Abstract
Objectives:
Captopril is a commonly used therapeutic agent in the management of renovascular hypertension (high blood pressure), congestive heart failure, left ventricular dysfunction following myocardial infarction, and nephropathy. Captopril has been found to interact with proteins that are significantly associated with bladder cancer (BLCA), suggesting that it could be a potential medication for BLCA patients with concurrent hypertension.
Methods:
DrugBank 5.0 was utilized to identify the direct protein targets (DPTs) of captopril. STRING was used to analyze the multiple protein interactions. TNMPlot was used for comparing gene expression in normal, tumor, and metastatic tissue. Then, docking with target proteins was done using Autodock. Molecular dynamics simulations were applied for estimate the diffusion coefficients and mean-square displacements in materials.
Results:
Among all these proteins, MMP9 is observed to be an overexpressed gene in BLCA and its increased expression is linked to reduced survival in patients. Our findings indicate that captopril effectively inhibits both the wild type and common mutated forms of MMP9 in BLCA. Furthermore, the LCN2 gene, which is also overexpressed in BLCA, interacts with captopril-associated proteins. The overexpression of LCN2 is similarly associated with reduced survival in BLCA. Through molecular docking analysis, we have identified specific amino acid residues (Tyr179, Pro421, Tyr423, and Lys603) at the active pocket of MMP9, as well as Tyr78, Tyr106, Phe145, Lys147, and Lys156 at the active pocket of LCN2, with which captopril interacts. Thus, our data provide compelling evidence for the inhibitory potential of captopril against human proteins MMP9 and LCN2, both of which play crucial roles in BLCA.
Conclusion:
These discoveries present promising prospects for conducting subsequent validation studies both in vitro and in vivo, with the aim of assessing the suitability of captopril for treating BLCA patients, irrespective of their hypertension status, who exhibit elevated levels of MMP9 and LCN2 expression.
Highlights
• MMP9 overexpression in BLCA, linked to reduced survival, is effectively inhibited by captopril.
• Molecular docking identifies key amino acid residues for captopril interaction, suggesting potential therapeutic use for BLCA.
• Elevated LCN2 levels, associated with poor BLCA survival, also interact with captopril-related proteins.
• Captopril’s inhibitory effects on both MMP9 and LCN2 hold promise for treating BLCA patients with hypertension.
Introduction
Captopril is commonly used in the treatment of renovascular hypertension, congestive heart failure, left ventricular dysfunction following myocardial infarction, and nephropathy. Captopril is an effective, competitive inhibitor of the angiotensin-converting enzyme (ACE), which transforms angiotensin I (ATI) into angiotensin II (ATII). 1 ATII is a strong vasoconstrictor hormone, that means it constricts the blood vessels, leading to an increase in blood pressure. Therefore, Captopril is used to treat high blood pressure by blocking the production of ATII. By blocking this hormone. Thus, captopril helps to relax blood vessels and reduce blood pressure. Captopril has been also shown to have preventive benefits against organ damage brought on by hypertension (high blood pressure). According to 96 studies involving 1570 patients, captopril was found to be superior to placebo and equivalent to beta-blocker or diuretic in the treatment of hypertension. 2 Furthermore, it was found that Captopril prevents insulin-dependent diabetic nephropathy from worsening renal function. 3 Another study showed that patients on long-term captopril therapy may experience unexpected advantages from its antiangiogenic properties. 4 Thus, Captopril plays an important role in the treatment of hypertension.
Importantly, Hypertension has also been found to be associated with an increased risk of developing certain types of cancer and cancer-related mortality rates. 5 The association between hypertension and cancer diseases are multidimensional, and they have common risk factors such as smoking, diabetes mellitus, and obesity. 6 The prevalence of hypertension in patients with cancer was similar in the general adult population. 7 Research demonstrated that Captopril may have some anticancer properties. Notably, Captopril could be a promising option in the treatment of lung cancer. 8 It was found that Captopril could be used to prevent or reduce the severity of radiation-induced pulmonary toxicity in patients, radiation treatment for lung cancer. 9 Moreover, several studies in prostate cancer showed that captopril could inhibit the growth and proliferation of prostate cancer cells in vitro and in vivo. 10 Thus, Captopril medication may have a protective effect against the development of prostate cancer. 11
In this study, we introduced an integrated analysis as an effective approach, which could analyze and prioritize candidate genes through the public databases. Firstly, we discovered direct protein targets (DPTs) of captopril in DrugBank 5.0. Subsequently, we analyzed the protein-protein interaction (PPI) network and signaling pathways of captopril DPTs. Refined analysis of KEGG (Kyoto Encyclopedia of Genes and Genomes) pathways showed that captopril is linked to various types of cancer, particularly Bladder cancer (BLCA). Another study also indicates that captopril can help maintain bladder structure. 12 Next, we identified 2 genes (MMP2 and MMP9) as the most significant target genes of captopril in BLCA. Simultaneously, we analyzed the expression of these genes in BLCA, comparing them with normal samples, and examined the survival of BLCA patients associated with these genes. Finally, molecular docking and molecular simulation were employed to analyze the interaction pattern of captopril with the proteins associated with the poor survival of BLCA. Consequently, this study aims to elucidate the potential use of captopril in managing hypertension or heart failure in BLCA patients.
Material and Methods
Recognition for direct protein targets (DPTs) of captopril
DrugBank is a web-based database that contains detailed molecular data on drugs, their mechanisms of action, interactions, and targets. 13 We found 5 target proteins for the drug captopril using DrugBank, which may have implications in the treatment of various diseases.
