Abstract
N6-methyladenosine (m6A) is the most common reversible internal modification in mammalian RNA. Changes in m6A levels have been implicated in a variety of cellular processes, including nuclear RNA export, control of protein translation, and protein splicing, and they have been linked to obesity, cancer, and other human diseases. METTL3 and METTL14 are N6-adenosine methyltransferases that work more efficiently in a stable METTL3-METTL14 heterodimer complex (METTL3-14). ALKBH5 is an m6A-RNA demethylase that belongs to the AlkB family of dioxygenases. We report the development of radioactivity-based assays for kinetic characterization of m6A-RNA modifications by METTL3-14 complex and ALKBH5 and provide optimal assay conditions suitable for screening for ligands in a 384-well format with Z′ factors of 0.78 and 0.77, respectively.
Introduction
RNA methylation at the N6 position on adenine (m6A) was originally identified in the 1970s,1–3 and was shown to be one of the most abundant internal mRNA modifications. 4 These initial reports showed that approximately 0.1% to 0.4% of the adenosines in cellular RNA are m6A modified. 4 The m6A modification has been identified in many species including viruses, plants, yeast, mammals, and flies (reviewed in refs. 5 and 6). Furthermore, m6A modification has been shown to occur on many RNAs including noncoding, ribosomal, and polyadenylated. 5 In eukaryotic cells, m6A accounts for more than 80% of all RNA methylation. 7 The m6A modification on mRNA mainly occurs on exonic regions along with 3′-untranslated regions (3′-UTR) and tends to mark unstable transcripts.7,8 It has been reported that for a large number of transcripts in mouse embryonic stem cell (mESCs), m6A methylation inversely correlates with mRNA stability and gene expression. 9 Beyond mESC development, m6A modification has been linked to many physiological processes such as synaptic signaling, cancer, sperm development, yeast meiosis, plant development, RNA splicing, and circadian periods.5,9–15 Not surprising given the many correlations of m6A and physiological processes, m6A methylation levels vary in different tissues and cell lines with higher enrichment in liver, kidney, and brain. 4
METTL3 (methyltransferase like 3) and METTL14 (methyltransferase like 14) catalyze m6A methylation (Scheme 1). These two enzymes with 43% amino acid identity recognize the GGACU sequence on mammalian nuclear RNA. 16 METTL3 and METTL14 form a stable heterodimer complex that functions to drive m6A methylation more efficiently than either enzyme does individually. 16 Although the heterodimer formation greatly increases the enzymatic activity of METTL3-14, two other binding partners have been recently identified. The binding of Wilms tumor 1-associating protein (WTAP) and KIAA1429 to the heterodimer complex further simulate the enzymatic activity. WTAP appears to greatly increase the binding affinity of METTL3 for RNA.17,18 However, WTAP is not needed for the heterodimer activity in vitro, and its presence appears to affect which m6A sites can be specifically methylated, suggesting WTAP-dependent and -independent methylation sites. 18 WTAP is also responsible for the localization of METTL3 and METTL14 to nuclear speckles enriched with pre-mRNA processing factors and for catalytic activity of the m6A methyltransferase in vivo. 17 Loss of m6A has been reported to promote self-renewal and hinder differentiation of mouse embryonic stem cells. 8 However, depletion of METTL3 and METTL14 in HeLa cells led to decreased cell viability and apoptosis. 16
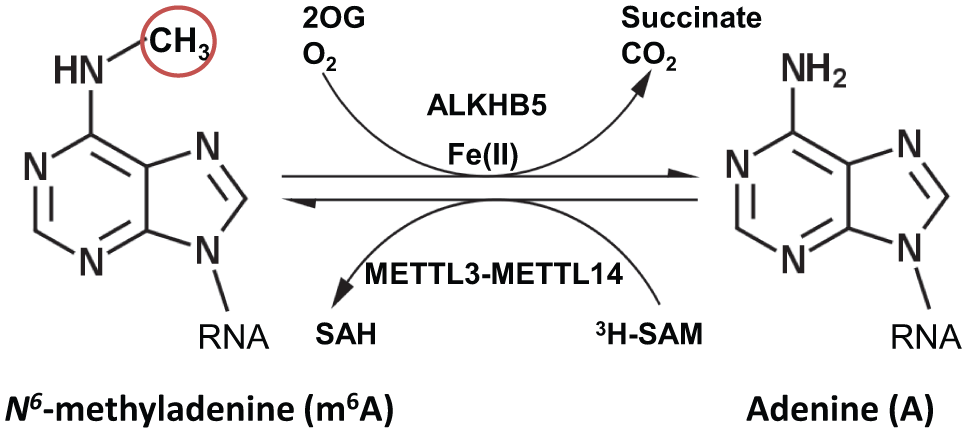
Reversible m6A modification in RNA. METTL3-14 complex methylates and ALKBH5 demethylates N6-adenine of RNA.
