Abstract
Introduction:
DNA barcode, a molecular marker, is used to distinguish among the closely related species, and it can be applied across a broad range of taxa to understand ecology and evolution. MaturaseK gene (
Result:
The family Dipterocarpaceae comprising of 15 genera is under threat due to some factors, namely, deforestation, habitat alteration, poor seed, pollen dispersal, etc. Species of this family was grouped into 6 clusters for
Conclusion:
Through the analysis of inter-generic, inter/intra-specific variation and phylogenetic data, it was found that both selection and mutation played an important role in synonymous codon choice in these genes, but they acted inconsistently on the genes, both
Introduction
Phylogenetic analysis is the big deal in biology, because it provides basic information of background of an organism especially about the status and modes of their existence. The phylogenetic analysis of Dipterocarpaceae has yet not been extensively studied. A very few major phylogenetic analysis on this family has been reported based on DNA barcode,
The parameter for plant barcodes success
First, geographical constraints generally make a high level of distinctive species discrimination.14,15 In contrast, the species diversity decreases as one moves toward dense populations which lead to shared barcodes among the coexisting species.16,17 Second, sufficient time is required for speciation driven by mutation or drift to form a set of genetic constituent which isolates conspecific individual together and separate them from other species. Barcode sequence represents the deficiency of proper species discrimination, due to slow rate mutation (
In the study of molecular evolution of individual genes, it is important to know the synonymous codon usages. Synonymous codon usages are not randomly used20,21 which have been influenced by factors such as CpG islands,
22
gene length,
23
gene expression,
24
protein secondary structure,
25
gene density,26,27 and so on. Two important models, ie, both mutational bias and natural selection, determine independently the codon usages variations. It is necessary to consider more codon usage patterns because no such unified theory for codon usages has been established. The chloroplast genome is found to be the most effective in the study of plant molecular evolution due to its small size, simple structure, and high copy number, which is closely similar to a bacterial genome. Recently, many more chloroplast genomes have been sequenced with the help of advanced DNA sequencing techniques (http://www.ncbi.nlm.nih.gov/genomes/GenomesGroup.cgi?taxid=2759&opt=Plastid#pageTop). The codon usage pattern of chloroplast genes (like
Transition rather than transversion, which is favored by natural selection, causes the biochemical advantage.32-34 The introductions of new alleles through the transition are several folds higher than that of transversion, and therefore, nucleotide transition is common in molecular evolution.
35
This pattern of amino acid replacement often supports the effect of selection on the ground that the transition is more conserved in their effect on protein supported by reviewing more than 8 published reports.
36
The selective hypothesis has proposed that conservative effect on biochemical factors by transition mutation over transversion is correlated to the pattern of evolutionary divergence.
37
One obvious question is that the changes fixed in the evolution of an organism are favored toward survival effect because natural selection encourages positive adaptive changes whether it happens through transitions or transversion. Every nucleotide side (eg, G) may experience one type of transition (G to A) at a rate X and 2 types of transversion (G to C, G to T) at rate Y. The aggregate rate ratio of transition to transversion has a null expectation of R = X/2Y = 0.5 = 50% (>R indicates that it is being fit or divergent in accordance with value and <R shows opposite results to earlier) leading to transition bias relative to a null model of equal rates. This study emphasizes on 2 major objectives. The primary objective is to analyze codon bias of
Materials and Methods
Data collection
The family Dipterocarpaceae comprises of 15 genera (www.theplantlist.org/1.1/browse/A/Dipterocarpaceae/). The entire coding region of
Indicator of codon usage
Codon usage pattern in the core barcode region was analyzed by using codon W 1.4.2. Relative synonymous codon usage (RSCU) is the ratio of the observed frequency of synonymous codons for a particular amino acid to the expected frequency. 38 Thus, RSCU values close to 1 indicate the lack of bias for codon usage where as the value >1 or <1 means preference and avoidance of that particular codon. The effective number of codon (ENC) is used to show the extent of codon bias of a gene and to quantify the absolute codon usage bias of a coding sequence. 39 The values of ENC always remain in-between 20 (a gene with extreme codon bias uses only one codon per amino acid) and 61 (a gene with no codon bias uses all the synonymous codon). 40 In general, 35 or less and 50 or higher ENC values of a gene are considered to have a strong and low codon bias, respectively. 41 The expected ENC values from GC3s under no selection in accordance with null hypothesis have been calculated according to equation (1), where S = GC3s.

