Abstract
Our study searched all available sequences of
Introduction
The single region 5ʹ end of cytochrome c oxidase 1 (
In this study, we aim to evaluate all available submitted sequences relating to
Materials and Methods
Collected sequences of Paphiopedilum from GenBank
All sequences belonged to
The number of sequences and the number of species between loci were different from each other. Among of them, 7 loci which have contained only 1 to 2 sequences, ie,
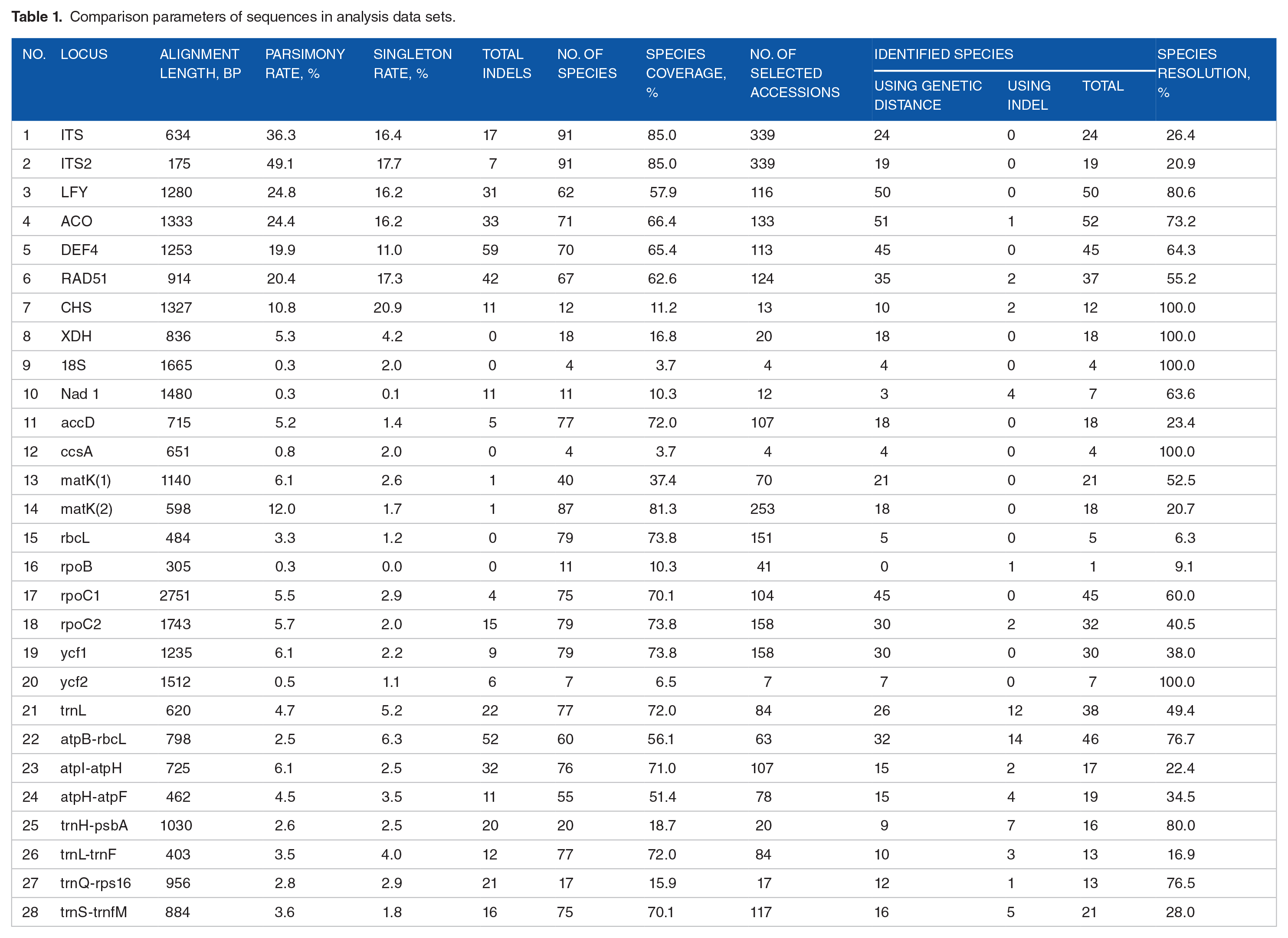
Comparison parameters of sequences in analysis data sets.
Multiple sequence alignment
Sequences at each locus were aligned using SeaView software version 4.4.0. 11 Alignments were then manually optimized, especially with noncoding regions in chloroplast genome and genes in nuclear genome which are highly divergent and contain many indel fragments. We manually noted all these information as they are also helpful in species resolution. The parsimony and singleton calculations for each data set were performed by Mega 7 program. 12 In the process of alignment, sequences that were too short and too divergent were removed from data sets. The output of alignment analyses is shown in Table 1.
