Abstract
For a therapeutic antibody to succeed, it must meet a range of potency, stability, and specificity criteria. Many of these characteristics are conferred by the amino acid sequence of the heavy and light chain variable regions and, for this reason, can be screened for during antibody selection. However, it is important to consider that antibodies satisfying all these criteria may be of low frequency in an immunized animal; for this reason, it is essential to have a mechanism that allows for efficient sampling of the immune repertoire. UCB’s core antibody discovery platform combines high-throughput B cell culture screening and the identification and isolation of single, antigen-specific IgG-secreting B cells through a proprietary technique called the “fluorescent foci” method. Using state-of-the-art automation to facilitate primary screening, extremely efficient interrogation of the natural antibody repertoire is made possible; more than 1 billion immune B cells can now be screened to provide a useful starting point from which to identify the rare therapeutic antibody. This article will describe the design, construction, and commissioning of a bespoke automated screening platform and two examples of how it was used to screen for antibodies against two targets.
Keywords
Introduction
The requirements and characteristics for therapeutic antibodies are now many fold. The biochemical and biophysical properties of antibodies need to be matched by potent therapeutic activity. These characteristics combine to increase the chances of the therapeutic antibody being a success in the clinic. These increased requirements have meant that in many situations, particularly when target molecules are challenging to make antibodies against, more and more antibodies have to be sampled to find those antibodies that are suitable to become a therapeutic entity. Using hybridoma to make monoclonal antibodies1,2 is limited by the poor efficiency of the fusion process, which limits the size of the antibody repertoire that can be screened. 3 Phage display systems can screen large libraries of antibody genes 4 but lose the natural cognate pairing of the antibody heavy and light chains that is selected for in vivo. Despite these shortcomings, both hybridoma and phage panning methods have been automated to increase the number of antibodies screened.5–7 The inefficiency of the hybridoma and phage methods provided us with an opportunity to change the way we make monoclonal antibodies. The Antibody Discovery Group at UCB developed a B cell culture method,8,9 linked to homogeneous assays and a sensitive PCR process to clone out the cognate antibody V region genes. 10 This methodology has given us a greater ability to screen the antibody repertoire of an immunized animal, allowing the screening of more than a billion B cells in a campaign over 2 weeks, to select a panel of antibodies, that after further in vitro and in vivo secondary screening, have the potential to be therapeutic antibody candidates.
In this article, we describe the automation of UCB’s early core antibody discovery process, a platform built around high-throughput memory B cell culture screening, following immunization. The automation covers the very early stages of the antibody discovery process, the primary screening of B cell cultures to identify antigen binders. The automation of the primary screening has increased throughput, increased the number of B cells that can be screened at this stage, and the number of targets that can be prosecuted in parallel, while reducing the project timelines. Hardware automation has been linked together with sophisticated software that provides a highly intuitive, bespoke user interface and allows the process to be linked to UCB’s Laboratory Information Management System (LIMS), providing a seamless movement of data.
The automated screening platform is used to set up B cell cultures, screen these culture supernatants using a homogeneous fluorescence binding assay to identify “binders,” pick off and consolidate positive supernatants onto “master plates,” and then prepare B cells for subsequent cryostorage.
Following the automated screening, the consolidated supernatants are further tested in a range of carefully designed in vitro and in vivo secondary screening assays to select antibodies with desired characteristics (e.g., high-affinity, functional activity binding to a specific epitope, etc.). These are assays that are not currently automated due to their complexity. Following secondary screening, antigen-specific B cells are recovered from interesting wells using the fluorescent foci method. 10 A micromanipulator is used to pick specific, single B cells from which the antibody variable region gene sequences are directly recovered. Messenger RNA from the single B cells is converted to cDNA, and antibody variable region genes are amplified by PCR, cloned, and prepared for transient expression to confirm activity. Antibodies with affinities (KD) in the low pM range against a range of therapeutic targets are frequently recovered using this process.
Our new automated platform replaces a disconnected process that was limited to individual workstations serviced by operators manually moving plates between the various liquid-handling robots and plate readers. The informatics systems were not fully integrated with the automation, requiring many user interventions within the process. The high number of interactions by the user with the system was severely limiting in terms of throughput of plates and the ability of the system to expand. We have now optimized many aspects of our biological methodologies and introduced bespoke state-of-the-art automation to increase the reliability and efficiency of our novel platform. When combined with the rest of the antibody discovery process, it is hoped that this automation will then facilitate both first-in-class and best-in-class drugs for patients suffering with serious disease.
