Abstract
Recently, DNA barcoding based on mitochondrial cytochrome c oxidase subunit I (COI) has gained wide attention because of simplicity and robustness of these barcodes for species identification including birds. The current GenBank records show the COI barcodes of only one species, chukar partridge (
Keywords
Introduction
Partridges are non-migratory birds of dry, open and often hilly terrains and belong to
DNA barcoding using mitochondrial cytochrome c oxidase subunit I (COI) sequences has enormous potential of discriminating closely related species across diverse phyla in the animal kingdom.1,2 Using a large dataset of North American birds, it has been concluded that DNA barcoding can be effectively applied across the geographical and taxonomic expanse of bird species. 3 Even a single DNA barcode has been suggested as a rapid tool to discover monophyletic lineages within a metapopulation that might represent undiscovered cryptic species. 4 For evaluating the discriminatory power of a barcode, it would be more appropriate that all members of a genus be examined, rather than a random sample of imprecisely defined close relatives, and taxa to be included from more than one geographic region. 5
In the recent years, DNA barcoding has been utilized for species identification of birds from different regions of the world.3,6–10 However, the barcodes of Saudi Arabian birds have not been firmly established. In the present investigation, we have sequenced the 694 bp region of COI gene of Arabian partridge and Philby's rock partridge and compared these sequences with previously published sequences of another species (chukar partridge) of the same genus. Arabian partridge is a resident, endemic to mountain areas of Saudi Arabia, Oman and Yemen, and found from 250–2800 m height. The Philby's partridge is a native to mountain area (1500–3000 m high) of southwestern Arabia and northern Yemen. This species is related to chukar, red-legged and barbary partridges and sometimes considered to be a race of the rock partridge. Although COI barcodes of chukar partridge are available in the GenBank, this is the first study reporting the barcodes of Arabian partridge and Philby's rock partridge.
Materials and Methods
The blood samples were collected from 3 specimens of Arabian partridge and 2 specimens of Philby's rock partridge. The taxonomic classification of these birds is as follows: Kingdom-Animalia, Phylum-Chordata, Class-Aves, Order-Galliformes, Family-Phasianidae, Genus-
COI sequences were amplified using the primer pair of BirdF1 and BirdR1 3 and FideliTaq PCR master mix (GE Healthcare) in a reaction volume of 30 μl. The PCR conditions included a denaturation step (1 min at 94 °C) followed by six cycles of 1 min at 94 °C, 1.5 min at 45 °C, and 1.5 min at 72 °C, followed in turn by 35 cycles of 1 min at 94 °C, 1.5 min at 55 °C, and 1.5 min at 72 °C, and a final extension for 5 min at 72 °C. The PCR products were electrophorsed on a 1% agarose gel stained with ethidium bromide. The PCR products were purified using MicroSpin S300 columns (GE Healthcare) before being sequenced using BigDye Terminator Cycle Sequencing Kit (Applied Biosystems, USA) on 3130XL genetic analyzer (Applied Biosystems). For each sample, two sets of sequencing reactions were performed using the forward and reverse primers for high accuracy. All the nucleotide sequences obtained in this study have been deposited in the GenBank with the following accession numbers: Arabian partridge specimens 1 to 3 (HQ168027 to HQ168029) and Philby's rock partridge specimens 1 and 2 (HQ168030 and HQ168031).
For comparative evaluation of barcode sequences of this study with previously published sequence data of other species form Alectoris genus, we searched the GenBank nucleotide database and found only 6 barcode records of a single species,
We also compared the COI-based phylogenetic inference with another suitable mitochondrial gene, control region (CR), which often evolves faster than rest of the mitochondrial genome and is highly variable in birds.
11
This variability has led to the expanding usage of CR sequences to examine questions ranging from population structure to phylogenetic relationship. We downloaded 9 records (3 for each species) of mitochondrial control region (CR) genes of
The sequences were aligned by ClustalW 12 and the alignment file was saved in appropriate formats (MEGA and PHYLIP). The aligned sequence data were subjected to unweighted pair group method with arithmetic mean (UPGMA) and maximum likelihood (ML) methods for looking into phylogenic inference. The UPGMA analysis was performed using MEGA4 software and the bootstrap consensus trees inferred from 1000 replicates were taken to represent the evolutionary history of the taxa analyzed.13,14 The software, Tree Puzzle was used for ML analysis 15 and the resulting phylogenetic trees were viewed by TreeView software. 16
Results
Although all the 9 sequences of COI gene segment (5 from this study and 4 from GenBank) were of equal length (694 bp), the alignment showed disparity in 2 nucleotides at both the ends of sequences. The
The pair-wise sequence comparison of COI gene showed 53 (7.66%) variable sites across all the species whereas within-species variable sites were found to be 4 (