Evaluation of species resolution
Phylogenetic tree for each locus were generated by Neighbor-Joining method using Mega 7 software with Kimura-2-parameter (K2P) model. 12 Evaluation of species resolution was performed using tree-based method and a combination of different measurements. First, species with just only one accession were considered to be distinguishable if the sequence was unique from others. In the phylogenetic tree, this accession would be shown as monophyletic branch. 13 Second, when multiple accessions were collected per species, these all accessions would be grouped into one monophyletic branch in the phylogenetic tree. Third, in converse, if conspecific individuals were not grouped together but separated in paraphyletic branches, then the species was considered as identification failure. A further description of insertions, deletions, and repeats should be included as if there was any difference between them that could help to authenticate them from the others.14,15 Finally, in case of undiscrimination by K2P distance, different species were grouped in the same branch in the phylogenetic tree. This means that sequences of these accessions were identical. These heterospecies sequences in the same branch would be also more observed with indel information.
In barcoding studies as well as phylogenetic researches, the identification of orthologs and paralogs give much influence on the exact species resolution. Although orthology is one of the most important criteria to evaluate the relationship between individuals, paralogous sequences which are the results of duplicate events and divergence are not correlated with speciation. The presence of paralogs in the same sample may lead to an overestimation of the number of unique species under identification. 16 In practice, this problem can be eliminated by identifying and avoiding paralog contamination before sequence analysis. First, if paralogs really exist in different size, the polymerase chain reaction products using universal primers will be expressed as ghost bands on gel in the electrophoresis step. Second, even the paralogs have the same length as the real gene, and we cannot recognize them under electrophoresis but the sequencing would be shown with ambiguities, double peaks, or noise. These are 2 simple ways to remove paralogs from our analysis. In this in silico study, all the accessions were selected from the available source of GenBank. In case of interspecies comparative, it was considered that the sequences were orthologs. In case of intraspecies relationship analysis, the accessions might be orthologs or paralogs, especially between different clones of the same samples. Instead of deleting the clone sequences, we decided to keep all of them and use as one of the factors affecting the species resolution. The locus with high variation between either intraspecies orthologs or homologous paralogs would result in low species resolution and could not be used as candidate molecular identification markers.
Results and Discussions
Multiple sequence alignment and divergence of the loci
The nuclear loci ITS, ITS2,

Species resolution of single loci of
To have a good discrimination effect, first, the selected sequences should have a proper length that is not too long for simple amplification and not too short for containing enough divergence information. Second, these sequences should be easy to be amplified, sequenced, and aligned. Third, the divergence of the sequence should be high enough for distinguishment at the species level but not too variable at underspecies level.
3
Finally, the high species resolution is a critical criterion. In our study, as the sequences were selected from GenBank, we did not analyze the amplification and sequencing rates. However, the sequence length, the alignment capability, the divergence range, and the resolution rate were all taken into account. The
The coding-gene regions in the chloroplast genome were easy to be aligned. The noncoding intergenic spacers in chloroplast and the nuclear loci took more time for alignment due to their great indels and polynucleotide repeats inside their sequences. However, they were all successfully aligned. Exceptionally, the
The capability of species identification with single regions
Some of the tested regions achieved 100% species resolution, ie,
Although the number species of
In the nuclear genome
The ITS2 locus has been highly noticed as an alternative barcode instead of the full-length ITS region due to ITS high divergence and short length for easy to amplify, to sequence, and to align.17,18 In this study, indeed ITS2 with a really short sequence (175-282 bp) could achieve just a little lower resolution than ITS (634 bp; Table 1). However, because the identification capability of ITS was still higher than ITS2, and the length of ITS (about 634 bp) was still suitable for amplification and sequencing, and moreover we did not meet any problem in alignment of this region, we still favored the whole ITS rather than the ITS2 in obtaining better resolution.
Among the analyzed regions in the nucleus, ITS and ITS2 loci gave the lowest resolution rate (26.4% and 20.9%). The highest ability was of

Comparison of resolution between ITS region and each of
In the chloroplast genome
To the
Among the chloroplast regions, the intergenic spacer

Comparison of resolution with the same species between pairs of regions
In the study of Parveen et al,
5
The use of indel information in species identification
In the chloroplast genome, the noncoding regions seemed to have more complexity than the coding regions based on high number of insertions and deletions. These characteristics contributed to the more difficulty in sequence alignment. However, because we could pass all the alignments in previous step, indel fragments on contrast became useful information for discrimination at species level. Specifically, in our study,
Capability of species identification with multiple region combinations
As no single locus could discriminate all species of
Comparison of species resolution of single-locus sequences and 2-locus combinations.