Similar to other methylation marks, mRNA methylation can be reversed through the enzymatic activity of demethylases (Scheme 1). Demethylation of the m6A mark is carried out by fat mass and obesity protein (FTO) and ALKBH5, both of which belong to the ALKB family that includes ALKBH1-8 and FTO.11,19,20 Both FTO and ALKBH5 are Fe(II)- and 2-oxoglutarate (2-OG)–dependent enzymes that remove m6A marks through an oxidative process. 21 Knockdown of ALKBH5 was shown to affect splicing in tissue culture cells and displayed a sterility phenotype in male mice. 11 FTO has been linked to obesity and type 2 diabetes.22,23 Specifically, loss of FTO in mice has been linked to a reduction in adipose tissue, decreased lean body mass, and postnatal growth retardation, 24 whereas Alkbh5-deficient mice have impaired fertility, a result of apoptosis during spermatocyte formation. 11 FTO and ALKBH5 are the only two m6A-RNA demethylases identified to-date.
Methyltransferase activity of METTL3-14 complex and demethylase activity of ALKBH5 have been previously reported.11,16,25 Here, we used a radioactivity-based assay to fully characterize the m6A methyltransferase activity of the METTL3-14 complex and used this complex to prepare the 3H-m6A RNA substrate to further characterize ALKBH5 activity. We further optimized these assays for screening in a 384-well format, which enables discovery of inhibitors for these key enzymes in the maintenance of m6A mark.
Materials and Methods
Iron (II) sulfate (cat. No. F7002), 2-OG (cat. No. K1875), ascorbic acid (cat. No. A5960), S-(5′-adenosyl)-L–methionine (SAM; cat. No, NET155V250UC), and S-adenosylhomocysteine (SAH; cat. No. A9384) were purchased from Sigma. 3H-SAM and a 384-well FlashPlate coated with streptavidin (cat. No. SMP410A001PK) were purchased from Perkin Elmer. Biotin-labeled single-strand RNA (ss-RNA; sequence: 5′-UACACUCGAUCUGGACUAAAGCU GCUC-biotin-3′) was ordered from Integrated DNA Technologies. The biotinylated RNA was about 90% pure according to quality control documents provided along with the product.
Protein Expression and Purification
The METTL3 and METTL14 clones were kindly provided by Dr. Chuan He (University of Chicago). The expression and purification were carried out as previously reported with some modifications.
16
The FLAG-tagged constructs were expressed by using Expi293 cells. The recombinant protein complex was purified by incubating the cleared lysate with equilibrated anti-FLAG M2 affinity agarose gel (Sigma, cat. No. A2220) for 1.5 h rotating, followed by washing with 10 CV TBS (50 mM Tris-HCl pH 7.4, 150 mM NaCl, and 10% glycerol). The recombinant protein complex was eluted by competitive elution with a solution containing 100 µg/mL FLAG peptide (Sigma, cat. No. F4799) in TBS, 1 mM dithiothreitol (DTT), and 10% glycerol. The purity of the complex was judged by sodium dodecyl sulfate polyacrylamide gel electrophoresis (
The ALKBH5 gene (amino acids 74 to 294) was expressed in Escherichia coli BL21 (DE3) V2R-pRARE and purified by DE52 and Ni-NTA columns (Qiagen) as previously described.19,26 Briefly, the lysate was passed through a DE52 column at 500 mM NaCl to filter DNA and other nonprotein contaminants followed by the Ni-NTA column. Resin was washed with binding buffer (20 mM Tris, pH 7.5, 500 mM NaCl, 5% glycerol, and 5 mM imidazole) followed by washing with wash buffer (binding buffer with 30 mM imidazole) and eluted with elution buffer (binding buffer with 250 mM imidazole). The His-tag was cleaved overnight with pTEV, while dialyzing in 10 mM Tris-HCl pH 7.5, 150 mM NaCl, followed by passing the sample through a second Ni-NTA column to filter the free tags out. Cut protein was then loaded onto a SuperDex 200 (26/60) column using AktaExplorer in 10 mM Tris-HCl pH 7.5, 150 mM NaCl, 1 mM DTT. After pooling selected fractions, ALKBH5 was then subjected to ion exchange using AktaFPLC, on a 1 mL Resource S column. Pure protein was eluted around 220 mM NaCl and was pooled, concentrated, aliquoted, and flash frozen (
METTL3-14 Assay
The methyltransferase activity of METTL3-14 complex was measured using a radioactivity-based assay. In this assay, tritiated S-adenosyl-L-methionine (3H-SAM) and unmethylated N6-adenine (A) ss-RNA were used as substrates. After reaction, the biotinylated ss-RNA (m6A) was captured in a FlashPlate coated with streptavidin/scintillant. The amount of methylated m6A ss-RNA was quantified by scintillation counting (count per minute [CPM]). All assays were performed using 20 mM Tris pH 7.5, 1 mM DTT, 0.01% Triton X-100, 40 U of RNaseOUT (cat. No. 10777019, Life Technologies)/100 µL buffer. For determination of kinetic parameters, protein concentrations and reaction time were optimized to obtain linear initial velocities. The METTL3-14 complex concentration was 10 nM. To determine the Kmapp values for SAM and ss-RNA, each was titrated into the assay mixture containing the enzyme at the saturating concentration of the other substrate (0.5 µM of SAM or 200 nM ss-RNA). The Km values were calculated using GraphPad Software.