Materials and Methods
System Design
The automated screening system is divided into three work cells ( Fig. 1 ), each within its own class II safety cabinet, providing a clean air environment and protection to users. The first work cell is used to fill barcoded 96-well cell culture plates with B cells suspended in a novel supplemented growth medium. The second work cell is used to screen the B cell culture plates for the presence of target-specific antibody, whereas the third work cell is used to pick off positive cell culture supernatants from the B cell culture plates into master plates. The hardware and process scheduling are controlled by Overlord software via PAA’s Harmony graphical user interface. The interface takes the user through a succession of pages using a touch screen that allows the user to describe what he or she wishes to do, selecting experimental parameters from drop-down menus. The system has its own Oracle database that allows direct communication with the UCB LIMS.
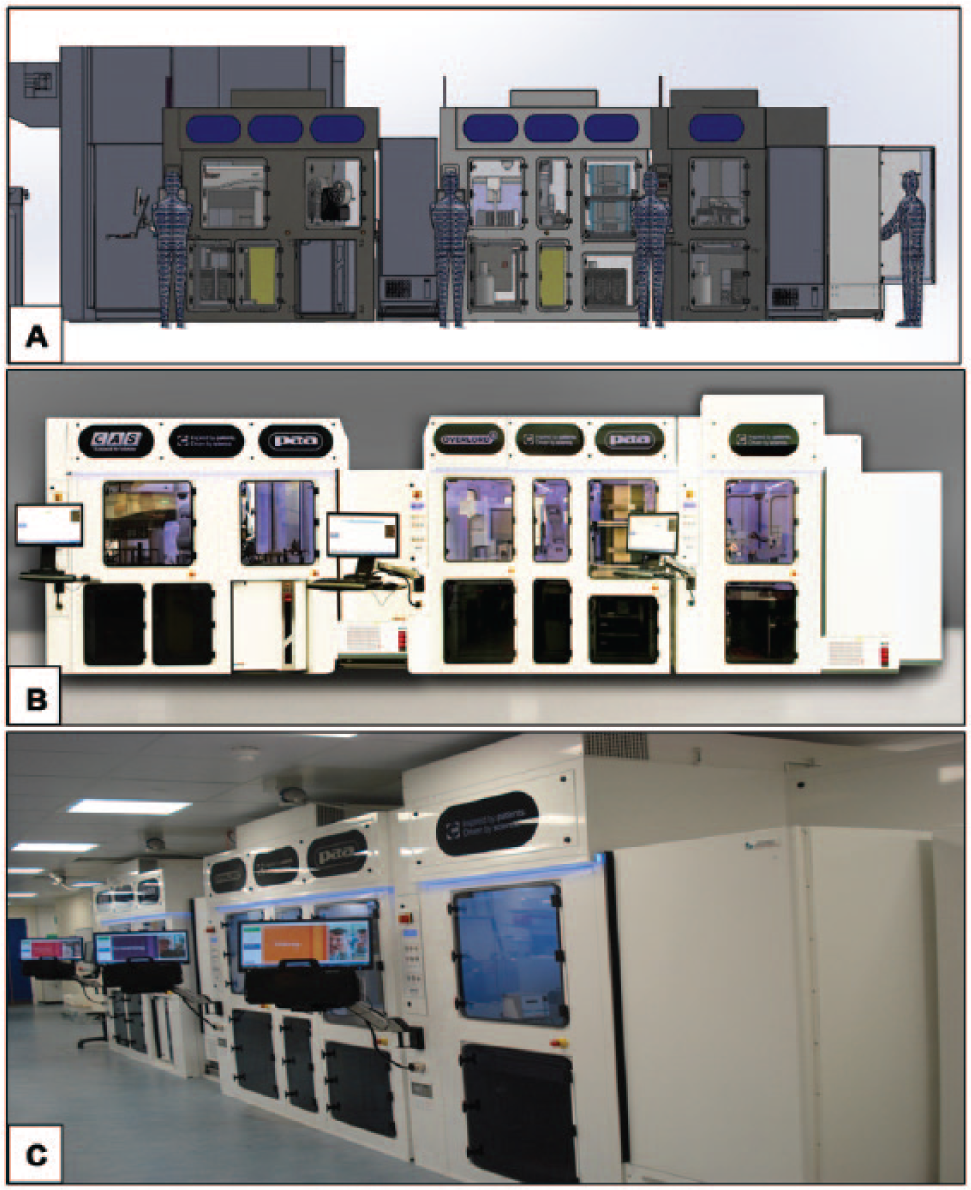

Automated antibody screening platform. (
The filling work cell consists of four Liconic STX 500 incubators. Each incubator has capacity to hold five hundred 96-well cell culture plates. At present, a 500-plate experiment, with approximately 108 B cells being cultured, is considered to be the maximum experiment size that can be handled in a single day. The movement of plates to different positions within the work cell is done by a Mitsubishi robotic arm that moves plates from the incubators to one of two Thermo Combi Multidrop dispensers. The Multidrop Combi dispensers dispense media containing B cells into the wells of the cell culture plates. There are two dispensers that allow for two different cultures, with a different immunization sample, for example, to be dispensed simultaneously. The cell cultures are kept in suspension at 37 °C in a water bath with a magnetic stirrer while filling occurs. Plate barcodes are uploaded to the LIMS, together with a record of the source of the B cells and the immunizing antigen.
The screening work cell contains a Liconic LPX 200, plate and tip box hotel, two Thermo Multidrop Combi dispensers, two TTP Labtech Mirrorball plate readers, and an Agilent Bravo liquid-handling robot. The cell culture plates are delidded and placed onto the Bravo, which transfers 10 µL of cell culture supernatant from four cell culture plates to one 384-well assay plate. The Multidrop Combi dispensers then add assay reagents to the assay plate before being incubated for 1 h and then read in the Mirrorball. The robot arm then transfers the cell culture plates into one of two Liconic STX 500 holding incubators between the screening and the picking work cell. The cell culture plates stay in these incubators until they are required for hit picking.