Protein-protein interaction (PPI) network and signaling pathways of captopril DPTs analysis
STRING (search tool for the retrieval of interacting genes) resource is available online at https://string-db.org/ STRING database contains all known relationships between proteins, including both physical interactions and functional associations between them. 14 Therefore, we also used STRING database to identify multiple proteins associated with captopril, which are obtained from DrugBank. Initially, the search module was employed to find multiple proteins by their names (“DPTs”) and the organism (“Homo sapiens”). Subsequently, the following settings were configured: the active interaction source was set to “Text mining and experimentation,” the interaction score required was set to “Medium confidence (0.400),” the maximum number of interactions to display was set to “No more than 50 interactors,” and the default options were used for all other settings. We also used this STRING database to identify the KEGG pathways of the target genes of captopril. 15
To investigate the linkage between captopril and its associated genes in Bladder cancer (BLCA), we used 2 web-based tools: STITCH and NetworkAnalyst. STITCH (http://stitch.embl.de/ ) integrates data on metabolic pathway interactions and drug-target relationships to create a network that can be explored interactively or used as the foundation for more extensive studies. 16 Similarly, NetworkAnalyst (https://www.networkanalyst.ca/) is a comprehensive web-based application that allows bench researchers to perform standard and advanced meta-analyses of gene expression data via an intuitive web interface.17,18 Thus, these 2 tools helped us to understand the interaction network between captopril and its associated genes in BLCA.
mRNA expression analysis of target genes BLCA
We utilized TNMplot (http://tnmplot.com/analysis/, accessed on September 28, 2023), a web tool for comparing DPTs gene expression between normal and tumor tissue in the bladder. 19 Mann Whitney was used for the statistical test between groups. UALCAN (http://ualcan.path.uab.edu) and GEPIA (http://gepia2.cancer-pku.cn/) web-based tools were also used to validate the expression results of DPT-related genes.
Survival analysis of target genes of captopril in BLCA
The online database Kaplan-Meier (available at https://kmplot.com/analysis/) was utilized to examine the relationship between target genes and the cumulative survival rates of cancer patients. The analysis was conducted using data from the TCGA-BLCA cohort. 20 The patients were classified into 2 groups, high and low expression, according to the median level of captopril-associated gene’s mRNA expression. Log-rank P values, hazard ratios, survival analysis plots, and 95% confidence intervals were analyzed in order to assess overall survival (OS) in BLCA patients. A total of 405 patients with BLCA were included in the study.
Identification of overexpressed genes in BLCA patients
GEO2R is a web tool that permits consumers to match 2 or more groups of samples in a GEO Series to find genes that are differentially expressed across experimental conditions. Using the GEO2R (https://www.ncbi.nlm.nih.gov/geo/geo2r/) online tool, we analyzed the differential gene expression between BLCA and normal samples. 21 The tool compared sample groups within a GEO Series (GSE65635) and examined gene symbols, where transcriptome analysis of normal and cancerous bladder tissues was performed. Malignant tumor samples were collected from patients after surgery, while non-cancerous control samples were obtained from autopsies of healthy donors who died in road accidents. A pathologist examined both tumor and normal tissues to confirm the diagnosis and determine tumor cell counts. All tumor samples used in this study contained at least 80% tumor cells. To control false positives, adjusted P values were used together with the Benjamini and Hochberg false discovery rate method. The cutoff criteria were adjusted P value .05 and |log Fold Change (FC)| ⩽ 2, ensuring that genes with significant differential expression in bladder cancer were identified.
Molecular docking
The wild type structures of the human proteins ACE, MMP9, and lipocalin 2 (LCN2) were retrieved from the AlphaFold Protein Structure Database (AlphaFold DB) using their UniProt accession codes I3UJJ7, P14780, and X6R8F3, respectively. 22 The wild MMP9 protein structure served as the template for generating the following mutated structures: F145L, C230W, H231P, S308F, and K566R, using the Swiss Model server. 23 Validation and energy minimization were achieved through the SAVES 24 and YASARA 25 webservers. Polar hydrogen atoms and Kollman charges were added prior to molecular docking using the AutoDock tools.
The structures of captopril, and other experimental/investigational drugs with similar chemical structures including epicaptopril, N-acetylproline, miridesap, and GPI-1485 were retrieved from the DrugBank database. 26 The PRODRG server was used for energy minimization. 27 Gasteiger charges were added using the AutoDock tools.
The CASTp server was used to predict the active binding pockets of ACE, MMP9, and LCN2. 28 The results were validated and were found consistent with the available information on the active site residues of these protein structures.29,30 AutoDock Vina 31 was used for molecular docking of the drugs at the active binding pockets of the proteins, opting a grid-based docking protocol. 32 The Protein-Ligand Interaction Profiler (PLIP) webserver 33 was used to analyze the protein-ligand interactions.
Molecular dynamics simulations
Molecular dynamics (MD) simulations of the unbound proteins and the protein-ligand complexes were carried out using the GROMACS software (version 2019) 34 with GROMOS96 43a1 force field parameters. 35 The unbound proteins and the protein-ligand complexes were solvated using the TIP3P water model within a periodic boundary box of distance 1.0 nm. 36 Based on the rebalancing charges, the whole system was neutralized by adding an accurate concentration of Na+/Cl− ions.37,38 Steepest descent method was used for energy minimization. 39 The thermostat coupling was set with a reference temperature of 300 K using the Berendsen thermostat, 39 and the pressure coupling was set with a reference pressure of 1 bar using the Parrinello-Rahman barostat. 39 The long-range interactions were estimated using the Particle-Mesh Ewald (PME) method. 40 The energy-minimized whole systems were equilibrated for 1000 ps at 300 K and 1 bar pressure in the NVT and NPT ensembles, respectively.39,41 Molecular dynamics simulations were run for a timescale of 120 ns to understand the dynamic behavior and stability of the unbound proteins and the protein-ligand complexes. The root mean square deviations (RMSD) and the root mean square fluctuations (RMSF) of the unbound proteins and the protein-ligand complexes were estimated for MD simulation analysis. In addition, principal component analysis (PCA) was performed on the MD trajectories to explore the conformational motions and stability of the unbound proteins and the protein-ligand complexes.
Results
Identification of captopril’s DPTs
Captopril is an ACE inhibitor as mentioned previously. It is described as extremely weak basic, cardiovascular agent, antihypertensive, renal protective agent, anti-inflammatory, antioxidant effect in output DB01197 from DrugBank 5.0. By using DrugBank, we identified 5 primary direct protein targets (DPTs) of captopril such as ACE, MMP2, MMP9, LTA4H, BDKRB1 (Table 1). The identification of DPTs can provide valuable insights into the specific pathways that may be modulated by captopril.
Identification of direct targets of captopril using DrugBank.

Captopril-regulated pathways in humans
In this study, the protein-protein interaction (PPI) network and signaling pathways associated with 5 DPTs of captopril were constructed using the STRING database (Figure 1A). Additionally, analyses using STITCH and NetworkAnalyst revealed direct interactions between captopril and specific proteins, namely LTA4H, ACE, MMP2, and MMP9 (Figure 1B and C). The visualizations generated by these tools provided valuable insights into the potential molecular associations that are relevant to the mechanism of action of captopril.