ALKBH5 Assay
The ALKBH5 substrate, m6A-ss-RNA (5′-UACACUCGA UCUGG(m6A)CUAAAGCUGCUC-3′-biotin) was generated using the METTL3-14 complex to completely monomethylate N6A ss-RNA. Of the METTL3-14, 500 nM was incubated with 5 µM of biotin-labeled ss-RNA and 5 µM of 3H-SAM in 20 mM Tris-HCl pH 7.5, 1 mM DTT, 0.01% Triton X-100, 40 U of RNaseOUT/100 µL buffer for 1 h at 23 °C. Then, the reaction mixture was heat shocked at 85 °C for 15 min to denature the methyltransferase METTL3-14 complex, after which the sample was left on ice to cool down. To determine the kinetic parameters, the demethylase assays were performed in 25 nM ALKBH5, 10 µM 2-OG, 100 µM ascorbate, 20 µM Fe (II), and varying concentrations of monomethylated ss-RNA. An equal volume of 7.5 M guanidine-HCl was used to quench ALKBH5 activity after 1 h reaction at 37 °C. Sixty microliters of 20 mM Tris-HCl, pH 8.0, was added into the quenched wells. The quenched reaction mixtures were transferred to a streptavidin/scintillant-coated microplate (FlashPlate Plus, Perkin Elmer Life Sciences), allowed to bind for at least 1 h, and then the decrease of radioactive signals was detected on a Top-Count NXT HTS (Perkin Elmer).
Z′ Factor Determinations
To assess the quality of METTL3-14 or ALKBH5 assays for screening in a 384-well format, Z′ factors 27 were determined. In these assays, the reactions were run in 8 complete rows (half plate) with or without a known inhibitor. SAH was used as an inhibitor for METTL3-14, and IOX1 (A broad-spectrum inhibitor of most 2-OG oxygenases) was used for ALKBH5. The Z′ factors were calculated as described by Zhang et al. 27
Results
Methylation of N6-adenine on ss-RNA (m6A) is a reversible modification that is maintained by methyltransferase activity of METTL3 and METTL1416 and demethylase activity of ALKBH5 and FTO (Scheme 1).11,28 Their roles in diverse biological processes and possible disease associations necessitate developing robust assays to enable screening with higher throughput to discover potent and selective inhibitors that could be used to further study the consequences of changes in their activities in cells. We chose to develop radioactivity-based assays for both the METTL3-14 complex and ALKBH5, which are amenable to screening in a 384-well format and suitable for kinetic characterizations.
Similar to the histone methylatransferase activity assays we previously reported, 29 the mRNA methyltransferase activity of the METTL3-14 complex was assayed by measurement of the enzymatic transfer of tritiated methyl groups from 3H-SAM to an N6 of adenine in ss-RNA (m6A). We then used the METTL3-14 complex at an optimized condition to generate m6A–ss-RNA (methylated) substrate to assay ALKBH5. ALKBH5 activity was measured by monitoring the removal of a 3H-methyl group from m6A–ss-RNA (Scheme 1).
Assay Optimization for METTL3-14 Complex
It was previously shown that METTL3 and METTL14 form a stable active complex. 16 Using 3H-SAM and ss-RNA, we were able to monitor m6A–ss-RNA production and tested the activity of the complex with respect to reducing agents ( Fig. 1A ), KCl ( Fig. 1B ), NaCl ( Fig. 1C ), MgCl2, ( Fig. 1D ), Triton X-100 ( Fig. 1E ), EDTA ( Fig. 1F ), DMSO ( Fig. 1G ), glycerol ( Fig. 1H ), and buffer pH ( Fig. 1I ). The complex was most active at pH 7.5. Reducing agents DTT and Tris (2-carboxyethyl) phosphine (TCEP) are widely used to keep enzymes active. Conversely, the addition of DTT or TCEP dramatically inhibited the activity of METTL3-14 in a concentration-dependent manner ( Fig. 1A ). We tried to avoid reducing agents; however, it was essential to use the RNase inhibitor (RNaseOUT) in the assay mixture, which requires the presence of DTT to function. Therefore, we kept the DTT at low concentration (≦1 mM). The ionic strength of the buffer also significantly affected METTL3-14 activity. The enzyme activity was inhibited as the ionic strength increased, inactivating the protein complex at 100 mM of KCl ( Fig. 1B ) or NaCl ( Fig. 1C ) and 1 mM of MgCl2 ( Fig. 1D ). Triton X-100 (up to 1%), EDTA (up to 1 mM), and glycerol (up to 5%) had no significant effect on METTL3-14 complex activity ( Fig. 1E , F , and H ). DMSO as high as 5% had little effect on the activity of the complex ( Fig. 1G ).