The relationship between nucleotide content and codon usage by NC-plot is investigated to reveal the relationship among them. Wright 39 has suggested that NC-plot (ENC plotted against GC3s) is used to explain the pattern of synonymous codon usage. The codon choices of a gene influenced by a (G + C) mutation constrain usually lie on or just below the curve of the predicted value. 39 Codon adaptation index (CAI), a measurement of the expression of the gene, is used to estimate the extent of bias toward codon and its values range between 0 to 1.0, where in the CAI, a higher value means a stronger codon usage and a higher expression level. 42
Correspondence analysis by using codon W 1.4.2
Correspondence analysis (COA) is an ordination technique that identifies the major trends in the variation of the data where genes with their degree of variation are arranged along the continuous axes. It represents continuous variation accurately. The first axis captures most of the variation of genes and each of the subsequent axes shows a diminishing variation.
Chemical properties
The physiochemical properties like molecular weight (MW), theoretical isoelectric point (PI), percentage of positive and negative charged amino acid instability index, grand average of hydropathicity (GRAVY), etc were determined by using ProtParam (http://web.expasy.org/protparam/) (Supplemental Tables S3 and S4). The instability index provides an estimation of stability of a protein in vitro.
Sequence analysis
Gene sequences (
Phylogenetic analysis
The nucleotide frequency and transition/transversion bias was computed by Molecular Evolutionary Genetics Analysis (MEGA 7.0).
43
DVADIST program from PHYLIP was used to analyze the distance between the clades. The phylogenetic data were validated by re-sampling sequence data using bootstrap, performed by NJ-plot in Phylogeny Inference Package (PHYLIP).
44
Clustering of individuals was made on the basis of their position on the phylogenetic tree (clusters I, II, III, IV, V, and VI for
Results
Advancement of molecular biology and DNA sequencing of the genome of various organisms rapidly provide valuable information regarding their genetic makeup and function. In this study, changes in the nucleotide sequence of
Phylogenetic analysis
Monophyly individuals of all the clusters showed poor barcode (

Evolutionary relationship of taxa based on
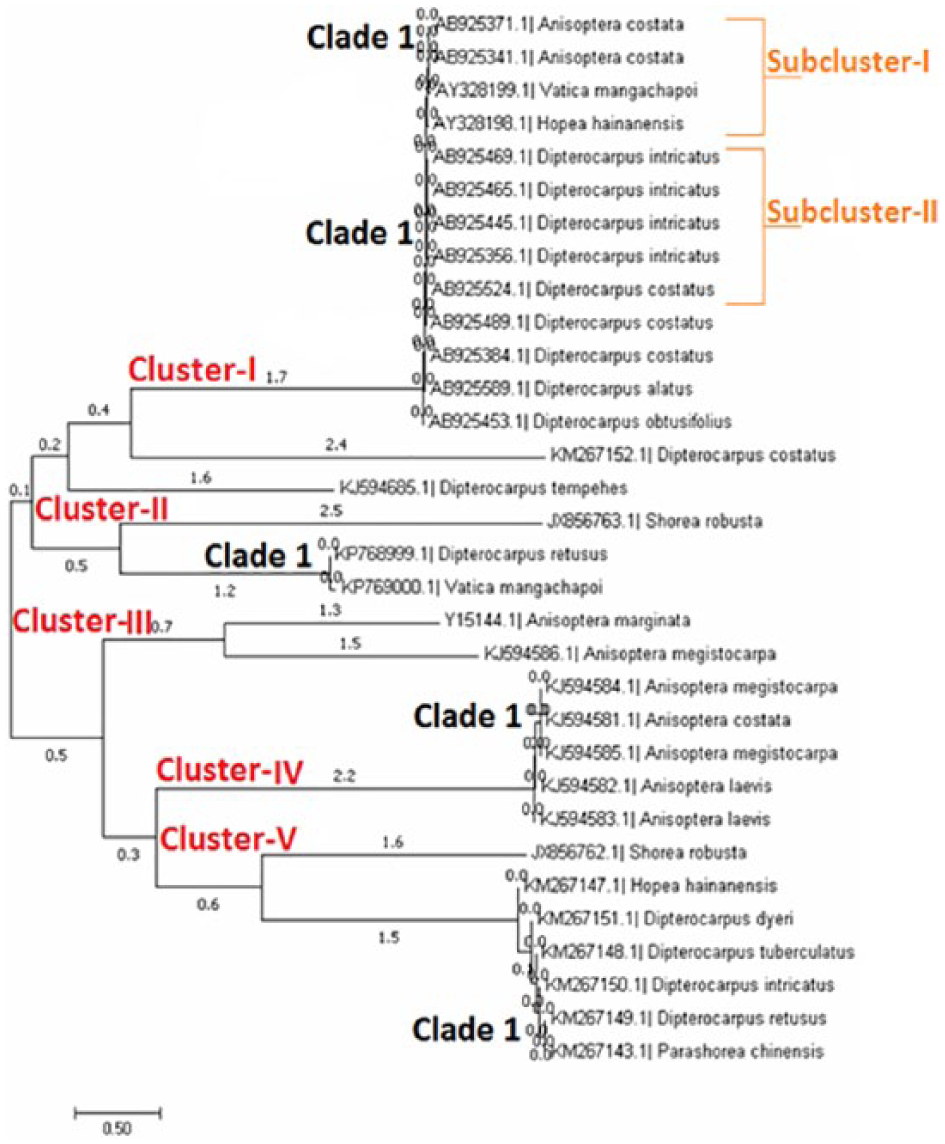
Evolutionary relationship of taxa based on
Codon usage
Average GC content among all the clusters ranged from 32.80% to 34.4% for
GC content in the all the clusters of


