Abstract
NBS-LRR (nucleotide-binding site and leucine-rich repeat) is one of the largest resistance gene families in plants. The completion of the genome sequencing of wild tomato
Introduction
Plants are surrounded by a wide variety of pathogens such as viruses, bacteria, fungi, nematodes, and aphids during their growth and development. 1 Some pathogens have successfully invaded crops and have caused severe damage to agricultural production and the quality of crops. To cope with disease attacks, the plants have evolved a series of sophisticated defense mechanisms to defend against various pathogens. Previous studies have shown that disease resistance (R) proteins of plants play an essential role in direct or indirect recognition of corresponding pathogens. 2 The plant-pathogen interaction model is regarded as the “gene-for-gene” interaction hypothesis. 3 In this hypothesis, an incompatible interaction of host R gene protein products with pathogen Avr proteins produces a defense response termed the hypersensitive response, which impedes pathogen progression via a variety of mechanisms, including localized programmed cell death and correlated immune responses. 4 Currently, numerous disease R genes of plants have been cloned, which not only confer resistance to a wide range of pathogens but also play a vital role in resistance to abiotic stress.1,5,6
Presently, researchers had divided R genes into at least 5 diverse classes of families, including NBS-LRR (Nucleotide Binding Site and Leucine-Rich Repeat domains), LRR-TM (Leucine-Rich Repeat plus Transmembrane Receptor), STK (Serine-Threonine Kinase), RLK (Receptor-Like Kinase), and SA-CC (Signal Anchor plus Coiled-Coil).
7
Among them, the NBS-LRR family is the largest class of known R proteins in the plant kingdom,1,7 whose encoded proteins are involved in an important part of the plant defense system. In
It was well known that the NBS-LRR protein encodes 3 main domains—N-terminal, NBS, and LRR domains.5,9,10 There are 2 structures in the N-terminal, one is the TIR (Toll/Interleukin-1 receptor) structure, and the other is the non-TIR structure, usually known as CC (coiled-coil). In
The NBS domain, composed of ~300 amino acid sequences, is the main structural domain of the NBS-LRR R genes. Eight distinct conserved motifs—P-loop, RNBS-A, Kinase2, RNBS-B, RNBS-C, GLPL, RNBS-D, and MHDV—have been confirmed in this domain.5,16 However, these 8 motifs are not completely conserved in each subfamily. It has been proved that motifs P-loop, GLPL, Kinase2, and GLPL have high similarity in the TNL and CNL subfamilies, while the similarity levels of the RNBS-A, RNBS-D, and RNBS-C motifs are lower in the TNL and CNL of
The NBS-LRR superfamily, accounting for the largest gene family among plant genomes, has become the core of the resistance research field. Currently, NBS-LRR genes have been studied in many monocotyledon and dicotyledon plants, including
Tomato (
Materials and Methods
Identification of NBS-LRRs in S. pimpinellifolium
The NBS-LRR genes of
Subsequently, these candidates were submitted to the online software PFAM (http://pfam.sanger.ac.uk/) to determine whether there are TIR, NBS, and LRR domains. Meanwhile, the CC domain may contain some smaller individual motifs or too divergent proteins; hence, it was further identified using COILS Serve (http://www.ch.embnet.org/software/COILS_form.html). Finally, all NBS-LRR genes were classified according to the presence or absence of TIR, CC, NBS, or LRR domains.
Chromosome mapping of the NBS-LRR genes in S. pimpinellifolium
Information on the physical position on the chromosomes of each NBS-LRR gene from the
Phylogenetic analysis of the NBS-LRR genes
All NBS-LRR genes with CNL and TNL domains were selected for phylogenetic analysis. The P-loop to GLPL motifs of these chosen NBS-LRR members were extracted to perform multiple sequence alignment using Clustalx 1.83. Then, phylogenetic analysis of the NBS-LRR gene family was performed using the neighbor-joining (NJ) and maximum likelihood (ML) methods. For NJ analysis, MEGA X software was selected. 67 The parameters of the phylogenetic tree were set as follows: 1000 bootstrap replications; p-distance model; and pairwise deletion gap. For ML analysis, ProtTest (version 2.4) software was used for model selection 68 and PhyML (version 3.0) software was used to construct ML trees with the Whelan and Goldman amino acid substitution model, γ-distribution, and 100 nonparametric bootstrap replicates. 69
Homologous comparison and phylogenetic analysis were also investigated between amino acid sequences of
Detection of the conserved motif
The NBS-encoding genes could be divided into CNL and TNL subfamilies based on phylogenetic analysis. To investigate the structural features of these genes, the sequences and distribution of the conserved motifs were analyzed individually using Multiple Expectation Maximization for Motif Elicitation (MEME) (http://meme-suite.org/tools/meme), and the parameters were set as follows: the maximum and minimum lengths of the conserved motif were 50 and 6, respectively, and the largest number of conserved motifs was 20; other parameters used the default settings. 70 Conservation or variation of each motif among NBS-LRR members was presented.