The picking work cell contains a Beckman Biomek NXP liquid-handling robot, a Thermo Alps 3000 plate sealer, and a Liconic STR 240 incubator. Cell culture plates are transferred from the buffer incubators into the picking work cell. The robotic arm transfers both cell culture plates and master plates onto the Biomek NXP™. The list of positive wells to be picked is provided as a simple text file through Spotfire to the Biomek. The Biomek transfers positive supernatants from cell culture plates to the master plate. The plates can either be sent into the −80 °C store after the addition of freezing media or be returned to the buffer incubator for immediate foci picking.
Running in parallel with the hardware is the scheduling software. As well as controlling the hardware, the software tracks all the plates within the system, the transfers of supernatants between plates, and interfaces with UCB’s LIMS. The software has to connect to six databases within the LIMS, pushing and pulling data in and out of the databases. The system has its own internal database that allows these transfers to be done directly, database to database, without the need to move text or XML files. The system is controlled via a PAA Harmony interface; this provides a simple, easy-to-follow graphical user interface, which means minimal training is required for the users to operate the system.
The software records the experiment parameters; the cell culture plate, assay plate, and master plate barcodes; the immunization samples; and the screening reagents. The software can e-mail notifications to the users during a run, requesting that they replenish cultures or change to a different B cell immunization sample. Overlord also controls the selection of protocols used by the Bravo and the Mirrorball. The upload of data from the Mirrorballs into the Activity Base is also controlled by Overlord. Assay data from the Mirrorballs is backed up daily into an Agilent ECM database. For a simple singleplex assay, the user has the option to input a threshold value directly into Overlord without having to create a picking list; this can be done before all the cell culture plates have been screened. This “picking on the fly” mode uses the assay data stored in the systems internal database controlled by Overlord and can speed up the throughput of plates through the system.
Immunizations
For experiment 1, three New Zealand White rabbits were immunized with 500 µg purified antigen A, and for experiment 2, four Sprague Dawley rats were immunized with 4 µg per shot of immunogen B (contained mostly antigen B but not 100% pure [data not shown]). The immunizations were performed using Freund’s adjuvant as described previously. 10 At termination, single-cell suspensions from the spleens were prepared and frozen in 10% DMSO in fetal calf serum at −80 °C.
