Abstract
The nucleotide-binding site (NBS)–leucine-rich repeat (LRR) gene family is crucially important for offering resistance to pathogens. To explore evolutionary conservation and variability of NBS-LRR genes across grass species, we identified 88, 107, 24, and 44 full-length NBS-LRR genes in sorghum, rice, maize, and
Introduction
Plants have developed sophisticated mechanisms to recognize and guard against pathogens, including pathogen-associated molecular pattern-triggered immunity (PTI) and effector-triggered immunity (ETI).
1
PTI is based on the actions of a basal immune system, which can be activated by generic signals of a pathogen, such as bacterial flagellins, lipopolysaccharides, and elongation factors. ETI is based on the actions of an adaptive immune system, which has a specific recognition of plant disease resistance (R) genes and pathogen avirulence (
To date, >100
The NBS-LRR genes are unevenly distributed in plant genomes and most of them reside in clusters. There are two types of NBS-LRR gene clusters: monophyletic cluster and mixed cluster. The monophyletic cluster contains genes with close relationship and high sequence similarity, while the mixed cluster contains clustered genes with diverged relationship and low sequence similarity. The distribution of
The objectives of this study are to (1) identify the full-length NBS-LRR genes from grass species using InterProScan, an integrated protein domain recognition method; (2) perform a comprehensive analysis on classification, genome organization, evolution, expression, and regulation of these NBS-LRR genes using sorghum as a representative of grass species; and (3) conduct a comparative analysis of NBS-LRR gene number, cluster, duplication, and targeting site of miRNA across four grass species. This comprehensive bioinformatics analysis of the full-length NBS-LRR genes will shed light on how NBS-LRR genes evolve and will provide foundation for
Methods
Identification, classification, and genomic cluster of full-length NBS-LRR genes
Genome assemblies and predicted gene models for sorghum (v2.1),
11
maize (v6a),
12
rice (
Linked NBS-LRR genes were grouped into clusters when they were interrupted by less than eight open reading frames not encoding NBS-LRR proteins following the same definition of NBS-LRR gene cluster described previously.
18
Monophyletic clusters were composed solely of genes belonging to monophyletic clades based on phylogenetic analysis.
5
If the linked NBS-LRR genes belong to different clades in the phylogenetic tree, they are considered as a mixed cluster. The expected number (μ) of mixed clusters in whole sorghum genome was calculated using the following formula
18
: μ = (33,032 -
Gene duplication and gene conversion
Duplication events of NBS-LRR genes across whole sorghum genome were identified by conducting BLASTP search of NBS-LRR gene sequences against the annotated sorghum protein sequences. The hits of NBS-LRR genes were filtered using the following threshold: expectation (
Coding DNA sequences (CDSs) of each cluster and duplication family were retrieved from sorghum CDSs and aligned in each cluster and family, respectively. Alignments were analyzed using GENECONV version 1.81a
19
according to the protocol outlined by Drouin
20
to detect gene conversion. Alignments containing only two sequences were analyzed using the
Phylogenetic analysis of NBS-LRR genes and exonintron configuration
A phylogenetic tree was constructed using the Molecular Evolutionary Genetics Analysis software version 6.0 (MEGA 6.0)
21
following the method described by Hall.
22
NBS domains coevolved with other protein domains, including the N terminal and LRR regions, which may contain unique information on a subgroup of NBS-LRR genes. To cover all sequence information, we used the complete protein sequence of NBS-LRR genes to construct a maximum likelihood phylogenetic tree, including the chromosome of origin (by sequence name), intron–exon configurations, gene classification, and expression sequence tag (EST) representatives. In brief, protein sequences of all NBS-LRR genes in sorghum were aligned using MUSCLE with default settings.
23
After the alignment, the best substitution model was chosen using the feature
The exon/intron positions and phases of the NBS-LRR genes in sorghum were extracted from sorghum gene GFF file. Intron phases are classified based on Sharp's description 24 : phase-0 introns lie between two codons, phase-1 introns interrupt a codon between the first and the second nucleotides in the codon, and phase-2 introns interrupt a codon between the second and the third nucleotides in the codon. The exon/intron structures were obtained using the online Gene Structure Display Server (http://gsds.cbi.pku.edu.cn).