Results
Identification and classification of the NBS-LRR gene family in S. pimpinellifolium
Previous studies showed that the NBS-LRR gene family in plants contained different conserved domains, and the NBS-LRR gene family was further divided into 6 different subfamilies according to the domains.
5
In this study, we identified a total of 245 NBS-LRR resistance genes in
Number and classifications of NBS-LRR genes.

Abbreviations: CC, coiled-coil; LRR, leucine-rich repeat; NBS, Nucleotide-binding site; TIR, toll/interleukin-1 receptor.
Among them, both of CNL and TNL were the most typical subfamilies, with 78 and 15 NBS-LRR members, respectively. The numbers of the NBS-LRR genes in the N and CN subfamilies were 62 and 54, respectively. Also, 29 genes were identified as belonging to the NL subfamilies, and only 7 genes were predicted to encode the TIR domain; hence, it belonging to the TN subfamily. Meanwhile, the TN subfamily was the smallest NBS-LRR subfamily among these 6 subfamilies. Based on the above result, we can conclude that these 245 NBS-LRR genes were unevenly located on the 6 subfamilies. Together, high genetic variation was observed in
Comparative analysis of the NBS-LRR genes between S. pimpinellifolium and S lycopersicum
In addition to comparing the number of genes of different subfamilies in
In this context, we further analyzed the number and distribution of the NBS-LRR gene family on the chromosome between

Chromosomal locations and duplications of the paralogous NBS-LRRs on S.
Distribution and characteristic of nucleotide-binding site and leucine-rich repeats on different chromosomes between

Gene cluster and tandem duplication of the NBS-LRR genes in S. pimpinellifolium
The distribution of NBS-LRR genes across the chromosomes was used to further analyze the evolutionary patterns of gene expansion (Figure 1). Six different colors represent the genes from the 6 subfamilies of NBS-LRRs in Figure 1. We found that NBS-LRRs of the CNL type were spread across all chromosomes, while TNL genes were selectively distributed on chromosomes 3, 6, 8, 10, and 12. It has been previously proved that most of the NBS-encoding genes are arranged in clusters on chromosomes.5,25 Also, the gene cluster was previously determined by the following criteria: a cluster of NBS-LRR proteins was described as the distance between neighboring homologous genes less than 200 kb and fewer than 8 non-NBS-encoding genes between TNLs and CNLs.5,19,28 Based on the above criteria, NBS-LRR gene clusters were carried out (Figure 1). A total of 49 gene clusters, including 146 NBS-LRR genes, were identified in wild tomato
Tandem duplication of NBS-LRRs was analyzed in
Phylogenetic analysis of the NBS-LRR genes in S. pimpinellifolium
To explore the evolutionary relationships of NBS-LRR genes, we conducted phylogenetic analyses with an alignment of the NBS domain from these 2 species using neighbor-joining (NJ) and maximum likelihood (ML) methods. ML analysis showed that proteins from different species cluster together in clades with high support values (not shown), with support from NJ analysis for most results. Therefore, in the present study, the phylogenetic tree of the NBS-LRR genes from the TNL and CNL subfamilies constructed using the NJ method was selected for analyses. Four members (Sopim04g008170.0.1, Sopim11g069660.0.1, Sopim11g043070.0.1, and Sopim10g055080.0.1) were excluded due to the presence of incomplete NBS domains. Both of the CNL and TNL subfamilies were separated from each other in the phylogenetic tree, and CNL was further divided into eight small branches, namely CNL1 to CNL8, respectively (Figure 2). Inversely, the TNL subfamily remained as one branch owing to fewer gene numbers. Moreover, we found that the CNL1 and CNL7 branches each contained 16 NBS-LRRs, which were from seven different chromosomes, and these 2 branches had the largest gene numbers in the phylogenetic tree. In contrast to CNL1 and CNL7, the CNL5 branch only contained 2 genes from chromosomes 8 and 11, with the lowest number of genes (Figures 1 and 2).