B Cell Culture
The B cell culture process is similar to that described previously.8,9 The culture uses an X-ray-treated mutant EL4 feeder cell line, together with an optimized proprietary supplement to support B cell expansion and activation in vitro. Two experiments were set up. Experiment 1 used B cells harvested from spleens taken from three rabbits immunized with a soluble antigen A. Splenocytes were plated out onto 96-well cell culture plates at the following densities: rabbit 1, 48 plates at 1500 cells/well; rabbit 2, 52 plates at 2400 cells/well; and rabbit 3, 52 plates at 1500 cells/well (152 plates in total). Experiment 2 used B cells harvested from spleens taken from four rats immunized with a soluble antigen B. Each spleen was plated out onto seventy-five 96-well cell culture plates, 300 plates in total. Rat 1 was plated at 5000 cells/well, rat 2 at 3800 cells/well, rat 3 at 4000 cells/well, and rat 4 at 5500 cells/well. The dispensed volume was fixed at 200 µL per well. The cell plating density was determined according to animal sera titers, to ensure a hit rate of less than 10%, which ensures the clonality of the hit. The B cell culture plates were incubated for 7 d; during this time, sufficient antibody was secreted into the media to allow the assaying for antigen binding.
Primary Screening Assay
A simple homogeneous antigen binding assay 11 was used to screen the cell culture supernatants. Both experiments were screened using the biotinylated target antigen bound to the surface of a Superavidin coated 10 µm polystyrene sphere (Bangs Laboratories, Fishers, IN). A 10 µL sample of cell culture supernatant was added to each well of a 384-well assay plate. To this sample, 10 µL of antigen-coated beads was added together with 10 µL of an AlexaFluor 488-conjugated goat anti-rabbit IgG Fcγ-specific reagent (Jackson ImmunoResearch, West