Expression analysis of sorghum NBS-LRR genes
Representations of EST for each sorghum NBS-LRR gene were studied by searching the NCBI EST database using the predicted cDNA sequence of each NBS-LRR gene. As of January 3, 2015, this database contained a total of 199,401 sorghum EST sequences, which included all EST sequences from multiple tissues of sorghum updated in the database by different researchers. All sorghum ESTs with the best match to NBS-LRR genes and sequence identity >80% were counted as representations. 25
About 105 M 100 bp single-end reads (FASTQ files) of sorghum and
miRNA databases of each species were used to predict miRNA-targeting sites of NBS-LRR genes. There were 241, 713, 321, and 464 miRNAs preloaded in psRNATarget (http://plantgrn.noble.org/psRNATarget/) for sorghum, rice, maize, and
Identification of paralogs/orthologs and synteny analysis
Paralog and ortholog groups were identified by using BLASTP with the same criteria as those in the gene duplication study (
For each gene pair of orthologs, 100 flanking genes (50 genes from each end) surrounding orthologous gene pairs were selected to study syntenic relationships. If flanking pairs with
Results
Identification, classification, and genomic cluster distribution of NBS-LRR genes
A total of 88 full-length NBS-LRR genes (Supplementary Table 1) were identified from sorghum genome by using InterProScan 5.
15
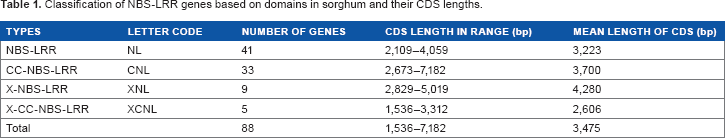
The number of these NBS-LRR genes on each chromosome varied from 1 on chromosome 4 to 21 on chromosome 5 (Fig. 1), indicating an uneven distribution of NBS-LRR genes in sorghum. Based on the combinations of different protein domains, including coiled-coil (CC), NBS, LRR, and X (other domains except CC, NBS, and LRR), the 88 NBS-LRR sorghum genes were classified into four types: NBS-LRR (NL), CC-NBS-LRR (CNL), X-NBS-LRR (XNL), and X-CC-NBS-LRR (XCNL) (Table 1). The majority of these genes were classified as genes with two domains, NBS-LRR (46.6%), and as genes with three domains, CC-NBS-LRR (37.5%). For each type, the mean and range of CDS length were calculated. On average, NBS-LRR gene CDSs contained 3475 nucleotides, which was much longer than that of the remaining genes in sorghum (1202 nucleotides).
Physical location of NBS-LRR genes on sorghum chromosomes. Classification of NBS-LRR genes based on domains in sorghum and their CDS lengths.
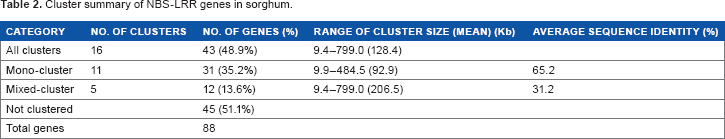
Cluster summary of NBS-LRR genes in sorghum.
Duplication and gene conversion of NBS-LRR genes
Duplication of NBS-LRR genes in sorghum, rice, maize, and

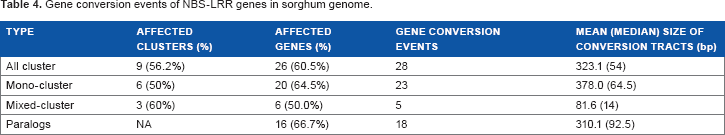
Gene conversion events of NBS-LRR genes in sorghum genome.