Phylogenetic relationship of nucleotide-binding site and leucine-rich repeats from CNL and TNL subfamilies in
To date, some NBS-LRR-encoding genes have been cloned, like
Apart from the members of the CNL and TNL subfamilies, NBS-LRR genes from other subfamilies also showed high homology with known functional genes. As shown in Supplemental Table S2, all subfamilies had homologous genes with known resistance genes, except for the TN subfamily. Twenty-one known resistance genes, involving multiple resistances to various pathogens, had as high as 80% similarity with the NBS-LRR genes in
Conserved motif analysis of the NBS-LRR genes
To uncover the structural characteristics of the NBS-LRR gene family, MEME was applied to analyze the structure and distribution of the conserved motifs among the TNL and CNL subfamilies. Twenty distinct motifs were determined in each subfamily (Tables 3 and 4). All of the conserved motifs displayed a diversity distribution in their respective subfamilies (Supplemental Figures S1-TNL and S2-CNL1-4, respectively).
Conserved motifs of the TNL subfamily in

Abbreviations: LRR, leucine-rich repeat; NBS, nucleotide-binding site; TIR, toll/interleukin-1 receptor.
Conserved motifs of the CNL subfamily in

Abbreviations: CC, coiled-coil; LRR, leucine-rich repeat; NBS, nucleotide-binding site.
In the TNL subfamily, there were 4, 7, and 9 motifs identified in the TIR, NBS, and LRR domains, respectively (Table 3 and Supplemental Table S3). The motifs in the TIR domain were named T-1 to T-4, and the motifs in the LRR domain were named L-1 to L-9. The motifs of the NBS domain were named following previous studies. 5 All of the 14 TNL members contained 4 motifs of the TIR domain, except that Sopim09g092410.0.1 lacked motif T-1 (Supplemental Figure S1 and Supplemental Table S3). The motif RNBS-A was not found in the NBS domain. Two novel motifs (TNBS-1 and TNBS-2) were identified in most of the TNL subfamily. Both of Kinase-2 and RNBS-B existed in Motif 5, as these 2 proteins were so close to each other. The motifs of the NBS domain had higher conservation in the 14 TNL genes, except TNBS-2, which was missed in some of the genes. Also, the motif compositions of the NBS-LRR genes provided further support for the grouping of phylogenetic branches. For example, 3 NBS-LRR genes (Sopim09g092410.0.1, Sopim04g056570.0.1, and Sopim07g052770.0.1) were locat-ed in the adjacent branches of the phylogenetic tree, and they did not contain the motif TNBS-2.
In the CNL subfamily, there were 4, 11, and 5 motifs in the CC, NBS and LRR domains, respectively (Table 4 and Supplemental Table S4). A low degree of conservation of motifs was observed from CC to LRR. Of them, 3 out of the 4 conserved motifs (C-1, C-2, and C-3) had lower conservation in the CC domain compared with the NBS and LRR domains (Supplemental Figure S2). In other words, most of the genes in the CNL subfamily lost motifs C-1, C-2, and C-3. Besides, most of the NBS-LRRs from the CNL4 to 8 branches lacked motif L-4 (Supplemental Table S4). In the NBS domain, most of the genes from CNL 6 missed the RNBS-D and CNBS-3 motifs. The remaining conserved motifs were detected in most of the NBS-LRR genes. Overall, the motifs of the NBS domain were relatively conservative compared with that in the N-terminal domain (Supplemental Figure S2).
When compared with the TNL and CNL subfamilies, some differences regarding the motif compositions were observed. For example, the MHDV motif was unique to the CNL family (Table 4). Besides, 2 conserved motifs (TNBS-1 and TNBS-2) were identified as novel members in the TNL subfamily, while 3 unique motifs (CNBS-1, CNBS-2, and CNBS-3) were only found in the CNL subfamily, with more diversity (Supplemental Table S3 and Table 4). In the TNL and CNL subfamilies, the conserved motifs of the LRR domain had relatively high diversity.