Percentage of frequently synonymous codon usage among the clusters: (A)
Synonymous codon usage pattern
Almost all the clusters of
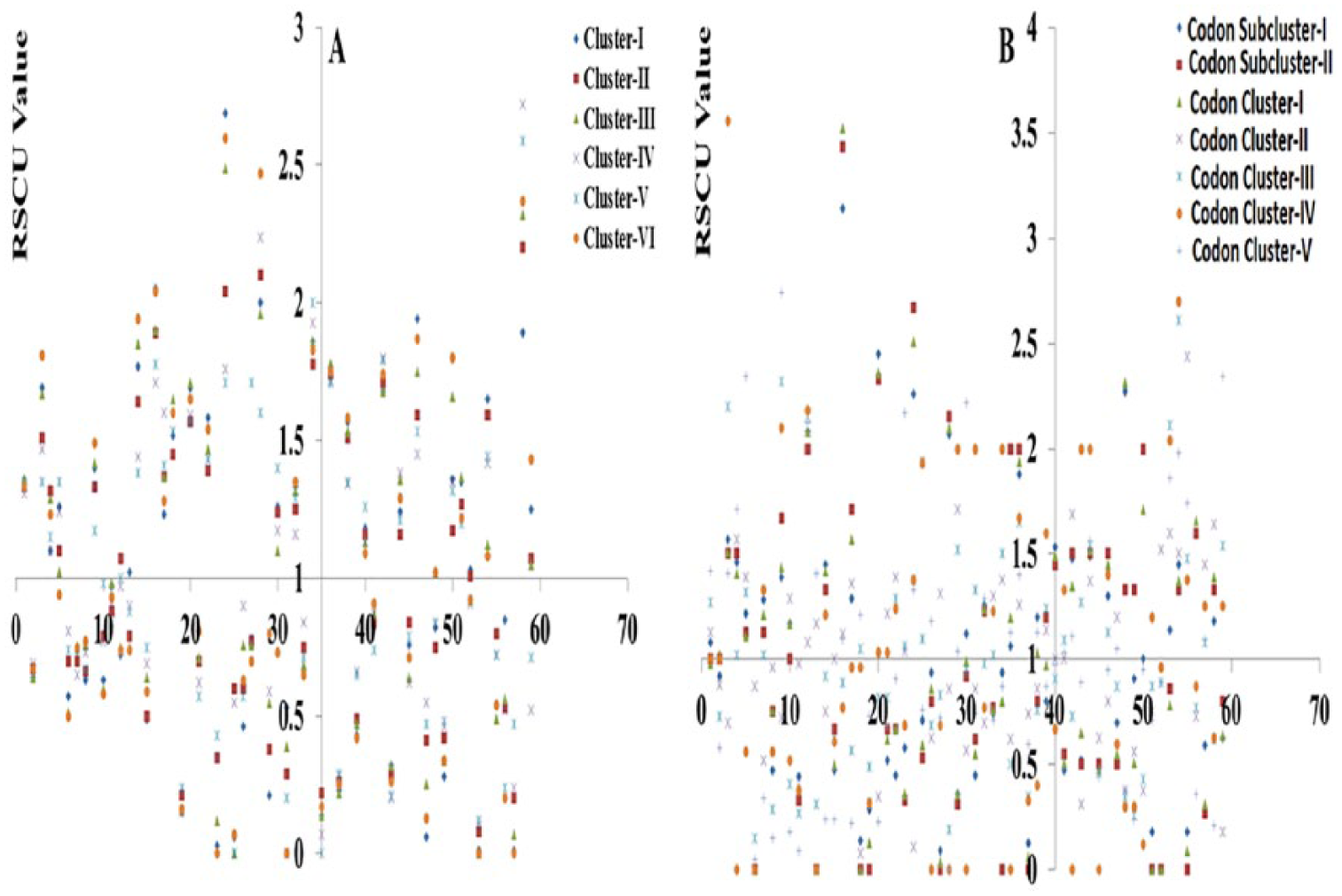
Codon usage pattern among the group members of this family based on RSCU value: (A)
Nucleotide content in 3 codon positions of gene
The GC content in 3 codon positions (GC1, GC2, and GC3) of