Phylogenetic analysis of sorghum NBS-LRR genes and exon–intron configurations
A maximum likelihood phylogenetic tree of 88 NBS-LRR genes was constructed to illustrate the evolutionary relationship of these NBS-LRR genes (Fig. 2A). The tree showed a mixture of the four types of NBS-LRR genes: NL with NBS-LRR domains, CNL with CC-NBS-LRR domains, XNL with X-NBS-LRR domains, and XCNL with X-CC-NBS-LRR domains. This phylogenetic mixture suggested that coevolving or exchange of genetic information may happen among the four different types of NBS-LRR genes. Most of the NBS-LRR genes on the same chromosome were grouped in the same clades with a few exceptions (Fig. 2A), indicating a more recent local duplication than ectopic duplication. The NBS-LRR genes from the same clades tended to have similar exon–intron configuration, suggesting high conservation of exon–intron configuration during evolution. The average rate of amino acid substitution showed an ascending order with the domain number increasing in the four types of NBS-LRR genes (Fig. 2B). Since the number of mutations per amino acid in a protein increases almost linearly with evolutionary time,
32
the XCNL-type genes may be divergent from their ancestor (A) A phylogenetic tree of NBS-LRR genes was constructed using MEGA 6.0. Branch numbers represent percentage of bootstrap values in 1000 bootstrapping replicates, and the scale indicates branch length. Chromosomal location of genes is included in its name (005G in Sobic.005G197600.1 showed that this gene is located on chromosome 5). Introns and exons are drawn to scale with the full encoding regions of their respective genes. Boxes indicate the exon, and lines indicate the intron. 0, phase-0 intron; 1, phase-1 intron; 2, phase-2 intron. Information on domains and number of supporting ESTs for each gene is also shown in the last two columns. (B) The average branch length of the four types of NBS-LRR genes. (C) The percentage of genes with introns and the percentage of genes with phase-0 introns in the four types of NBS-LRR genes.
Expression analysis of sorghum NBS-LRR genes
Expression of the 88 NBS-LRR genes in sorghum was evaluated by identifying the sorghum EST hits in EST GenBank with ~200,000 entries, using BLAST program. Only 32 NBS-LRR genes (36.3%) had ESTs detected with an average of 8.4 ESTs for each gene. Twenty-eight of the 33 expressed NBS-LRR genes had four or more EST representatives per gene. Fifty-five NBS-LRR genes were not detected in this depth of EST sampling.
The expression of the 88 NBS-LRR genes in sorghum was further evaluated by aligning 104 million high-quality reads of a set of RNA-seq data
26
with the genomic sequences of the 88 genes. Sixty-eight of the 88 NBS-LRR genes had an FPKM value of ≥0.05, indicating certain level of expression. The average FPKM value of the 88 NBS-LRR genes in sorghum was 5.4, which is much lower than the expression levels at the whole genome level (17.6), suggesting relatively low basal expression of NBS-LRR genes. Compared to the control, 553 sorghum genes were differentially expressed, and one of them was an NBS-LRR gene (Sobic.002G104400.1) (Fig. 3). The expression of this NBS-LRR gene was significantly reduced at 12 hpi and then it increased but with a lower level than the control at 24 hpi, suggesting the critical function of this gene for the host during the interaction with this pathogen. Further analysis indicated that this gene encoded an RPP13-like protein and its ortholog in rice contained WRKY- and DNA-binding domains. Further analysis is needed to validate the function of this NBS-LRR gene in disease resistance.
Gene expression (in FPKM) plot showing differences of one NBS-LRR gene across three conditions. CK, control, no inoculation; t12, 12 hours postinoculation; t24, 24 hours postinoculation.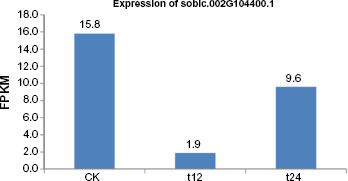
Out of the 88 NBS-LRR genes, 6.8% were identified as targets of miRNA by using the miRNA of sorghum, which were significantly higher than the remaining genes in the genome with the targeted percentage of 2.2% (Fig. 4 and Supplementary Table 3).