Haplogram of COI gene showing the variable sites among the 9 individuals from 3 different species of
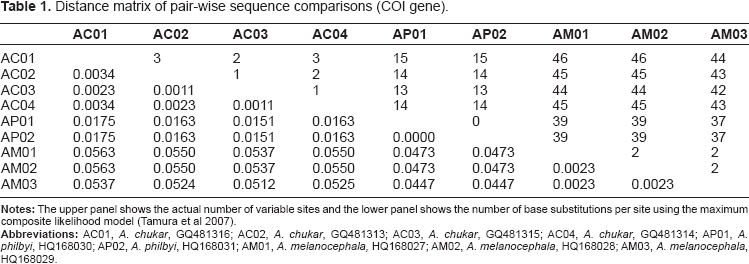
Distance matrix of pair-wise sequence comparisons (COI gene).

Haplogram of CR gene showing the variable sites among the 9 individuals from 3 different species of
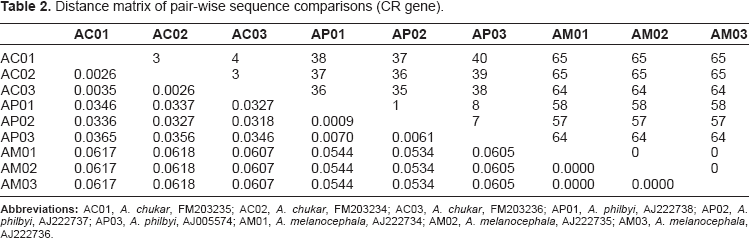
Distance matrix of pair-wise sequence comparisons (CR gene).
Phylogenetic analysis using 692 bp nucleotide segment of COI clearly discriminated the 3 species of Alecoris genus, while

Phylogenetic trees (based on 692 base pairs nucleotides sequence of COI gene) showing the relationship among three different species of

Phylogenetic trees (based on 1152–1155 base pairs nucleotides sequence of CR gene) showing the relationship among three different species of
The translation of nucleotide sequences (codon starts at position 2) revealed identical sequences of amino acids for all the taxa except one individual (