GC1, GC2, and GC3 among all the clusters: (A)
According to neutrality analysis, it is found that mutational pressure presumably influences the codon bias if the correlation between GC12 and GC3 is statistically significant and the slope of the regression line is close to 1. Conversely, a narrow distribution of GC content and the nonsignificant correlation between GC12 and GC3 are caused by selection.45,46 In vitro stable protein encoding individuals within each cluster of both
Correlation coefficient (

NA, not applicable, because sample size is 2 which is invalid for correlation.
Relation between ENC and GC3s
ENC and GC3s values for

ENC vs GC3s (the portions where observed ENC value occur on the standard curve): (A)
Mutational bias analysis
In the case of mutational bias, generally, GC or AT is used proportionally among the degenerate codon groups in a gene. On the contrary, GC or AT is not proportionally used for codon choice by natural selection.
48
The relationship among G, C, A, and T content in 4 degenerated codon families are analyzed to know whether these codon bias choices are restricted to the high bias genes or not (Figure 7A and B). Figure 7A showed codon bias for

Comparative ratio of A3/A3 + T3 and G3/G3 + C3: (A)
Chemical properties
Physiochemical properties of completely sequenced
Correspondence analysis
In this study, we investigated on the synonymous codon usage variation among the clusters and the individuals of each cluster of

Schematic representation of correlation axis 1 vs CAI or ENC or Fop or GC or GC3s for each cluster of
Correlation (

Abbreviations: CAI, codon adaptation index; ENC, effective number of codon; Fop, frequency of optimal codon; IN-VSPEI, in-vitro stable protein-encoded individual; NS, non-significant correlation.
NA, not applicable, because sample size is not more than 2. Not identified means codonW could not produce data of 4 axis. * and ** means significant at 0.01 and 0.05 probability levels, respectively.
Sequence analysis
Substitution bias (transition/transversion) ratio at codon position for each cluster revealed evolutionary trend in accordance with their values. The inference was made for both the genes on the basis of overall substitution bias value on their entire codon position (1st + 2nd + 3rd nucleotide). According to selective hypothesis on substitution bias (at 1st + 2nd + 3rd position), clusters I, II, and V of
Substitution bias at codon position of

NA, not applicable, because more than 2 samples are required to calculate substitution bias in MEGA 7.0.
Discussion and Conclusion
Codon usage bias is a complex and important issue regarding evolution in both prokaryotes and eukaryotes. There are some hypotheses that have been proposed to explain the origin of codon usage bias. Neutral theory
49
and the selection-mutation-drift balance model38,50 are one of the best representatives among them (hypothesis of the origin of codon usage bias). According to the neutral theory, random synonymous codon choices are the results of mutation at degenerate coding positions. The selective-mutation-drift model explains that the codon bias is supposed to be determined by the stability among mutational pressures, genetic drift, and selection. However, with the advancement of genome projects in the recent years, these 2 hypotheses are not sufficient for the explanation of codon usage bias. Several parameters like gene length,
23
GC content,51,52 recombination rate,51,53,54 gene expression level,23,53,55 RNA structure,24,56,57 protein structure,
58
intron length,
59
population size,
60
evolutionary age of genes,
61
environmental stress,
62
hydrophobicity and aromaticity of encoded proteins,63,64 and so on may influence the codon usage bias. In this study, gene expression level and gene compositional constraint have been given primary focus. In vitro stable protein of both
GC rich organisms such as bacteria, archea, fungi,
Phylogenetic analysis helped to identify the variations, patterns, transition/transversion bias, and codon bias in nucleotide sequence. Genome-based phylogeny is found to be effective in this concern, and it has been practiced in bacterial system (due to smaller genome size). In angiosperm, whole genome phylogeny is being challenged, because of very hard processing of so large massive information unlike bacterial genome. However, it is quite obvious to looking at DNA barcode. A software, MEGA, provided information about inter- and intra-specific relationship of the family Dipterocarpaceae. Phylogenetic tree analysis showed that cluster IV of
Supplemental Material
Supplemental_tables – Supplemental material for In Silico Analyses of Burial Codon Bias Among the Species of Dipterocarpaceae Through Molecular and Phylogenetic Data
Supplemental material, Supplemental_tables for In Silico Analyses of Burial Codon Bias Among the Species of Dipterocarpaceae Through Molecular and Phylogenetic Data by Raju Biswas, Anindya Sundar Panja and Rajib Bandopadhyay in Evolutionary Bioinformatics
Footnotes
Acknowledgements
Raju Biswas is thankful to CSIR for Junior Research Fellowship (File No: 09/025(0216)/2015-EMR-I). Authors are thankful to UGC-Center of Advanced Study and DST-FIST at Department of Botany, The University of Burdwan for pursuing research activities.
Funding:
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
Both RB designed the work. RB conducted the work. Both RB and ASP analysed the data. RB and ASP wrote the paper. RB checked the paper. All authors finalized and submitted the paper.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