Grove, PA) for experiment 1 or AlexaFluor 488-conjugated goat anti-rat IgG Fcγ-specific reagent (Jackson ImmunoResearch) for experiment 2. In both cases, the final dilution of labeled antibody was 1 in 5000. The cell culture supernatant, antigen-coated beads, and labeled antibody were incubated for 1 h before being read on the Mirrorball ( Fig. 2 ).
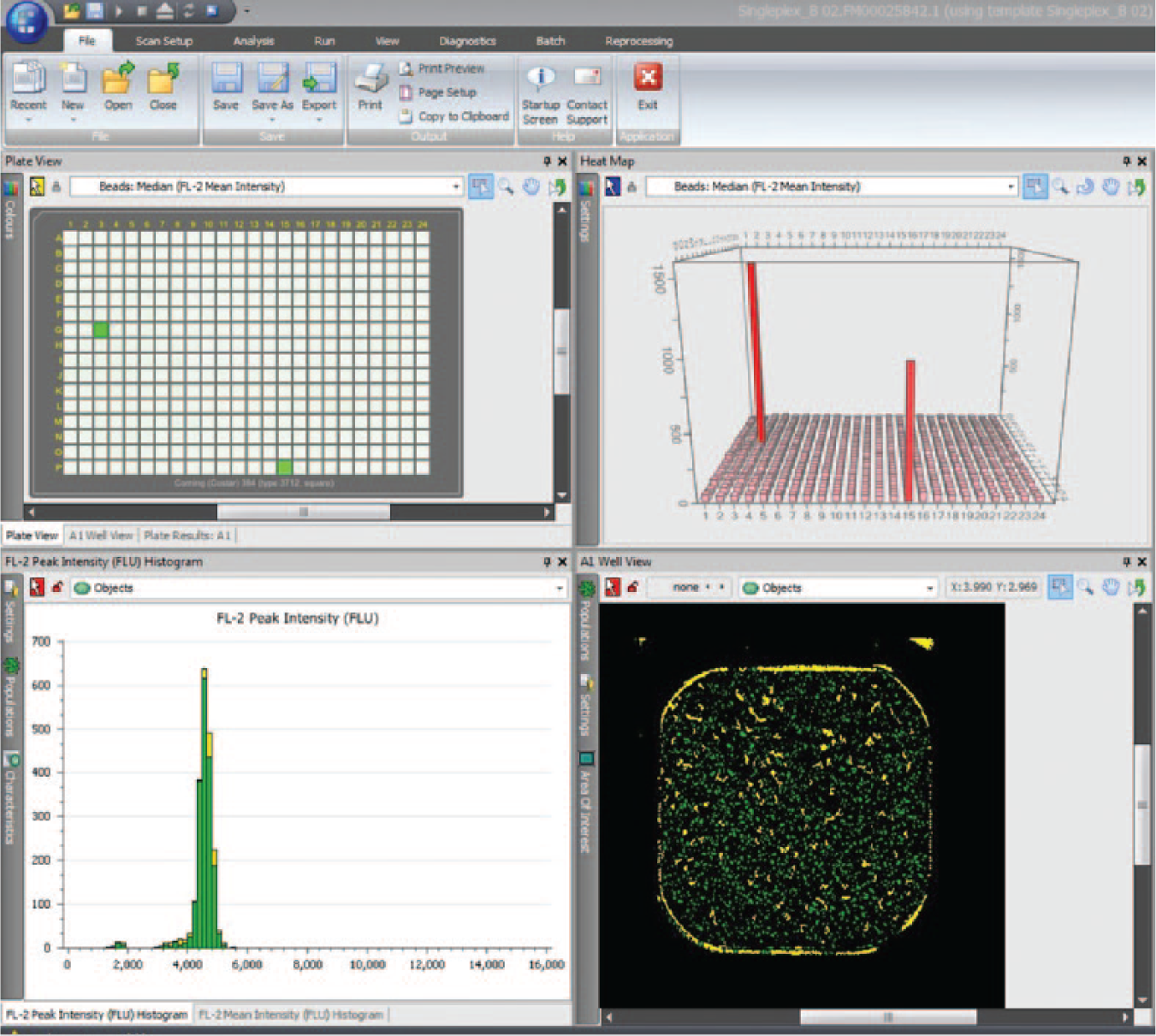
Screenshot of Mirrorball Cellista software showing data from screening assay. The two upper pains show two hits on the 384-well assay plate. The lower left pain shows the peak fluorescent intensity for well G3. The lower right pain shows the scanned image from well G3.
The data from the assay was analyzed using Spotfire and used to generate a picking list of positive wells. The positive supernatants were transferred onto a 96-well master plate, and the cells were cryopreserved in situ at −80 °C. The supernatants on the master plate were used for secondary assays to look for antibody function, affinity, cross-reactivity, and other properties.
Results
The data presented in this article describes experiments undertaken during the commissioning phase of the robotic system. Prior to screening the B cell cultures, serum from the immunized animals was tested in a simple antigen-binding enzyme-linked immunosorbent assay ( Fig. 3 , Fig. 4 ). The sera titers in experiment 1 were considered to be relatively good (binding detected out to a dilution of 1:106), indicating an effective and robust immune response. However, the sera titers in experiment 2 were moderate in comparison (binding detected out to a dilution of 1:10 000). This provided us with two different sources of B cells from which to screen against the two targets. Based on the sera titers, we would expect to see significantly higher hit rates in experiment 1, and indeed this is what we observed ( Table 1 ).