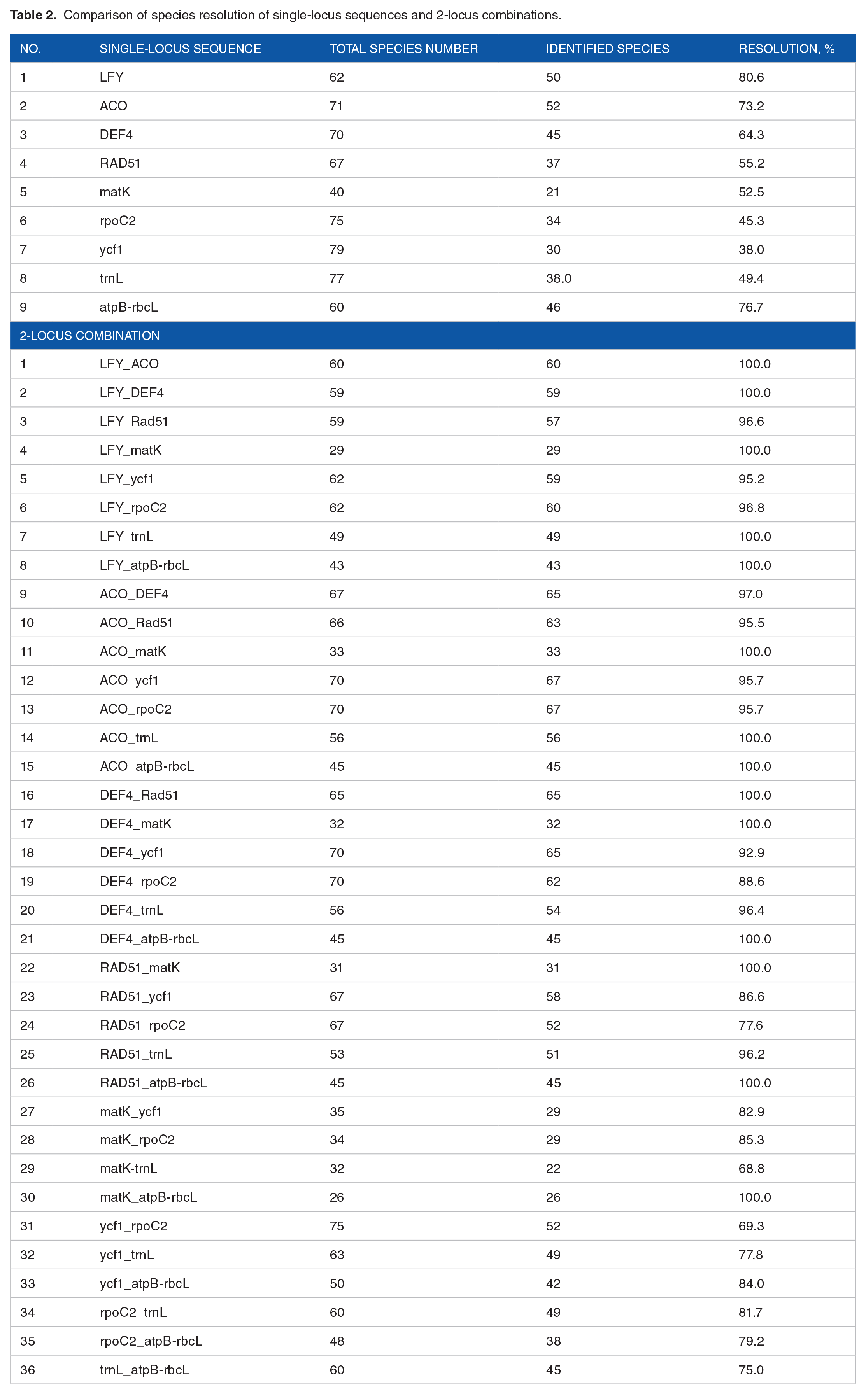
The resolutions of combination sequences were all higher than that of the single loci. In 2-locus combination, 14/36 data sets could be resolved completely (100%) with interspecies relationships (Figure 4).

Species resolution of single loci and 2-locus combinations. Percent species resolutions showed more potential in a combination of sequences for identification study.
Conclusions
Our study revealed that the 4 loci,
Because of the limitation of available sequence information for
Supplementary Material
Supplementary Material, Supplementary_Material – In Silico Study on Molecular Sequences for Identification of Paphiopedilum Species
Supplementary Material, Supplementary_Material for In Silico Study on Molecular Sequences for Identification of
Footnotes
Acknowledgements
The computer resources were provided by Computational Biology Center of International University, Vietnam National University.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This project was funded by the Asian Office of Aerospace Research and Development (AOARD) under grant FA2386-17-1-4032.
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
H-TV analyzed and interpreted the results and was a major contributor in writing the manuscript. PH performed the bioinformatics calculation. H-DT gave advice on the manuscript. LL gave advice and gave final approval of the version to be published. All authors read and approved the final manuscript.
Availability of Data and Materials
All the sequences used in the study were downloaded from GenBank. Details of accession numbers are given in Additional file 3.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