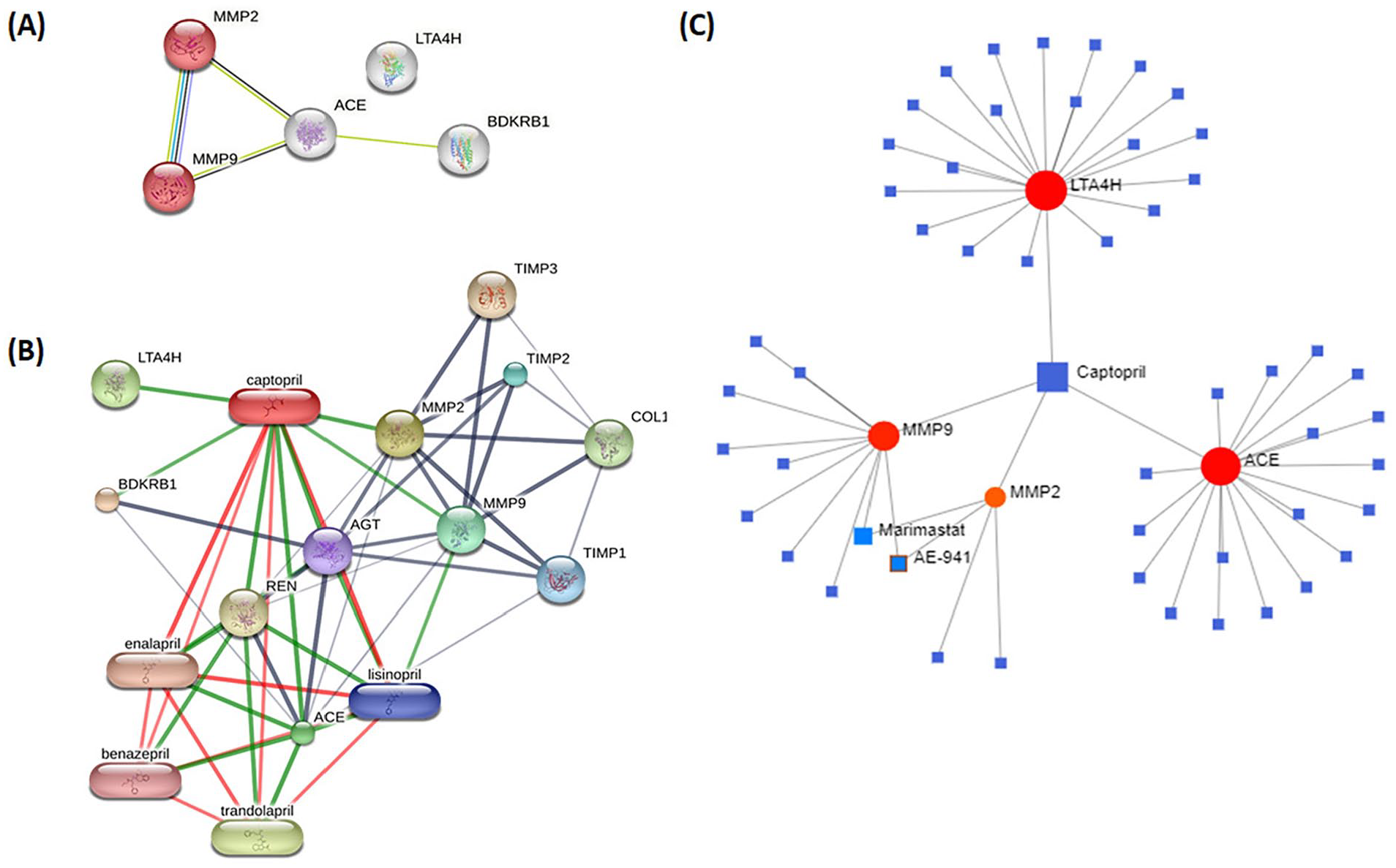
Relationship between captopril and captopril-associated genes. (A) Protein-protein interaction (PPI) network of captopril target genes (ACE, MMP2, MMP9, BDKRB1, LTA4H) from STRING. (B) STITCH and (C) NetworkAnalyst visualization of direct interactions between Captopril and LTA4H, ACE, MMP2, and MMP9.
Moreover, the study identified the top 5 significant KEGG pathways associated with these DPTs, which include bladder cancer, endocrine resistance, leukocyte transendothelial migration, relaxin signaling pathway, fluid shear stress, and atherosclerosis (Table 2). Notably, bladder cancer (BLCA) was identified as the most relevant solid tumor linked to captopril, and 2 genes, MMP2 and MMP9, were found to be involved in BLCA, supported by the STRING analysis.
List of KEGG pathway in captopril associated DPT.