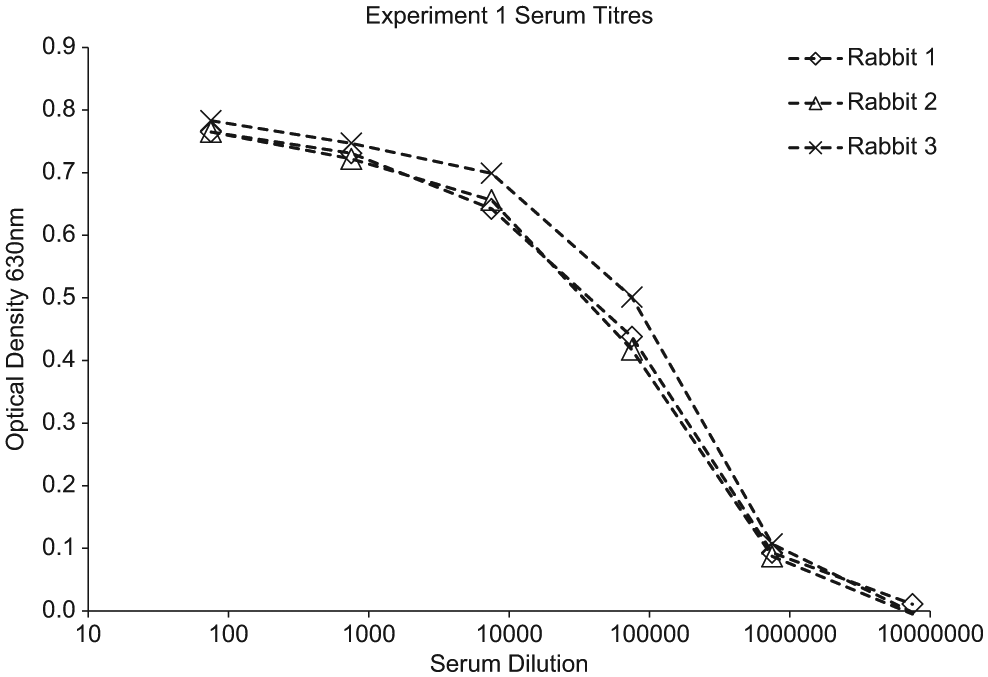
Data from antigen-binding enzyme-linked immunosorbent assay (ELISA) on sera from immunized rabbits. Antigen was coated onto Nunc Maxisorb ELISA plates at 5 µg mL in carbonate coating buffer, 100 µL per well overnight at +4 °C. The plates were blocked with 1% PEG 2000 (VWR, Radnor, PA) and 0.1% Tween20 in phosphate-buffered saline (PBS). Sera were diluted in PBS/PEG/Tween and put onto the blocked plate 100 µL per well for 1 h ambient temperature. The plate was washed in PBS + 0.1% Tween before the addition of a peroxidase labeled goat anti-rabbit IgG antibody (Jackson ImmunoResearch) diluted 1 in 5000 in PBS/PEG/Tween. The conjugate antibody was incubated for 1 h at room temperature before the plate was washed again. The bound antibody was detected using 3,3′,5,5′-Tetramethylbenzidine (TMB) liquid substrate (Sigma-Aldrich, St. Louis, MO). The plate was read at 630 nm.
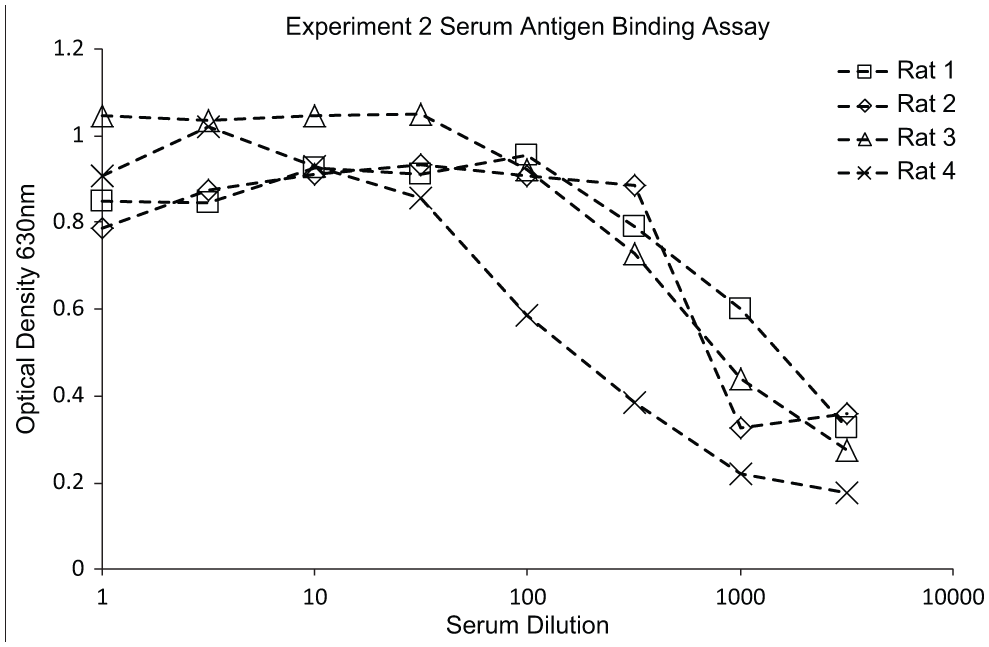
Data from antigen-binding enzyme-linked immunosorbent assay (ELISA) on sera from immunized rats. Antigen was coated onto Nunc Maxisorb ELISA plates at 5 µg mL in carbonate coating buffer, 100 µL per well overnight at +4 °C. The plates were blocked with 1% PEG 2000 (VWR) and 0.1% Tween20 in phosphate-buffered saline (PBS). Sera were diluted in PBS/PEG/Tween and put onto the blocked plate 100 µL per well for 1 h ambient temperature. The plate was washed in PBS + 0.1% Tween20 before the addition of a peroxidase labeled goat anti-rat IgG antibody (Jackson ImmunoResearch) diluted 1 in 5000 in PBS/PEG/Tween. The conjugate antibody was incubated for 1 h at room temperature before the plate was washed again. The bound antibody was detected using 3,3′,5,5′-Tetramethylbenzidine (TMB) liquid substrate (Sigma-Aldrich). The plate was read at 630 nm.