Evolutionary comparison of the NBS-LRR genes between S. pimpinellifolium and Arabidopsis
Previous findings have revealed that the NBS-LRR genes of

Phylogenetic relationship of the TNL subfamily between
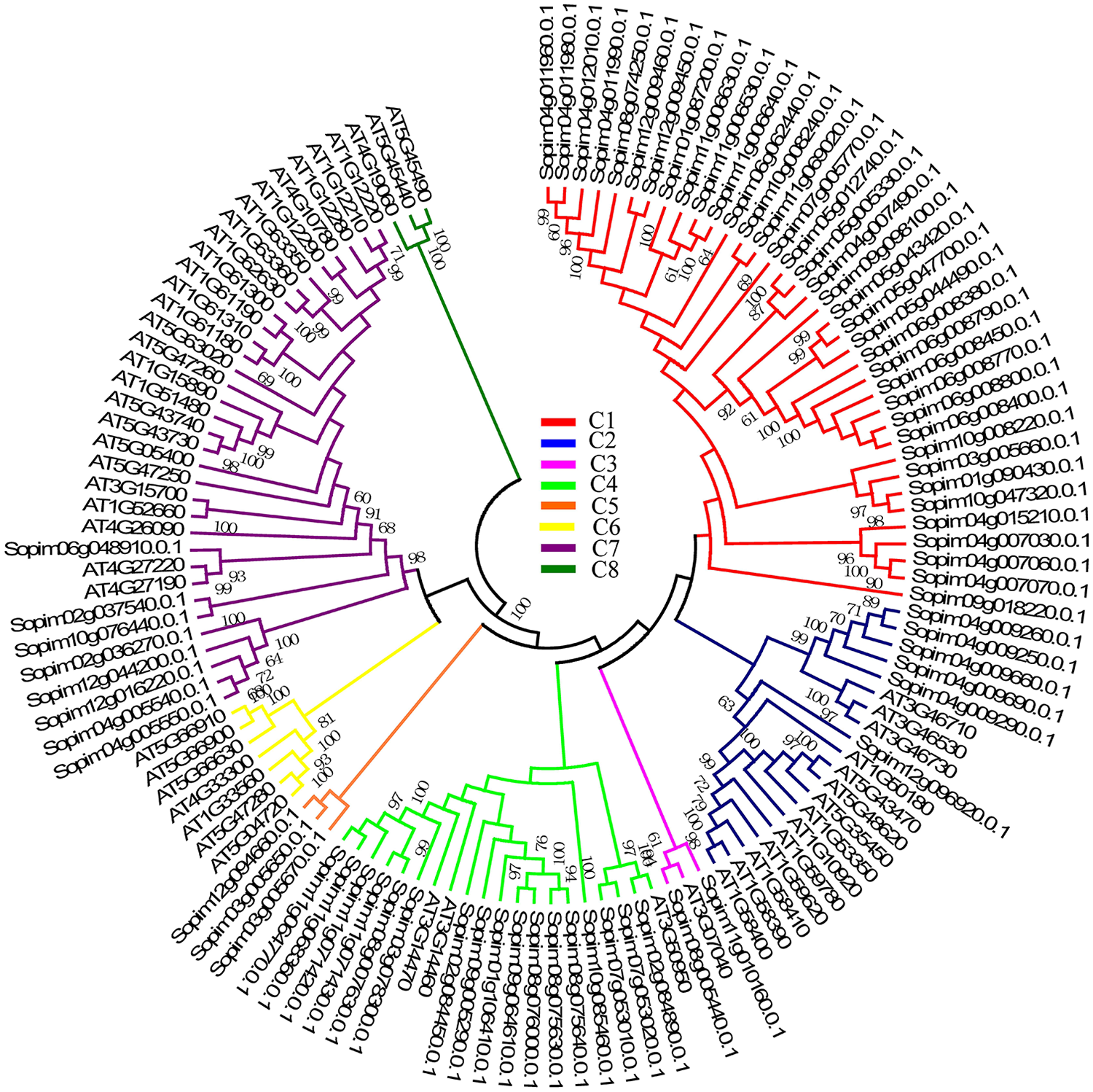
Phylogenetic relationship of the CNL subfamily between
The phylogenetic tree of the TNL genes from the
In the CNL subfamily, 53 NBS-LRRs in
Discussion
Wild species, as an important component of germplasm resources, contain resistance to disease and abiotic stress genes and play a key role in the hereditary improvement of the cultivated species. In disease-resistant tomato cultivars, a crowd of resistance genes was derived from the wild species.