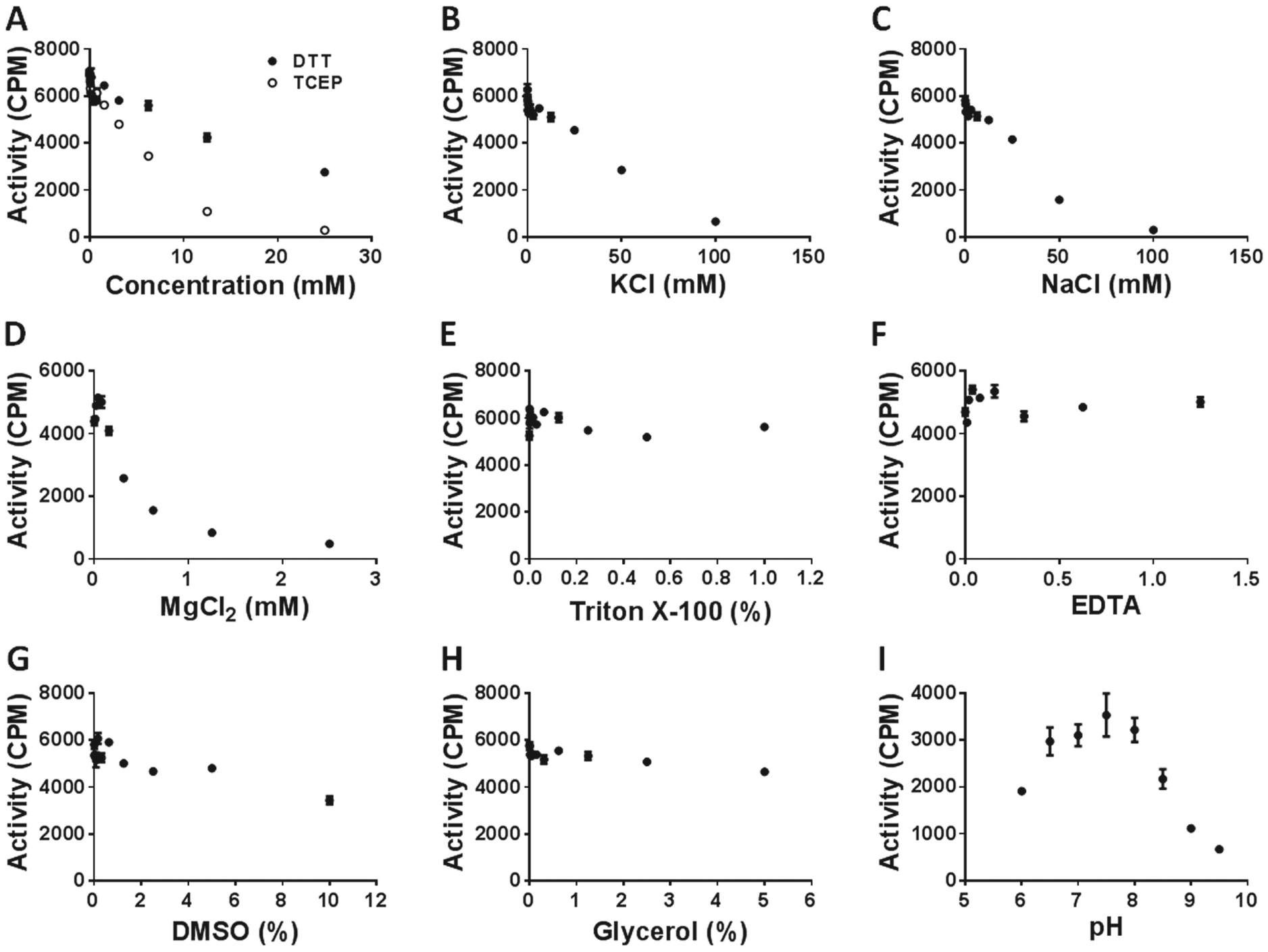
Assay optimization for the METTL3-14 complex. The methyltransferase activity was measured to determine the optimum assay conditions with respect to reducing agents (
Based on these observations, the optimal assay condition (20 mM Tris-HCl, pH 7.5, 0.01% Triton X-100, 1 mM DTT, and 40 U RNaseOUT/100 µL) was used for all kinetic parameter determination experiments.