The number of positive wells from each identified by the primary screen and the number of wells elected for picking from experiments 1 and 2.
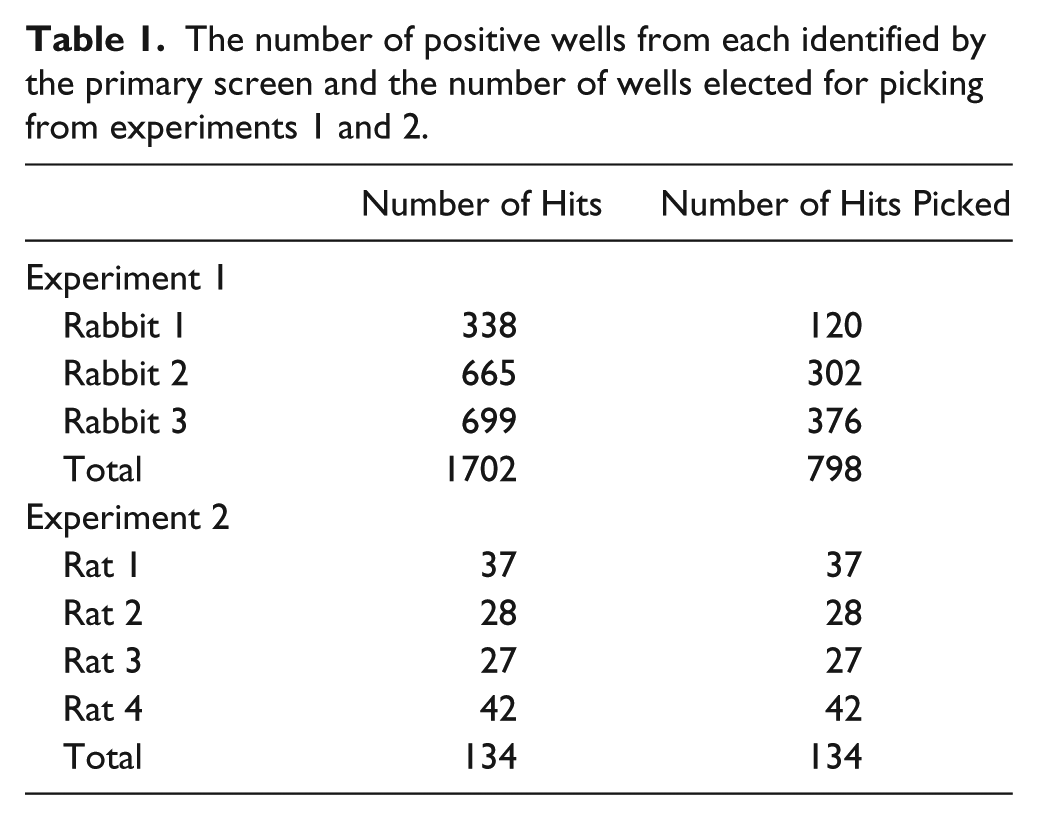
From the 152 plates screened in experiment 1, 1702 wells were deemed to be positive, exceeding a fluorescence threshold value of >100 in the Mirrorball assay ( Fig. 2 ). This represented a hit rate of 11.6% (percentage of wells containing specific antibody). Because of the relatively high hit rate observed in this screen, only the top 798 hits were selected for picking onto master plates. Experiment 2 had a lower number of hits, with only 134 positive wells being identified (hit rate of 0.46%). The screening hit rate varied between each rat ( Fig. 4 ). However, the hit rates were broadly consistent with the observed serum titers from each rat; this hit rate reflects the antigen-specific IgG+ memory B cell frequency within the total B cell repertoire. For this experiment, all the positives from the primary screen were selected for picking into master plates.
