryos under hypoxia,
E2 and T levels
The concentrations of E2 and T in whole embryos varied under normoxic and hypoxic conditions and with or without overexpression and knockdown of zHIF-1α (Figure 5). The concentrations of E2 and T in whole embryos under hypoxia were 2.2-fold less and 1.8-fold greater than those of the normoxic controls (Figure 5A), which resulted in a 3.9-fold greater T/E2 ratio (from 3.0 ± 0.42 (normoxia) to 11.73 ± 1.28 (hypoxia) (Figure 5B). The concentrations of both E2 and T were lower in embryos overexpressing zHIF-1α under normoxic conditions and in zHIF-1α knockdown embryos under hypoxic conditions (Figure 5A). The T/E2 ratio in the zHIF-1α morphants under hypoxia was similar to that of the control embryos under normoxia. However, the T/E2 ratio in embryos overexpressing zHIF-1α under normoxia was 10.3-fold lower than those maintained under hypoxia (Figure 5B).
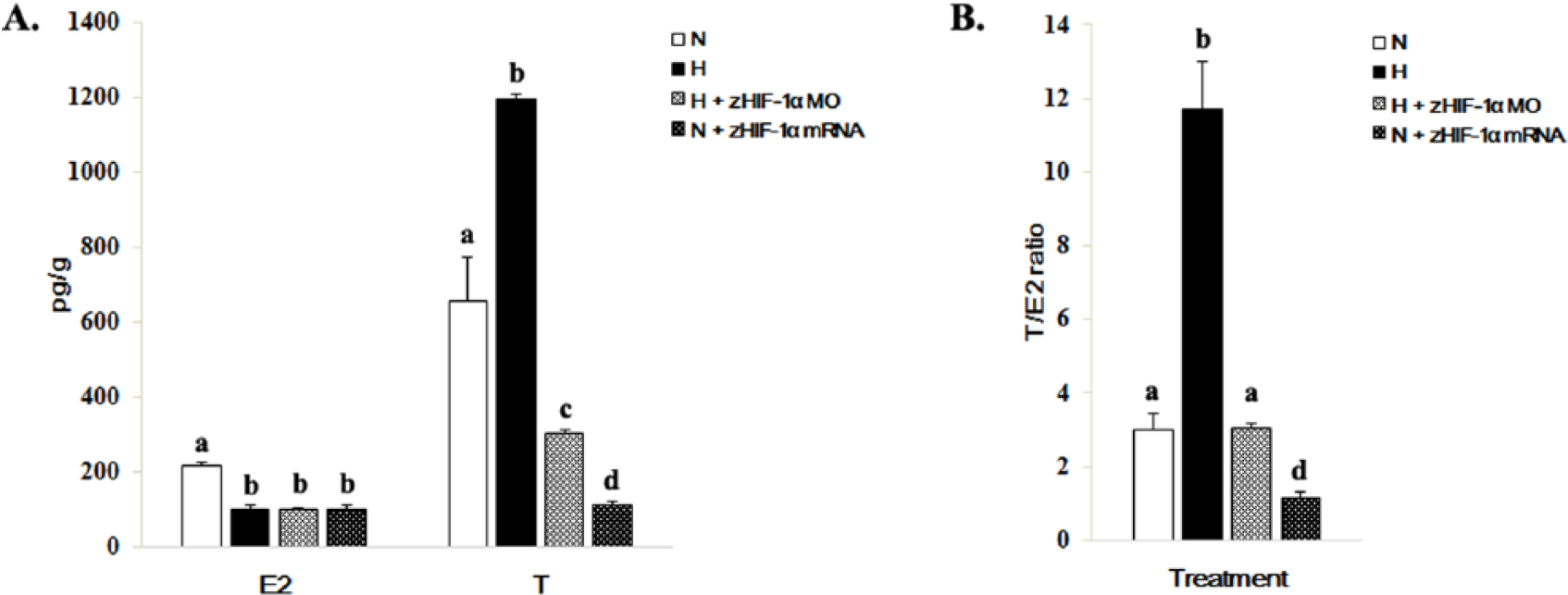
Effects of hypoxia and overexpression of zHIF-1α on E2 and T levels in zebrafish embryos. (A) Effects of hypoxia and zHIF-1α expression on E2 and T levels. Methods for hypoxic exposure and zHIF-1α overexpression and knockdown are detailed in Figures 1 to 3, respectively. Hormone levels in whole zebrafish embryos (5 replicates of 60 pooled embryos each) were measured by the ELISA method. The data are presented as the mean concentration of hormone (in picograms per gram of embryo wet weight) ± SD. The different letters above the bars indicate statistically significant differences between the indicated groups for the same hormone (
Discussion
There is increasing evidence that the HIF-1 signaling system plays essential roles in embryonic development in vertebrates through the activation of genes that regulate energy metabolism and survival. Previous studies have reported the expression of HIF-1α mRNA in the brain, blood vessels, somites, notochord, retina and optic tectum in zebrafish embryos,
32
and HIF-1α-deficient mouse embryos (HIF-1α−/−) show defects in the formation of blood vessels and the neural fold as well as malformations in the cardiovascular system.33,34 HIF-1 has also been demonstrated to affect the development of zebrafish embryos through the regulated expression of
There is ample evidence that hypoxia impairs the sexual and reproductive development of fish through perturbation of the HPG axis at multiple levels, including enzymes that control steroidogenesis.5,10 A qualitative comparison of the results from this study on the expression patterns of steroidogenic genes in zebrafish embryos exposed to hypoxia, embryos overexpressing zHIF-1α mRNA, and zHIF-1α knockdown embryos is provided in Table 2. Changes in the expression of genes encoding steroidogenic enzymes, including
Comparison of steroidogenic gene expression patterns in hypoxic, zHIF-1α overexpressing, and zHIF-1α knockdown embryos.
