Abstract
Novel tissue injury biomarkers have recently been identified that outperform or add value to the conventional safety biomarkers. These novel biomarkers have enhanced sensitivity and/or specificity in monitoring drug-induced tissue injury in a variety of tissues, included liver, kidney, and skeletal muscle. Among these novel biomarkers, microRNAs (miRNAs) are one type in particular that have received much attention in recent years. These microRNAs are short, endogenous noncoding nucleic acids that are involved in modulation and regulation of mRNA transcripts. Other attributes of miRNAs are that they exist in tissues at high abundance, and individual miRNAs can be highly tissue-specific. These miRNAs can be readily assayed in blood, urine, or cerebral spinal fluid, making them attractive as accessible biomarkers of tissue injury. Further, the miRNA processing involves embedding the miRNA within a protein complex, making them stable in plasma upon leakage from injured tissues. This review article will highlight the discovery of tissue-specific miRNAs and their evolution as novel toxicity biomarkers in recent years.
Keywords
Introduction
MicroRNAs (miRNAs) are short, single-stranded noncoding nucleic acids that are found in animals, plants, and some viruses. 1,2 Following processing within the cell, mature miRNAs are packaged within a protein complex, known as the RNA-induced silencing complex (RISC), which functions to modulate mRNA translation through repression and targeted degradation. 1 –3 Hundreds of unique miRNA species exist within cells, with some individual miRNAs expressed at hundreds of thousands copies per cell. 4 Biologically, miRNAs are involved in a variety of cellular functions, including cell differentiation, proliferation, and cell survival, 5,6 although attributing functions to some individual miRNAs has proven challenging. 6
Early identification of miRNAs annotated them based on sequence and used a numbering system to classify. 1 Further, it was also noted that within certain tissues, some of the individual miRNAs were present in a tissue-specific or predominant manner. 7,8 Another noteworthy observation is that individual miRNA sequences are highly conserved across species, with the RNA sequence the same in a range of mammalian species including human. 9 Taken together, miRNAs have become attractive novel accessible biomarker candidates for monitoring tissue injury in blood, urine, or cerebral spinal fluid, due to their high abundance, tissue specificity, relative stability (in part due to the fact that the miRNAs are embedded with the protein RISC), and easily measured using quantitative polymerase chain reaction (qPCR) methodologies that translate across species.
Early Demonstration of Circulating miRNAs as Biomarkers of Tumorigenesis
Following the discovery of miRNAs in 1993 in
Initial Observation of miRNA Tissue Specificity and Use as Toxicity Biomarkers
In the early 2000s, a critical observation was that certain miRNAs existed in a tissue-specific manner. 7 Notably, miR-122 was one of the first to be identified that was only located within liver hepatocytes. 7,21 Research employing targeted qPCR and sequencing approaches in humans and nonclinical species identified several additional candidates that were present only within a given tissue, expanding the list of tissue-specific miRNAs. 8,22 –25 Table 1 contains a selection of representative tissue-specific miRNAs.
Representative Tissue-Specific or Predominant miRNAs.