Kinetic Parameters for METTL3-14 Complex
Using the optimized assay conditions, the kinetic parameters were determined for the METTL3-14 complex with ss-RNA substrate containing a consensus sequence of GGACU (Km values of 102 ± 15 nM and 22 ± 2 nM for SAM and ss-RNA, respectively, and kcat value of 18 ± 2 h–1; Table 1 ). The catalytic efficiency (kcat/Km) of the METTL3-14 complex was 818 h–1 µM–1 ( Table 1 ; Fig. 2A , B ). SAH showed strong inhibitory effects on METTL3-14 activity with an IC50 value of 0.9 ± 0.1 µM ( Fig. 2C ). The quality of the high-throughput screen (HTS) was examined, and the Z′ factor of 0.78 ( Fig. 2D ) for screening in a 384-well format was obtained, indicating that this assay is very suitable for HTS.
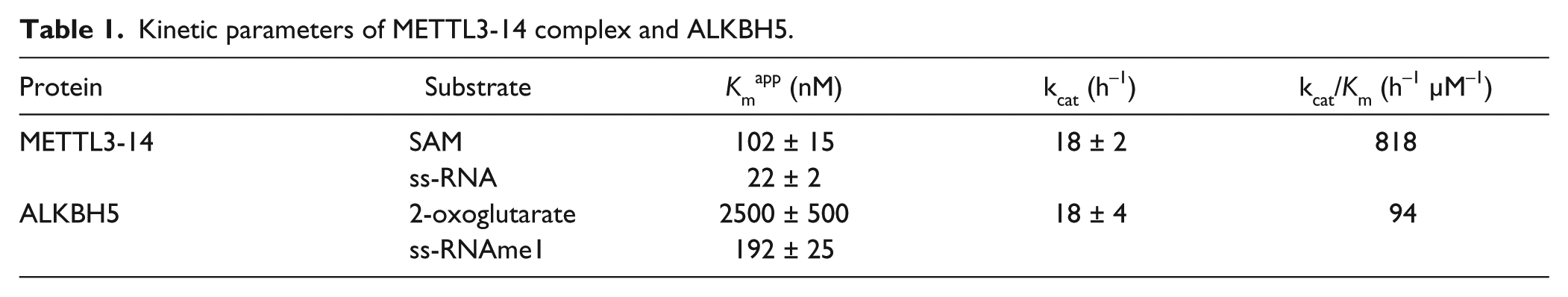
Kinetic parameters of METTL3-14 complex and ALKBH5.
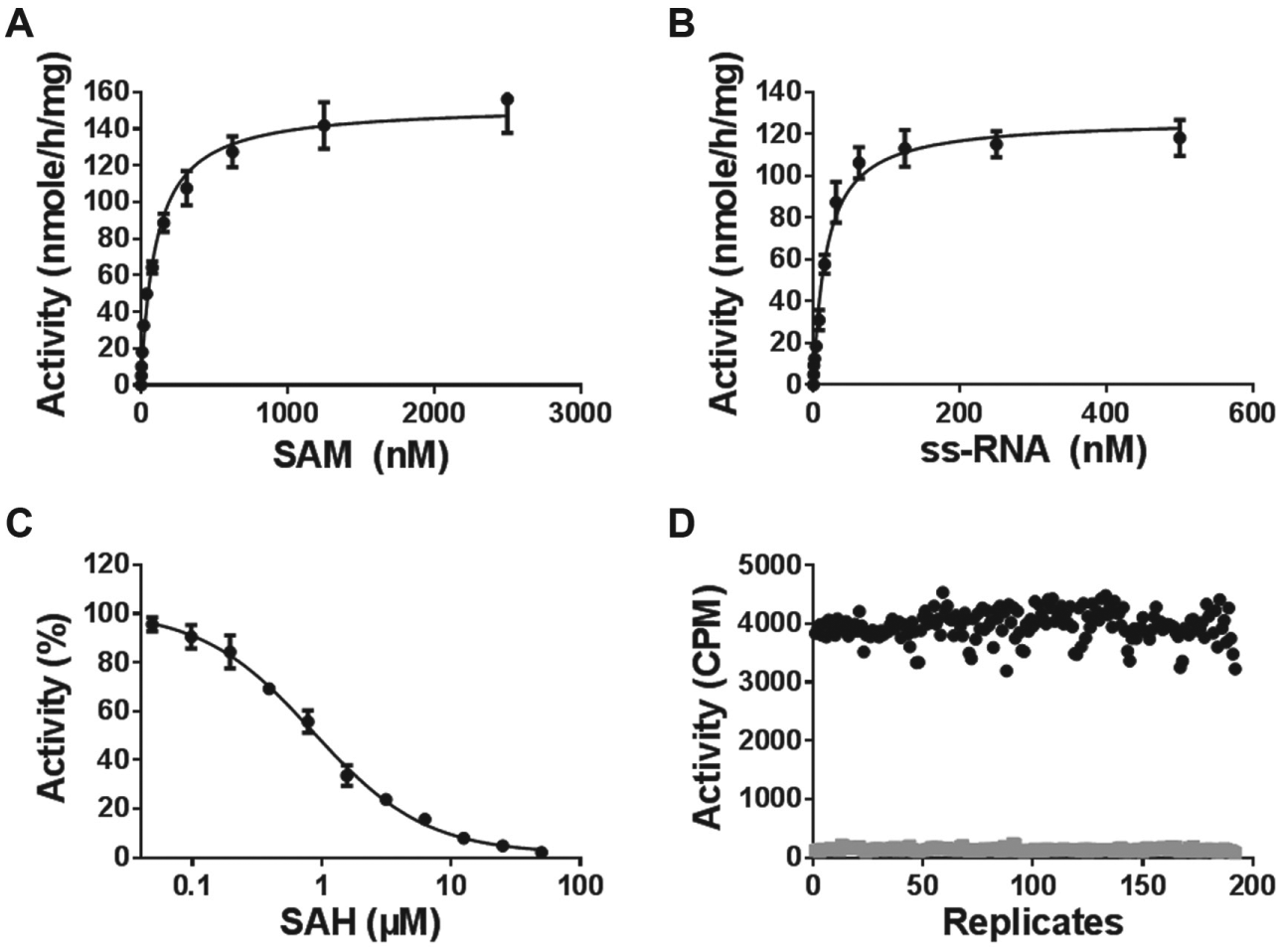
Kinetic parameter determination of the METTL3-14 complex. Km values for the METTL3-14 complex were determined for SAM (
Assay Optimization for ALKBH5
m6A–ss-RNA demethylase activity of ALKBH5 is essential to regulate the level of m6A modification for normal functions. To develop a radioactivity-based activity assay for ALKBH5, we needed a 3H-m6A–ss-RNA substrate that was not commercially available. We therefore employed our optimized METTL3-14 assay to generate 3H-m6A–ss-RNA using biotinylated ss-RNA and 3H-SAM. The efficiency of methylation of ss-RNA was evaluated using the commercially available 3H-biotin, indicating that 5 µM of biotinylated ss-RNA was completely methylated in 1 h by using 250 nM of METTL3-14 complex (