77
Among them, the wild tomato
In previous studies, the numbers of NBS-LRR gene families and members of each subfamily were different in distinct plants. In this study, a total of 245 NBS-LRR genes were identified, among which 78 genes belonged to the CNL subfamily and 15 genes belonged to the TNL subfamily (the ratio of CNL/TNL was about 5:1). A similar phenomenon has been found in other plants. For example, the number of CNL subfamilies in potato is 4.7 times that of that in the TNL subfamily,
19
being the closest ratio with
There was no significant difference in the NBS-LRR disease resistance genes between
The identified 245 NBS-LRR genes in
In addition, there were lots of singleton genes in these 2 plant species. Approximately 42.9% (12/28) of singleton NBS-LRR genes shared a homologous relationship with the cloned resistance genes (Supplemental Table S2). Several singletons were homologous genes on other chromosomes, such as singletons Sopim08g074250.0.1, Sopim03g005660.0.1, and Sopim04g056570.0.1, and some of them seem to have evolved independently.
Two subfamilies, CNL and TNL, were identified through phylogenetic tree construction. The conserved motifs were used to distinguish the difference of protein sequences of N-terminal and NBS domains between these 2 subfamilies (Tables 3 and 4). Most of the conserved motifs were selectively distributed within a subfamily in the phylogenetic tree, implying that structural and functional similarities existed among NBS-LRRs within the same clade. In the TNL subfamily, all motifs were detected in all of the analyzed genes, except that the novel identified the TNBS-2 motif of the NBS domain was missed in 3 genes (Sopim09g092410.0.1, Sopim07g052770.0.1, and Sopim04g056570.0.1) (Supplemental Table S3 and Supplemental Figure S1). In contrast to the TNL subfamily, motifs of the CNL subfamily had much higher diversity (Supplemental Table S4 and Supplemental Figure S2). Some motifs were specific to each branch within the CNL subfamily, such as Motifs 03, 10, and 11 found in CNL4, while these motifs were not observed in CNL6. Whether this discovery reflected the more ancient origin of the CNL subfamily during plant evolution was unclear. We also found that some conserved motifs only existed in a particular clade, for example, Motif 19 and Motif 20 existed in the CNL2 and CNL3 branches, respectively. The motif analysis of the NBS-LRR genes in
Although the functions of most of these motifs have not been identified, it is plausible that some probably involved a crucial role. For instance, previous reports demonstrated that 3 domains (LRR, TIR, and CC) regulated downstream signaling events through intramolecular interactions.81-83 Proteins homologous to plant NBS-LRR proteins play a role in mammalian defense responses. In these mammalian proteins, the N-terminal domain is involved in downstream signaling partners through protein-protein interactions, the NBS hydrolyzes ATP functions as a regulatory domain, and LRR binds to upstream regulatory factors.84,85
When compared with the TNL and CNL subfamilies between
Conclusions
In this study, a comprehensive and systematic analysis of the NBS-LRR genes of
Supplemental Material
Figure_S1_xyz330564316c3b6 – Supplemental material for Genomic Organization and Comparative Phylogenic Analysis of NBS-LRR Resistance Gene Family in Solanum pimpinellifolium and Arabidopsis thaliana
Supplemental material, Figure_S1_xyz330564316c3b6 for Genomic Organization and Comparative Phylogenic Analysis of NBS-LRR Resistance Gene Family in Solanum pimpinellifolium and Arabidopsis thaliana by Huawei Wei, Jia Liu, Qinwei Guo, Luzhao Pan, Songlin Chai, Yuan Cheng, Meiying Ruan, Qingjing Ye, Rongqing Wang, Zhuping Yao, Guozhi Zhou and Hongjian Wan in Evolutionary Bioinformatics
Supplemental Material
Figure_S2_xyz33056e5992d7c – Supplemental material for Genomic Organization and Comparative Phylogenic Analysis of NBS-LRR Resistance Gene Family in Solanum pimpinellifolium and Arabidopsis thaliana