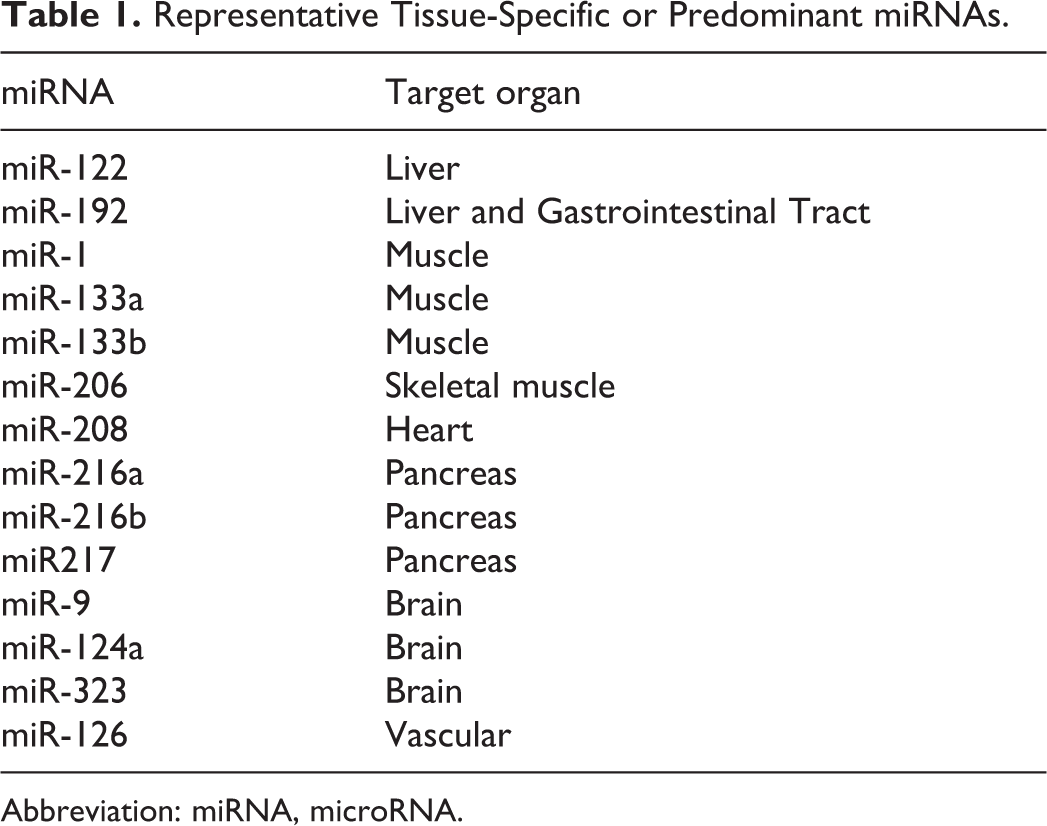
Abbreviation: miRNA, microRNA.
Building on the miRNAs’ tissue specificity and monitorability in blood, researchers started using circulating miRNAs to monitor drug-induced tissue injury (defined as tissue degeneration and necrosis), where highly abundant, tissue-specific miRNAs leak into circulation following cell injury (Figure 1). The first publication demonstrating the potential value of circulating miRNAs as toxicity biomarkers involved measuring the liver-specific miR-122 and other miRNAs in plasma following liver injury induced by acetaminophen in mice. 26 These data provided supporting evidence that miRNAs were as sensitive as conventional liver biomarkers, notably alanine aminotransferase (ALT). A subsequent report further expanded this observation by evaluating miR-122 along with muscle-specific miRNAs, miR-133a, and miR-133b in rats, following administration of either a liver or muscle toxicant. 