Similar to METTL3-14, the higher the concentration of reducing agents DTT or TCEP, the less active ALKBH5 was ( Fig. 3A ). In contrast, the presence of SAH, NaCl, MgCl2, DMSO, and Triton X-100 ( Fig. 3B–F ) had no significant effects on ALKBH5 activity. The optimum pH was 7 to 7.5 ( Fig. 3G ). ALKBH5 activity was ferrous iron dependent ( Fig. 3H ), which was further activated by low concentrations of ascorbate ( Fig. 3I ).
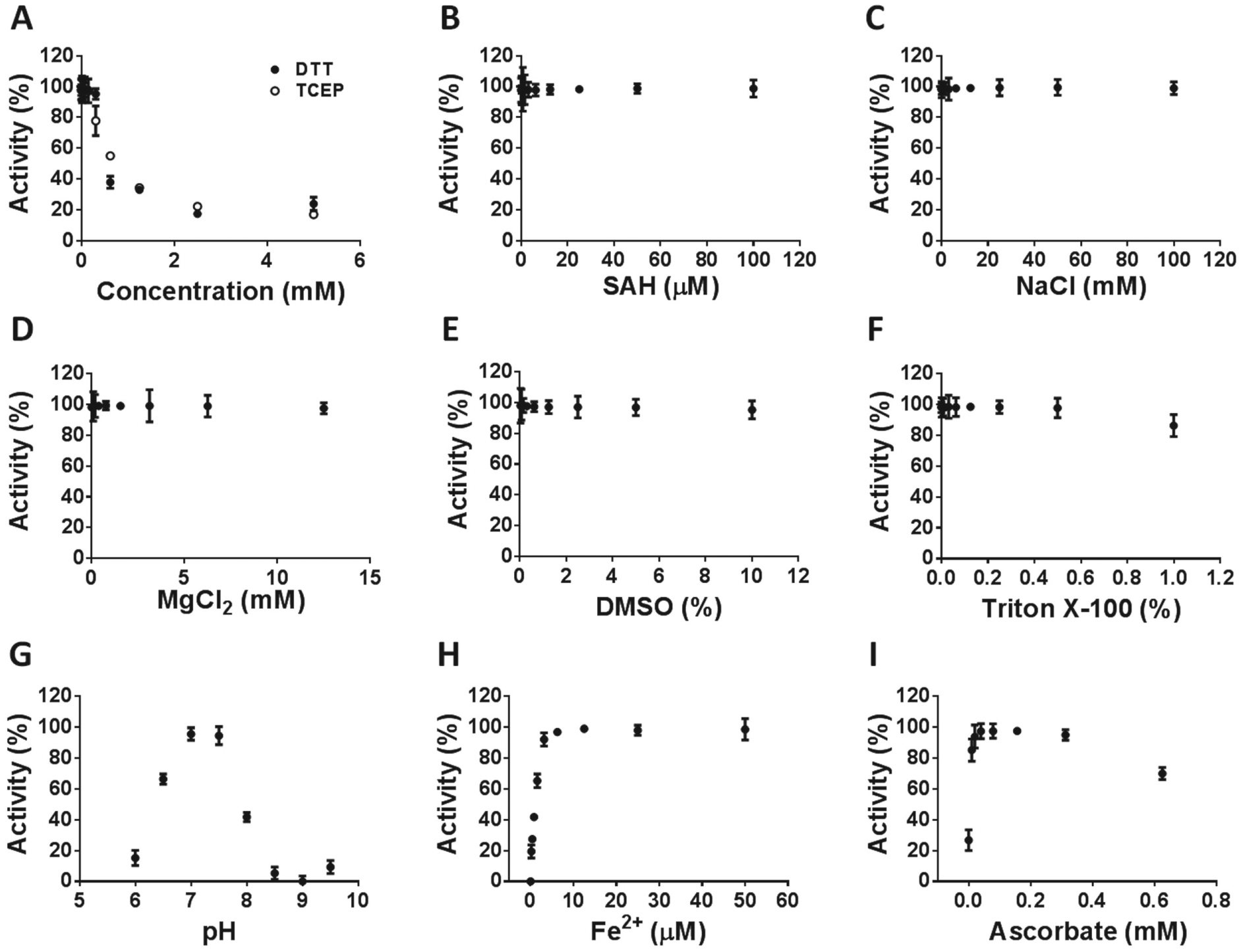
Effects of buffer additives and pH on ALKBH5 activity. The demethylase activity was measured to determine the optimum of reducing agents DTT and TCEP (
Kinetic Parameters for ALKBH5
Optimized assay conditions (20 mM Tris-HCl pH 7.0, 100 µM ascorbate, 20 µM Fe[II], 0.01% Triton X-100) were used for determination of kinetic parameters for ALKBH5 ( Table 1 ). The Km values were 2.5 ± 0.5 µM and 192 ± 25 nM ( Fig. 4A , B ) for 2-OG and m6A–ss-RNA, respectively, and the kcat value was 18 ± 4 h–1. IOX1, a broad-spectrum 2-OG oxygenase inhibitor, also inhibited ALKBH5 activity in a cofactor 2-OG competitive manner, with IC50 values of 0.5 ± 0.1 µM, 1.8 ± 0.5 µM, and 2.9 ± 0.3 µM at 2 µM, 5 µM, and 20 µM of 2-OG, respectively ( Fig. 4C ). The optimized assay was suitable for screening in a 384-well format with a Z′ factor of 0.77 ( Fig. 4D ).
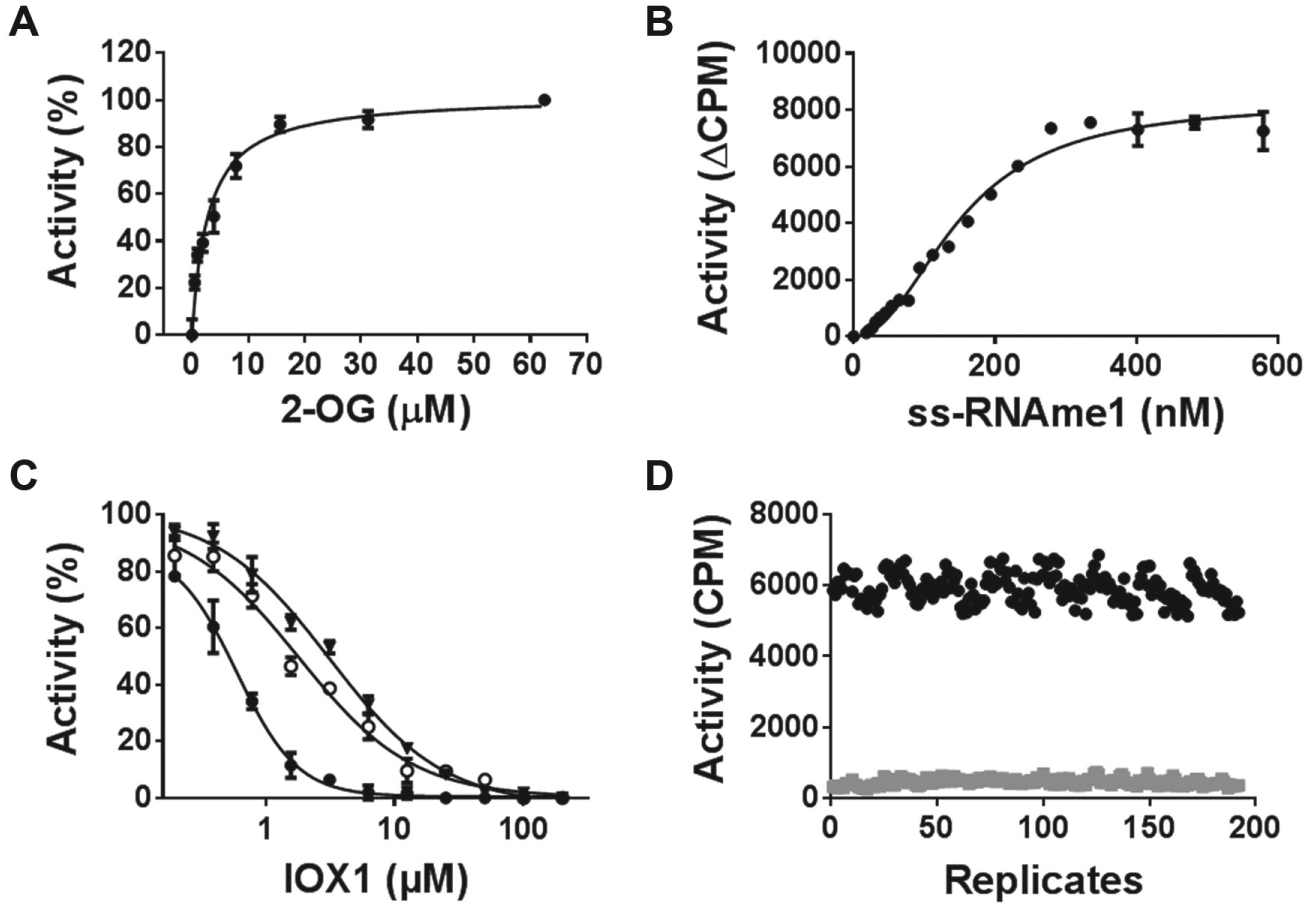
Kinetic parameter determination of ALKBH5. Km values of ALKBH5 were measured for cofactor 2-OG (
Discussion