Percentage of genes targeted by miRNA of each species in NBS-LRR and the other genes (not NBS-LRR genes) in the genomes of the four grass species. There were 241, 713, 321, and 464 miRNAs preloaded in psRNATarget for sorghum, rice, maize, and 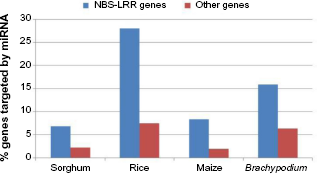
Identification of paralogs/orthologs and syntenic relationships
In total, 24 NBS-LRR genes belonging to eight paralogous groups were identified from the 88 NBS-LRR genes (Supplementary Table 4). These eight groups have an average of 72.7% protein sequence identity. Twenty-one NBS-LRR paralogs (87.5%) are located in clusters. In order to study the divergence of these paralogs, Ka and Ks were calculated for CDSs of NBS domain, LRR domain, and complete gene sequence (Fig. 5A). On average, the Ka values of complete gene sequence, NBS domain, and LRR domain were 0.18, 0.15, and 0.51, respectively. The Ka values were not significantly different between complete gene sequence and NBS domain, except for the LRR domain, as revealed by the Divergence of NBS-LRR genes: (A) divergence of different classes of NBS-LRR genes in sorghum and (B) divergence of NBS-LRR genes in grass species.
In total, four ortholog groups of NBS-LRR genes (with an average of 81.1% protein sequence identity) (Supplementary Table 5) were identified among sorghum, rice, maize, and Phylogenetic tree of 16 orthologs in grass species. NBS-LRR genes and their clustering, duplication, miRNA targeting, and SSRs in four grass species.
The density of NBS-LRR genes is calculated as: 1000 × the total number of NBS-LRR genes in each species/the total number of genes in each species. The density of SSRs is calculated as: the total number of SSRs in each species/the number of NBS-LRRs with SSRs in each species.
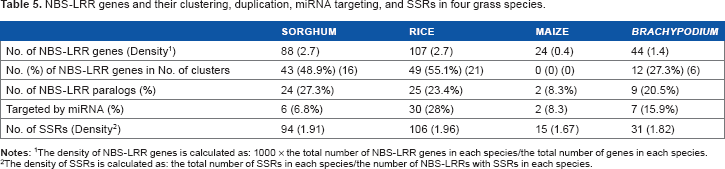
Comparative analysis of NBS-LRR genes in grass species
In sorghum, rice, maize, and
The number of syntenic genes identified out of 100 flanking genes surrounding four NBS-LRR gene groups of orthologs.


Syntenic relationships surrounding NBS-LRR orthologs in grass species.
Discussion
Identification of NBS-LRR genes based on a protein domain recognition method
In this study, we used InterProScan, a software packet with an integration of different protein domain recognition methods and a comprehensive database, to identify full-length NBS-LRR genes, a large gene family in plant genomes involved in diverse pathogen recognition. The numbers of NBS-LRR genes in this study were much less than the numbers reported before, such as 160 genes in rice, 175 in
The accurate set of NBS-LRR genes is the foundation for further structure, distribution, and evolutionary analyses. To ensure that the number of NBS-LRR genes identified in this study is not a significant underestimation due to systemic and bioinformatics errors, we compared the 88 NBS-LRR genes identified from the sorghum genome with the more than 200 NBS-LRR genes reported previously by Mace.