25 Here, not only did the miRNAs reflect drug-induced liver or muscle injury but provided additional value related to specificity of tissue injury. In these representative studies with liver or muscle injury, both ALT and aspartate aminotransferase (AST) were significantly increased with both liver and muscle injury, whereas miR-122, miR-133a, and miR-133b were only increased in a tissue injury–specific manner. These early efforts highlighted the utility of miRNAs as circulating biomarkers of tissue injury, with both sensitivity and specificity for monitoring drug-induced tissue degeneration and necrosis. Several additional investigations over the years have further demonstrated the added value of the liver and muscle-specific biomarkers and potential use in drug development. 27
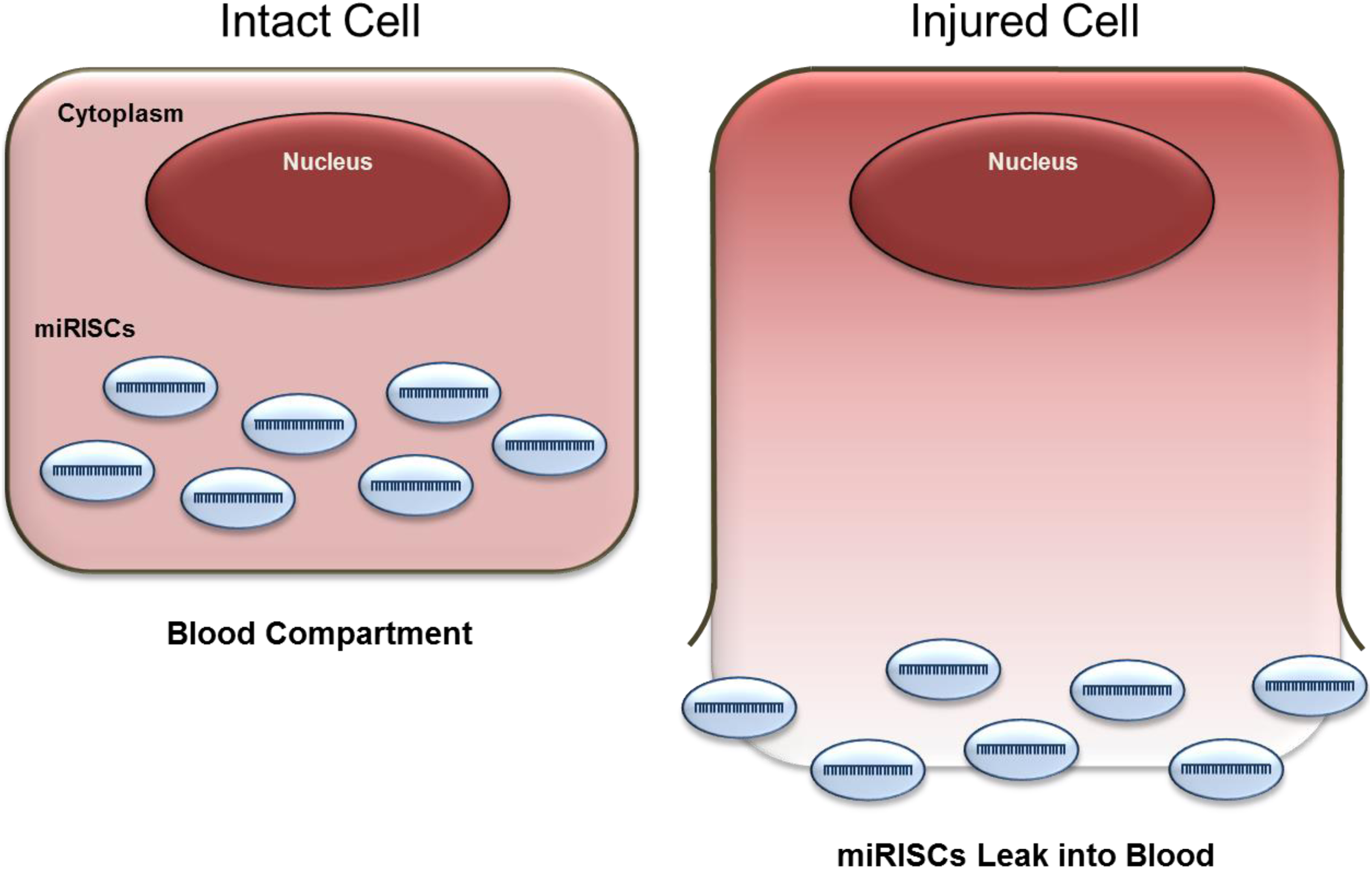
miRNA leakage into blood following cell injury. miRNAs embedded in protein miRISC. miRISC indicates RNA-induced silencing complex; miRNA, microRNA.