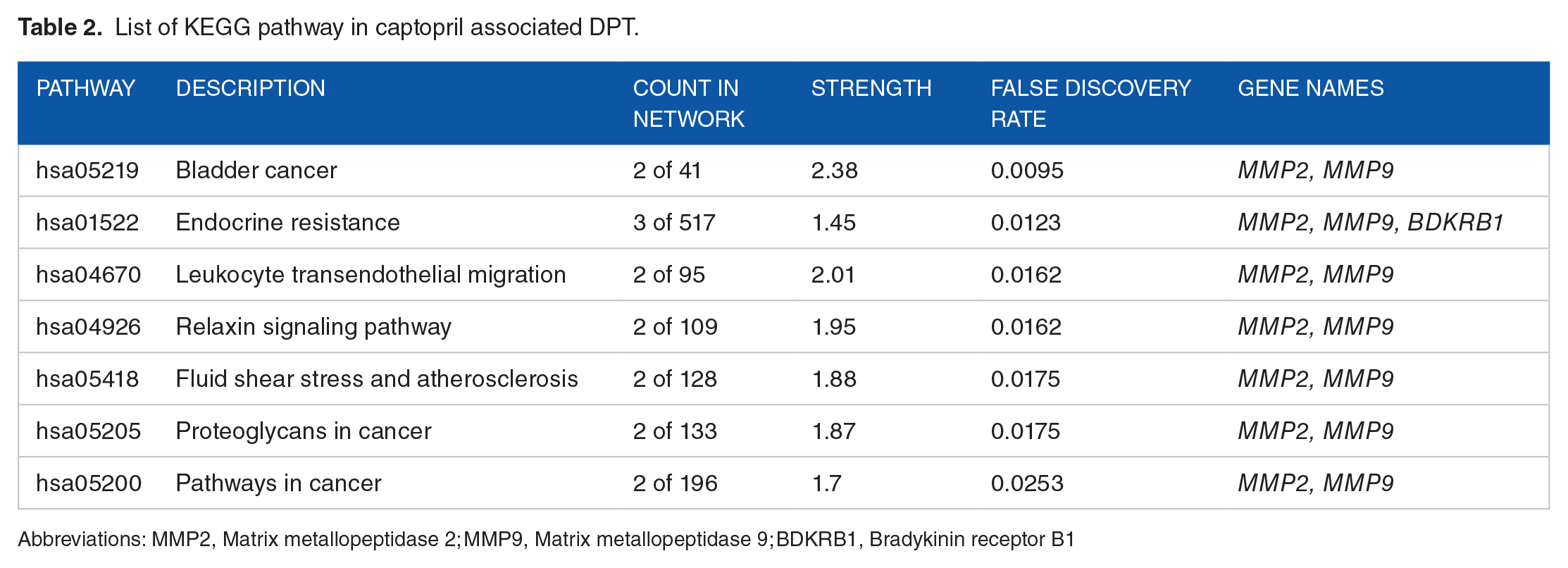
Abbreviations: MMP2, Matrix metallopeptidase 2; MMP9, Matrix metallopeptidase 9; BDKRB1, Bradykinin receptor B1
mRNA expression profile of captopril associated genes in BLCA
In light of the substantial regulatory impact of Captopril on BLCA, we conducted an expression analysis of the genes associated with the direct target proteins DPTs that interact with Captopril specifically in BLCA. The findings are presented in Figure 2, which showcases the results obtained from the TNMplot web tool, focusing on ACE, MMP2, MMP9, LTA4H, and BDKRB1 mRNA expression levels in both normal and tumor specimens of BLCA. Notably, it was observed that the expression of MMP9 was significantly elevated in BLCA patients compared to normal samples (Figure 2C). This outcome was further supported by results obtained from ULCAN and GEPIA, as depicted in Supplemental Figures 1 and 2, respectively. Additionally, through analysis using the ULCAN web tool, it was discovered that patients with extreme obesity, who are at a high risk of developing hypertension, exhibited heightened MMP9 expression in BLCA (Figure 2F). Collectively, these results suggest a potential association between MMP9 expression and BLCA, particularly in patients at a heightened risk for hypertension and severe obesity.
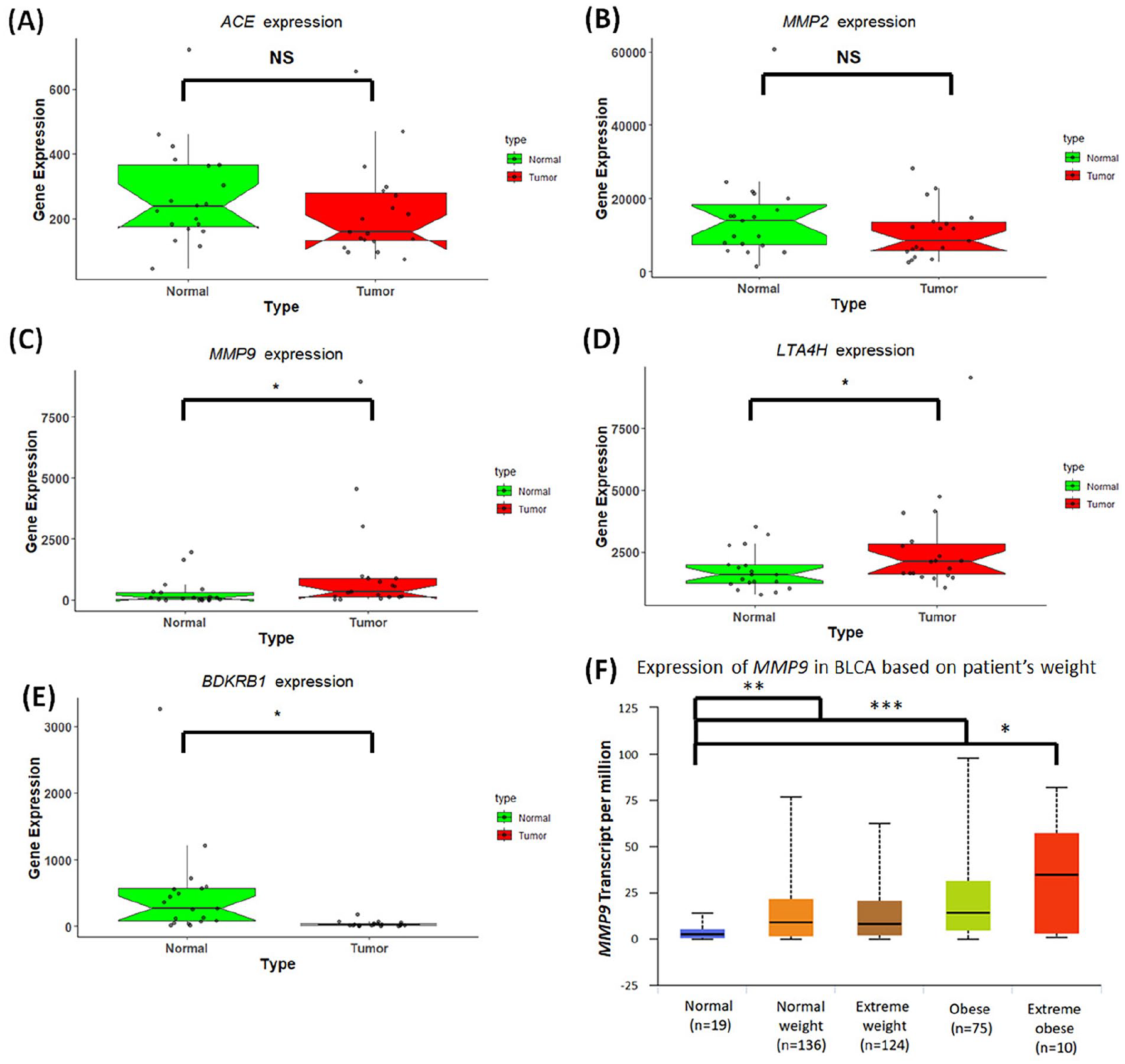
mRNA expression profile of captopril associated genes in BLCA. Box plot showing the expression of genes (A) ACE (B) MMP2 (C) MMP9 (D) LTA4H and (E) BDKRB1. RED color indicates bladder cancer and GREEN color means non-cancer/Normal. The X-axis of the plot shows normal and tumor samples and the Y-axis shows gene expression. The thick line in the middle represents the median, and the upper and lower limits of the box represent the third and first quartile respectively. Statistical analysis is Mann-Whitney U test (P values < .05 = *, <.005 = **, <.0005 = ***). (F) Expression (Transcript per million) of MMP9 in Normal weight, Extreme weight, Obese and Extreme obese. Normal weight = BMI greater than equal to 18.5 and BMI less than 25; Extreme weight = BMI greater than equal to 25 and BMI less than 30; Obese = BMI greater than equal to 30 and BMI less than 40; Extreme obese = BMI greater than 40. Statistical analysis is Mann-Whitney U test (P values < .05 = *, <.005 = **, <.0005 = ***).