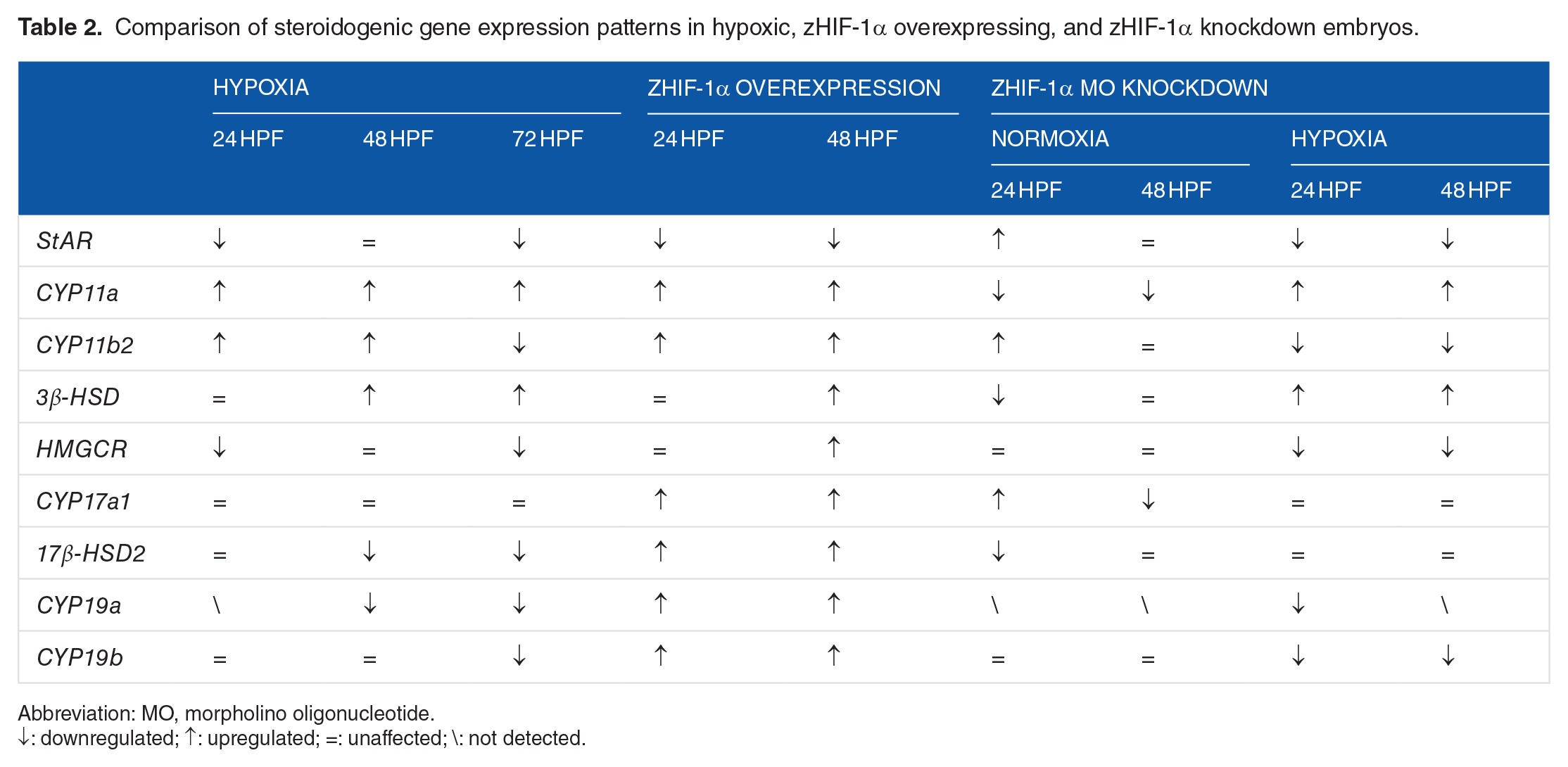
Abbreviation: MO, morpholino oligonucleotide.
↓: downregulated; ↑: upregulated; =: unaffected; \: not detected.
The results of downregulating of
Specifically, we showed that hypoxia and zHIF-1α overexpression significantly downregulated the expression of
Conclusions
In this study, it is demonstrated that zHIF-1 regulates the expression of several steroidogenic enzyme genes in zebrafish embryos. Specifically, the regulated expression of
Footnotes
Acknowledgements
The authors thank Professor SH Cheng’s group for providing zebrafish embryos and for using microinjection facilities in her lab.
Peer Review:
Five peer reviewers contributed to the peer review report. Reviewers’ reports totaled 1240 words, excluding any confidential comments to the academic editor.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by a grant from the General Research Fund (PJ9041727) from the Research Grants Council of Hong Kong Special Administrative Region, People’s Republic of China.
Declaration of Conflicting Interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
TT conceived and designed the experiments; collected, analyzed, and interpreted the data; wrote the first draft of the manuscript. RMKY performed literature review, contributed to the writing and made critical revisions of the manuscript. RSSW contributed to the writing of the manuscript and approved the final version. RYCK designed the study and performed a critical review of the manuscript. All authors reviewed and approved the final manuscript.