In rodents, approximately 50% of splenocyte cells are thought to be B cells. 12 Therefore, the 300-plate rat experiment sampled approximately 7 × 107 B cells. Assuming the rabbit spleen is similarly populated with B cells, the 150-plate rabbit experiment sampled approximately 2 × 107 B cells. This deep mining approach allowed for the identification of several hundred binders from both rats and rabbits from which further antibody characterization could be performed. The very-early-stage filling, screening, and hit picking process was fully automated and provided minimal user interaction. The examples provided here were well within the limits of the robot’s capability, with an experiment size of 500 plates being possible without experiencing overlap with other scheduled experiments.
Discussion
The experiments described here show two markedly different examples of antibody screening in two relevant animal species and against two different target antigens. In experiment 1, the immunizations produced a good serum response in the rabbits. This was expected due to the divergence in homology between the host rabbit and the human target antigen amino acid sequence (i.e., no tolerance issues to overcome) and the availability of large amounts of purified protein for immunization. The good sera titer translated to a relatively high hit rate in the primary screen. Experiment 2, on the other hand, is an example of a much more challenging target antigen. This target was immunized at a relatively low dose due to availability of the sample (4 µg rather than the target dose of 50 µg), and the preparation was thought to contain impurities (data not shown). The protein is also relatively well conserved between species (85–90%). As expected, the serum titers from the immunized rats were relatively low, and this was reflected in the primary screen hit rate. The automated system is ideally suited to finding antibodies to both these target antigens, especially those exemplified by antigen 2 where the immune response is weak and the hit rate low. When the desired antibody is very rare, the system allows for a large number of B cell culture plates to be screened rapidly. The system requires little user intervention and continues to run outside of normal working hours. The automated platform can effectively mine out an animal’s entire immune B cell repertoire to enable subsequent discovery of the highest quality antibodies.
The work described here provides a description of a novel antibody discovery platform that combines a novel high-throughput B cell culture screening method and the identification and isolation of single, antigen-specific IgG-secreting B cells through a proprietary technique called the fluorescent foci method. Using state-of-the-art automation to facilitate primary screening, extremely efficient interrogation of the natural antibody repertoire is made possible; in fact, more than 1 billion immune B cells can now be screened within a 2-wk period. This automated screening provides a starting point in the antibody discovery process, to identify a panel of target antigen, binding antibodies. These antibodies can be further screened, through carefully designed secondary and tertiary assays, and engineered to further refine their profiles and facilitate their development into a drug.
UCB has designed, commissioned, and installed a series of linked, automated work cells into a new antibody discovery laboratory at its Research Centre of Excellence in Slough, Berkshire, U.K. By automating the early steps of the antibody discovery process (primary screening), UCB has been able to take advantage of efficiency savings, increased throughput, and increased capacity, allowing us to prosecute more projects in parallel and affect a broader spectrum of disease mediators. This also results in reduced project cycle times, helping us more rapidly deliver treatments to patients. The automation gives improved process consistency and accuracy, ensuring quality in our screening campaigns.
The integration of the software with the LIMS has also had a beneficial effect on the screening process, with fewer user interventions to move data, thereby minimizing data handling errors and inconsistencies. The automated platform provides impressive sampling power, with the ability to sample 108 B cells in a single experiment.
We believe this integrated approach to antibody screening using our novel discovery processes has significant advantages over other methods of antibody discovery.
Footnotes
Acknowledgements
The authors would like to thank Mike Collins, Ian Pye, Jon Newman-Smith, and Jeremié Fizet at Peak Analysis and Automation for the design and build of the robotic system and the scheduling software. We would also like to thank Charles Casterlain and Mike Kent from UCB’s IT Department for their work on the software integration and to thank Anthony Hammett from UCB’s Informatics Department who worked on the data handling.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