Expansion of Tissue-Specific miRNAs as Toxicity Biomarkers of Other Tissues
After demonstrating that miR-122 and miR-133 were sensitive and specific biomarker of liver and muscle injury, respectively, additional research has expanded the use of other tissue-specific miRNAs as biomarkers of drug-induced injury. For example, plasma miR-124 has been studied to monitor brain 25,28 and retinal 29 injury, and the heart-specific miR-208 has been studied as a biomarker of drug-induced cardiac injury in rats, 30 miR-202 as a circulating biomarker of testicular toxicity 31 and miR-216a, miR-216b, and miR-217 to monitor drug-induced pancreas injury. 32,33 Kidney is another tissue where miRNAs have been used to monitor drug-induced injury 34 –36 ; however, a majority of the miRNAs evaluated are not tissue-specific. Since generally the assessment of kidney injury is performed in urine, tissue-specific miRNAs may not be as critical. Further investigations also demonstrated the potential to use specific miRNAs to provide mechanistic insight by association with regional tissue injury, utilizing patterns of miRNAs measured in urine to differentiate glomerular versus tubular degeneration. 36 These representative examples demonstrate the expanding list of tissues where accessible miRNAs can be monitored to assess drug-induced tissue toxicity in rodents, dogs, and nonhuman primates.
Translation of miRNAs to the Clinic
As the supporting nonclinical data builds, the translation of miRNAs as circulating biomarkers of toxicity has also expanded into clinical settings. Early clinical efforts have focused on liver-specific miR-122 as a biomarker of liver injury, 37 –39 most notably in patients with acetaminophen-induced liver injury, but miRNAs have also show promise for other tissue injury, including doxorubicin-induced cardiotoxicity, 40 pancreatitis, 41 and kidney injury. 42,43 With RNA-sequencing technologies allowing for cheaper and efficient analyses of the full panel of miRNAs, rather than a targeted qPCR assay, researchers have started to employ miRNA profiles or “liquid biopsy” approaches in the patients with tissue injury, as well as in disease populations. 37,43 –45 These approaches not only help identify additional miRNA candidates but also provide a pattern of miRNAs associated with both certain disease states and types of injury.
Current Gaps and Limitations
As research continues to demonstrate the added value of circulating miRNA as novel biomarkers of toxicity and tissue injury, one area of focus remains on the analytical methodologies for measuring miRNAs in biological fluids. 46,47 Unlike enzymatic assays that are standardized across automated analyzers or protein assays using commercially available kits, miRNA methods can vary from researcher to researcher, including the isolation of total RNA from the biological sample, the qPCR platform, and the normalization of the end point miRNA signal. These analytical differences make it challenging to directly compare data across independent studies and emphasize the need for a standardized and consistent manner of miRNA measurements.
Although some of the miRNAs highlighted here are highly tissue-specific, not all tissues have examples of tissue-specific or even enriched miRNAs. Individual cell populations within tissues may also lack candidate miRNAs to monitor induced toxicity or provide mechanistic insight into the nature of the induced injury within a given organ. Perhaps as miRNA-sequencing approaches become more prevalent, patterns of miRNA increases in blood may provide additional insight into mechanism or the localization of the injury in the tissue. Additionally, the miRNA stability in circulating blood may not be comparable to other protein biomarkers (eg, miR-122 vs glutamate dehydrogenase to monitor liver injury 27 ), where a relatively shorter half-life could make sample collection timing critical for the assessment of drug-induced injury.
Summary
The discovery of miRNAs and their attributes such as tissue specificity, high tissue abundance, and sequence homology across species has positioned miRNAs as the next generation of novel-specific and novel-sensitive biomarkers of tissue injury. Within the 10 years since the initial observations of miR-122 as a circulating biomarker of liver injury, the list of tissue-specific miRNAs has expanded greatly and their performance to monitor tissue toxicity has been demonstrated in dozens of nonclinical toxicity studies. Translation into clinical settings will further expand their value added in drug development and gain further acceptance as sensitive and specific biomarkers of drug-induced tissue injury.
Footnotes
Acknowledgments
The authors would like to thank Dr Jose Lebron and Dr Frank Sistare, and Dr Katerina Vlasakova for their critical evaluation of this article.
Authors’ Contributions
All authors drafted manuscript, critically revised manuscript, gave final approval,and agreed to be accountable for all aspects of work ensuring integrity and accuracy.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