Although the m6A mark on RNA was reported many years ago,1–3 recent progress in the discovery of components of methyltransferase complex responsible for methylation of N6-adenine on RNA,16,17 m6A-RNA readers, 13 and two m6A-RNA demethylases, FTO 20 and ALKBH5,11,28 created greater interest in investigating the roles that this reversible m6A-RNA modification plays in cells. Increasing evidence shows that m6A in mRNA affects mRNA splicing, trafficking, translation, and stability; thus, the reversible m6A modification in mRNA plays broad and critical roles in fundamental biological processes and disease development.5,30 METTL3 and METTL14 methyltransferase activities have been examined using mass spectrometry–based assays, which indicated that they form a stable heterodimer that is more active than each component individually. 16 Similar methods have also been used to assay demethylases. 11 A radioactivity-based cell assay was also used by tracing the mRNA modifications through analyzing the composition of the methyl-labeled nucleosides. 1
Discovery of potent and selective inhibitors of METTL3-14, ALKBH5, and FTO would be important for studying the consequences associated with the dysregulation of their activities and the m6A-RNA mark in cells. The natural product rhein was reported as an inhibitor of FTO with an IC50 value of 30 µM using m6A-containing 15-mer ss-RNA as substrate and a high-performance liquid chromatography (HPLC)–based assay. 31 Huang et al. 32 identified meclofenamic acid (MA) as a highly selective inhibitor of FTO (IC50: 8 µM) over ALKBH5 (no inhibition) using HPLC-based assays. 32 They also used a fluorescence polarization–based assay to test the displacement of ssDNA by MA. In this study, we reported on the full kinetic characterization of the METTL3-14 complex and the development of a radioactivity-based assay that is suitable for screening in