Supplemental material, Figure_S2_xyz33056e5992d7c for Genomic Organization and Comparative Phylogenic Analysis of NBS-LRR Resistance Gene Family in Solanum pimpinellifolium and Arabidopsis thaliana by Huawei Wei, Jia Liu, Qinwei Guo, Luzhao Pan, Songlin Chai, Yuan Cheng, Meiying Ruan, Qingjing Ye, Rongqing Wang, Zhuping Yao, Guozhi Zhou and Hongjian Wan in Evolutionary Bioinformatics
Supplemental Material
Table_S1_xyz33056b5977b4d – Supplemental material for Genomic Organization and Comparative Phylogenic Analysis of NBS-LRR Resistance Gene Family in Solanum pimpinellifolium and Arabidopsis thaliana
Supplemental material, Table_S1_xyz33056b5977b4d for Genomic Organization and Comparative Phylogenic Analysis of NBS-LRR Resistance Gene Family in Solanum pimpinellifolium and Arabidopsis thaliana by Huawei Wei, Jia Liu, Qinwei Guo, Luzhao Pan, Songlin Chai, Yuan Cheng, Meiying Ruan, Qingjing Ye, Rongqing Wang, Zhuping Yao, Guozhi Zhou and Hongjian Wan in Evolutionary Bioinformatics
Supplemental Material
Table_S2_xyz33056ee822a92 – Supplemental material for Genomic Organization and Comparative Phylogenic Analysis of NBS-LRR Resistance Gene Family in Solanum pimpinellifolium and Arabidopsis thaliana
Supplemental material, Table_S2_xyz33056ee822a92 for Genomic Organization and Comparative Phylogenic Analysis of NBS-LRR Resistance Gene Family in Solanum pimpinellifolium and Arabidopsis thaliana by Huawei Wei, Jia Liu, Qinwei Guo, Luzhao Pan, Songlin Chai, Yuan Cheng, Meiying Ruan, Qingjing Ye, Rongqing Wang, Zhuping Yao, Guozhi Zhou and Hongjian Wan in Evolutionary Bioinformatics
Supplemental Material
Table_S3_xyz330562220ba64 – Supplemental material for Genomic Organization and Comparative Phylogenic Analysis of NBS-LRR Resistance Gene Family in Solanum pimpinellifolium and Arabidopsis thaliana
Supplemental material, Table_S3_xyz330562220ba64 for Genomic Organization and Comparative Phylogenic Analysis of NBS-LRR Resistance Gene Family in Solanum pimpinellifolium and Arabidopsis thaliana by Huawei Wei, Jia Liu, Qinwei Guo, Luzhao Pan, Songlin Chai, Yuan Cheng, Meiying Ruan, Qingjing Ye, Rongqing Wang, Zhuping Yao, Guozhi Zhou and Hongjian Wan in Evolutionary Bioinformatics
Supplemental Material
Table_S4_xyz33056bbc1857b – Supplemental material for Genomic Organization and Comparative Phylogenic Analysis of NBS-LRR Resistance Gene Family in Solanum pimpinellifolium and Arabidopsis thaliana
Supplemental material, Table_S4_xyz33056bbc1857b for Genomic Organization and Comparative Phylogenic Analysis of NBS-LRR Resistance Gene Family in Solanum pimpinellifolium and Arabidopsis thaliana by Huawei Wei, Jia Liu, Qinwei Guo, Luzhao Pan, Songlin Chai, Yuan Cheng, Meiying Ruan, Qingjing Ye, Rongqing Wang, Zhuping Yao, Guozhi Zhou and Hongjian Wan in Evolutionary Bioinformatics
Footnotes
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was financially supported by the National Key Research and Development Program of China (2017YFE0114500), the State Key Laboratory Breeding Base for the Zhejiang Sustainable Pest and Disease Control (2010DS700124-ZZ1903 and 2010DS700124-ZZ1807), the National Natural Science Foundation of China (31772294), the Zhejiang Provincial major Agricultural Science and Technology Projects of New Varieties Breeding (2016C02051), the Zhejiang Provincial Natural Science Foundation of China (LY18C150008), the General Program from the National key research and development program (2017YFD0101902, 2018YFD1000800), and the earmarked fund for China Agriculture Research System (CARS-23-G-44).
Declaration of Conflicting Interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
HJW and HWW conceived and designed the research. JL, QWG, LZP, SLC, YC, MYR, QJY, and RQW performed the experiments. ZPY and GZZ analyzed the data and wrote the manuscript. HJW revised the manuscript. All authors read and approved the final manuscript.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