UPGMA tree (based on 230 amino acid residues sequence) showing the relationship among three different species of
Discussion
The results of this study clearly demonstrated the discriminatory power of COI barcodes for species identification. The sequences from the two samples of Philby's rock partridge were found to be identical whereas only 3 and 4 within-species variable sites were observed in the samples of Arabian partridge and chukar partridge respectively (Fig. 1). Both COI and CR genes exhibited nearly equal rates of divergence (7.66% versus 7.54%) in pair-wise sequence comparison, indicating that both the genes evolved at similar speeds, at least for the taxa of our study. This is not surprising as a previous study looking at the CR of 68 species of birds found that even cytochrome b evolved at equal speed or faster than the CR in the majority of cases. 17 There was a symmetrical distribution of variable sites in COI barcode (Fig. 1) as compared to skewed distribution in CR gene, where most of the variable sites occurred at the ends while the middle region (400–800 bp) represented most of the conserved sites (Fig. 2).
Phylogenetic analysis using nucleotide sequences of COI gene separated the three species into three different clades with high bootstrap support (Fig. 3); similar phylogenetic inference was also observed using CR gene (Fig. 4). However, the COI protein tree did not appear to be phylogenetically informative, apparently due to the use of closely related species (Fig. 5). Nevertheless protein trees are virtually more advantageous for intraspecific discrimination among distantly related organisms. We used two different methods for creating the phylogenetic trees; a distance method (UPGMA) and a character-based method (ML). Earlier, we have noticed that UPGMA could be a better alternative to maximum parsimony (MP) method for phylogenetic inference using mitochondrial sequences. 18 Moreover, ML and NJ methods have been shown to be nearly equally efficient and generally more efficient than the MP method. 19
Due to their faster evolutionary rates compared to ribosomal RNA genes, the mitochondrial protein-coding genes (such as COI) and noncoding CR are regarded as powerful markers for genetic diversity analysis at lower categorical levels, including families, genera and species.
20
However, COI barcodes are able to identify entities below the species level that may constitute separate conservation units or even species units.
21
Recently COI barcodes have been utilized for various purposes. Fleischer et al
22
have conducted DNA analysis of seven museum specimens of the endangered North American ivory-billed woodpecker (
Several investigators have also used mitochondrial CR sequences for molecular diversity and phylogenetic analyses of bird species.25–28 Escalante et al 29 have studied the phylogenetic relationship based on CR together with other two mitochondrial genes to trace the ancestry of the avian genera Oporornis and Geothlypis. Sequencing of the CR has also supplied valuable information on the ancestry of chukar partridge populations. 30 Huang et al 31 have analyzed CR from 180 individuals from 13 populations of the partridges and suggested the existence of a geographical structure among Chinese bamboo partridge populations, resulting from the synergistic affect of Pleistocene climatic variations. Qu et al 32 have observed a close relationship between CR-based phylogeny and phylogeographical structuring, while the level of genetic diversity in all avian populations studied were associated with the wide ecological distributions and niche variation. Huang et al 33 have utilized mitochondrial CR for studying population differentiation in context with phylogeographic structure of rusty-necklaced partridges. The mitochondrial CR sequences have suggested the existence of a phylogeographical structuring among rock partridge populations, resulting from genetic divergence in southern refugia and subsequent postglacial colonization of northern mountain areas. 34 Barbanera et al 35 have used CR in combination with cytochrome b to determine the genetic structure of Mediterranean chukar populations to aid management decisions. Moreover, female-mediated gene flow determined by CR haplotyping could be an important consideration for captive-breeding programs for threatened birds. 26
Although partridges have not acquired the threatened status at the global level, it is likely that local vulnerable population exists within the species range, mainly due to habitat loss and hunting that may warrant special attention. 36 Barilani et al 37 have reported widespread introgressive hybridisation, suggesting that released captive-bred partridges have reproduced and hybridized in nature polluting the gene pool of wild rock partridge populations in Greece. A clear understanding of evolutionary history and phylogeography provides valuable information about the influence of physical barriers and habitat preference on gene flow in birds. 38 High levels of mitochondrial genetic diversity in combination with genetic differentiation among subgroups within regions and between regions highlight the importance of local population conservation to preserve maximal levels of genetic diversity. 26 Genetic data have also suggested that patterns of speciation and population diversification of Przewalski's rock partridge have been affected by the stability of the climate, natural selection, and human intervention. 39 Thus, application of COI barcoding for molecular diversity and phylogenetic analysis of partridges could provide valuable information about species identification, population structure, evolutionary history and molecular conservation.
In conclusion, COI barcoding is a powerful tool for species identification and phylogenetic inference. The nucleotide sequence of partial segment of COI gene effectively discriminated 3 species of the genus
Disclosure
This manuscript has been read and approved by all authors. This paper is unique and is not under consideration by any other publication and has not been published elsewhere. The authors and peer reviewers of this paper report no conflicts of interest. The authors confirm that they have permission to reproduce any copyrighted material.
Footnotes
Acknowledgments
This study was supported by a grant from HRH Prince Sultan Bin Abdulaziz Research Chair for Environment and Wildlife, Riyadh, Saudi Arabia. We sincerely thank the authorities of the NWRC for supporting our research and making arrangements for sample collection. The technical assistance of Mr. Anis Ahamed is highly appreciated.