Identification of overall survival in BLCA based on gene expression
Subsequently, we assessed the relationship between the survival of BLCA patients and the expression levels of genes corresponding to the DPTs associated with captopril. Figure 3 depicts the overall survival (OS) rate of patients with BLCA based on the expression levels of ACE, MMP2, MMP9, LTA4H, and BDKRB1. Notably, the data shows that patients with high MMP9 expression have a significantly lower overall survival rate than those with low expression (as shown in Figure 3C).

Association of captopril associated genes with overall survival in BLCA patients. Survival risk curves are shown for captopril DPT genes (A) ACE (B) MMP2 (C) MMP9 (D) LTA4H (E) BDKRB1 expression in BLCA using KM-plotter. The expression ranges of the probes for ACE, MMP2, MMP9, LTA4H, and BDKRB1 are 26-2345, 160-98 392, 2-63 2587, 828-7249, and 1-1862, respectively. The cutoff values for these genes were 241, 8323, 207, 2195, and 22. Low and high expression level of genes are drawn in black and red respectively. X-axis shows per months or time and Y-axis shows probability of survival. In KM-plotter the threshold for the high and low gene expression cohorts is automatically calculated.
Common SNPs and structural alterations of MMP9 in BLCA
In order to assess the viability of targeting MMP9 with captopril in BLCA patients, a thorough analysis was carried out to identify all potential mutations within the MMP9 gene, focusing specifically on BLCA. Six single nucleotide polymorphisms (SNPs) in the MMP9 gene were discovered using cBioportal data on BLCA, 3 of which are located in the protein’s fibronectin type-II domain (H231P, C230W, and S308F)
Analysis of overexpressed genes in BLCA for evaluating the therapeutic benefits of captopril
We conducted an extensive analysis of captopril’s DPTs and their associated proteins using the STRING database. Subsequently, we validated and visualized the interactions using Cytoscape, as depicted in Figure 4A. The PPI network of captopril DPTs was generated on STRING, applying a cutoff criterion that allowed for a maximum of 20 interactors in layers 1 and 2.

Identification of common DPTs of captopril and overexpressed genes in BLCA for assessing the benefits of captopril use. (A) Interaction of DPTs captopril with other proteins. (B) Common proteins between overexpressed genes in BCLA and interacting proteins with captopril DPTs. (C) Expression of LCN2 gene in BLCA and normal tissue. (P < .05). (D) Survival risk curves are shown for LCN2 expression in BLCA using KM-plotter. Expression range of the probe is 0 to 228 457 and cut-off is 707.
Furthermore, we identified overexpressed genes in bladder cancer (BLCA) by analyzing the GEO database (GSE656351) through GEO2R. Specifically, genes with adjusted P-values less than .05 and LogFC greater than 2 were considered significantly overexpressed in BLCA. We then utilized a Venn diagram to identify the common genes between captopril’s direct or indirect DPTs and the overexpressed gene in BLCA, as shown in Figure 4B. LCN2 was the only one common gene.
The overexpression of LCN2 in BLCA patients was further confirmed using the TNMPlot analysis, which demonstrated higher expression levels in paired tumor and adjacent normal tissues, as presented in Figure 4C. Notably, our findings revealed an intriguing association between the overexpression of LCN2 and poor survival outcomes in BLCA patients, as illustrated in Figure 4D.
Interaction of captopril and similar experimental/investigational drugs with human ACE, MMP9, and LCN2
In the present study, we investigated the inhibitory prospects of captopril and other similar experimental/investigational drugs from DrugBank, including epicaptopril, N-acetylproline, miridesap, and GPI-1485 against the human proteins ACE, MMP9, and LCN2. Molecular docking of the drugs was performed at the active binding pockets of these proteins. The binding energy scores have been detailed in Table 3. It was evident from our analysis that captopril, a known inhibitor of human ACE 40 exhibited significantly higher (P < .01) binding potential with a binding energy score of −8.2 ± 0.03 kcal/mol, in comparison to the other similar experimental/investigational drugs (Table 3).
Binding energy scores (kcal/mol) of MMP9, LCN2 and ACE with captopril and similar drugs.

Abbreviations: ACE, angiotensin-converting enzyme; MMP9, matrix Metallopeptidase 9; LCN2, Lipocalin 2.
Captopril also produced the most promising binding energy score of −7.2 ± 0.01 kcal/mol with the human MMP9 protein, as compared to the other drugs (Table 3). Interaction profile (Figure 5A and B) revealed that captopril formed a hydrogen bond with Tyr423, a hydrophobic interaction with Pro421, a π-stacking with Tyr179 and a salt bridge with Lys603, at the active binding pocket of human MMP9. We also investigated the inhibitory potential of captopril against the mutated MMP9 structures involving the mutations that are implicated in BLCA. Interestingly, captopril returned decent binding energy scores of −7.18 ± 0.02 kcal/mol, −7.20 ± 0.01 kcal/mol, −7.20 ± 0.01 kcal/mol, −7.19 ± 0.01 kcal/mol, and −7.20 ± 0.02 kcal/mol with the F145L, C230W, H231P, S308F, and K566R mutated structures of human MMP9, respectively (Supplemental Table 3).