10
We were able to retrieve the sequences of 194 NBS-LRR genes reported in Mace's article. The InterProScan indicated that only 53 of them were NBS-LRR genes, while 140 genes contained only NBS domain or just the LRR domain and one gene contained neither of the two domains. Furthermore, 10 of the 140 NBS genes were randomly chosen and manually checked using Pfam (http://pfam.xfam.org/), which is a publicly accepted method to determine protein domains. The results from Pfam confirmed that none of those 10 genes contained both of the domains. In addition, we compared the 44 NBS-LRR genes identified from the genome of
Plasticity of NBS-LRR genes in clusters
Although the number of NBS-LRR genes in clusters varies depending on species, NBS-LRR gene clustering is a common feature in plant genomes. For example, nearly 50% of NBS-LRR genes reside in clusters in
The clusters provide a reservoir of genetic variation of NBS-LRR genes through mechanisms such as duplication, gene conversion, and diversifying selection. 2 Of the 24 sorghum NBS-LRR parologs, 21 were in clusters, and 27 out of 43 genes in clusters were duplicated. Local duplication in clusters could be the major evolutionary mechanism of NBS-LRR gene expansion and could potentially determine the number of varieties of NBS-LRR genes in grass species. The majority of the genes in clusters were affected by gene conversion, especially for monophyletic clusters with 64.5% affected genes. LRR domain was involved in the recognition of pathogen ligands and is usually highly variable, while NBS domain was involved in signaling and included highly conserved and strictly ordered motifs. 10 The Ka and Ks values in this study showed that the diversity of NBS-LRR genes mainly came from LRR domains, while the NBS domain might be under purifying selection.
Compared to mixed clusters, monophyletic clusters contained NBS-LRR genes with a high sequence similarity and small cluster size, indicating that the two types of clusters may have different originating mechanisms. Monophyletic clusters might result from local duplication of NBS-LRR genes on the chromosome, while mixed clusters might result from ectopic recombination, in which heterozygous NBS-LRR genes combined with a physical cluster. In addition, the number of mixed clusters (5) is greater than expected at a genome-wide level (2.1), indicating that the genes in mixed clusters promote certain functions of NBS-LRR genes. In fact, an emerging theme was reported that two NBS-LRR genes functioned together to mediate disease resistance and many of them were in genomic clusters (eg,
Regulation mechanisms of NBS-LRR gene expression
Most of the previously investigated species (except papaya with a relatively even distribution) showed varying number of NBS-LRR genes and an uneven distribution of genes on chromosomes.
3
Grass species, such as sorghum, rice, maize, and
miRNA plays an important role in RNA silencing and posttranscriptional regulation of gene expression. For example, many transcription factors and development-related genes have been reported as targets of these regulatory small RNAs. The dosage effect of NBS-LRR genes could be balanced through miRNA regulation as more NBS-LRR genes were predicted to be targets of miRNA in these species with high NBS-LRR duplication. Our results showed that there was a higher percentage of NBS-LRR genes targeted by miRNA than the remaining genes in the genomes of the four grass species. miRNA might serve as a regulator, controlling the expression levels of NBS-LRR genes. When no pathogen attacks, the host plant would regulate the expression of NBS-LRR genes at a low level to reduce its possible detrimental impact, especially for those NBS-LRR genes with high dosage. Targeting sizes were frequently located at NBS or other conserved domains (63.2%). Expression of NBS-LRR genes needs to be regulated to limit their metabolic costs and detrimental effects. 3
Conclusion
In summary, we have identified and characterized full-length NBS-LRR genes in sorghum, rice, maize, and
Author Contributions
Conceived the study: JW, XY. Analyzed the data: XY. Wrote the article: XY, JW. Both authors read and approved the final article.
Supplementary Materials
Supplementary Table S1
Full length NBS-LRR genes in grass species.
Supplementary Table S2A
85 genes in 16 NBS-LRR clusters of sorghum.
Supplementary Table S2B
58 R-like genes in 16 NBS-LRR clusters of sorghum.
Supplementary Table S3
NBS-LRR genes from grass species targeted by miRNA.
Supplementary Table S4
Twenty four NBS-LRR genes belonging to eight paralogous groups in sorghum.
Supplementary Table S5
Four ortholog groups of NBS-LRR genes in grass species.
Supplementay Figure 1
Lengths (bp) of gene conversion tracts.in mono-cluster, mixed-cluster, and paralogs.
Footnotes
Acknowledgments
The authors thank Ze Peng, Dev Paudel, Johnny Molestina, and Yang Zhao at the Agronomy Department, University of Florida, for reviewing this article.