a 384-well format. We also used the METTL3-14 complex to generate m6A-RNA substrate for ALKBH5 to determine its kinetic parameters and optimize a 384-well format screening assay. The kinetic parameters we reported here for ALKBH5 are comparable with those reported by He’s lab. 11 These assays can be used in screening large libraries of compounds in a 384-well format, which could have a higher throughput in comparison with mass spectrometry–based or HPLC-based assays. Radioactivity-based assays typically have fewer false-positives compared with fluorescence-based assays and are less laborious compared with mass spectrometry–based assays. Even in our academic setting, these assays allow for the screening of 4000 compounds per person per day. The kinetic parameters provided in this study as well as the highly reproducible assays pave the way to higher-throughput screening and identification of more potent inhibitors of METTL3-14 and ALKBH5 to further investigate their roles in cells.
Footnotes
Acknowledgements
The authors thank Dr. Chuan He for providing the METTL3 and METTL14 clones.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The SGC is a registered charity (No. 1097737) that receives funds from AbbVie, Bayer Pharma AG, Boehringer Ingelheim, Canada Foundation for Innovation, Eshelman Institute for Innovation, Genome Canada, Innovative Medicines Initiative (EU/EFPIA; ULTRA-DD grant No. 115766), Janssen, Merck & Co., Novartis Pharma AG, Ontario Ministry of Economic Development and Innovation, Pfizer, São Paulo Research Foundation-FAPESP, Takeda, and the Wellcome Trust.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