Interaction profile of captopril-MMP9 and captopril-LCN2 complexes. (A) Molecular docking of captopril with MMP9. Blue and brown spheres represent MMP9 and captopril, respectively. (B) Molecular interaction of captopril-MMP9 complex. Hydrophobic interactions are represented as gray dashed lines and hydrogen bonds are shown as blue lines. Green dashed lines show π-stacking interactions. Yellow dashed lines represent salt bridges. (C) Molecular docking of captopril with LCN2. Gray and brown spheres represent LCN2 and captopril, respectively. (D) Molecular interaction of captopril-LCN2 complex. Hydrophobic interactions and hydrogen bonds are represented as gray dashed lines and blue lines, respectively.
Molecular docking revealed that the drug miridesap displayed the most encouraging binding energy score of −7.9 ± 0.02 kcal/mol with the human LCN2 protein, followed by captopril which produced a binding energy score of −7.4 ± 0.02 kcal/mol. Interaction profile of the captopril-LCN2 complex (Figure 5C and D) revealed that captopril formed hydrogen bonds with the amino acid residues Tyr106, Lys147, and Lys156, and hydrophobic interactions with the amino acid residues Tyr78 and Phe145, at the active pocket of human LCN2.
The investigational drug miridesap displayed a significantly higher (P < .01) binding energy score with LCN2 as compared to captopril. However, the binding energy scores were significantly lower (P < .01) for miridesap-ACE and miridesap-MMP9 complexes as compared to captopril-ACE and captopril-MMP9 complexes, respectively (Table 3). The aim of our study was to screen drugs that may show inhibitory potential against hypertension as well as BLCA. Captopril, the known inhibitor of ACE involved in hypertension, displayed encouraging binding energy scores with the human proteins MMP9 and LCN2, that are implicated in BLCA.42,43 Accordingly, the captopril-MMP9 and captopril-LCN2 complexes were selected for further molecular dynamics simulations.
Analysis of molecular dynamics simulations
The unbound proteins MMP9 and LCN2, and the complexes of captopril with these proteins were subjected to molecular dynamics (MD) simulations for a timescale of 120 ns. The captopril-MMP9 and captopril-LCN2 complexes returned average RMSD values of 1.1 and 0.9 Å, respectively. The captopril-MMP9 and captopril-LCN2 complexes experienced initial fluctuations in RMSD values of Cα atoms and attained stability after 97 (Figure 6A) and 103 ns (Figure 6C), respectively, and remained stable thereafter. On the contrary, the unbound proteins MMP9 and LCN2 were found to display more fluctuations throughout the course of MD simulations displaying higher average RMSD values of 1.5 and 1.1 Å, respectively, than their corresponding captopril-bound complexes (Figure 6A and C). A low RMSD value is consistent with higher stability of a protein-ligand complex. 44
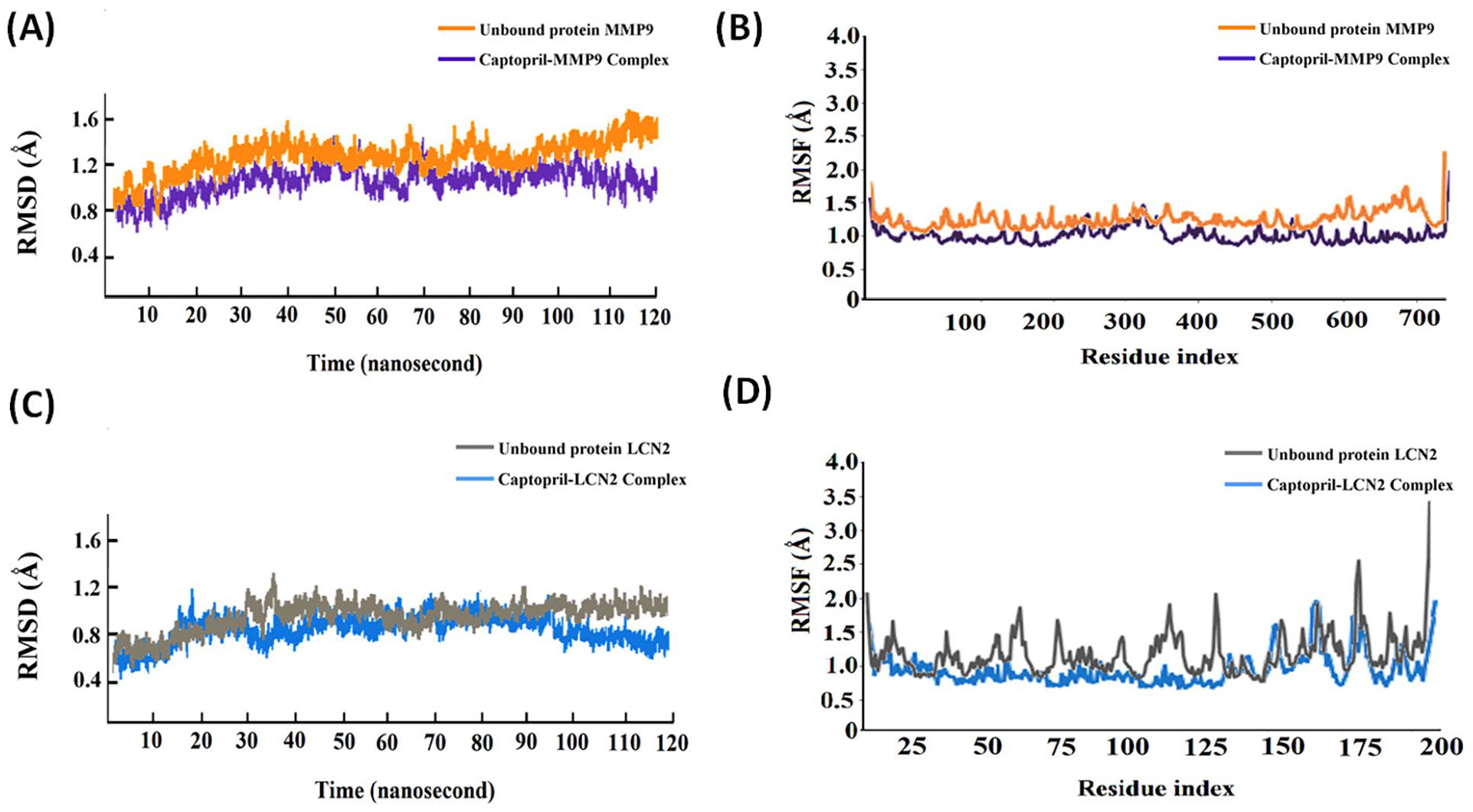
MD simulations of unbound protein MMP9, captopril-MMP9 complex, unbound protein LCN2, and captopril-LCN2 complex. (A) RMSD analysis of unbound MMP9 and captopril-MMP9 complex for 120 ns of MD simulations. (B) RMSF analysis of of unbound MMP9 and captopril-MMP9 complex for 120 ns of MD simulations. (C) RMSD analysis of unbound LCN2 and captopril-LCN2 complex for 120 ns of MD simulations. (D) RMSF analysis of unbound LCN2 and captopril-LCN2 complex for 120 ns of MD simulations.
RMSF analysis reveals the fluctuations in a protein–ligand complex during MD simulations and provides insights into conformational stability. 45 A thorough RMSF analysis of the unbound proteins MMP9 and LCN2, and their respective complexes with captopril revealed reduced fluctuations with lower average RMSF values among the complexes as compared to the unbound proteins (Figure 6B and D). The captopril-MMP9 and captopril-LCN2 complexes displayed average RMSF values of 1.2 and 1.4 Å, respectively (Figure 6B and D). However, the unbound proteins MMP9 and LCN2 produced average RMSF values of 1.6 and 1.8 Å, respectively. Interestingly, the residues at the active pockets of the proteins MMP9 and LCN2 that interacted with captopril suffered minimal fluctuations (Figure 6B and D). Our findings accentuate the conformational stability of the complexes.
Principal component analysis (PCA) was performed on the MD trajectories to explore the functional motions and conformational stability of the unbound proteins MMP9 and LCN2, and their complexes with captopril. The determination values were calculated from the atomic motions based on the eigenvectors. PCA analysis based on the principal components 1 and 2 (PC1 and PC2) have been depicted in Figure 7. It was evident from our analysis that the complexes of MMP9 and LCN2 with captopril (Figure 7B and D, respectively) were more compact in phase space in comparison to the unbound proteins MMP9 (Figure 7A) and LCN2 (Figure 7C). Compactness in phase space reflects narrow energy fluctuations and signifies higher stability. 39 PCA analysis reinforced the observation that binding of captopril to MMP9 and LCN2 imparted significant stability to the complexes.

Principal component analysis (PCA) on the MD trajectories of unbound proteins and protein-ligand complexes in the phase spaces. (A) unbound protein MMP9. (B) captopril-MMP9 complex. (C) unbound protein LCN2. (D) captopril-LCN2 complex.
Discussion
Captopril may be a good medication for a patient with bladder cancer (BLCA) who also has hypertension because it has many beneficial properties. This is especially important for BLCA patients who may be receiving chemotherapy or other treatments that might have negative cardiovascular side effects. Furthermore, captopril has been shown to have renal protective properties, which may help to prevent or delay the progression of kidney damage caused by BLCA or its treatment. 46 This is due to the presence of the primary direct protein target of captopril, ACE, in the renin-angiotensin-aldosterone system, which regulates kidney and blood pressure. 46 Furthermore, it has been discovered that captopril’s other 2 main direct protein targets, MMP2 and MMP9, are involved in a variety of BLCA-related processes such as leukocyte transendothelial migration, the relaxin signaling pathway, fluid shear stress, and atherosclerosis. 47 By blocking these targets, captopril might have additional anti-inflammatory and anti-cancer effects, which may be beneficial for BLCA patients. One study reported that captopril exhibits inhibitory effects on colon and prostate cancer cells. Captopril impedes cellular proliferation and cancer cell migration, potentially inducing apoptosis. These findings suggest that captopril could serve as an effective cancer therapeutic with reduced side effects. 10 Another study found that captopril, when combined with cyclophosphamide, enhanced the antitumor effects of cyclophosphamide in mice with Lewis lung carcinoma. However, administering captopril 5 days prior to cyclophosphamide treatment did not alter its antitumor activity. These findings indicate that captopril could be a potential treatment for cancer. 48
Our study found that MMP9 expression is significantly higher in BLCA, and high MMP9 expression is linked to a lower overall patient survival. Furthermore, MMP9 expression is increased in obese patients with BLCA who are at a higher risk of developing hypertension. MMP9 has been shown to promote tumor invasion by degrading the extracellular matrix and promoting tumor cell migration into nearby tissues. 49 MMP9 also activates inflammatory signaling pathways, promotes inflammatory cell migration to the arterial wall, and contributes to hypertension and other cardiovascular diseases. 50 As a result, measuring MMP9 levels in patients with hypertensive BLCA may aid in predicting prognosis and making treatment decisions. Overall, MMP9 is a key player in the progression of BLCA, particularly in obese-related hypertensive patients, and might be a promising target for the development of new diagnostic tools and therapies.
Our study also revealed the overexpression of LCN2 in BLCA and its association with the protein targets affected by captopril. Furthermore, a separate study indicated an interaction among LCN2, SLC22A17, and MMP9 genes, suggesting their cooperative role in cancer. Notably, the heightened expression of both LNC2 and MMP9 genes was observed in 16 tumor types, including BLCA, thus underscoring their involvement in tumor development. 51 High-molecular-weight matrix metalloproteinases (MMPs), like the LCN2-MMP-9 complex, have also been shown in studies to be independent predictors of metastatic phenotypes in a variety of cancers, including bladder and prostate cancer. 52 A recent study found that the LCN2/LOXL2/MMP9 ternary complex promotes cancer cell migration and invasion, as well as malignant tumor progression, making it a potential therapeutic target. 53 LCN2 and MMP9 form a complex, which protects MMP9 from degradation and thus increases its enzymatic activity. 54 This interaction is critical for modulating cancer cell metastatic behavior and is linked to the aggressive nature of neoplastic cells in a variety of cancers. 43 Thus, we proceeded to conduct further analysis to assess the ability of captopril and similar structures to inhibit MMP9 and LCN2.
Molecular docking analysis revealed promising inhibitory potential of captopril against human MMP9 protein. Captopril was found to interact with the Tyr179, Pro421, Tyr423, and Lys603 amino acid residues at the active pocket of MMP9. The catalytic domain residues Pro421 and Tyr423 of MMP9 have been reported to be instrumental in binding the reverse hydroxamate inhibitor through hydrogen bonds. 55 In addition, captopril displayed encouraging binding energy scores with the mutated MMP9 structures involving mutations that are implicated in BLCA.56,57
Captopril also returned promising binding energy score with the human LCN2 protein. It was found to interact with the Tyr78, Tyr106, Phe145, Lys147, and Lys156 residues at the active pocket of LCN2. Tyr78 and Tyr106 residues of LCN2 are imperative in ligand recognition and binding. 30 Our data from molecular docking were consistent with the results of MD simulations. The RMSD and RMSF trajectories for 120 ns of MD simulations revealed that the captopril-MMP9 and captopril-LCN2 complexes had more stable conformations in comparison to the unbound proteins MMP9 and LCN2. To further investigate the stability of the complexes, we compared their interaction profiles prior to and after MD simulations. The interaction profiles of the complexes after 120 ns of MD simulations have been provided as (Supplemental Figure 3). It was evident that the captopril-MMP9 (Figure 5B and Supplemental Figure 3A) and captopril-LCN2 (Figure 5D, Supplemental Figure 3B) complexes did not experience any change in interacting residues, following MD simulations.
Conclusion
Our comprehensive analysis reveals a significant association between the progression of bladder cancer BLCA and obesity-related comorbidities, with MMP9 emerging as a crucial mediator. These findings underscore the critical role of MMP9 in the development and progression of BLCA in the context of obesity, emphasizing the need to address these comorbidities for effective management and treatment of BLCA patients. Our findings also provide robust evidence of captopril’s inhibitory potential against 2 critical human proteins, MMP9 and LCN2, which play a crucial role in BLCA. Remarkably, our results indicate that captopril exhibits equal efficacy in inhibiting both wildtype and mutated forms of MMP9, highlighting its broad applicability in treating BLCA patients with diverse genetic backgrounds. Nonetheless, further validation through in vitro and in vivo studies is imperative to consolidate and substantiate these promising findings. These investigations will facilitate the translation of captopril’s therapeutic potential into clinical settings for the benefit of BLCA patients.
Supplemental Material
sj-docx-1-cix-10.1177_11769351241276759 – Supplemental material for Computational Insights into Captopril’s Inhibitory Potential Against MMP9 and LCN2 in Bladder Cancer: Implications for Therapeutic Application
Supplemental material, sj-docx-1-cix-10.1177_11769351241276759 for Computational Insights into Captopril’s Inhibitory Potential Against MMP9 and LCN2 in Bladder Cancer: Implications for Therapeutic Application by Sanjida Kabir Annana, Jannatul Ferdoush, Farzia Lamia, Ayan Roy, Pallab Kar, Monisha Nandi, Maliha Kabir and Ayan Saha in Cancer Informatics
Supplemental Material
sj-docx-2-cix-10.1177_11769351241276759 – Supplemental material for Computational Insights into Captopril’s Inhibitory Potential Against MMP9 and LCN2 in Bladder Cancer: Implications for Therapeutic Application
Supplemental material, sj-docx-2-cix-10.1177_11769351241276759 for Computational Insights into Captopril’s Inhibitory Potential Against MMP9 and LCN2 in Bladder Cancer: Implications for Therapeutic Application by Sanjida Kabir Annana, Jannatul Ferdoush, Farzia Lamia, Ayan Roy, Pallab Kar, Monisha Nandi, Maliha Kabir and Ayan Saha in Cancer Informatics
Supplemental Material
sj-docx-3-cix-10.1177_11769351241276759 – Supplemental material for Computational Insights into Captopril’s Inhibitory Potential Against MMP9 and LCN2 in Bladder Cancer: Implications for Therapeutic Application
Supplemental material, sj-docx-3-cix-10.1177_11769351241276759 for Computational Insights into Captopril’s Inhibitory Potential Against MMP9 and LCN2 in Bladder Cancer: Implications for Therapeutic Application by Sanjida Kabir Annana, Jannatul Ferdoush, Farzia Lamia, Ayan Roy, Pallab Kar, Monisha Nandi, Maliha Kabir and Ayan Saha in Cancer Informatics
Supplemental Material
sj-docx-4-cix-10.1177_11769351241276759 – Supplemental material for Computational Insights into Captopril’s Inhibitory Potential Against MMP9 and LCN2 in Bladder Cancer: Implications for Therapeutic Application
Supplemental material, sj-docx-4-cix-10.1177_11769351241276759 for Computational Insights into Captopril’s Inhibitory Potential Against MMP9 and LCN2 in Bladder Cancer: Implications for Therapeutic Application by Sanjida Kabir Annana, Jannatul Ferdoush, Farzia Lamia, Ayan Roy, Pallab Kar, Monisha Nandi, Maliha Kabir and Ayan Saha in Cancer Informatics
Supplemental Material
sj-docx-5-cix-10.1177_11769351241276759 – Supplemental material for Computational Insights into Captopril’s Inhibitory Potential Against MMP9 and LCN2 in Bladder Cancer: Implications for Therapeutic Application
Supplemental material, sj-docx-5-cix-10.1177_11769351241276759 for Computational Insights into Captopril’s Inhibitory Potential Against MMP9 and LCN2 in Bladder Cancer: Implications for Therapeutic Application by Sanjida Kabir Annana, Jannatul Ferdoush, Farzia Lamia, Ayan Roy, Pallab Kar, Monisha Nandi, Maliha Kabir and Ayan Saha in Cancer Informatics
Footnotes
Acknowledgements
Not applicable
Declaration Of Conflicting Interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding:
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Author Contributions
All authors contributed to the study conception and design. Material preparation, data collection and analysis were performed by Sanjida Kabir Annana, Farzia Lamia, Ayan Roy, Jannatul Ferdoush, Pallab Kar, Monisha Nandi, Maliha Kabir and Ayan Saha. The first draft of the manuscript was written by Sanjida Kabir Annana and all authors commented on previous versions of the manuscript. Whole project was done under the supervision of Ayan Saha and Jannatul Ferdoush. All authors read and approved the final manuscript.
Availability of Data and Materials
Data are available from the corresponding author on reasonable request.
Ethics Approval and Consent to Participate
Not applicable
Consent for Publication
Not applicable.
Authors’ Note
Monisha Nandi is now affiliated to Department of Biomedical Science, College of Medicine, Kyung Hee University, Republic of Korea.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
